Abstract
The κ opioid receptor (κOR) is a member of G-protein-coupled receptors and is considered as a promising drug target for treating neurological diseases. κOR selective 6′-GNTI was proved to be a G-protein biased agonist, whereas 5′-GNTI acts as an antagonist. To investigate the molecular mechanism of how these two ligands induce different behaviors of the receptor, we built two systems containing the 5′-GNTI-κOR complex and the 6′-GNTI-κOR complex, respectively and performed molecular dynamics simulations of the two systems. We observe that transmembrane (TM) helix 6 of the κOR rotates about 4.6o on average in the κOR-6′-GNTI complex. Detailed analyses of the simulation results indicate that E2976.58 and I2946.55 play crucial roles in the rotation of TM6. In the simulation of the κOR-5′-GNTI system, it is revealed that 5′-GNTI can stabilize TM6 in the inactive state form. In addition, the kink of TM7 is stabilized by a hydrogen bond between S3247.47 and the residue V691.42 on TM1.
Similar content being viewed by others
Introduction
Opioid receptors (ORs) are important members of G-protein-coupled receptors (GPCRs) and the main targets for analgesics1,2,3. There are three subtypes of ORs, namely - the μOR, δOR and κOR. Among the three subtypes, the κOR is a promising drug target for alleviating pain with a possible lower abuse potential4,5,6. As a result, the κOR subtype represents an excellent target to treat neurological disorders7,8,9,10,11,12,13. Selective agonists targeting the κOR have been developed to treat related disorders14 and antagonists selectively targeting the κOR have proved to be effective on curing depressant and anxiolytic diseases15,16,17,18.
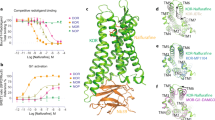
Among all the known ligands targeting the κOR, 5′-GNTI (5′-Guanidinonaltrindole) and 6′-GNTI (6′-Guanidinonaltrindole) are an interesting pair (Fig. 1). These two compounds are derivatives of Naltrindole (NTI), a highly potent δOR-selective antagonist19,20. The difference between 5′-GNTI and 6′-GNTI is only in the substitution site of the guanidinium group. However, these two compounds lead to distinct behaviors of the κOR. 5′-GNTI works as an antagonist to block or dampen the activation of the κOR21,22, whereas 6′-GNTI has proved to be a G-protein biased agonist to trigger the activation of the receptor23,24. It has been verified that the guanidinium group in 5′-GNTI can interact with the residue E2976.58 (Ballesteros/Weinstein numbering25) on transmembrane helix (TM) 620. It is interesting that the guanidinium group of 6′-GNTI has also been suggested to be interacting with E2976.58. Upon the binding of 6′-GNTI to the κOR, a rotation of TM6 of the receptor is likely triggered which leads to the subsequent activation behaviors of the receptor. Unfortunately, all these interaction patterns and the related activation mechanisms were surmised mostly based on homology models, which largely limit our understanding on the behaviors of the κOR.
Thanks to Stevens and his co-workers, the crystal structure of the human κOR in complex with its selective antagonist JDTic26 and the crystal structures of the δOR were reported recently27,28. These structures provide us atomic structural information on opioid receptors29,30. Unfortunately, all these crystal structures were obtained with the receptors in the inactive states. Furthermore, for new analgesics design, it is indispensable for us to exhaustively grasp the opioid activation mechanisms31. Therefore, to further study the behaviors of the κOR induced by different kinds of ligands, large amount of work still needs to be done.
To explicate the activation mechanism of the κOR triggered by 6′-GNTI, as well as the interactions between the antagonist 5′-GNTI and the κOR, we carried out sub-microsecond unbiased molecular dynamics (MD) simulations of systems with the κOR in the apo state and with 5′-GNTI-κOR and 6′-GNTI-κOR complexes, respectively, to provide active and inactive models of the GPCR based on the action of the so-called “molecular switches“32,33 buried in the receptor. TM6 of the κOR was observed to rotate about 4.6o on average in the κOR-6′-GNTI complex. Helices TM7 and TM2 highly responded to the activation signal in 6′-GNTI systems too. In the simulation of the κOR-5′-GNTI system, 5′-GNTI could stabilize TM6 in the inactive state form.
