Abstract
Bony fishes are the most numerous and phenotypically diverse group of vertebrates inhabiting our planet, making them an ideal target for identifying general principles of tissue development and function. However, lack of suitable experimental platforms prevents the exploitation of this rich source of natural phenotypic variation. Here, we use a zebrafish strain lacking definitive hematopoiesis for interspecific analysis of hematopoietic cell development. Without conditioning prior to transplantation, hematopoietic progenitor cells from goldfish stably engraft in adult zebrafish homozygous for the c-mybI181N mutation. However, in competitive repopulation experiments, zebrafish hematopoietic cells exhibit an advantage over their goldfish counterparts, possibly owing to subtle species-specific functional differences in hematopoietic microenvironments resulting from over 100 million years of independent evolution. Thus, our unique animal model provides an unprecedented opportunity to genetically and functionally disentangle universal and species-specific contributions of the microenvironment to hematopoietic progenitor cell maintenance and development.
Similar content being viewed by others
Introduction
The hematopoietic system has deep roots in animal evolution (for recent review, see1). This observation has stimulated the study of other vertebrate species apart from the traditional chicken and mouse models, most notably the zebrafish, to examine general and species-specific aspects of hematopoietic development and differentiation2,3,4,5. The zebrafish provides us with an ideal model system and is notable both for its ease of handling and the availability of a highly diversified genetic tool-box6. The zebrafish is a teleost and belongs to the most species-rich group of vertebrates inhabiting our planet; indeed, teleosts make up almost half of all vertebrates and are notable for their vast diversity in form and their ability to occupy the most extreme ecological niches7,8. Although it is conceivable that the analysis of diverse fish hematopoietic systems will shed light on the general principles and requirements of hematopoietic differentiation, with potentially great translational impact, no suitable experimental system has yet been developed to achieve this goal.
We recently described a zebrafish model for allogeneic hematopoietic cell transplantation9. It utilizes adult zebrafish homozygous for a missense mutation in the c-myb gene (c-mybI181N); this mutation causes failure of definitive hematopoiesis, thus obviating the requirement for pre-conditioning prior to transplantation of allogeneic hematopoietic cells10. Here, we have explored the possibility of using this model for xenogeneic transplantation as a first step towards examining the phenotypic diversity of fish hematopoietic systems. To this end, we chose goldfish as a species for our proof-of-principle studies. The goldfish is a member of the cyprinidae, the largest family of vertebrates, comprising more than 2,000 fresh-water species. Like carp, the goldfish Carassius auratus belongs to the subfamily of Cyprininae, whereas the zebrafish Danio rerio belongs to the subfamily of Danioninae. Recent phylogenetic analyses suggest that the last common ancestor of Cyprininae and Danioninae lived more than 100 million years ago11. This long period of independent evolution has led to widely diverging ecological preferences, the most notable of which is water temperature; whereas zebrafish adapted to tropical conditions with an optimal water temperature of 28.5 °C, goldfish tolerates water temperatures down to near freezing point. We therefore considered goldfish as an appropriate candidate to examine the feasibility of xenogeneic transplantation of hematopoietic cells.
There is considerable interest in examining the properties of vertebrate hematopoietic stem cells (HSCs). In particular, knowledge about the requirements for self-renewal and maintenance of HSCs could be harnessed for several therapeutic settings. The zebrafish has been successfully used for in vivo chemical screens aimed at identifying the molecular pathways regulating the number and maintenance of HSCs (for a recent example, see 12). However, while the zebrafish is clearly suitable for carrying out large-scale screens, it is not clear whether potential biological differences between the hematopoietic cells of fish and mammals may hinder the direct translational application of such results. Hence, it is important to explore the possibility of carrying out transplantations of xenogeneic HSCs into a zebrafish background with a view to generating an in vivo complement of existing in vitro models. As a first step in this direction, we explored the possibility of establishing a xenogeneic hematopoietic system in zebrafish. Our results indicate that the hematopoietic environment of the zebrafish supports the maintenance of xenogenic hematopoietic progenitor cells, opening up unprecedented opportunities for comparative studies on vertebrate hematopoiesis.
