Abstract
Bacterial persisters, usually slow-growing, non-replicating cells highly tolerant to antibiotics, play a crucial role contributing to the recalcitrance of chronic infections and treatment failure. Understanding the molecular mechanism of persister cells formation and maintenance would obviously inspire the discovery of new antibiotics. The significant upregulation of Mycobacterium tuberculosis Rv3290c, a highly conserved mycobacterial lysine ε-aminotransferase (LAT) during hypoxia persistent model, suggested a role of LAT in persistence. To test this, a lat deleted Mycobacterium smegmatis was constructed. The expression of transcriptional regulator leucine-responsive regulatory protein (LrpA) and the amino acids abundance in M. smegmatis lat deletion mutants were lowered. Thus, the persistence capacity of the deletion mutant was impaired upon norfloxacin exposure under nutrient starvation. In summary, our study firstly reported the involvement of mycobacterium LAT in persister formation and possibly through altering the intracellular amino acid metabolism balance.
Similar content being viewed by others
Introduction
Bacterial persistence, a ubiquitous bacterial physiological state phenotypically tolerant to antibiotics, is increasingly recognized as the culprit of the intractability of chronic and relapsing infections. Bacterial persisters are specialized survivors genetically identical to nontolerant kins but under a non-growing or extremely slow-growing, non-replicating dormant state1. Sequestration of the targets instead of targets mutation contribute to the antibiotics tolerance of persister2. Persister cells are stochastically developed during mid-exponential phase3 under favorable conditions4. Despite more than 70 years after the first report of persisters5, the molecular mechanisms underlying bacterial persistence remain largely elusive. Several pathogens, such as Mycobacteria, Pseudomonas and Staphylococcus6,7,8, are remarkable persisters.
Mycobacteria have two notorious pathogens, M. tuberculosis (Mtb) for tuberculosis and M. leprae for leprosy and nontuberculous Mycobacterium (NTM)9. Many M. tuberculosis genes have been reported to be associated with persister formation and essential for persistence in mouse model, such as those involved in energy metabolism10,11,12,13,14, RelA (ppGpp synthase)15, M. tuberculosis PhoY2, a homolog of E.coli PhoU16,17, RpsA (S1 protein), a protein involved intrans-translation, proteasome PrcBA18,19, Toxin-antitoxin (TA) modules20,21,TA locus vapBC22, RelE20, other 88 putative TA system candidates in M. tuberculosis genome and 30 TA modules23,24,25,26,27. The differential expression of Rv3290c in persisters27,28 implicates an important role in persistence.
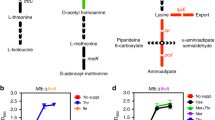
Lysine ε-aminotransferase (LAT) is a pyridoxal 5′-phosphate(PLP)-dependent enzyme that converts l-lysine to α- aminoadipate-δ-semialdehyde and glutamate, which is subsequently converted to Al-piperideine-6-carboxylic acid29,30. In the β-lactam-producing Actinomycetes, LAT has been shown to catalyze the first steps of β-lactam antibiotic biosynthesis pathway31. Rv3290c, encoding a Lysine ε-aminotransferase (LAT) in M. tuberculosis, was upregulated over 40-fold in nutrient-starved persistence models32. In this study, we initially reported the involvement of LAT in mycobacterium persister formation.
Materials and Methods
Antibiotics
Ampicillin, kanamycin, hygromycin, norfloxacin were bought from Sangon Biotech Co. and their stock solutions were freshly prepared, filter-sterilized and used at indicated concentrations.
Bacterial culture and starvation conditions
The bacterial strains and plasmid used in this study are shown in Table 1. E.coli strains were grown on LB broth agar or in LB broth, Mycobacterium smegmatis mc2155 was grown in 7H9 liquid medium (Difco) supplemented with 0.05% w/v Tween 80, 0.5%glycerol and 0.5%glucose or were grown on 7H10 agar supplemented with 1% glycerol and 0.5% glucose. The starvation culture condition as described32,33 with minor modifications. In brief, exponential phase cultures were pelleted and washed twice with 1 × PBS before being resuspended in 1 × PBS, transferred to standing flasks or microwell and incubated at 37 °C, 110 rpm. For viability determination during starvation, bacteria were cultured in 50 ml volumes in 250 ml bottles (Shuniu) and the number of cfu/ml was determined by plating serial dilutions onto 7H10 agar from triplicate cultures at several time points (0 h, 24 h and 72 h).
When required, the following antibiotics were used at the final concentration: ampicillin,100 μg/ml; kanamycin, 500 μg/ml for E.coli or 200 μg/ml for M. smegmatis; hygromycin, 50 μg/ml.