Results
Initial interaction modes obtained from molecular docking
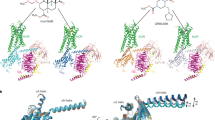
The docking poses of 6′-GNTI and 5′-GNTI are almost identical with each other as shown in Fig. 2. Strong electrostatic interaction is formed between the protonated nitrogen on the heterocyclic ring and D1383.32 in each complex. The phenolic hydroxyl group of both ligands interacts with H2916.52 bridged by two water molecules and the cyclopropyl group interacts with W2876.48. Meanwhile, Y1393.33 and Y3207.43 also contribute to the stabilization of the two ligands at the orthosteric site. Because of the difference in the substitution site of the guanidinium group in 6′-GNTI and 5′-GNTI, different interaction patens were observed when the two ligands bind to the κOR. The guanidinium group of 5′-GNTI is located in the cavity formed by the residues on TM6 and TM7 and forms three hydrogen bonds with E2976.58, whereas the guanidinium group of 6′-GNTI is positioned in a cavity formed by the residues on TM5 and TM6 and forms only one hydrogen bond with the residue E2976.58. In addition, the docking poses of the two GNTIs are almost identical with the conformation of NTI adopted in the crystal structure of the NTI-δOR complex. With the structures of 5′-GNTI or 6′-GNTI in the orthosteric pocket of the κOR and the κOR in the apo state, we built three systems, 5′-GNTI-κOR, 6′-GNTI-κOR and the κOR in the apo state, to be used for the following simulations.
Overview of the MD simulation results
In our simulations, the systems reached the equilibrium at about 100 ns as indicated from the root mean square deviation (RMSD) values in the simulations (Fig. 3A). The RMSD values of the κOR with respect to its crystal structure range from 2.0 to 3.6 Å (Fig. 3A). The ligands undergo comparably small conformational changes as indicated by their RMSD values, which range from 0.2 to 1.2 Å (Fig. S1). In addition, we also calculated the root mean square fluctuations (RMSFs) of the systems to analyze the fluctuations of the receptor. The loop section of the receptor exhibits a much larger fluctuation as compared to the conserved TM section (Fig. 3B). IL (intracellular loop) 2, EL (extracellular loop) 2 and IL3, show significant fluctuations, which are related to the fact that there are many residues on these three loops. The large fluctuation of the loop section reflects the significant conformational changes of the residues adjacent to these loops.
Ligand cluster analysis was performed on the two 600 ns GNTIs-κOR systems and two sub-state conformations were obtained under a 0.2 Å RMSD cut-off (Table 1). The main diversity between the two clusters was the tilt direction of indole ring of GNTIs (Fig. S2). One direction was outward if viewed from the extracellular side, whereas the other one was inward. Interestingly, the tilt direction of indole ring was inward among Cluster 1 of 6′-GNTI-κOR system, while that was outward in 600 ns 5′-GNTI-κOR system. Meanwhile, the population of Cluster 1 ran up to a half in 6′-GNTI-κOR system, whereas that was in the majority, amount to 81.4% in 600 ns 5′-GNTI-κOR complexes. Given the corresponding changes of guanidinium group (Fig. S2) and their important roles in GNTIs binding, we think the above-mentioned differences were mainly due to the diverse performances of guanidinium group between the two GNTIs-κOR systems.
Movements of the transmembrane helices
Comparisons of GPCR crystal structures in the inactive and active states have revealed several conserved switches to explain the inhibition or activation mechanisms of GPCRs34. One of the conserved switches is the large-scale rearrangements of the TM helices, including the rotation of TM6, the movement of TM5, the slight rotation and upward movement of TM3 and the inward movements of TM7 and TM135. TM6 and TM3 are at the heart of any common activation pathways, because they are coupled with critical conserved residues and are in direct contact with other helices except for TM136. We thus analyzed the conformational changes of TM6 and TM3.
A sharp kink of TM6 has been observed in the crystal structure of the κOR as well as in other family A GPCR structures with P6.50 as the pivot point36,37. Such a kink is preserved in our MD simulations (Fig. 4B). However, the kink angle of TM6 with P2896.50 as the pivot point varies with the simulation time. It changes from 160o to 140 o at the initial stage of the simulation and then increases to about 159o again (Fig. 4B and Table S1). Besides the kink of TM6, TM6 in the 6′-GNTI-κOR complex counterclockwise rotates about 4.6o on average, even 11.4o at 600 ns conformation viewed from the extracellular side of the receptor (Fig. 4A). This observation is consistent with the published results35. In contrast, the TM6 kink in the 5′-GNTI-κOR complex or the apo κOR form was found to be stabilized at about 160o and the counterclockwise rotation of TM6 was not observed from our simulations. We thus believe that the variation of the kink angle and the counterclockwise rotation of TM6 can be used to explain the different behaviors of the κOR triggered by 5′-GNTI and 6′-GNTI.
Normal mode analysis (NMA)38 is an efficient method for predicting inherent flexibilities in biological macromolecules. We performed NMA on the typical structures of the initial crystal structure, principal component and final 600 ns state in Fig. 4A, to detect the intrinsic motions of κOR. The low-frequency modes of κOR produced by the NMA reflected the global motions of the receptor and were often related to biological functions39. The first two lowest-frequency motion modes (modes 1 and 2) on the final 600 ns state and principal component were relevant to the transition from active-like to inactive (Fig. 5). Interestingly, NMA of the initial crystal structure revealed that the inactive state had an intrinsic potential to change back to the active-like conformation. However, the modes relevant to this transition were only among the fifth and third lowest-frequency motion modes. Therefore, unless an external force or ligand was present, κOR would favour the inactive state within the circuit. The consistency between the results of MD and NMA supports the efficiency of both methods in studying the large-scale motions of TM6 domains.