Results
Transplantation of goldfish whole kidney marrow into c-myb mutant zebrafish
In a first set of experiments, we compared the extent of hematopoietic reconstitution after transplantation of zebrafish or goldfish cells over a period of several weeks (Fig. 1A). Following transplantation of approximately 4 × 105 zebrafish kidney marrow cells per 6-week-old adult c-myb mutant recipient10, hematopoietic reconstitution was readily observable after two weeks, as described previously10 (see also Supplementary Fig. 4A). After transfer of approximately the same number of goldfish cells (3.3 ± 0.9 × 105; n = 7), evidence of reconstitution became apparent at a later time point (Fig. 1B); nonetheless, in week 4 after transplantation, c-myb mutant recipients exhibited a reddish complexion (Fig. 1C), had lost their heart edema as a sign of reversal of anemia, and, as a consequence of improved tissue oxygenation, could swim much faster than their unmanipulated mutant siblings. When the cellular composition of the head kidney marrow was analyzed, the first distinct signs of repopulation were also evident at 4 weeks and reconstitution was essentially complete at 6 weeks after transplantation (Fig. 1B). Of note, a shift in the relative distributions of different cell types - identifiable by their distinct forward and side light scatter characteristics - was evident; the most obvious change was the paucity of cells in the myelomonocytic gate (marked in green in Fig. 1B) relative to the original cell composition (Fig. 1B). Pulse-labelling with the nucleoside analogue EdU five weeks after transplantation (Fig. 1D) indicated that vigorous proliferation of hematopoietic cells occurred in the c-myb mutant recipients after transplantation of both zebrafish and goldfish cells, but not in unmanipulated c-myb mutant animals (Fig. 1E). Proliferation occurred primarily in progenitor and lymphocyte-like cells.
Reconstitution of hematopoiesis in adult c-mybI181N zebrafish mutants after transplantation of goldfish kidney marrow cells.
(A) Schematic outline. A total of 23 transplantations were carried out and analyzed at various time points; representative data are shown for each time point. (B) Flow cytometric analysis of whole kidney marrow cell populations of adult fish of various genotypes and species (top panels) and after transplantation of goldfish marrow cells into mutant zebrafish (lower panels); the time points indicate time periods after transplantation. Wild type zebrafish (zf wt), c-myb mutant (c-myb−/−), goldfish (gf) and c-myb mutant transplanted with goldfish whole kidney marrow cells (gf > c-myb−/−). The different cell populations in zebrafish kidney are identified by their forward (size) and side (granularity) light scatter characteristics: red, erythrocytes; blue, lymphocytes; green, myelomonocytic cells; magenta, progenitor cells. (C) Macroscopic representation of kidney of wild type zebrafish (left), zebrafish mutants (middle) and zebrafish mutants reconstituted with goldfish hematopoietic cells (4 weeks after transplantation; right), all 10 weeks old. The locations of the head kidneys are indicated by ovals. (D) Schematic outline of proliferation assay. (E) Flow cytometric analysis of whole kidney marrow cells isolated from fish of the indicated genotypes pulse-labelled with EdU, creating a red-fluorescent signal in cells. No signal is observed in control mock-pulsed cells (two leftmost columns). Proliferating cells are detectable only in wild type zebrafish kidney marrow and in mutant zebrafish reconstituted with either zebrafish or goldfish cells, but not in unmanipulated zebrafish mutants (third column from the left). The light scatter characteristics of EdU-positive cells (red) are superimposed on EdU-negative cells (blue) (second and fourth columns from the left). Representative data of corresponding biological replicates are shown (zebrafish wild type, n = 7; c-myb−/−, n = 5; zf > c-myb−/−, n = 3; gf > c-myb−/−, n = 3).
