Abstract
Functional interplays of microbial activity, genetic diversity and contaminant transformation are poorly understood in reactors for mineralizing halogenated aromatics anaerobically. Here, we investigated abundance and distribution of potential microbes and functional genes associated with pentachlorophenol (PCP) anaerobic mineralization in a continuous-flow cylindrical reactor (15 cm in length). PCP dechlorination and the metabolite (phenol) were observed at segments 0–8 cm from inlet, where key microbes, including potential reductive dechlorinators (Dehalobacter, Sulfurospirillum, Desulfitobacterium and Desulfovibrio spp.) and phenol degraders (Cryptanaerobacter and Syntrophus spp.), as well as putative functional genes, including putative chlorophenol reductive dehalogenase (cprA) and benzoyl-CoA reductase (bamB), were highly enriched simultaneously. Five types of putative cprAs, three types of putative bamBs and seven types of putative nitrogenase reductase (nifHs) were determined, with their copy numbers decreased gradually from inlet to outlet. Distribution of chemicals, bacteria and putative genes confirmed PCP dechlorination and phenol degradation accomplished in segments 0–5 cm and 0–8 cm, respectively, contributing to a high PCP mineralization rate of 3.86 μM d−1. Through long-term incubation, dechlorination, phenol degradation and nitrogen fixation bacteria coexisted and functioned simultaneously near inlet (0–8 cm), verified the feasibility of anaerobic mineralization of halogenated aromatics in the compact reactor containing multiple functional microbes.
Similar content being viewed by others
Introduction
Halogenated aromatic compounds (HACs) are noteworthy for their characteristics of high toxicity, persistence and bioaccumulation. They have led to the serious contamination upon the extensive applications1. Twenty-three types of halogenated benzenes or phenols have been recorded as the priority pollutants by the US Environmental Protection Agency, which occupied 17% of the total list2. Seeking for the efficient remediation strategies became an urgent and tough assignment to completely remove the toxiticity of HACs in worldwide.
The use of mciroorganisms in detoxification of HACs has addressed the increasing attention as a promising and cost-effective bioremedaition approach3. Several aerobic isolates, such as Pseudomonas or Burkholderia spp., could mineralize HACs through oxidative or hydrolytic dechlorination and oxygenolysis; meanwhile, the related enzymatic genes were also well elucidated3,4. The biodegradation of HACs, has been extensively investigated in both bioreactors and batch cultures5,6. With pentachlorophenol (PCP) as an example, under aerobic condition, PCP could be oxidized to chlorohydroquinone accompanied with the removal of chloride and aromatic ring cleavage and then completely mineralized to CO27. Under anaerobic conditions, reductive dehalogenation by replacing the halogen with hydrogen atom played a vital role for converting HACs to the less/none halogenated ones8,9. Several isolates have been reported that possessed the reductive dehalogenation activities, like Desulfitobacterium spp., Dehalobacter spp., Desulfovibrio spp., Dehalococcoides spp. and Dehalobacter spp.10. However, after reductive dehalogenation, the less or none halogenated metabolites still carry aromatic rings and toxicity. Anaerobic oxidative degradation of these metabolites to CO2 is another obligatory bioprocess to completely remove the toxicity of HACs anaerobically11.
Hence, artificial combination of two bioprocesses including reductive dehalogenation and anaerobic oxidative degradation is an essential means to realize the complete bio-mineralization of HACs anaerobically. Previously, some anaerobic mineralization systems have been constructed by synthesizing consortia for reductive dehalogenation and oxidative degradation, which acquired the favorable and efficient mineralization activities11,12,13,14. However, the two bioprocesses, reductive dehalogenation and oxidative degradation, went through the distinct metabolic pathways and nutrient requirements and their working mechanism during HACs transformation were little known. So identification of the interplays of responsible bacteria, functional genes and organic chemicals, are vital to better understand the competitive or cooperative interrelations of functional bacteria. The study would give suggestions for the adaptability and regulation strategy for complete mineralization of HACs by alignment of bacteria and nutrients with different functions in contaminated sites or bioreactors.
In this study, a long-term maintained cylindrical reactor for pentachlorophenol (PCP) mineralization utilizing lactate as the sole external nutrient was disassembled. The spatial ecophysiological characterizations of microbes and genes at different cylindrical segments were carried out. The abundance and distribution of organic compounds (PCP, lactate and their corresponding organic metabolites), involved bacteria (reductive dechlorinators and oxidative degraders) and functional genes (chlorophenol reductive dehalogenase cprA, phenol anaerobic degradation benzoyl-CoA reductase/ring-cleavage hydrolase genes bamB, bzdN, bcrC and bamA and nitrogenase gene nifH) inside the cylindrical reactor, were extensively investigated. The objects of this study were to (i) reveal the distribution and diversity of the functional bacteria and genes associated with the mineralization of HACs in the constructed reactor and (ii) give understanding on the occurrence and functional mechanisms under the roles of potential bacteria and genes during HACs-mineralization process.