NMA profiles of TM6 conformational changes of the initial crystal structure, principal component and final 600 ns state (Fig. 4A) in 6′-GNTI-κOR system.
The arrows mean the directions of these conformational changes.
The conformational changes of TM3 and TM7 in the simulations can also be used to disclose the role of 5′-GNTI and 6′-GNTI in the activation of the κOR. The binding of GNTIs causes large conformational change of TM3 in the GNTIs−κOR complexes as shown in our simulations. In the simulation of the 5′-GNTI-κOR complex, we observed a clockwise rotation of TM7 if viewed from the extracellular side (Fig. S4A). We attribute the clockwise rotation of TM7 to the fluctuation of ECL3 (Video S1A). ECL3 is mainly composed of hydrophilic residues, such as Ser or His. These residues form strong interactions with the guanidino group of 5′-GNTI, which is responsible for the clockwise rotation of TM7.
Comparison of the binding interactions of GNTIs with the κOR
Hydrogen bonds between the residues D1383.32, Y1393.33, H2916.52 and E2976.58 on the receptor and the ligands are formed for most of the time in the simulations of the 5′-GNTI-κOR and 6′-GNTI-κOR complexes (Table 2 and Fig. 2). E2976.58 forms two HBs with the ligands and contributes significantly to the stabilization of the ligands at the orthosteric site. In addition, the protonated nitrogen in either 6′-GNTI or 5′-GNTI is not able to form any HBs with D1383.32 (Table 2). This can be considered as a common binding mechanism of opioid towards ORs40.
Contribution of I2946.55 to the conformational change of TM6
Detailed analysis of the trajectory from the simulation of the 5′-GNTI-κOR system showed that the minimum distance between Cα of I2946.55 and 5′-GNTI is mainly stabilized at about 5.5 Å (Fig. 6 and Table S1). The corresponding distance in the 6′-GNTI-κOR complex fluctuates between 4 and 6 Å, with the average at about 5.2 Å. The average minimum distance between Cα of I2946.55 and 6′-GNTI is about 0.3 Å shorter than that between Cα of I2946.55 and 5′-GNTI (Fig. 6A and Video S1B). The shorter distance between Cα of I2946.55 and 6′-GNTI can raise the steric clash between 6′-GNTI and I2946.55. The existence of the steric clash between I2946.55 and 6′-GNTI can be further indicated from the temporal evolutions of an angle between residue I2946.55 and TM6 (Fig. 6B). Because of the steric clash, this angle undergoes a significant fluctuation between 40o and 120o, with an average value at about 63o. On the contrary, the corresponding angle is stabilized at about 81o in the 5′-GNTI-κOR complex. The RMSF values also reflect that the fluctuation of the angle in the 6′-GNTI-κOR complex is larger than in the 5′-GNTI-κOR complex (Fig. 3B). In the simulation of the κOR in the apo state, this angle fluctuated between 40o and 100o freely (Fig. S5A).
The strong steric clash between I2946.55 and 6′-GNTI has a significant influence on the conformational states of E2976.58 (Fig. 6C). The angles between residue E2976.58 and TM6 is mainly stabilized at about 77o in the 6′-GNTI-κOR complex, whereas in the 5′-GNTI-κOR complex it is at about 86o on average. The large influence could also be reflected from the diverse performances of ligand cluster among 6′-GNTI-κOR interaction (Table 1 and Fig. S2).
Conformational change of TM7
A sharp kink on TM7 was observed in the crystal structure of the κOR26 with P3277.50 as the pivot point36,37. Although this kink was preserved in our simulations, the kink angles behave differently in the three systems. Temporal evolutions of the kink angle of TM7 indicate that the kink in the 5′-GNTI-κOR system is more stable than that in the other two systems. The kink angle is stabilized at about 150o in the 5′-GNTI-κOR complex (Fig. 7 and Table S1). However, the kink angle in the 6′-GNTI-κOR complex or in the apo form shows an obvious deviation. Specifically, this kink angle in the 6′-GNTI complex changed from about 150o to 130o at the initial stage of the simulation. The temporal evolution of the kink angles could be used to explain the responses of TM7 caused by GNTIs.
A detailed analysis of the simulation results reveals that the kink is stabilized by a hydrogen bond formed between S3247.47 and V691.42 (Table 2). The occupancy rate of this hydrogen bond is 24% from the simulation of the 5′-GNTI-κOR system, which is much higher than that in the 6′-GNTI-κOR system (2.7%) and in the apo κOR (6.1%). A recent study on CRF1R also proved the significance of the kink in explaining the antagonistic mechanism41.