No reactivation of endogenous hematopoiesis after transplantation
Next, we investigated whether the presence of wild type hematopoietic cells might stimulate c-myb mutant progenitor cells to undergo normal hematopoietic maturation, contributing to reconstitution after transplantation of goldfish hematopoietic cells. To exclude this possibility, we generated c-myb mutant fish additionally transgenic for gata1:DsRed13 and ikaros:eGFP14 reporters; the former transgene marks red blood cells, whereas the latter directs GFP expression in hematopoietic precursors and lymphoid cells. In gata1:DsRed; ikaros:eGFP double-transgenic c-myb wild type fish, numerous green fluorescent cells are clearly visible at 1 day after fertilization (dpf), whereas fewer fluorescent cells could be detected in c-myb mutants; this is compatible with the observation of failing definitive hematopoiesis in c-myb mutants10 (Supplementary Fig. 1A). At the same time point, red-fluorescent cells were detectable in both wild type and mutant fish, compatible with the presence of unimpaired primitive hematopoiesis10 (Supplementary Fig. 1A). In older fish, expression of gata1:DsRed and ikaros:eGFP reporters was confined to c-myb wild type fish, but absent in c-myb mutants; importantly, cellular fluorescence was not observed after transplantation of wild type zebrafish kidney marrow cells (Supplementary Fig. 1B and 1C). After the hematopoietic system had been fully reconstituted (Supplementary Fig. 1D), we used flow cytometry to examine the whole kidney marrow for the presence of red or green (i.e. mutant host) cells; however, none could be detected (Supplementary Fig. 1E–G). In adult gata1:DsRed and ikaros:eGFP wild type fish, gata1 expression is particularly strong in the progenitor population, whereas ikaros expression predominates in cells with the light scatter characteristics of lymphocytes (Supplementary Fig. 1E). As expected, these cell populations are absent in adult c-myb mutants (Supplementary Fig. 1F). Similarly, no such cells could be found in c-myb mutants fully reconstituted with non-transgenic wild type kidney marrow cells (Supplementary Fig. 1G). These observations indicate that the c-myb mutation results in cell-intrinsically defective hematopoietic stem cell (HSC) maintenance and/or differentiation, preventing their contribution to hematopoiesis even after transplantation with wild type donor tissue. From an experimental point of view, this observation indicates that reconstituted hematopoietic and immune systems will consist entirely of descendants of transplanted HSCs.
Multilineage hematopoietic reconstitution by goldfish cells
In order to determine the relative contribution of goldfish cells in cell suspensions prepared from whole kidney marrow, we developed a simple PCR-based DNA typing assay; a small fragment of the highly conserved activin beta A genes of zebrafish and goldfish could be amplified using the same primer pair and readily distinguished by presence (goldfish) or absence (zebrafish) of an AvaII restriction site (Fig. 2A; Supplementary Fig. 2A). The results of this assay suggested the presence of goldfish cells in the repopulated zebrafish kidney marrow (Fig. 2B). We carried out detailed histological analyses to provide further evidence for the presence of ongoing hematopoiesis in c-myb mutants after transplantation of goldfish cells (Fig. 2C; Supplementary Fig. 3). Notably, our analyses demonstrated colonization of the thymus with strong rag1 expression indicative of ongoing T cell development; similarly, B cell poiesis (as revealed by rag1 expression in the kidney marrow) is readily detectable after transplantation (Fig. 2C). Finally, the pleomorphic appearance of peripheral blood cells (including red blood cells, lymphocytes and myeloid cells) was indicative of multilineage reconstitution (Fig. 2C; Supplementary Fig. 3). Cytological analysis indicated the presence of cells with the characteristically larger size of goldfish cells (Supplementary Fig. 2B); this size difference is also evident in the flow cytometric profiles (see Fig. 1B). Collectively, these results strongly suggest that transplantation of goldfish kidney marrow cells results in multilineage hematopoietic reconstitution of adult c-myb mutant zebrafish.
Multilineage hematopoietic reconstitution of c-myb mutant zebrafish after transplantation of goldfish whole kidney marrow cells.
(A) Genotyping of whole kidney marrow cells of fish of indicated genotypes. The schematic at the top highlights the AvaII restriction site distinguishing zebrafish and goldfish sequences of the activin beta A gene (c.f. Supplementary Fig. 2A). Amplicons are shown before and after AvaII digestion of wild type zebrafish (zf wt), c-myb mutant zebrafish (zf−/−) and goldfish (gf). The AvaII digest of an amplicon generated from a 1+1 mixture of wild type zebrafish and goldfish cells is also shown on the right for reference. (B) Genotyping of whole kidney marrow cells of c-myb mutant zebrafish reconstituted with goldfish whole kidney marrow cells (gf > c-myb−/−) at several time points (in weeks) after transplantation. (C) Histological analysis of reconstituted c-myb mutant animals. Sections of the thymus (top panels), kidney (middle panels) and heart (bottom panels) are shown; insets represent 4× higher magnifications (haematoxylin/eosin staining) with the origin of selected regions indicated by broken squares (see also Supplementary Fig. 3). The fourth column indicates the results of RNA in situ hybridization using a rag1 probe, detecting lymphoid progenitor cells actively rearranging antigen receptor loci. Wild type zebrafish (zf wt), c-myb mutant (c-myb−/−) and c-myb mutant transplanted with goldfish whole kidney marrow cells 6 weeks after transplantation (gf > c-myb−/−). All fish are 12 weeks old. Anatomical landmarks are indicated (G, gills; T, thymus; K, kidney; H, heart; A, atrium of the heart; V, ventricle of the heart). Scale bars, 50 μm.