Results
Spatial distributions of PCP, lactate and their organic metabolites
Concentration distributions of PCP, lactate and their organic metabolites were shown in Fig. 1. PCP concentration decreased quickly and became below the detection limit in segment 5–8 cm from the inlet and while, phenol was detected at a trace concentration less than 5 μM between 0–8 cm (Fig. 1A), indicating the occurrence of reductive dechlorination between 0–5 cm. Besides phenol, none of the other possible metabolites, like 3-chlorophenol (3-CP), 3,5-dichlorophenol or 2,3,5-trichlorophenol, was detectable in the whole reactor segments (Fig. 1A; data not shown). Above segment 8 cm, no phenol was detected, indicating phenol complete degradation accomplished between 0–8 cm. Concentration of lactate decreased quickly between 0–8 cm, where its metabolites, acetate and propionate, were frequently detetced in the range of 0–3 μM and 0–1.8 μM, respectively. Acetate and propionate disappeared in the segment 8–11 cm and 13–15 cm, respectively (Fig. 1B).
Spatial distributions of PCP, lactate and their corresponding organic metabolites in segments of 0–2 cm, 2–5 cm, 5–8 cm, 8–11 cm, 11–13 cm and 13–15 cm, respectively, along with the cylindrical reactor.
(A) Concentration changes of PCP, 3-chlorophenol and phenol; (B) Concentration changes of lactate, acetate and propionate.
Spatial distribution of potential reductive dehalogenators and phenol anaerobic degraders
Figure 2 represented the distribution of 16S rRNA genes of the genera of potential dehalogenators, Desulfovibrio, Sulfurospirillum, Dehalobacter and Desulfitobacterium spp., as well as the potential fermentative or syntrophic degraders of phenol/3-CP, Cryptanaerobacter and Syntrophus spp. along with the cylindrical reactor. The highest bacterial density appeared in the inlet segment (0–2 cm) (about 108 copies mL–1) and dropped to about 106 copies mL–1 near the outlet segment (13–15 cm) (Fig. 2A). Archaea was detected with the density ranged from 103 to 106 copies mL–1, except for the outlet segment (13–15 cm).
Spatial distributions of Bacteria, Archaea, potential reductive dehalogenators and phenol oxidative degraders in segments of 0–2 cm, 2–5 cm, 5–8 cm, 8–11 cm, 11–13 cm and 13–15 cm, respectively.
(A) 16S rRNA genes copies of Bacteria and Archaea; (B) 16S rRNA genes copies of possible reductive dehalogenase; (C) 16S rRNA genes copies of possible phenol degraders.
Desulfovibrio and Sulfurospirillum spp., known as the facultative dechlorinators15,16, were evenly distributed among the whole reactor and occupied 0.2–30.3% and 0.2–8.3% of the copy number of Bacteria, respectively. Both of their highest densities were appeared at the segment 8–11 cm and while no obvious density decrease was observed where PCP was disappeared above 5 cm of segment (Fig. 1A). In contrast, Dehalobacter and Desulfitobacterium spp., the two dehalogenators17,18, occupied the much lower properties of bacterial composition (0.02–0.2% and 0.002–0.01% of the copy number of Bacteria, respectively). However, their density profiles were consistent with the distribution of PCP (Fig. 1A). Both of them showed a great decrease of wider than 10 times in the segments from 8 cm and were below the detection limit above 13 cm and 8 cm, respectively (Fig. 1A). Correlation analysis showed Dehalobacter and Desulfitobacterium spp. were positively correlated with the concentration of PCP, with the pearson correlation constant r of 0.22 and 0.94, respectively. However, no obvious correlation was observed for Desulfovibrio and Sulfurospirillum spp. It is inferred Dehalobacter and Desulfitobacterium spp. were involved in reductive dechlorination process and while, Desulfovibrio and Sulfurospirillum spp., were probably participated in the other bioprocesses, such as fermentation or nitrogen fixation17,18.
Copy numbers of the two possible phenol degraders, Cryptanaerbacter and Syntrophus spp., were gradually decreased from the inlet to outlet and occupied 0.1–0.3% and 0.4–5.4% of the copy number of Bacteria, respectively (Fig. 2C). None of them was detected in the outlet segment of 13–15 cm. Changes in numbers of Cryptanaerbacter and Syntrophus spp. were consistent with the distribution of Archaea, indicating their latent syntrophic relationships. Also, gene copy numbers of both Cryptanaerbacter and Syntrophus spp. were significantly correlated with the determined concentration of phenol (r = 0.63 and 0.51, respectively). So it is inferred Cryptanaerbacter and Syntrophus spp. were in associated with phenol syntrophic degradation.
PCR amplification and phylogenetic analysis of cprA genes
The presence of cprA genes was examined using the extracted DNA from the inlet segment 0–2 cm with the designed primers. About 400–500 bp lengths of products were successfully amplified with the designed primer pairs, except for a primer pair of f5-m3-g2 and r3-m3-g2 (SI Table S4). Through clone library analysis, genes that possessing the following two traits were judged as the putative cprA reductive dehalogenases: that is (i) possessed two-conserved iron-sulfur cluster binding motifs and (ii) shared the highest similarity with the reported cprA genes from the BLAST program of GenBank nucleotide sequence database compared other functional genes. Over 90% of amplified genes were regarded as the putative cprA genes, indicating the validity of the designed primers used. No positive amplicon was observed with the specific primer sets for cprA3, cprA4 and cprA5 (SI Table S4) identified in Desulfitobacterium hafniense PCP-117, suggesting the lack of these genes in the reactor.