Knockout mutant construction and complementation
The lat gene of M. smegmatismc2155 was disrupted using specialized transduction previously described34. PCR and sequencing of latMsm was used to confirm the deletion. For knockout mutant complementation, the M. tuberculosis H37Rv Rv3290c coding region was amplified by polymerase chain reaction using the primers: 5′-CGCGGATCCGCGTCCTGCTATCATAGCGTCATG-3′, bearing a BamH I restriction site followed by the start codon of Rv3290c; and 5′-GGAATTCCATATGGAATTCCGGCTGCCTTACGTCACCAC-3′, consisting of the final three C-terminal amino acids of Rv3290c and a TAA termination codon followed by a Nde I restriction site. The gel-purified polymerase chain reaction product was digested with BamH I and Nde I, yielding a 1395-base pair BamH I/Nde I fragment. The fragments were ligated into the plasmid pALACE digested by BamH I and NdeI, to produce the plasmid pALACE-latMtb. Sequencing of pALACE-latMtb confirmed the correctness of the constructed fragment. Competent cells of latMsm mutant were prepared as described34 and pALACE containing latMtb gene was used to transform latMsm mutant competent cells. This was followed by electroporation35 into the mutant as previously described34. Transformed cells were streaked on 7H10 plates containing 100 μg/ml ampicillin and 50 μg/ml hygromycin. The desired complemented strain was identified by bacterial PCR and Western Blotting. To detect His-tagged LATMtb, bacterial pellets were harvested and disrupted by ultrasonication. Samples were then subjected to SDS-PAGE and the His-tagged LATMtb protein was detected by mouse anti-His antibodies (TIANGEN, China).
Amino acids determination
Overnight cultures of M. smegmatis mc2155 were diluted 1:100 in M9 medium and incubated at 37 °C on shaker (PEIYING DHZ-CA, TAI CANG SHI YAN SHE BEICHANG) at 110 rpm. Exponential phase cultures were harvested by centrifugation at 8000 rpm, 4 °C for 15 min. Harvested cells were washed three times with ddH2O and resuspended in 5 ml of ddH2O. Then the bacteria were pipetted into the dialysis tube and dialyzed in ddH2O at 4 °C for 24 h. Cells were collected and transferred to freeze-dried reagent bottle for freeze dehydration by vacuum pump. The procedure was from reference36 with slight modification. Briefly, 50 mg dried sample was put into a 15 * 150 mm testtube and then 6 ml of 6 M HCl were added into the testtube containing bacterial cells. The upper part in the testtube was removed and the testtube was sealed after 10 min vacuumization. The treated testtube was hydrolyzed for 22 hours in a 110 °C ± 1 °C oven. The testtube was taken out and cooled to room temperature, mixed and filtered.1 ml of filtrate was put into a 50 ml beaker and waterbathing evaporated at 60 °C, 2-fold diluted by adding 0.02 M HCl, the sample was filtered by 0.22 um membrane and loaded into a Hitachi L-8800 amino acid analyzer. The analysis cycle is 53 min, using two columns during the analysis process: (1) Separation column: (4.6 mm × 60 mm) Eluent flow rate is 0.4 ml/min, the column temperature was 70 °C, column pressure was 11.627 MPa. (2) Reaction column: Ninhydrin and ninhydrin buffer flow was 0.35 ml/min, the column temperature was 135 °C, column pressure was 1.078 MPa.
MIC assay and drug treatment of cultures
The MICs of antibiotics were determined by using serial two-fold dilution of the antibiotics in 7H9 medium as previously described in reference37. The initial cell densities were 108cfu/ml of exponential culture and the samples were incubated for 3 days at 37 °C.The MIC was recorded as the minimum drug concentration abolishing visible growth. For drug treatment assay, 50 ml M. smegmatis was cultured under starvation conditions in 250 ml bottles. At 0, 24 h and 72 h time point, one milliliter of a starvation culture of M. smegmatis was diluted 10 times in 1 × PBS and 1 ml was aliquoted per well of a 48-well plate, every sample with 3 repeats. Norfloxacin was added to duplicate wells of cultures at final concentration of 20 μg/ml. Control wells for cultures received no drug or no cells. Cultures were incubated with or without drug at 37 °C for 24 h and 48 h, followed by serial diluting and plating on 7H10 agar to determine bacterial viability.
RNA Isolation and reverse transcription-PCR (RT-PCR)
M. smegmatis cultured under starvation conditions in 50 ml volumes in 250 ml bottles (Shuniu).Three 50 ml cultures were harvested by centrifugation after 24 h and 72 h of starvation. Control samples were prepared by washing log-phase cultures twice with PBS then resuspended in PBS, as described for the preparation of starved bacteria and harvesting 50mlculture by centrifugation at time zero (t = 0). Pellets were pulverized in liquid nitrogen and homogenized in Trizol solution (Invitrogen) and RNA was isolated according to the manufacturer’s instructions. The subsequent steps were performed according to the reference38.
Real-Time PCR
1 μg of total RNA was reversely transcribed to cDNA using a first strand cDNA synthesis kit (Roche) according to the manufacturer’s instructions. Resultant cDNA was used for real-time PCR. Advanced SYBRGreen Supermix (BIO-RAD) were used to quantify cDNA in a 20 μL reaction containing 10 μM each primer, 10 μl supermix (2X), 4 μL cDNA. Primers used are listed in Table 1. Copy numbers of MSMEG_1764 mRNA were normalized with copy numbers of sigA mRNA. Each reaction was run in triplicate in Bio-Rad CFX-96 Real-Time Detection System with the following parameters: 95 °C for 2 Min, 40 cycles of 95 °C for 10 Sec and 64 °C for 40 Sec.