Serial transplantation of xenogeneic hematopoietic progenitor cells
Next, we assessed whether the hematopoietic system in c-myb mutants originating from goldfish cells would also support reconstitution after a second transplantation. To this end, we transferred all kidney marrow cells from one primary recipient to a secondary host (Fig. 3A). We established that recovery of hematopoiesis in secondary recipients took place with similar kinetics to those during primary reconstitutions; notably, the proportions of different cell lineages in the kidney are indistinguishable from those of the recipients of primary transplants (Fig. 3B–D). Moreover, the repopulated thymi displayed ongoing T cell development, as indicated by rag1 expression (Fig. 3E,F). Collectively, multilineage reconstitution after secondary transplantation suggests that progenitor cells survive in the zebrafish kidney marrow environment.
Serial transplantation of goldfish hematopoietic progenitors.
(A) Schematic of successive transplantation experiment. A total of 8 transplantations were carried out and analyzed at various time points; representative data are shown for each time point. The first transplantation (gf > c-myb−/−) was carried out when c-myb mutants had reached the age of 6 weeks; the secondary transplantation (gf > gf > c-myb−/−) was carried out 4 weeks later. (B) Flow cytometric analysis of whole kidney marrow cells. The profile of the cellular source material for secondary transplant is indicated on the left (gf > gf > c-myb−/− [start]); the other panels indicate the profiles for cells at various time points after the second transplantation. The different cell types are tentatively assigned (c.f. legend to Fig. 1). (C) Genotyping assay indicating the presence of goldfish cells in the whole kidney marrow derived from secondary transplantation. (D) Macroscopic view of kidney after secondary transplantation. The location of the head kidney is indicated by oval. (E) Colonization of the thymus (T) after secondary transplantation (haematoxylin/eosin staining); G, gills. (F) T cell development in the thymus (T) after secondary transplantation (RNA in situ hybridization with rag1 probe). Scale bars, 50 μm.
Interspecific competition of hematopoietic cells
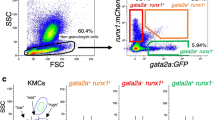
The above experiments clearly indicate that under non-competitive conditions, goldfish hematopoiesis can be maintained in the zebrafish kidney marrow environment. However, the perturbed ratios of different cell types (i.e., lymphocyte vs. myelomonocytic cells) indicate possible species-specific differences in microenvironmental cues influencing the quantitative outcome of transplantation. If so, then such differences should become more pronounced in situations of competitive repopulation. To illustrate this potential use of the c-myb transplantation model, we carried out reconstitution experiments using transplants consisting of different ratios of zebrafish and goldfish cells. To monitor the kinetics of the reconstitution process, we employed zebrafish cells carrying the ikaros:eGFP transgenic marker, which is expressed in lymphoid and myelomonocytic cells. When we used a 1+1 mixture of zebrafish and goldfish whole kidney marrow cells, we established that hematopoietic reconstitution was eventually dominated by descendants of zebrafish HSCs, as indicated by the diagnostic proportion of GFP-positive lymphocytes (Supplementary Fig. 4). However, it is evident from a comparison of pure zebrafish transplants (Supplementary Fig. 4A) with the 1+1 mixtures (Supplementary Fig. 4B) that zebrafish hematopoiesis becomes established at a later time point and that goldfish myelomonocytic cells are at a disadvantage as compared to their zebrafish counterparts. Whereas zebrafish myelomonocytic cells had essentially outcompeted their goldfish counterparts 4 weeks after transplantation, lymphopoiesis was dominated by zebrafish cells only after 6 weeks (Supplementary Fig. 4C). When a 1+9 mixture of zebrafish and goldfish kidney marrow cells was used, reconstitution by zebrafish cells was even more delayed (Fig. 4A,B). However, goldfish cells proliferate alongside zebrafish cells in the initial phases of reconstitution, as indicated by the composition of the respective genomic DNA of kidney marrow cells (Fig. 4C). This observation argues against an immune-mediated depletion of either cell type and rather supports the notion that direct competition for niche space and/or growth and inductive factor activities contribute to the distorted outcome of chimeric reconstitution.