Allosteric action mediated by the sodium ion
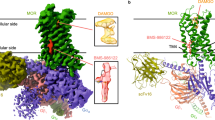
The role of the sodium ion in the function of some GPCRs has become an attractive topic recently. The crystallographic structure of the adenosine A2A receptor (A2AR) at an ultra-high resolution provides us with the first evidence of the binding of the sodium ion to the GPCR, followed by the crystallographic structures of the β1-adrenergic receptor and δOR28,42,43. In the δOR, the sodium ion is positioned around the negatively changed D2.50 and coordinated by two polar residues (N1313.35 and S1353.39) and two water molecules28. Many studies have been carried out to investigate the allosteric activation of GPCRs mediated by the sodium ion44. A recent molecular dynamics simulation of the μOR suggested that a sodium ion penetrates into the allosteric pocket from the extracellular side of the receptor to perform its allosteric effects on the receptor40. The dynamic properties of the sodium ion in the apo-κOR have also been described in the latest work of Filizola et al.45.
In the crystal structure of the κOR, the sodium ion is not co-crystallized with the κOR. We thus placed a sodium ion in the allosteric pocket of the κOR by referring to the position of the co-crystallized sodium ion in the crystal structure of the NTI-δOR complex28. In our simulations, the sodium ion is stablized in the allosteric pocket, reflecting that the position of the sodium ion is reasonable (Fig. 8A).
In our simulations, although the sodium ion is stablized in the allosteric pocket, the residues coordinating the sodium ion undergo conformational changes and result in the fluctuation of the distances between the sodium ion and the residues (Fig. 8A). For example, the distance between the sodium ion and D1052.50 increased significantly during 180–300 ns in the simulation of the 6′-GNTI-κOR system due to the conformational change of D1052.50 (Fig. 8B); D1052.50 deviated from the allosteric pocket and N3227.45 was involved in the direct interaction with the sodium ion during 180–280 ns.
The 3-7 lock
Formation of a hydrogen bond between D1383.32 and Y3207.43, which is named as the 3–7 lock in the following text, is suggested to play a key role in the activation of the κOR. To investigate the 3–7 lock in detail, we monitored the temporal evolution of the distance between D1383.32 and Y3207.43 (Fig. 8C). In the 5′-GNTI-κOR system, a hydrogen bond between D1383.32 and Y3207.43 formed at about 80 ns and broke at about 500 ns of the simulation. The occupation rate of the hydrogen bond was 69.6%. In the simulation of the 6′-GNTI-κOR system, a hydrogen bond between D1383.32 and Y3207.43 formed at about 340 ns and was preserved until the end of the simulation, resulting in an occupation rate of 44.4%. On the contrary, the 3–7 lock was not found in the whole simulation of the κOR in the apo form. We thus contribute the formation of the hydrogen bond between D1383.32 and Y3207.43 to the binding of 5′-GNTI or 6′-GNTI to the orthosteric pocket of the κOR.
The antagonistic effects of 5′-GNTI
To investigate whether 5′-GNTI can trigger a change of the conformation of the κOR induced by 6′-GNTI to the inactive conformation, we docked 5′-GNTI to the last snapshot the κOR generated in the simulation of the 6′-GNTI-κOR system (Fig. S6) and performed another MD simulation with a time scale of 100 ns. Interestingly, we found that TM6 gradually returned to the antagonistic state under the action of 5′-GNTI (Fig. 9A) as indicated by the kink angle of TM6. This angle changed from about 160o to 145o at the beginning of the simulation and then increased to 157 ± 4o (Fig. 9B and Table S1). Structural alignments of TM6 also indicated that 5′-GNTI triggered the conformational change of the κOR induced by 6′-GNTI to the inactive state gradually. In addition, the tilt direction of the indole ring was outward in Cluster 1 of the additional 5′-GNTI, which was identical with the 600 ns 5′-GNTI-κOR system (Table 1 and Fig. S2).
The conformational changes of TM6 in additional 5′-GNTI system.
(A) Structural alignment of κORs with initial crystal structure (gray), principal component of additional 5′-GNTI-κOR complex (yellow) and 600 ns state of 6′-GNTI-κOR complex (blue). (B) Evolution of the kink angle of TM6 in additional 100 ns 5′-GNTI system.
Discussion
In our simulations, we found that E2976.58 in the κOR is directly interacting with 6′-GNTI. This interaction, together with the steric effect from I2946.55, contributes to the rotation of TM6 as well as the movements of other TMs. In contrast, this ligand shows no agonistic effect towards the μOR and δOR23. From sequence alignments of the κOR, μOR and δOR, we can easily find that E6.58 in the κOR corresponds to K6.58 in the μOR and to W6.58 in the δOR46. On the other hand, it has been suggested that both K6.58 in the μOR and W6.58 in the δOR are not directly interacting with the guanidinium group of 6′-GNTI, which leads to the absence of the agonistic effect of this ligand towards the μOR and δOR23. All these observations demonstrate that the direct interaction between the residue E2976.58 and 6′-GNTI plays an indispensible role in the activation of the κOR and the residues E2976.58 and I2946.55 function as a critical pair in the activation of the κOR.