Results
lat is conserved among Mycobacteria
lat is conserved among M. tuberculosis, M. marinum, M. leprae, M. bovis BCG, M. smegmatis, M. canettii and Rhodococcus erythropolis PR4 by BLAST analysis. The amino acid identity between M. tuberculosis LAT and its homologs is greater than 59% in all cases, its neighboring gene lrpA is conserved too. Most neighboring genes of M. tuberculosis, M. marinum, M. leprae and M. bovis BCG are highly conserved (Fig. 1).
latMsm is upregulated under nutrient starvation in M. smegmatis
MSMEG_1764 is the homolog of Rv3290c in M. smegmatis and shares 77% identity with Rv3290c. Since lat was highly upregulated in M. tuberculosis under nutrient starvation, it is interesting to know whether this is the case in M. smegmatis. To this end, a nutrient starvation model according to Betts et al.39 was established. The transcription of lat in M. smegmatis at 0 h, 24 h and 72 h was measured. lat was upregulated 20 and 23-fold in M. smegmatis undergoing starvation after 24 and 72 hours, respectively (Fig. 2). The results showed that the expression pattern of M. smegmatis lat under nutrient starvation is the same as that in M. tuberculosis. Given M. smegmatis is a well-recognized facile surrogate to address M. tuberculosis biology40, in particular to study the persistence under nutrient starvation41, M. smegmatis was adopted as a model in our study.
The expression of M. smegmatis lat under nutrient starvation.
Cells of M. smegmatis were initially grown in 7H9 medium to log-phase and then washed by 1 × PBS and resuspended in 1 × PBS, incubate at 37 °C, 110 rpm. RNAs were extracted from bacteria harvested at indicated time points. RT-PCR was performed as described in Materials and Methods. Data are means ± s.d. of triplicates in one of at least three experiments.
lat (MSMEG_1764) is nonessential and can be deleted in M. smegmatis mc2155
To determine the effect of lat mutation on the persistence of mycobacterium, a lat knockout mutant was constructed by Xer site-specific recombination as described in the Materials and Methods section. M. smegmatis hygromycin-resistant colonies were selected and cultured consecutively for five generations. Primer lat-F1 and lat-R1 were then used to confirm the mutant genotype, with the wild-type strain as a control. The 1565 bp (1365 bp of the wild type latMS and extra 200 bp from the genome) fragment can be amplified from the wild-type, while there was only 449 bp (249 bp of ΔlatMS and extra 200 bp from the genome) amplicon from the latMsm knockout mutant (Fig. 3A). The latMsm knockout mutant was further confirmed by the absence of transcription product (Fig. 3B), indicating that the Δlat mutant of M. smegmatis was successfully constructed. Blastp shows that MSMEG_1764 is the homolog of Rv3290c in M. smegmatis and shares 77% identity with Rv3290c. Hence, we use M. tuberculosis Rv3290c gene for complementation assays. The recombination vector pALACE-Rv3290c in which Rv3290c is under the control of an ACE promoter was transformed into the Δlat mutant strain to get the complementary strain. Bacteria PCR result shows that a 1350 bp band was amplified in the complementary strain and no band for the parental strain (Fig. 3C).Western Blotting analysis using the anti-His antibody further confirmed the presence of the expressed ~52 kDa LATMtb-His fusion protein in the cell lysates of the complemented strain, while absent in the parental strain (Fig. 3D).
Construction of MSMEG_1764 knockout mutant and complement strains.
(A) PCR verification of the construction MSMEG_1764 knockout strain. Lanes: 1.Wild-type MS; 2. MSMEG_1764 knockout strain. (B) Verify the transcription of MSMEG_1764 knockout strain by RT-PCR. Wild type and ΔlatMsm strains were grown at 37C in MB 7H9 liquid medium to an OD600 of 0.8–1.0. Total bacterial RNA was isolated and subjected to RT-PCR to detect the expression of the latMsm gene. (C) Construction of Δlat-Rv3290c strain; Lanes: 1. Knockout strain complement with pALACE-Rv3290c; 2. Knockout strain complement with pALACE plasmid. (D) Western-blotting to confirm the expression of Rv3290c.Lysates were prepared from bacterial cells cultured as in (B), after 16 h induction and subjected to Western blotting to detect His-tagged Rv3290c protein using mouse anti-His antibody.
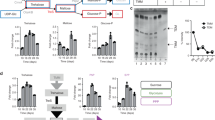
lat deleted M. smegmatisis is hypersensitive to norfloxacin and shows declined persistence under nutrient starvation
LAT was profoundly upregulated during M. tuberculosis starvation persistence model. The effect of lat deletion on M. smegmatis persistence remains elusive. To test this, we first measured the MIC of norfloxacin, rifampicin and isoniazid, the MIC value showed that the deletion of lat has no effect upon the antibiotic sensitivity for the M. smegmatis wild-type and complement strain (2 μg·ml−1 for norfloxacin, 8 μg·ml−1 for rifampicin and 4 μg·ml−1 for isoniazid). The deletion of genes involved in persistence usually has no effect upon MIC42,43. To test the effect of lat deletion on the MIC, the strains were exposed to high concentration antibiotics. The survival rate of ΔlatMsm strain is lower than wild-type and complement strain under 20 μg·ml−1 norfloxacin exposure (Fig. 4A).