Competitive repopulation assay with excess goldfish cells.
(A) Flow cytometric profiles after transplantation of a 1+9 mixture of zebrafish and goldfish cells, with different cell populations indicated (c.f. legend to Fig. 1). The starting material derived from c-myb wild type zebrafish transgenic for ikaros:eGFP reporter (1 part) and goldfish cells (9 parts) is shown in the top left panel; very few cells in the lymphocyte gate (blue) are GFP-positive (lower left panel). Analyses of whole kidney marrow cells at several time points after transplantation are also shown, indicating that complete reconstitution has not yet occurred after 6 weeks. Representative data for a total of five (1+9) transplantations are shown. (B) Time-resolved reconstitution of lymphocyte compartment, as measured by the percentage of GFP-positive cells over time after transplantation (in weeks). Summary of transplantation experiments for zebrafish whole kidney marrow cells only (1+0) and 1+1 or 1+9 mixtures of zebrafish and goldfish cells; each symbol represents the data of a single transplanted fish. (C) Genotyping assay indicating the presence of goldfish cells in whole kidney marrow 4 weeks after competitive transplantation of a 1+9 (zebrafish + goldfish) mixture.
Discussion
Maintenance of HSCs and their differentiation into the various blood cell lineages is regulated by a plethora of extrinsic cues such as growth factors and cytokines, some of which are known to be particularly species-specific in their action. In this context, it is surprising to find that hematopoietic reconstitution from transferred goldfish cells occurs in the zebrafish microenvironment. However, the relative paucity of myeloid cells in the xenotransplants relative to their frequency in the goldfish kidney marrow is one indication that the conditions for differentiation and/or survival of goldfish myeloid cells provided by the zebrafish microenvironment are not optimal. Nonetheless, our results demonstrate a surprising degree of functional conservation of hematopoietic supportive factors in species that have evolved independently for more than 100 Mya and that inhabit very different ecological niches. More specifically, our results predict that the homeostatic chemokines and cytokines supporting niche occupancy, colonization of primary hematopoietic (kidney) and lymphoid (kidney, thymus) organs and also differentiation of the major cell lineages are functionally compatible with the respective goldfish receptors. Our findings therefore provide important information concerning the molecular evolution of intercellular communication systems once the appropriate genomic and transcriptomic resources have been developed for the goldfish. Collectively, our observations suggest that the c-mybI181N transplantation model described here could be developed for the study of hematopoietic systems of other fish species, facilitating comparative analyses of their cellular and functional characteristics. Our success bodes well for future attempts at transplanting into c-mybI181N zebrafish hematopoietic cells from more distantly related organisms, such as jawless fishes and mammals, including humans.
So far, transplantation of non-tumorigenic xenogeneic cells into zebrafish was mostly performed using embryonic hosts to avoid the complications arising from immunological incompatibility (for instance, see15). Alternatively, hosts with compromised immune functions were used (for instance, see16,17). By contrast, the c-mybI181N transplantation model does not require conditioning prior to transplantation. It is possible that the unmanipulated microenvironment contributes to the robust and stable engraftment of allogeneic9 and xenogeneic (this paper) hematopoietic cells. One interesting potential application of our model is the definition of the relative contribution of individual microenvironmental factors underlying the species-specific differences observed here. This could be done for instance by replacing zebrafish genes, individually or in combination, with orthologous sequences from other fish species or even more distantly related vertebrates; a second, but not mutually exclusive, alternative is the use of pharmacological or genetic screens to examine possible epistatic effects of the many different hematopoietic factors. Moreover, it should be possible, without having to consider interference by hematopoietic cells of the host, to examine the relative fitness of co-transplanted hematopoietic cell types, each differing in particular functionalities. Similarly to what was previously demonstrated for immunocompromised zebrafish hosts, our model could also be used for the transplantation of allogeneic and xenogeneic16,18,19 tumour cells to facilitate studies on their general biological properties and drug sensitivity profiles.