Besides the nontrivial function of E2976.58 and I2946.55 in the activation of the κOR, large conformation changes of TM7 and TM2 were also identified in our simulations as indicated by the kink angles of TM7 with P3277.50 as the pivot point (Fig. 7) and TM2 with D1052.50 as the pivot point (Fig. 8B). Such conformational changes can further help us understand the mechanism of κOR activation. In addition, D1052.50 is the key residue in positioning the sodium ion in the allosteric pocket. Therefore, we believe that the sodium ion also contributes to the activation of the κOR.
S3247.47 is located adjacent to the pivot point P3277.50 on TM7 and forms a hydrogen bond with a residue on TM1 in the simulation of the 5′-GNTI-κOR system (Fig. 7 and Table 2). Because of the formation of the hydrogen bond, the conformational change of TM7 is hampered. As a result, TM7 is stabilized in its initial conformation and leads to that the κOR adopts the inactive conformation. We thus suggest S3247.47 plays an important role in stabilizing the κOR at the inactive state.
The 3-7 lock is found to be formed in our simulations of the 5′-GNTI- κOR and 6′-GNTI-κOR systems. In contrast, such a lock is breaking-down in the crystal structure of the JDTic-κOR complex. Both 5′-GNTI and 6′-GNTI are derivatives of morphinan, while the ligand JDTic47 is a pure antagonist with (3R,4R)-3,4-dimethyl-4-(3-hydroxyphenyl)piperidine as the scaffold. We can thus use the ligand in the orthosteric pocket to explain the formation and breaking-down of the 3-7 lock in the κOR - the derivatives of morphinan can induce the formation of the 3-7 lock, while a pure antagonist results in the breaking-down of such a lock in the κOR.
In addition, we observed a large rotation of TM6 around the pivot point P2896.50 in the simulation of the 6′-GNTI-κOR system, which is an indispensible step for the activation of the κOR. In our opinion, the receptor needs to visit a set of intermediate states for its activation from the inactive state to the fully active state48. The rotation of TM6 allows the receptor to visit one of the key intermediate states for the activation of the κOR. However, we did not observe full activation of the receptor probably due to the limitation of the available computational resources.
Conclusions
In this work, we studied the molecular switches of the κOR triggered by 5′-GNTI and 6′-GNTI using molecular dynamics simulations. We observed about 4.6o rotation of TM6 on average in the κOR-6′-GNTI complex. Detailed analysis of the simulation results revealed that E2976.58 and I2946.55 play a crucial role for the rotation of TM6. On the other hand, we found that the hydrogen bond between S3247.47 and residue V691.42 on TM1 contributes to the stabilization of the κOR in the inactive state as revealed from the simulation of the κOR-5′-GNTI complex.
Materials and Methods
Protein preparations
The published crystal structure of human inactive JDTic-bound κOR (PDB code: 4DJH), was obtained from an engineered κOR mutant protein where part of the intracellular loop between transmembrane helices TM5 and TM6, was replaced by T4 lysozyme (T4L)26. In order to do molecular dynamics simulations on the wild type receptor based on the crystal structure of the T4L mutant κOR dimer, we removed the T4L from the mutant receptor structure, the monomer B and other unnecessary parts. Then we reconstructed the loop of the remaining chain A by adding the missing residues S262 and T302–S303–H304–S305–T306 using the loop refinement protocol in Discovery Studio 3.5; 10 loops were generated and the most reasonable one was chosen for receptor construction. According to the latest crystal structure of inactive NTI-complexed δOR, ion Na+ was exactly located at the allosteric pocket position28.
The above three-dimensional integrated κOR structure was then imported into the Schrödinger software package. The protein structure was prepared with Protein Preparation Tool (ProPrep) in the Schrodinger 2012 suit software. Asn, Gln and His residues checked for protonated states automatically in ProPrep. Hydrogen atoms were added into the κOR crystal structure at the physiology pH environment by the PROPKA tool in Maestro with optimized hydrogen-bond network. No non-standard protonation state of the amino acids was found.
Molecular docking
By referring to the conformation of ligand NTI in crystal structures of δOR, κOR-selective antagonist 5′-GNTI and κOR-selective agonist 6′-GNTI were sketched in Maestro and subjected to a Monte Carlo Μultiple Minimum conformational search using the OPLS_2005 force field and water as implicit solvent (Surface Generalized Born (SGB) model). The output conformations were used as the starting point for the docking experiments.
The pocket Grid file of the κOR was generated by 20 Å around residue D1383.32 with the Receptor Grid Generation module and docked with Glide Docking module (Glide 5.8)49,50. The Van der Waals (vdW) scaling was set to 0.8 for nonpolar atoms of receptor and ligand. During docking, the number of docking output was set as 10000 poses per docking run at most. The most reasonable conformation was picked out for molecular dynamics simulations.