Four clone libraries, PCP-1, PCP-2, PCP-3 and PCP-4, were constructed with the gene fragments amplified by primer sets of f4-m3-g1 and r3-m3-g1, f4-m3-g2 and r3-m3-g2, f5-m3-g1 and r4-m1 and f5-m3-g2 and r4-m1, respectively (SI Table S4). Representative genes were majorly classified into five types (Fig. 3) and their highest similarities with the reported cprA genes were shown in SI Table S5. Gene f4r3g1-1 was the most abundant in PCP-1 and possessing the highest similarity with the o-position chlorophenol reductase in Desulfitobacterium chlororespirans clone 1256 (75.7%) harboring two iron-sulfur clusters. Genes of f4r3g2-1 and f5r4g2-1 in clone libraries of PCP-2 and PCP-4, respectively, were of 100% similarity with each other and the most associated with the determined o-chloro/bromophenol cprA of Desulfitobacterium dehalogenans. F4r3g1-3 and f5r4g1-1 in clone libraries of PCP-1 and PCP-3 possessed less than 80% of similarities with o-chloro/bromophenol cprA of D. dehalogenans, cprA3 of Desulfitobacterium hafniense PCP-1 and putative cprA genes of Dehalobacter sp. CF, respectively. F5r4g1-2, f4r3g1-2, f4r3g2-2 and f5r4g2-2 were over 98% similar with each other and the most related to cprA genes specific for o-chloro/bromophenol in D. dehalogenans. Genes in group four, f4r3g1-4 and f4r3g2-3, harbored over 99% of the similarity with each other and both of them possessed the highest similarity with cprA genes determined in Dehalobacter restrictus.
Phylogenetic tree of the putative cprA genes determined from the four groups of clone libraries PCP-1 to PCP-4 in the inlet segment (0–2 cm) of the reactor (SI Table S4) and their closest relatives, reductive dehalogenase cprA genes.
The branch in bold represents the gene types, with the appearance frequency as shown in brackets. The numbers at the nodes indicate the percentages of times that nodes appeared in 1000 bootstrap analyses. The scale bar represents an estimated difference of 10% in nucleotide sequence positions.
PCR amplification and phylogenetic analysis of bcr and bamA genes
DNAs extracted on the inlet segment (0–2 cm) and outlet segment (13–15 cm) were examined for the genes associated with the central intermediate of benzoyl-coenzyme A (CoA) pathway in phenol degradation, including the benzoyl-coenzyme A reductases (BCR) of bamB, bzdN and bcrC and the benzoyl-CoA ring-cleaving hydrolase of bamA19. Positive amplicons with the correct size were detected for bamB gene at 0–2 cm of the segment, but no amplicon was found for bzdN, bcrC, or bamA genes in both the inlet and outlet segments (SI Table S6), indicating the lack of these genes there. The primer set for bamB genes was rather conserved19 and over 95% of the picked colonies shared the highest similarities with the identified bamB genes through clone library analysis. As bamB codes for BCR subunit in obligate anaerobes and while bzdN or bcrC targets for the facultative anaerobes19,20, the results further confirmed that phenol degradation was carried out through the obligate anaerobes13.
Clone library showed the amplified putative bamB genes were classified into mainly three types, BamB-23, BamB-47 and BmaB-77, with their closely related bamB genes as shown in phylogenetic tree (Fig. 4). Among them, BamB-47 was the most abundant (47.2%) and possessed 77% of similarity with bamB aldehyde dehydrogenase of Syntrophus aciditrophicus SB. BamB-23 and BamB-77 were closely related to bamB genes of Desulfotomaculum thermobenzoicum (75% of similarity) and Desulfotomaculum gibsoniae (72% of similarity), respectively. The determined putative bamB genes were mostly associated with the ones identified in either Syntrophus sp. or Desulfotomaculum sp. (Fig. 4), further revealed the syntrophic or fermentative features during phenol anaerobic degradation13.
PCR amplification and phylogenetic analysis of nifH genes
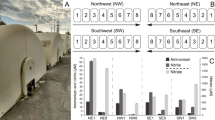
DNAs on both the inlet (0–2 cm) and outlet (13–15 cm) segments were examined for the presence of highly conserved nifH genes encoding iron protein21,22,23. Positive amplicons were obtained in both the inlet and outlet segments (SI Table S6) and over 95% of the picked colonies shared the highest similarities with the identified nifH genes. Clone library analysis showed five types (NifI-1, 2, 5, 22 and 47) of putative nifH genes in the inlet segment and four types (NifO-1, 2, 4 and 48) of putative nifH genes in the outlet segment, respectively (Fig. 5). Among of them, NifI-47, identical with NifO-1, was the most abundant in the inlet segment (35.4%) and possessed 76% of similarity with nitrogenase determined in Desulfotomaculum carboxydivorans strain CO-1-SRB, a benzoate degrader under sulfate reducing environment. The gene NifO-48 was the most abundant in the outlet segment (40.3%) and possessed 83% of similarity with nitrogenase determined in Methanoregula boonei strain 6A8. NifI-1 and NifO-2 were of 99% identical in sequence each other and closely related to the nitrogenase determined in Methanobacterium lacus strain (91% of similarity). The gene NifH-I-2 had the highest identity in DNA sequence with nifH genes in Bradyrhizobium japonicum clone (75% of similarity), a facultative nitrogen fixation bacterium. The genes, nifH-I-5 and nifH-I-22 possessed 78% and 75% similarities with genes in Dehalobacter sp. DCA and fermentation bacteria Clostridium sp. BNL1100, respectively. The gene nifH-O-4 possessed 69% similarity with nifH gene determined in Clostridium clariflavum DSM 19732.