Methods
Animals
Zebrafish (D. rerio) wild type strains (TLEK and TL), the transgenic ikaros:eGFP14, the transgenic gata1:DsRed13 (a generous gift of Dr. T. Schwerte, University of Innsbruck, Austria) and the mutant c-mybt25127 10 lines are bred at the Max Planck Institute of Immunobiology and Epigenetics. All procedures were conducted in accordance with institutional guidelines under a license (Az 35-9185.81/G-12/85) and approved by the local government (Regierungspräsidium Freiburg, Germany). In the past, homozygous c-myb mutants were housed in groups of 20 fish in separate tanks; recently, we have found that they can be maintained more easily by co-housing them with sentinel wild type fish, such as slc45a2 albino fish20. In some experiments, transparent21 c-myb mutants were used (a generous gift of Dr. L. Zon, Harvard University).
Transplantation procedure
Whole kidney marrow (WKM) cells from adult fish were harvested as described9 and finally resuspended at a concentration of 20,000 to 40,000 cells/μl in 0.9 × PBS supplemented with 5%FCS; cells were injected retro-orbitally in a volume of 10 μl per animal. After transplantation, zebrafish were kept at a water temperature of 22 °C (room temperature) instead of the usual 28.5 °C.
Flow cytometric analysis
Flow cytometric analysis was carried out as described; the identities of cell populations were assigned according to their light scatter characteristics22. Profiles were recorded with an Aria II instrument, with the exception of the data shown in Supplementary Fig. 1; in this case an AriaFusion instrument was used.
Fluorescence microscopy
Fluorescence microscopy was carried out as described9,14.
Proliferation assay
To detect proliferating cells, 7 μg of 5-ethynyl-2′-deoxyuridine (EdU) in 10μl PBS solution (Click-iT® EdU Flow Cytometry Assay Kits, Invitrogen Molecular Probes®) was injected retro-orbitally into wild type zebrafish, c-myb mutants and c-myb mutants carrying zebrafish and goldfish transplants, 4 weeks after primary transplantation. The analysis was performed 14–16 hours after EdU treatment. Staining was based on a click reaction using the Alexa Fluor 647® dye and performed according to the manufacturer’s instructions.
RNA in situ hybridization
This procedure was carried out as described10 using a zebrafish rag1 probe10, which has a 87% nucleotide identity with goldfish rag1 (c.f. Genbank accession numbers U71093.1 (zebrafish) und EF186007.3 (goldfish)).
Genotyping
Genotyping for the activin beta A genes was carried out using primers 5′-GGATTATTTTGGACTGGCTCC and 5′-TTCAGATGCAGCATGTTGAGG; these bind to sequences that are identical for both zebrafish and goldfish orthologs and amplify fragments of almost identical size (zebrafish, nucleotides 248–598 in Genbank accession number BC129208; goldfish, nucleotides 134–479 in Genbank accession number AF169032) (Supplementary Fig. 2A). Amplicons were digested with AvaII (New England Biolabs) according to the manufacturer’s instructions and the products resolved on agarose gels. Genotyping for the presence of the gata1:DsRed transgene was carried out with primers 5′-CAAGGAGTTCATGCGCTTCA and 5′-TTCACGCCGATGAACTTCAC (amplicon size 360 bp); genotyping for the presence of the ikaros:eGFP transgene was carried out with primers 5′-TGGTGCCCATCCTGGTCGAG and 5′-GTCCTCGATGTTGTGGCGGA (amplicon size 488 bp).
Additional Information
How to cite this article: Hess, I. and Boehm, T. Stable multilineage xenogeneic replacement of definitive hematopoiesis in adult zebrafish. Sci. Rep. 6, 19634; doi: 10.1038/srep19634 (2016).
References
Hartenstein, V. Blood cells and blood cell development in the animal kingdom. Annu. Rev. Cell Dev. Biol. 22, 677–712 (2006).
Stachura, D. L. & Traver, D. Cellular dissection of zebrafish hematopoiesis. Methods Cell Biol. 101, 75–110 (2011).
Iwanami, N. Zebrafish as a model for understanding the evolution of the vertebrate immune system and human primary immunodeficiency. Exp. Hematol. 42, 697–706 (2014).