Survival of strains under norfloxacin exposure in 1 × PBS buffer.
Wild type, complement strain and ΔlatMsm strain treated with norfloxacin after 0 h (A), 24 h (B) and 72 h (C) starvation. Data are shown as means ± SD of triplicate wells. Similar results were obtained in three independent experiments. (D) Starvation time and persister formation. Log phase cultures of wild type, complement and latMsm knockout strain were washed by 1 × PBS and then cultured under starvation condition. Data points are averages of 3 independent experiments. **P < 0.01, *P < 0.05, Student’s t test, significant difference from the ΔlatMS and the wild type strain.
Since LAT was significantly upregulated under nutrient starvation in M. tuberculosis. It is interesting to know whether the deletion of lat will compromise the survival of M. smegmatis under starvation. Therefore, we tested the viability of mutant strain in 1 × PBS buffer as described by Betts et al.39. As shown in Fig. 5, no difference can be seen between wild-type and ΔlatMsm strain after 72 hours starvation. To explore whether the deletion of latMsm only affect the persistence of mutant strain in high antibiotic concentrations, strains were subjected to antibiotic exposure and the ratio of persisters was determined. The 24 hours and 72 hours-starved cultures were exposed to norfloxacin for 48 h and the number of survived strains was assessed. We can see that 24 h and 72 h (Fig. 4B,C) starvation cultures of three strains exhibit higher level of norfloxacin persistence than 0 h time point. With the increase of starvation, wild type and complement strains persisted, but not ΔlatMsm strain (Fig. 4D).Complementation of the mutant with Rv3290c restored the phenotype of wild type strain. After 24 h and 72 h starvation, all strains showed a higher tolerance to norfloxacin than non-starvation cultures. This result indicated that deletion of latMsm failed to affect the viability under nutrient starvation, but can impair the persistence of the mutant strain under norfloxacin treatment.
Deletion of latMsm resulted in the decrease of intracellular amino acid content.
(A) Determination of intracellular amino acid content in WT and ΔlatMsm strain. (B) The transcription level of lysA and lrpA in wild type and ΔlatMsm strains. Data are shown as means ± SD of triplicate wells. Similar results were obtained in three independent experiments. n.s indicates no significant difference (P ≥ 0.1), *P < 0.05, Student’s t test, significant difference from the ΔlatMS and the wild type strain.
Inactivation of latMsm decreased the intracellular amino acids content
13C-isotope profiling of the persisters of Staphylococcus aureus revealed an active amino acid anabolism in this subpopulation, including Ala, Asp, Glu, Ser, Gly and His44. Dysregulation of intracellular amino acids level has been shown to lower the survival capability within macrophage exemplified by M. tuberculosis pknG mutants, which encodes protein serine/threonine kinase involved in the regulation of amino acids level45. To examine whether the inactivation of MSMEG_1764 has an effect on the amount of intracellular amino acids, the intracellular levels of the amino acids were determined for the wild-type, mutant and complemented strains. As shown in Fig. 5A, the concentration of glutamic acid, glycine, methionine and total amino acids content in the ΔMSMEG_1764 mutant strain was decreased, but without significant differences for the lysine content between the mutant strain and the wild-type strains. This might be due to the feedback repression of lysA (Diaminopimelate (DAP) - decarboxylase, the enzyme involved in the last step of lysine biosynthesis) by the excess of lysine. This shows that the rate of lysine biosynthesis was regulated by the intracellular lysine amount46. To test whether lysA control the lysine level in M. smegmatis, we measured the expression level of lysA in these strains. As expected, the transcription of lysA in ΔlatMsm strain was decreased (Fig. 5B). Genomic context analysis showed that there is a transcriptional regulator leucine-responsive regulatory protein (LrpA) upstream of lat (Fig. 2), a regulator capable of directly binding to the upstream region of lat47. To test whether the transcription of lrpA was also decreased, RT-PCR was applied. The result showed that lrpA transcription was also lowered in Δlat strain (Fig. 5B).The results suggested that the deletion of lat in M. smegmatis decreased the intracellular amino acids amount and downregulated the transcription of lrpA.