Phylogenetic tree of the nitrogenase putative nifH genes determined in the inlet segment (0–2 cm) (clone library NifI) and outlet segment (12–15 cm) (clone library NifO) of the reactor (SI Table S6), as well as their closest related nitrogenase genes.
Spatial distribution of putative functional genes: cprA, bamB and nifH
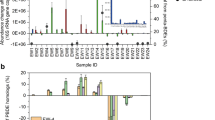
Spatial distributions of putative cprA, bamB and nifH genes along with the reactor segments were shown in Fig. 6. Gene copy numbers of putative cprA genes amplified from four pairs of primer sets, appeared the highest at the inlet segment 0–2 cm (104–106 copies mL–1) and decreased gradually as the distance increase from the inlet segment until below the detection limit at 13–15 cm. Distribution trend of the putative cprA genes was coincident with that of distribution of Dehalobacter and Desulfitobacterium spp. (Fig. 2B). In segments 0–13 cm, genes amplified with f5-m3-g1 and r4-m1 primers were numerically dominant (104–106 copies mL–1) except for 5–8 cm and while, genes amplified by f4-m3-g1 and r3-m3-g1 primers were the most frequently detected, although they possessed a relative low copy number (104–106 copies mL–1). Gene obtained by f4-m3-g2 and r3-m3-g2 primers ranged between 103–104 copies mL–1 at the segment 0–11 cm and below the detection limit in the segment 11–15 cm. Genes obtained by f5-m3-g2 and r4-m1 primers presented at a rather high copy numbers (104–106 copies mL–1), but, they were not detected between 8–15 cm. The correlation analysis showed the sum of determined cprA genes were highly correlated with concentration of PCP (r = 0.95), which further indicated that cprA genes played an important role during PCP reductive dechlorination process.
Spatial distributions of putative cprA, nifH and bamB genes in segments of 0–2 cm, 2–5 cm, 5–8 cm, 8–11 cm, 11–13 cm and 13–15 cm, respectively.
(A) quantitive distribution of cprA. F4r3g1, f4r3-g2, f5r4-g1 and f5r4-g1 indicated the amplified products with primer sets of f4-m3-g1/ r3-m3-g1, f4-m3-g2/r3-m3-g2, f5-m3-g1/r4-m1 and f5-m3-g2/r4-m1, respectively; (B) quantitive distribution of bamB-like genes; (C) quantitive distribution of nifH-like genes.
Putative bamB genes were distributed along with the whole segments, except for the segment 13–15 cm (Fig. 6B). Copy numbers of putative bamB genes did not show any big fluctuation in the segments 0–13 cm, being consistent with the distributions of the potential phenol syntrophic degraders (Fig. 2C). Copy numbers of putative bamB genes were comparable with the sum number of potential phenol syntrophic degraders, Cryptanaerbacter or Syntrophus spp. The significant correlation of bamB genes with concentration of phenol (r = 0.74) suggested its involvement in phenol degradation process.
Putative nifH genes were distributed widely among the whole reactor, with the highest copy number of appropriate 107 copies mL–1 at 2–5 cm and the lowest copy number of 9 × 105 copies mL–1 at 13–15 cm, respectively (Fig. 6C). The copy numbers of putative nifH genes showed a slight of decrease as the increase in the distance from inlet segment, which suggested the distribution of nitrogen fixing bacteria.
Discussion
The current study demonstrated the abundance and distribution of representative bacteria, functional genes as well as organic chemicals, which clearly revealed PCP mineralization process accomplished by the cooperative activities of the different bacteria (potential reductive dechlorinators and phenol degraders) and functional genes (putative functional genes of cprA, bamB and nifH genes). The distributions of PCP and phenol (Fig. 1A) suggested PCP dechlorination occurred at the segments 0–5 cm, confirmed by the highest densities of Dehalobacter, Desulfitobacterium spp. and putative cprA genes there (Figs 2B and 6A). Distributions of PCP and phenol revealed the anaerobic degradation of phenol took place at the segments 0–8 cm (Fig. 1A), verified by the distribution trends of both phenol potential degraders and putative bamB genes (Figs 2C and 6B). The gradual decrease in density of the potential PCP dechlorinators, phenol degraders and the corresponding genes at the segments longer than 5 cm and 8 cm from the inlet of the reactor (Figs 2B,C and 6A,B) reflected the gradual drop and lose of PCP dechlorination and phenol degradation activities at the middle and outlet segments of the reactor. Meanwhile, the wide distributions, the high copy numbers and diversities of putative nifH genes (Figs 5 and 6C) indicated the nitrogen fixation process throughout the reactor.