MD preparations
Two 5′-GNTI-κOR complexes, one 6′-GNTI-κOR complex and one apo-κOR system, were built for the simulations. A POPC (1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine) bilayer with the surface area of 75 × 75 Å2 on the X-Y plane was generated under Charmm36 force field by VMD program (Version 1.9.1). For each system, the receptor was first embedded into the POPC bilayer using our in-house program pre-aligned in the OPM (Orientations of Proteins in Membranes) database51,52,53,54. Thereafter, Pre-equilibrated 103 POPC lipids coupled with 11067 TIP3P water molecules in a box ~75 × 75 × 100 Å3 were used solvate the protein. Lipid molecules within 0.85 Å of the heavy atoms on the protein structure and water molecules in the bilayer were removed. 51 Na+ and 59 Cl- ions were used to produce neutral systems with 0.15 M NaCl in the water phase of GNTI systems, whereas the number of Cl- ions was 57 in apo-κOR system. We described the protein using CHARMM36 force field with cmap correction.
Ligand force field was generated by Paramchem webserver, a program coupled with CHARMM Force field55,56,57. Small molecules with a correct configuration were imported into Paramchem webserver and force field parameters of these compounds were then obtained through a script CHARMM General Force Field (CGenFF).
Molecular dynamics simulations
All simulations were performed using Gromacs V.4.6.558 and the CHARMM36 parameters for all compositions. In the first simulation step, the system was subjected to a 10000-step energy minimization with 1000.0 kJ/mol/nm as the force threshold. Then, the systems were gradually heated from 0 K to 310 K followed by 50 ps initial equilibration at constant volume and temperature at 310 K (NVT). An additional 1 ns equilibration was performed at constant pressure and temperature (NPT ensemble; 310 K, 1 bar) with two thermostats (stabilizing temperature independently for protein-ligand system and the lipids-water-ions system) at 0032 ps time steps. vdW and short-range electrostatic interactions were cut off at 12 Å. Long-range electrostatic interactions were computed by the Particle Mesh Ewald (PME) summation scheme. The MD simulations of the additional 5′-GNTI system was performed for 100 ns, while all the other systems were performed for about 600 ns under NPT conditions with Parrinello-Rahman pressure coupler methods and Nose-Hoover thermostat for temperature coupling. The time step for MD simulation was 2 fs and the integrator leap-frog algorithm was employed. While the MD simulations of the additional 5′-GNTI-κOR system was performed for 100 ns, all the other systems were performed for about 600 ns using the NPT ensemble with the Parrinello-Rahman pressure coupling and the Nose-Hoover temperature coupling methods.
Analysis of simulations
RMSD and RMSF calculations, ligand cluster statistics, hydrogen bond analysis, angle and distance evolutions were produced by the program Gromacs. The interval time of trajectory calculations, including RMSD, RMSF, angle, distance and principal component analysis, was 500 ps in 600 ns systems, whereas that in 100 ns 5′-GNTI systems was 100 ps. RMSD values were calculated through comparing to the initial simulation conformations. Principal components analysis was carried on through the g_covar tool. All the smooth curves in Figs 4, 7 and Fig. S3 were fitting groups, which were identical with corresponding Angle/RMSD calculations in color.
To determine the sub-state conformations that close ligands bound to κOR, structural clustering was performed using the g_cluster tool in Gromacs with the linkage algorithm based on the RMSD of each ligand molecule after alignment of the Cα atoms in the transmembrane helices in each frame to the starting structure59. A 0.2 Å RMSD cut-off was chosen because it best captured spatially distinct clusters and allowed the top clusters to be representative of the predominant binding sites explored. The clustering was performed on the simulation frames with 100 ps interval time. The populations of each cluster are given in Table 1. We examined the two most populated clusters and calculated the RMSDs for each ligand found in these clusters (Table 1).
Angle analysis was carried on by g_angle tool in Gromacs. The specific input parameters were as follows. The kink angle of TM6 was measured with atom Cα of V2856.46, P2896.50 and F2936.54; The angle between I2946.55 and TM6 was measured with atom Cδ and Cα of I2946.55 and atom Cα of W2876.48, while that angle between E2976.58 and TM6 was computed with atom Cδ and Cα of E2976.58 and atom Cα of F2936.54; The kink angle of TM7 was measured with atom Cα of Y3207.43, P3277.50 and A3317.54; The angle between D1052.50 and TM2 was measured with atom Cγ and Cα of D1052.50 and atom Cα of I982.43.
NMA was conducted using the ElNemo60 (http://www.igs.cnrs-mrs.fr/elnemo/index.html), a web interface to the elastic network model-based NMA.
Additional Information
How to cite this article: Cheng, J. et al. Molecular switches of the κ opioid receptor triggered by 6′-GNTI and 5′-GNTI. Sci. Rep. 6, 18913; doi: 10.1038/srep18913 (2016).