relAMsm was downregulated in latMsm deletion strain under nutrient starvation
The stress alarmones guanosine 3′,5′-bispyrophosphate ((p)ppGpp), a key molecule in antibiotic resistance48 and bacterial persistence49, was found to accumulate rapidly under starvation. The relationship between relA and lrpA has previously been noted50,51.The expression of lrp is positively controlled by ppGpp, namely the lower ppGpp level means fewer Lrp50. Moreover, the expression of lrp was significantly reduced in the strains failed to produce ppGpp50. RelMtb, the M. tuberculosis bifunctional enzyme responsible for both (p)ppGpp synthesis and hydrolysis52 and has been shown to play an important role in the survival of bacteria during nutrient starvation condition53. To test whether the downregulation of lrpA resulted from the decreased expression of Rel in M. smegmatis, the transcription level of relMsm, the relMtb homolog of M. smegmatis54, was measured at different intervals under starvation. No significant difference can be spotted between wild type and knockout strain at 0 h starvation (Fig. 6A). But relMsm was markedly downregulated in ΔlatMsm strain after 24 h starvation (Fig. 6B).Complementation of M. smegmatis ΔlatMSm with the M. tuberculosis lat Rv3290c can partially restore the expression of relMsm. This result suggested that knockout latMsm has no effect upon the intracellular (p)ppGpp content under normal conditions, but can influence the synthesis and hydrolysis of (p)ppGpp under nutrient starvation.
Discussion
Bactericidal antibiotics usually can sterilize most bacteria rapidly. However, a sub-population will survive and be reactivated upon the withdrawal of the antibiotics55. Genes involving in the persisters formation are intensively studied56 both for fundamental insights and translational medicine ends.
Here we showed that lat gene MSMEG_1764, the homologue of Mtb lat Rv3290c is involved in the persister formation (specifically tolerance to norfloxacin) via mediating the intracellular amino acid contents and altering the expression of ppGpp synthase expression in M. smegmatis.
Inactivation of the homologue of Rv3290c in M. smegmatis leads to more sensitive to norfloxacin than the wild type strain (Fig. 4), but without change to the MIC (Data not shown). Generally, mutation of genes involved in persister formation should not alter the MIC, but only affect the persistence42,43. LAT was previously showed to be involved in the L-lysine metabolism57. The data here showed that the knockout of latMsm rendered a decrease of intracellular total amino acid, but without discernable effect on the accumulation of L-lysine (Fig. 5A). LAT is just one of the several genes involved in M. tuberculosis glutamate metabolism58, given the diversity and redundancy of genes to control the robustness of this important amino acid homeostasis, it is not quite unexpected that the disruption of LAT only slightly decreased the glutamate content in M. smegmatis.
Leucine-responsive regulatory protein (Lrp) is a global transcriptional regulator widespread among prokaryotes and modulates the expression of a variety of genes involved in metabolism during starvation, especially in the amino acid catabolism and anabolism59,60. The knockout of lrp in E.coli downregulated the expression of the amino acid metabolism-related genes61. In our study, we found the expression of lrpA was downregulated in latMsm knockout strain (Fig. 5B), which can explain why the intracellular amino acid content was decreased in knockout mutants. Previous study showed that lrpA is profoundly up-regulated during nutrient starvation conditions characteristic of persistent/latent phase in M. tuberculosis39, as well as in M. smegmatis (Fig. 5B). The persistence of lrpA inactivated Mycobacterium fortuitum was attenuated in a murine infection model62. The expression of lrp is stimulated by (p)ppGpp and LRP can regulate the biosynthesis of amino acid50. Here we have shown that the deletion of latMsm downregulated the expression of relMsm (Fig. 6). M. tuberculosis failed to produce (p)ppGpp under starvation was defective in long-term survival both in vitro53 and in vivo15. In our study, the downregulation of relMsm is coincident with the diminished bacterial tolerance to antibiotics.
In conclusion, we have shown that LAT is involved in persister formation in mycobacteria. Under nutrition starvation, (p)ppGpp is produced by RelMsm and accumulates within bacteria, then upregulates the transcription of LrpA gene, which in turn upregulates the protein level of LAT and response for amino acid metabolism. Intracellular amino acid level regulates the accumulation of (p)ppGpp and (p)ppGpp controls the dormant persister formation (Fig. 7). The lat deleted mutant strain failed to replenish the amino acid pool, then downregulated the synthesis of (p)ppGpp. This is largely due to the positive regulation of the transcription of lrpA by (p)ppGpp. The determination of the M. tuberculosis H37Rv LAT crystal structure and the identification of its active sites63 will promisingly facilitate the discovery of novel LAT inhibitors64. Our finding has shown that LAT is a new player in mycobacterial persistence provided a potential drug target for inhibitors against persisters.
Additional Information
How to cite this article: Duan, X. et al. Mycobacterium Lysine ε-aminotransferase is a novel alarmone metabolism related persister gene via dysregulating the intracellular amino acid level. Sci. Rep. 6, 19695; doi: 10.1038/srep19695 (2016).
References
Lewis, K. Persister Cells. Annual Review of Microbiology 64, 357–372 (2010).
Keren, I., Kaldalu, N., Spoering, A., Wang, Y. & Lewis, K. Persister cells and tolerance to antimicrobials. FEMS Microbiol Lett 230, 13–18 (2004).
Balaban, N. Q., Merrin, J., Chait, R., Kowalik, L. & Leibler, S. Bacterial persistence as a phenotypic switch. Science 305, 1622–1625 (2004).