PCP dechlorination rate was determined as 5.89 μM d−1, calculated by multiplying the influent PCP concentration (50 μM) with the flow rate (0.003 mL min−1) and dividing by the effective volume where the reductive dechlorination occurred. The effective volume was calculated as 36.7 mL by multiplying the reactor pore volume (110 mL) with the effective length of reductive dechlorination occurred (5 cm) and dividing by the reactor length (15 cm). Therefore, the PCP mineralization in the segments 0–8 cm means 3.86 μM d−1 of the PCP mineralization rate, which is much higher than the previous report of 1.96 μM d−1 13, as well as the one achieved in an UASB reactor (0.13 μM d−1)24 and in a system with two combined columns in series (3.46 μM d−1)11. Lactate, the sole external nutrient, was completely fermented at the segment 0–5 cm and the metabolites of acetate and propionate were further decomposed in 0–13 cm (Fig. 1B). As calculated in the same way, lactate mineralization rate was 0.57 mM d−1 in this reactor.
It is noticed phenol degradation proceeded simultaneously with PCP reductive dechlorination at the segments 0–5 cm, confirmed by the low phenol concentration (less than 5 μM) and the decrease in concentration of PCP (Fig. 1A). The simultaneous reactions were further verified by the high density of both potential dechlorinating and phenol degrading bacteria and the corresponding functional genes: putative cprA and bamB genes (Figs 2C and 6B). Previously, the acclimatization of an anaerobic community capable of mineralizing HACs is considered challenging because the different redox niches required for multiple bioprocesses, especially reductive dechlorination and anaerobic oxidative degradation11,12,13,14. This study revealed the coexistence and cooperative work of dechlorinators and anaerobic phenol degraders in an acclimated consortium at 0–8 cm, which satisfied the requirement of simultaneous reduction and oxidation and avoided the internal nutrient competition. Besides, functions of nitrogen-fixing bacteria and lactate-fermenting bacteria13 should be mentioned (Figs 1B and 6C), which supplied organic nitrogen for microbial growth and electron (H2) for reductive dechlorination, respectively, although the distribution of their related populations and/or functional genes were not thoroughly elaborated.
The correlation analysis suggested Desulfitobacterium and Dehalobacter spp. were involved in reductive dechlorinating process and while, Syntrophus and Desulfotomaculum spp. were in associated with phenol degradation process (Figs 1 and 2). The functional bacterial composition was different from the previous constructed HAC anaerobic mineralization systems. Li et al.14 reported the anaerobic mineralization of tribromophenol, through fermentation, reductive debromination and oxidative degradation with Clostridium strain Ma13, Dehalobacter sp. and Desulfatiglans strain DS, respectively. A recent study reported anaerobic mineralization of the wide types of HACs in a long-term maintained anaerobic consortium and the involved potential dehalogenators were Dehalobacterium and Sulfurospirillum spp. and while, oxidative degraders were Geobacter spp.25 This indicated the diverse bacterial community structure for HAC mineralization system. However, the bacterial communities were analyzed by setting up the 16S rRNA clone library, which could only provide the restrictive community information. The recent developed high-throughput sequencing technics with the high taxonomic resolution are warranted to give the comprehensive evaluation of structure and diversity of functional bacteria and genes.
Through clone libraries, five types of cprA-like genes, three types of bamB-like genes and seven types of nifH-like genes were identified in the cylindrical reactor. The applied primers is based on the recent reports or synthesized according to the recently identified genes. So far, studies on identification and characterization of cprA and bamB genes were just on the beginning and the identified functional genes were still in limited genomes, i.e. cprA genes from the limited dechlorinating bacteria (Desulfitobacterium sp., Dehalobacter sp.) and bamB genes from the obligate anaerobes for aromatic-compounds degradation (Geobacter sp., Syntrophus sp.)17,18,19,20. There is the possibility that other types of functional genes might be present with the more diverse molecular. In fact, various type of reductive dehalogenase homologous have been recently reported existed in single isolates5,26, reflected that the identified genes with pairs of synthesized primers could not cover all the possible functional dehalogenases, bamB or nifH genes. Further study is required on the identification, characterization and genetic expression analysis of those functional genes, to extend the understanding of gene functioning mechanism in molecular aspect.
It should be mentioned that the determined putative nifH genes were originated from not only nitrogen fixation bacteria but also some other bacteria holding the nifH motifs, such as Dehalobacter sp., Methanoregula boonei and Bradyrhizobium sp. (Fig. 5). Nitrogen fixation activity has been reported by some reductive dechlorinators, aromatic-ring degraders or fermenters and while, their activities were enhanced through nitrogen fixation process23,27. So there is the possbility that the bacteria in charge of PCP mineralization also participated in nitrogen fixation process and simutanously supplied organic nitrogen for microbial growth. Compared with the inlet segment (0–2 cm), the outlet segment (13–15 cm) lacked putative nifH genes which originated from the potential dechlorinators and aromatic-ring degraders, caused by the lack of the related bacteria there (Fig. 2B).
Collectively, the study gave, for the first time, a clear image on the spatial distribution and diversity of potential reductive dechlorinators, anaerobic oxidative degraders and related functional genes (cprA, bamB and nifH) and represented a sustaining anaerobic bacterial community functioned cooperatively and sequentially to achieve PCP mineralization at the segment 0–8 cm from the column inlet. The study gave suggestions of designing a compact anaerobic HACs mineralization system by applying the multiple functional microbes in porous medium.