References
Lee, M. C., Wanigasekera, V. & Tracey, I. Imaging opioid analgesia in the human brain and its potential relevance for understanding opioid use in chronic pain. Neuropharmacology 84, 123–130 (2014).
Mehendale, A. W., Goldman, M. P. & Mehendale, R. P. Opioid overuse pain syndrome (OOPS): the story of opioids, prometheus unbound. J. Opioid Manag. 9, 421–438 (2013).
Candiotti, K. A. & Gitlin, M. C. Review of the effect of opioid-related side effects on the undertreatment of moderate to severe chronic non-cancer pain: tapentadol, a step toward a solution? Curr. Med. Res. Opin. 26, 1677–1684 (2010).
Riviere, P. J. Peripheral kappa-opioid agonists for visceral pain. Br. J. Pharmacol. 141, 1331–1334 (2004).
Cahill, C. M. et al. Does the kappa opioid receptor system contribute to pain aversion? Front Pharmacol. 5, 253 (2014).
Vanderah, T. W. Delta and kappa opioid receptors as suitable drug targets for pain. Clin. J. Pain 26 Suppl 10, S10–15 (2010).
Zhou, L. et al. Development of functionally selective, small molecule agonists at kappa opioid receptors. J. Biol. Chem. 288, 36703–36716 (2013).
Kivell, B. & Prisinzano, T. E. Kappa opioids and the modulation of pain. Psychopharmacology (Berl) 210, 109–119 (2010).
Eguchi, M. Recent advances in selective opioid receptor agonists and antagonists. Med. Res. Rev. 24, 182–212 (2004).
Wang, Y. H., Sun, J. F., Tao, Y. M., Chi, Z. Q. & Liu, J. G. The role of kappa-opioid receptor activation in mediating antinociception and addiction. Acta Pharmacol. Sin. 31, 1065–1070 (2010).
Neumeyer, J. L., Mello, N. K., Negus, S. S. & Bidlack, J. M. Kappa opioid agonists as targets for pharmacotherapies in cocaine abuse. Pharm. Acta Helv. 74, 337–344 (2000).
Bidlack, J. M. Mixed kappa/mu partial opioid agonists as potential treatments for cocaine dependence. Adv. Pharmacol. 69, 387–418 (2014).
Butelman, E. R., Yuferov, V. & Kreek, M. J. kappa-opioid receptor/dynorphin system: genetic and pharmacotherapeutic implications for addiction. Trends Neurosci. 35, 587–596 (2012).
Phan, N. Q., Lotts, T., Antal, A., Bernhard, J. D. & Stander, S. Systemic kappa opioid receptor agonists in the treatment of chronic pruritus: a literature review. Acta Derm. Venereol. 92, 555–560 (2012).
Metcalf, M. & Coop, A. Kappa Opioid Antagonists: Past Successes and Future Prospects. AAPS J. 7, E704–722 (2005).
Runyon, S. P. et al. Analogues of (3R)-7-hydroxy-N-[(1S)-1-{[(3R,4R)-4-(3-hydroxyphenyl)-3,4-dimethyl-1-piperidinyl ]methyl}-2-methylpropyl)-1,2,3,4-tetrahydro-3-isoquinolinecarboxamide (JDTic). Synthesis and in vitro and in vivo opioid receptor antagonist activity. J. Med. Chem. 53, 5290–5301 (2010).
Carroll, F. I. & Carlezon, W. A. Jr. Development of kappa opioid receptor antagonists. J. Med. Chem. 56, 2178–2195 (2013).
Urbano, M., Guerrero, M., Rosen, H. & Roberts, E. Antagonists of the kappa opioid receptor. Bioorg Med Chem Lett 24, 2021–2032 (2014).
Olmsted, S. L., Takemori, A. E. & Portoghese, P. S. A remarkable change of opioid receptor selectivity on the attachment of a peptidomimetic kappa address element to the delta antagonist, natrindole: 5′-[N2-alkylamidino)methyl]naltrindole derivatives as a novel class of kappa opioid receptor antagonists. J. Med. Chem. 36, 179–180 (1993).
Sharma, S. K., Jones, R. M., Metzger, T. G., Ferguson, D. M. & Portoghese, P. S. Transformation of a kappa-opioid receptor antagonist to a kappa-agonist by transfer of a guanidinium group from the 5′- to 6′-position of naltrindole. J. Med. Chem. 44, 2073–2079 (2001).
Jones, R. M. & Portoghese, P. S. 5′-Guanidinonaltrindole, a highly selective and potent kappa-opioid receptor antagonist. Eur J Pharmacol 396, 49–52 (2000).
Stevens, W. C., Jr. et al. Potent and selective indolomorphinan antagonists of the kappa-opioid receptor. J. Med. Chem. 43, 2759–2769 (2000).
Waldhoer, M. et al. A heterodimer-selective agonist shows in vivo relevance of G protein-coupled receptor dimers. Proc. Natl. Acad. Sci. USA. 102, 9050–9055 (2005).