Dorr, T., Lewis, K. & Vulic, M. SOS response induces persistence to fluoroquinolones in Escherichia coli. PLoS Genet 5, e1000760 (2009).
Bigger, J. Treatment of staphylococcal infections with penicillin by intermittent sterilisation. The Lancet 244, 497–500 (1944).
Gengenbacher, M. & Kaufmann, S. H. Mycobacterium tuberculosis: success through dormancy. FEMS Microbiology Reviews 36, 514–532 (2012).
Harrison, J. J., Turner, R. J. & Ceri, H. Persister cells, the biofilm matrix and tolerance to metal cations in biofilm and planktonic Pseudomonas aeruginosa. Environmental Microbiology 7, 981–994 (2005).
Chong, Y. P. et al. Persistent Staphylococcus aureus bacteremia: a prospective analysis of risk factors, outcomes and microbiologic and genotypic characteristics of isolates. Medicine 92, 98–108 (2013).
Cohen, N. R., Lobritz, M. A. & Collins, J. J. Microbial persistence and the road to drug resistance. Cell Host Microbe 13, 632–642 (2013).
Bryk, R. et al. Selective killing of nonreplicating mycobacteria. Cell Host Microbe 3, 137–145 (2008).
Dhiman, R. K. et al. Menaquinone synthesis is critical for maintaining mycobacterial viability during exponential growth and recovery from non‐replicating persistence. Molecular microbiology 72, 85–97 (2009).
Dhar, N. & McKinney, J. D. Mycobacterium tuberculosis persistence mutants identified by screening in isoniazid-treated mice. Proc Natl Acad Sci U S A 107, 12275–12280 (2010).
Deb, C. et al. A novel in vitro multiple-stress dormancy model for Mycobacterium tuberculosis generates a lipid-loaded, drug-tolerant, dormant pathogen. PLoS One 4, e6077 (2009).
McKinney, J. D. et al. Persistence of Mycobacterium tuberculosis in macrophages and mice requires the glyoxylate shunt enzyme isocitrate lyase. Nature 406, 735–738 (2000).
Dahl, J. L. et al. The role of RelMtb-mediated adaptation to stationary phase in long-term persistence of Mycobacterium tuberculosis in mice. Proc Natl Acad Sci U S A 100, 10026–10031 (2003).
Li, Y. & Zhang, Y. PhoU is a persistence switch involved in persister formation and tolerance to multiple antibiotics and stresses in Escherichia coli. Antimicrob Agents Chemother 51, 2092–2099 (2007).
Shi, W. & Zhang, Y. PhoY2 but not PhoY1 is the PhoU homologue involved in persisters in Mycobacterium tuberculosis. J Antimicrob Chemoth 65, 1237–1242 (2010).
Shi, W. et al. Pyrazinamide inhibits trans-translation in Mycobacterium tuberculosis. Science 333, 1630–1632 (2011).
Gandotra, S., Schnappinger, D., Monteleone, M., Hillen, W. & Ehrt, S. In vivo gene silencing identifies the Mycobacterium tuberculosis proteasome as essential for the bacteria to persist in mice. Nat Med 13, 1515–1520 (2007).
Singh, R., Barry, C. E. 3rd & Boshoff, H. I. The three RelE homologs of Mycobacterium tuberculosis have individual, drug-specific effects on bacterial antibiotic tolerance. J Bacteriol 192, 1279–1291 (2010).
Black, D. S., Irwin, B. & Moyed, H. S. Autoregulation of hip, an operon that affects lethality due to inhibition of peptidoglycan or DNA synthesis. J Bacteriol 176, 4081–4091 (1994).
Demidenok, O. I., Kaprelyants, A. S. & Goncharenko, A. V. Toxin-antitoxin vapBC locus participates in formation of the dormant state in Mycobacterium smegmatis. FEMS Microbiol Lett 352, 69–77 (2014).
Rosenberg, S. M., Ramage, H. R., Connolly, L. E. & Cox, J. S. Comprehensive Functional Analysis of Mycobacterium tuberculosis Toxin-Antitoxin Systems: Implications for Pathogenesis, Stress Responses and Evolution. PLoS Genetics 5, e1000767 (2009).
Hu, Y. et al. Detection of mRNA transcripts and active transcription in persistent Mycobacterium tuberculosis induced by exposure to rifampin or pyrazinamide. J Bacteriol 182, 6358–6365 (2000).
Muttucumaru, D. G., Roberts, G., Hinds, J., Stabler, R. A. & Parish, T. Gene expression profile of Mycobacterium tuberculosis in a non-replicating state. Tuberculosis (Edinb) 84, 239–246 (2004).
Kesavan, A. K., Brooks, M., Tufariello, J., Chan, J. & Manabe, Y. C. Tuberculosis genes expressed during persistence and reactivation in the resistant rabbit model. Tuberculosis (Edinb) 89, 17–21 (2009).
Keren, I., Minami, S., Rubin, E. & Lewis, K. Characterization and Transcriptome Analysis of Mycobacterium tuberculosis Persisters. mBio 2, e00100-00111-e00100-00111 (2011).