Materials and Methods
Preparation of the cylindrical reactor and the methods of sample storage
The cylindrical reactor used in this study was described previously13. Briefly, a glass beads-packed anaerobic cylindrical reactor (15 cm long) was provided by the inoculation of a consortium reductively dechlorinating PCP and introduction of solutions containing lactate (20 mM) and PCP (50 μM) under continuous up-flowing conditions. The cylindrical reactor performed the dechlorination of PCP to 3-CP and phenol with H2 as electron donor generated by lactate fermentation and then, 3-CP and phenol were further degraded without externally added electron acceptor after extending the hydraulic retention time. Nitrogen required for the microbial growth was supplied by microbial N2-fixation process. A 16S rRNA gene library analysis showed that the inlet segment harbored the potential dechlorinators, Dehalobacter, Desulfitobacterium, Sulfurospirillum and Desulfovibrio spp. and the phenol/3-CP fermentative or syntrophic degraders, Cryptanaerobacter and Syntrophus. Some possible nitrogen-fixers were detected in both the inlet and outlet segments.
After continuous running for 1.5 years with HRT of 25.3 days, the cylindrical reactor (15-cm long) was unloaded, crossed-cut into six segments under anoxic condition in a glove box (90% N2, 10% H2 atmosphere) and stored in −80 °C. Samples were collected as the following intervals from the bottom to the top of cylindrical reactor: 0–2 cm, 2–5 cm, 5–8 cm, 8–11 cm, 11–13 cm and 13–15 cm. The mixture of solid (soil and glass beads) and liquids of each segment was sampled with an autoclaved spatula. The samples were placed in 10 mL polypropylene tubes and stored at −30 °C before the chemical analysis and DNA extraction. The framework of the reactor and the analytical details after reactor decomposition was listed in SI Figure S1.
Chemical analysis
Initially, glass beads were gently removed from the frozen samples. PCP and the metabolites were determined by GC-MS (Shimadzu, Kyoto, Japan) equipped with the capillary column (DB-5MS, 30 m of length, 0.25 mm of inner diameter, J & W Scientific, Folsom, CA). Organic acids (lactate, acetate and propionate) were determined using a high-performance liquid chromatograph (CTO-10A; Shimadzu, Kyoto, Japan) equipped with an ultraviolet detector (UV 280 nm) and a Puresil C18 column (4.6 mm inner diameter, 250 mm length; Waters, Milford, MA, USA). The detailed method for measuring PCP, lactate and their metabolites were described in supplementary information (SI).
DNA extraction and the primer design
Initially, glass beads were gently removed from the frozen samples. DNA extraction from the samples (1.5 mL of the total volume of soil and liquid) was carried out using a DNA extraction kit, Isoplant II, according to the manufacturer’s instruction (Nippon Gene Co., Ltd., Toyama, Japan). Primers for the amplification of chlorophenol reductive dehalogenase cprA gene fragments, targeting for the binding motifs of the two iron-sulfur clusters were designed modified by combining the recently released cprA gene sequences18,28,29,30. The cprA gene sequences applied for primers design were listed in SI. Information of the sequences, positions and targeted genes of the designed primers were listed in supplementary information (SI) Table S1. Primer sets specific for gene sequences of cprA3, cprA4 and cprA5 in Desulfitobacterium hafniense DCB-217 were also utilized as listed in SI Table S1. Primer sets of bamB, bzdN, bcrC and bamA were applied for amplification of benzoyl-CoA reductase and ring-cleaving hydrolase involved in the anaerobic phenol degradation, with the detailed information as shown in SI Table S2. BamB targeted for active-site subunit of class II benzoyl-coenzyme reductases (BCRs) from obligate anaerobes19,20. BzdN and bcrC targeted for subunits of class I BCRs from facultative anaerobes (bcr referred to as genes coding for Thauera type and bzd referred to as genes coding for Azoarcus type)19. BamA targeted for ring-cleaving hydrolase of the benzoyl-CoA degradation genes20. The primer set utilized for the amplification of the highly conserved nitrogenase was NifH-f and NifH-r targeting nifH gene encoding the iron protein (SI Table S2)21,22. The amplification of putative cprA genes, nifH genes and benzoyl-CoA related genes were conducted with DNAs extracted from individual segments.
PCR, cloning and nucleotide sequencing analysis
PCR reactions for cprA genes, nifH genes and benzoyl-CoA related genes were performed with ExTaq DNA polymerase kit (Takara Bio Inc., Shiga, Japan). After PCR amplification and the product purification, clone libraries were set up using the pGem-T easy cloning kit (Promega, Madison, WI, USA). The detailed information for PCR program, the product purification and clone library setting up were described in in detail in SI. The clones were categorized on the basis of their distinct RFLPs. The representative clones with a unique RFLP pattern were sequenced with the BigDye terminatorv 3.1 cycle sequencing kit (Applied Biosystems, Foster, CA, USA) using an ABI 3100DNA sequencer (Applied Biosystems, Foster, CA, USA). The obtained nucleotide sequences were aligned using BioEdit Sequence Alignment Editor (7.0.5.3, Ibis Biosciences, Carlsbad, CA, USA) and classified into one type at a level of sequence similarity of more than 98%. The classification and analysis of nucleotide sequence were described in detail in SI. The nucleotide sequences obtained from clone library have been deposited in DNA Data Bank of Japan nucleotide sequence databases under accession numbers from LC033461 to LC033477.