Boatman, S. et al. Assaying hematopoiesis using zebrafish. Blood Cells Mol. Dis. 51, 271–276 (2013).
Ciau-Uitz, A., Monteiro, R., Kirmizitas, A. & Patient, R. Developmental hematopoiesis: Ontogeny, genetic programming and conservation. Exp. Hematol. 42, 669–683 (2014).
Detrich, H. W. III. In Methods Cell Biol Vol. 134 (Academic Press, Waltham, 2011).
Morrissey, J. & Sumich, J. L. Introduction to the Biology of Marine Life (Jones & Bartlett Learning, 2012).
Wootton, R. J. & Smith, C. Reproductive Biology of Teleost Fishes (Wiley-Blackwell, 2015).
Hess, I., Iwanami, N., Schorpp, M. & Boehm, T. Zebrafish model for allogeneic hematopoietic cell transplantation not requiring preconditioning. Proc. Natl. Acad. Sci. USA 110, 4327–4332 (2013).
Soza-Ried, C., Hess, I., Netuschil, N., Schorpp, M. & Boehm, T. Essential role of c-myb in definitive hematopoiesis is evolutionarily conserved. Proc. Natl. Acad. Sci. USA 107, 17304–17308 (2010).
Xu, P. et al. Genome sequence and genetic diversity of the common carp, Cyprinus carpio. Nat. Genet. 46, 1212–1219 (2014).
Li, P. et al. Epoxyeicosatrienoic acids enhance embryonic haematopoiesis and adult marrow engraftment. Nature 523, 468–471 (2015).
Yaqoob, N., Holotta, M., Prem, C., Kopp, R. & Schwerte, T. Ontogenetic development of erythropoiesis can be studied non-invasively in GATA-1:DsRed transgenic zebrafish. Comp. Biochem. Physiol. A Mol. Integr. Physiol . 154, 270–278 (2009).
Hess, I. & Boehm, T. Intravital imaging of thymopoiesis reveals dynamic lympho-epithelial interactions. Immunity 36, 298–309 (2012).
Orlova, V. V., Drabsch, Y., ten Dijke, P. & Mummery, C. L. Assessment of functional competence of endothelial cells from human pluripotent stem cells in zebrafish embryos. Methods Mol. Biol. 1213, 107–119 (2014).
Tang, Q. et al. Optimized cell transplantation using adult rag2 mutant zebrafish. Nat. Methods 11, 821–824 (2014).
Shayegi, N. et al. CXCR4 blockade and Sphingosine-1-phosphate activation facilitate engraftment of haematopoietic stem and progenitor cells in a non-myeloablative transplant model. Br. J. Haematol. 164, 409–413 (2014).
Zhang, B. et al. Quantitative phenotyping-based in vivo chemical screening in a zebrafish model of leukemia stem cell xenotransplantation. PloS One 9, e85439 (2014).
Eden, C. J. et al. Orthotopic models of pediatric brain tumors in zebrafish. Oncogene 34, 1736–1742 (2015).
Streisinger, G., Singer, F., Walker, C., Knauber, D. & Dower, N. Segregation analyses and gene-centromere distances in zebrafish. Genetics 112, 311–319 (1986).
White, R. M. et al. Transparent adult zebrafish as a tool for in vivo transplantation analysis. Cell Stem Cell 2, 183–189 (2008).
Traver, D. et al. Transplantation and in vivo imaging of multilineage engraftment in zebrafish bloodless mutants. Nat. Immunol. 4, 1238–1246 (2003).
Acknowledgements
This project has received funding from the Max Planck Society and the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007–2013) ERC grant agreement no. 323126. We thank Michael Schorpp for advice, Thorsten Schwerte for the gata1:DsRed transgenic line, Leonard Zon for the transparent zebrafish line and Dagmar Diekhoff and Monika Held for technical help.
Author information
Authors and Affiliations
Contributions
I.H. designed and carried out experiments, interpreted results and wrote the manuscript; T.B. conceived the study, designed experiments, interpreted results and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hess, I., Boehm, T. Stable multilineage xenogeneic replacement of definitive hematopoiesis in adult zebrafish. Sci Rep 6, 19634 (2016). https://doi.org/10.1038/srep19634
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep19634
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.