Voskuil, M. I., Visconti, K. C. & Schoolnik, G. K. Mycobacterium tuberculosis gene expression during adaptation to stationary phase and low-oxygen dormancy. Tuberculosis (Edinb) 84, 218–227 (2004).
Soda, K., Misono, H. & Yamamoto, T. L-Lysine - Alpha-Ketoglutarate Aminotransferase .I. Identification of a Product Delta1-Piperideine-6-Carboxylic Acid. Biochemistry 7, 4102-& (1968).
Soda, K. & Misono, H. L-Lysin - Alpha-Ketoglutarate Aminotransferase .2. Purification Crystallization and Properties. Biochemistry 7, 4110-& (1968).
Fujii, T., Narita, T., Agematu, H., Agata, N. & Isshiki, K. Characterization of L-lysine 6-aminotransferase and its structural gene from Flavobacterium lutescens IFO3084. J Biochem-Tokyo 128, 391–397 (2000).
Betts, J. C., Lukey, P. T., Robb, L. C., McAdam, R. A. & Duncan, K. Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence. Mol Microbiol 43, 717–731 (2002).
Albrethsen, J. et al. Proteomic profiling of Mycobacterium tuberculosis identifies nutrient-starvation-responsive toxin-antitoxin systems. Mol Cell Proteomics 12, 1180–1191 (2013).
van Kessel, J. C. & Hatfull, G. F. Mycobacterial recombineering. Methods Mol Biol 435, 203–215 (2008).
Parish, T. & Stoker, N. G. Electroporation of mycobacteria. In: Mycobacteria protocols (ed^(eds). Springer (1998).
Novikova, A. E. S. N., Sycheva, E. V. & Kolokolova, A. V. Identification and analysis of norvaline and norleucine in fermentation broths of E. coli strains. Sorption Chromatogr Proc 6, 796–806 (2006).
Ribeiro, A. L. et al. Analogous mechanisms of resistance to benzothiazinones and dinitrobenzamides in Mycobacterium smegmatis. PLoS One 6, e26675 (2011).
Du, Q., Long, Q., Mao, J., Fu, T., Duan, X. & Xie, J. Characterization of a novel mutation in the overlap of tlyA and ppnK involved in capreomycin resistance in Mycobacterium. IUBMB Life 66, 405–414 (2014).
Betts, J. C., Lukey, P. T., Robb, L. C., McAdam, R. A. & Duncan, K. Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence by gene and protein expression profiling. Mol Microbiol 43, 717–731 (2002).
Jacobs, W. R., Tuckman, M. & Bloom, B. R. Introduction of foreign DNA into mycobacteria using a shuttle phasmid. Nature 327, 532–535 (1987).
Smeulders, M. J., Keer, J., Speight, R. A. & Williams, H. D. Adaptation of Mycobacterium smegmatis to stationary phase. J Bacteriol 181, 270–283 (1999).
Hansen, S., Lewis, K. & Vulic, M. Role of global regulators and nucleotide metabolism in antibiotic tolerance in Escherichia coli. Antimicrob Agents Chemother 52, 2718–2726 (2008).
Manuel, J., Zhanel, G. G. & de Kievit, T. Cadaverine suppresses persistence to carboxypenicillins in Pseudomonas aeruginosa PAO1. Antimicrob Agents Chemother 54, 5173–5179 (2010).
Lechner, S. et al. Metabolic and transcriptional activities of Staphylococcus aureus challenged with high-doses of daptomycin. Int J Med Microbiol 304, 931–940 (2014).
Cowley, S. et al. The Mycobacterium tuberculosis protein serine/threonine kinase PknG is linked to cellular glutamate/glutamine levels and is important for growth in vivo. Mol Microbiol 52, 1691–1702 (2004).
Boy, E., Richaud, C. & Patte, J. C. Multiple regulation of DAP‐decarboxylase synthesis in Escherichia coli K‐12. FEMS Microbiology Letters 5, 287–290 (1979).
Reddy, M. C., Gokulan, K., Jacobs, W. R.,, Jr., Ioerger, T. R. & Sacchettini, J. C. Crystal structure of Mycobacterium tuberculosis LrpA, a leucine-responsive global regulator associated with starvation response. Protein Sci 17, 159–170 (2008).
Wu, J., Long, Q. & Xie, J. (p)ppGpp and drug resistance. J Cell Physiol 224, 300–304 (2010).
Maisonneuve, E., Castro-Camargo, M. & Gerdes, K. (p)ppGpp controls bacterial persistence by stochastic induction of toxin-antitoxin activity. Cell 154, 1140–1150 (2013).
Landgraf, J. R., Wu, J. & Calvo, J. M. Effects of nutrition and growth rate on Lrp levels in Escherichia coli. J Bacteriol 178, 6930–6936 (1996).
Tani, T. H., Khodursky, A., Blumenthal, R. M., Brown, P. O. & Matthews, R. G. Adaptation to famine: A family of stationary-phase genes revealed by microarray analysis. Proceedings of the National Academy of Sciences of the United States of America 99, 13471–13476 (2002).