Quantitative PCR (qPCR) analysis
QPCRs were performed using a Light Cycler system (Roche Diagnostics, Mannheim, Germany) and a Light Cycler Fast-start DNA Master SYBR green I kit (Roche Molecular Biochemicals, Indianapolis, IN), as described previously13. Bacteria, Archaea, Dehalobacter sp., Sulfurospirillum sp., Desulfitobacterium sp., Desulfovibrio sp., Cryptanaerobacter sp. and Syntrophus sp. were quantified using specific primer sets targeting for 16s rRNA genes as shown in SI Table S3. The putative cprA genes were quantified using the primer sets of f4-m3-g1/r3-m3-g1, f4-m3-g2/r3-m3-g2, f5-m3-g1/r4-m1 and f5-m3-g2/r4-m1, shown in SI Table S1. The putative nifH- genes and putative bamB genes were also quantified using the primer sets of NifH-f/NifH-r and BamB-f/BamB-r, respectively, as shown in SI Table S2. The detailed information on calibration curves and detection limits were described in detail in SI. Triplicate cultures were examined for each condition.
Additional Information
How to cite this article: Li, Z. et al. Spatial Abundance and Distribution of Potential Microbes and Functional Genes Associated with Anaerobic Mineralization of Pentachlorophenol in a Cylindrical Reactor. Sci. Rep. 6, 19015; doi: 10.1038/srep19015 (2016).
References
Sim, W. J., Lee, S. H., Lee, I. S., Choi, S. D. & Oh, J. E. Distribution and formation of chlorophenols and bromophenols in marine and river environments. Chemosphere. 77, 552−558 (2009).
US EPA (US Environmental Protection Agency). Ground water and drinking water. National Primary Drinking Water Standards, 2003. EPA-816-F-03-016. USEPA, Office of Water, Washington, DC, USA.
Field, J. A. & Sierra-Alvarez, R. Microbial degradation of chlorinated benzenes. Biodegradation. 19, 463−480 (2008).
Potrawfke, T., Timmis, K. N. & Wittich, R. M. Degradation of 1,2,3,4-tetrachlorobenzene by Pseudomonas chlororaphis RW71. Appl. Environ. Microb. 64, 3798−3806 (1998).
Fricker, A. D., LaRoe, S. L., Shea, M. E. & Bedard, D. L. Dehalococcoides mccartyi strain JNA dechlorinates multiple chlorinated phenols including pentachlorophenol and harbors at least 19 reductive dehalogenase homologous genes. Environ. Sci. Technol. 48, 14300−14308 (2015).
Huang, L. P. et al. Mineralization of pentachlorophenol with enhanced degradation and power generation from air cathode microbial fuel cells. Biotechnol. Bioeng. 109, 2211−2221 (2012).
Kazuhiro, K. et al. Aerobic mineralization of hexachlorobenzene by newly isolated pentachloronitrobenzene-degrading Nocardioides sp. strain PD653. Appl. Environ. Microbiol. 75, 4452−4458 (2009).
Bhatt, P., Kumar Suresh, M., Mudliar, S. & Chakrabarti, T. Biodegradation of chlorinated compounds−a review. Crit. Rev. Environ. Sci. Technol. 37, 165−198 (2007).
Hollinger, C., Wohlfarth, G. & Diekert, G. Reductive dechlorination in the energy metabolism of anaerobic bacteria. FEMS Microbiol. Rev. 22, 383−398 (1999).
Li, Z. et al. Involvement of Dehalobacter strains in the anaerobic dechlorination of 2,4,6-trichlorophenol. J Biosci. Bioeng. 116, 602−609 (2015).
Li, Z. L., Yang, S. Y., Inoue, Y., Yoshida, N. & Katayama, A. Complete anaerobic mineralization of pentachlorophenol (PCP) under continuous flow conditions by sequential combination of PCP-dechlorinating and phenol-degrading consortia. Biotechnol. Bioeng. 107, 775−785 (2010).
Yang, S., Shibata, A., Yoshida, N. & Katayama, A. Anaerobic mineralization of pentachlorophenol (PCP) by combining PCP-dechlorinating and phenol-degrading cultures. Biotechnol. Bioeng. 102, 81−90 (2009).
Li, Z. L., Inoue, Y., Suzuki, D., Ye, L. & Katayama, A. Long-term anaerobic mineralization of pentachlorophenol in a continuous-flow system using only lactate as an external nutrient. Environ. Sci. Technol. 47, 1534−1541 (2013).
Li, Z. L. et al. Anaerobic mineralization of 2,4,6-tribromophenol to CO2 by a synthetic microbial community comprising Clostridium, Dehalobacter and Desulfatiglans. Bioresour. Technol. 176, 225−232 (2015).
Drzyzga, D. & Gottschal, J. C. Tetrachloroethene dehalorespiration and growth of Desulfitobacterium frappieri TCE1 in strict dependence on the activity of Desulfovibrio fructosivorans. Appl. Environ. Microbiol. 68, 642−649 (2002).