Avarbock, D., Salem, J., Li, L. S., Wang, Z. M. & Rubin, H. Cloning and characterization of a bifunctional RelA/SpoT homologue from Mycobacterium tuberculosis. Gene 233, 261–269 (1999).
Primm, T. P., Andersen, S. J., Mizrahi, V., Avarbock, D., Rubin, H. & Barry, C. E.,, 3rd . The stringent response of Mycobacterium tuberculosis is required for long-term survival. J Bacteriol 182, 4889–4898 (2000).
Jain, V., Saleem-Batcha, R., China, A. & Chatterji, D. Molecular dissection of the mycobacterial stringent response protein Rel. Protein Sci 15, 1449–1464 (2006).
Spoering, A. L. & Lewis, K. Biofilms and planktonic cells of Pseudomonas aeruginosa have similar resistance to killing by antimicrobials. J Bacteriol 183, 6746–6751 (2001).
Fauvart, M., De Groote, V. N. & Michiels, J. Role of persister cells in chronic infections: clinical relevance and perspectives on anti-persister therapies. J Med Microbiol 60, 699–709 (2011).
Tripathi, S. M. & Ramachandran, R. Overexpression, purification and crystallization of lysine epsilon-aminotransferase (Rv3290c) from Mycobacterium tuberculosis H37Rv. Acta Crystallogr Sect F Struct Biol Cryst Commun 62, 572–575 (2006).
Cole, S. T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Calvo, J. M. & Matthews, R. G. The leucine-responsive regulatory protein, a global regulator of metabolism in Escherichia coli. Microbiol Rev 58, 466–490 (1994).
Brinkman, A. B., Ettema, T. J., de Vos, W. M. & van der Oost, J. The Lrp family of transcriptional regulators. Mol Microbiol 48, 287–294 (2003).
Traxler, M. F. et al. Discretely calibrated regulatory loops controlled by ppGpp partition gene induction across the ‘feast to famine’ gradient in Escherichia coli. Mol Microbiol 79, 830–845 (2011).
Parti, R. P. S. et al. A transposon insertion mutant of Mycobacterium fortuitum attenuated in virulence and persistence in a murine infection model that is complemented by Rv3291c of Mycobacterium tuberculosis. Microb Pathogenesis 45, 370–376 (2008).
Mani Tripathi, S. & Ramachandran, R. Direct evidence for a glutamate switch necessary for substrate recognition: crystal structures of lysine epsilon-aminotransferase (Rv3290c) from Mycobacterium tuberculosis H37Rv. J Mol Biol 362, 877–886 (2006).
Dube, D., Tripathi, S. M. & Ramachandran, R. Identification of in vitro inhibitors of Mycobacterium tuberculosis Lysine ε-aminotransferase by pharmacophore mapping and three-dimensional flexible searches. Medicinal Chemistry Research 17, 182–188 (2008).
Acknowledgements
This work was supported by National Natural Science Foundation (grant numbers 81371851, 81071316, 81271882, 81301394, 81511120001); New Century Excellent Talents in Universities (grant number NCET-11-0703); National Major projects for Key Infectious Diseases (grant number 2008ZX10003-006); Excellent PhD thesis fellowship of Southwest University (grant numbers kb2010017, ky2011003);The Fundamental Research Funds for the Central Universities (Nos XDJK2011D006, XDJK2012D011, XDJK2012D007, XDJK2013D003 andXDJK2014D040); Natural Science Foundation Project of CQ CSTC (grant number CSTC 2010BB5002); The Chongqing municipal committee of Education for postgraduates excellence program (No.YJG123104); The undergraduates teaching reform program (No. 2011JY052, 2013JY201); Graduate research and innovation project of graduate in Chongqing (XDJK2014D040).
Author information
Authors and Affiliations
Contributions
X.D., Y.L., Q.D., Q.H., M.X., Y.L. and S.G. performed the experiments. X.D. and J.X. analyzed the data. X.D., Z.L. and J.X. designed the study and wrote the paper. All authors have read and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Duan, X., Li, Y., Du, Q. et al. Mycobacterium Lysine ε-aminotransferase is a novel alarmone metabolism related persister gene via dysregulating the intracellular amino acid level. Sci Rep 6, 19695 (2016). https://doi.org/10.1038/srep19695
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep19695
This article is cited by
-
The pathogenic mechanism of Mycobacterium tuberculosis: implication for new drug development
Molecular Biomedicine (2022)
-
Transient drug-tolerance and permanent drug-resistance rely on the trehalose-catalytic shift in Mycobacterium tuberculosis
Nature Communications (2019)
-
Derailing the aspartate pathway of Mycobacterium tuberculosis to eradicate persistent infection
Nature Communications (2019)
-
Enantiomer discrimination in β-phenylalanine degradation by a newly isolated Paraburkholderia strain BS115 and type strain PsJN
AMB Express (2018)
-
Mycobacterium tuberculosis toxin Rv2872 is an RNase involved in vancomycin stress response and biofilm development
Applied Microbiology and Biotechnology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.