Renpenning, J. et al. Combined C and Cl isotope effects indicate differences between corrinoids and enzyme (Sulfurospirillum multivorans PceA) in reductive dehalogenation of Tetrachloroethene, but not Trichloroethene. Environ. Sci. Technol. 48, 11837−11845 (2014).
Bisaillon, A., Beaudet, R., Lepine, F. & Villemur, R. Quantitative analysis of the relative transcript levels of four chlorophenol reductive dehalogenase genes in Desulfitobacterium hafniense PCP-1 exposed to chlorophenols. Appl. Environ. Microb. 77, 6261−6264 (2011).
Smits, T. H. M., Devenoges, C., Szynalski, K., Maillard, J. & Holliger, C. Development of a real-time PCR method for quantification of the three genera Dehalobacter, Dehalococcoides, an Desulfitobacterium in microbial communities. J. Microbiol. Methods. 57, 369−378 (2004).
Loffler, C. et al. Occurrence, genes and expression of the W/Se-containing class ΙΙ benzoyl-coenzyme a reductases in anaerobic bacteria. Environ. Microbiol. 13, 696−709 (2011).
Kuntze, K., Vogt, C., Richnow, H. H. & Boll, M. Combined application of PCR-based functional assays for the detection of aromatic-compound-degrading anaerobes. Appl. Environ. Microb. 77, 5056−5061 (2011).
Burbano, C. S. et al. Predominant nifH transcript phylotypes related to Rhizobium rosettiformans in field-grown sugarcane plants and in Norway spruce. Environ. Microbiol. Rep. 3, 383−389 (2011).
Mehta, M. P., Butterfield, D. A. & Baross, J. A. Phylogenetic diversity of nitrogenase (nifH) genes in deep-sea and hydrothermal vent environments of the Juan de Fuca Ridge. Appl. Environ. Microb. 69, 960−970 (2003).
Ju, X., Zhao, L. & Sun, B. Nitrogen fixation by reductively dechlorinating bacteria. Environ. Microbiol. 9, 1078−1083 (2007).
Wu, W. M., Bhatnagar, L. & Zeikus, J. G. Performance of anaerobic granules for degradation of pentachlorophenol. Appl. Environ. Microbiol. 59, 389−397 (1993).
Li, Z. L., Nan, J., Yang, J. Q., Jin, X. & Katayama, A. Temporal distributions of functional microbes and putative genes associated with halogenated phenol anaerobic dehalogenation and further mineralzaition. RSC Adv. 5, 89157–89163 (2015).
Chow, W. L., Cheng, D., Wang, S. Q. & He, J. Z. Identification and transcriptional analysis of trans-DCE-producing reductive dehalogenases in Dehalococcoides species. ISME J. 4, 1020–1030 (2010).
Desai, M. S., Assig, K. & Dattagupta, S. Nitrogen fixation in distinct microbial niches within a chemoautotrophy-driven cave ecosystem. ISME J. 7, 2411−2423 (2013).
Behrens, S. et al. Monitoring abundance and expression of “Dehalococcoides” species chloroethene-reductive dehalogenases in a tetrachloroethene-dechlorinating flow column. Appl. Environ. Microbiol. 74, 5695−5703 (2008).
Van De Pas, B. A. et al. Purification and molecular characterization of ortho-chlorophenol reductive dehalogenase, a key enzyme of halorespiration in Desulfitobacterium dehalogenans. J. Biol. Chem. 274, 20287−20292 (1999).
Von Wintzingerode, F., Schlotelburg, C., Hauck, R., Hegemann, W. & Gobel, U. B. Development of primers for amplifying genes encoding cprA- and pceA-like reductive dehalogenases in anaerobic microbial consortia, dechlorinating trichlorobenzene and 1,2-dichloropropane. FEMS Microbiol. Ecol. 35, 189−196 (2001).
Acknowledgements
This study was supported by Grants-in-Aid for Scientific Research (B2: 23310055 and B2:26281040) from the Ministry of Education, Culture, Sports, Science and Technology of Japan, by the New Energy and Industrial Technology Development Organization, Japan, by National Science Foundation for Distinguished Young Scholars of China (Grant No.51225802), by the National Natural Science Foundation of China (31400104), by the China Postdoctoral Science Foundation (2014M550196), by the innovation fund of Harbin Institute of Technology, China (HIT.NSRIF.2015092), by the Postdoctoral Science Foundation supported by Heilongjiang province of China (LBH-Z14087).
Author information
Authors and Affiliations
Contributions
L.Z.L., W.A.J. and K.A., designed the experiments; L.Z.L., N.J., H.C., conducted the experiment, wrote the paper and prepared the figures and tables; L.B., L.W.Z., C.H.Y., helped to check the manuscript and analyzed part of the data; Z.C.F. and Z.D.D. maintained the reactor system; K.D.Y., K.K. and K.K., instructed the qPCR and functional gene data analysis. All authors contributed to the manuscript writing and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, ZL., Nan, J., Huang, C. et al. Spatial Abundance and Distribution of Potential Microbes and Functional Genes Associated with Anaerobic Mineralization of Pentachlorophenol in a Cylindrical Reactor. Sci Rep 6, 19015 (2016). https://doi.org/10.1038/srep19015
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep19015
This article is cited by
-
Pesticide Pollution in Agricultural Soils and Sustainable Remediation Methods: a Review
Current Pollution Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.