Rives, M. L., Rossillo, M., Liu-Chen, L. Y. & Javitch, J. A. 6′-Guanidinonaltrindole (6′-GNTI) is a G protein-biased kappa-opioid receptor agonist that inhibits arrestin recruitment. J. Biol. Chem. 287, 27050–27054 (2012).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods in neurosciences 25, 366–428 (1995).
Wu, H. et al. Structure of the human kappa-opioid receptor in complex with JDTic. Nature 485, 327–332 (2012).
Granier, S. et al. Structure of the delta-opioid receptor bound to naltrindole. Nature 485, 400–404 (2012).
Fenalti, G. et al. Molecular control of delta-opioid receptor signalling. Nature 506, 191–196 (2014).
Filizola, M. & Devi, L. A. Grand opening of structure-guided design for novel opioids. Trends Pharmacol. Sci. 34, 6–12 (2013).
Cox, B. M. Recent developments in the study of opioid receptors. Mol. Pharmacol. 83, 723–728 (2013).
Filizola, M. & Devi, L. A. Structural biology: how opioid drugs bind to receptors. Nature 485, 314–317 (2012).
Ahuja, S. & Smith, S. O. Multiple switches in G protein-coupled receptor activation. Trends Pharmacol. Sci. 30, 494–502 (2009).
Trzaskowski, B. et al. Action of molecular switches in GPCRs–theoretical and experimental studies. Curr. Med. Chem. 19, 1090–1109 (2012).
Katritch, V., Cherezov, V. & Stevens, R. C. Structure-function of the G protein-coupled receptor superfamily. Annu. Rev. Pharmacol. Toxicol. 53, 531–556 (2013).
Tehan, B. G., Bortolato, A., Blaney, F. E., Weir, M. P. & Mason, J. S. Unifying family A GPCR theories of activation. Pharmacol. Ther. 143, 51–60 (2014).
Venkatakrishnan, A. J. et al. Molecular signatures of G-protein-coupled receptors. Nature 494, 185–194 (2013).
Rosenbaum, D. M., Rasmussen, S. G. F. & Kobilka, B. K. The structure and function of G-protein-coupled receptors. Nature 459, 356–363 (2009).
Tirion, M. M. Large Amplitude Elastic Motions in Proteins from a Single-Parameter, Atomic Analysis. Phys. Rev. Lett. 77, 1905–1908 (1996).
Bahar, I., Atilgan, A. R. & Erman, B. Direct evaluation of thermal fluctuations in proteins using a single-parameter harmonic potential. Fold. Des. 2, 173–181 (1997).
Yuan, S., Vogel, H. & Filipek, S. The role of water and sodium ions in the activation of the mu-opioid receptor. Angew. Chem. Int. Ed. Engl. 52, 10112–10115 (2013).
Sun, X. et al. Residues remote from the binding pocket control the antagonist selectivity towards the corticotropin-releasing factor receptor-1. Sci. Rep. 5, 8066 (2015).
Liu, W. et al. Structural basis for allosteric regulation of GPCRs by sodium ions. Science 337, 232–236 (2012).
Miller-Gallacher, J. L. et al. The 2.1 Å resolution structure of cyanopindolol-bound beta1-adrenoceptor identifies an intramembrane Na+ ion that stabilises the ligand-free receptor. PLoS One 9, 1–9 (2014).
Katritch, V. et al. Allosteric sodium in class A GPCR signaling. Trends Biochem. Sci. 39, 233–244 (2014).
Shang, Y. et al. Mechanistic insights into the allosteric modulation of opioid receptors by sodium ions. Biochemistry 53, 5140–5149 (2014).
Cheng, J., Liu, G., Zhang, J., Xu, Z. & Tang, Y. Insights into subtype selectivity of opioid agonists by ligand-based and structure-based methods. J. Mol. Model. 17, 477–493 (2011).
Thomas, J. B. et al. Identification of the first trans-(3R,4R)-Dimethyl-4-(3-hydroxyphenyl)piperidine derivative to possess highly potent and selective opioid κ receptor antagonist activity. J. Med. Chem. 44, 2687–2690 (2001).
Dror, R. O. et al. Activation mechanism of the beta2-adrenergic receptor. Proc. Natl. Acad. Sci. USA 108, 18684–18689 (2011).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Halgren, T. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 47, 1750–1759 (2004).
Lomize, A. L., Pogozheva, I. D. & Mosberg, H. I. Anisotropic solvent model of the lipid bilayer. 2. Energetics of insertion of small molecules, peptides and proteins in membranes. J. Chem. Inf. Model. 51, 930–946 (2011).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res. 40, D370–376 (2012).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol .Graph. 14, 33–38, 27–38 (1996).
Vanommeslaeghe, K. et al. CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Vanommeslaeghe, K. & MacKerell, A. D. Jr. Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J. Chem. Inf. Model. 52, 3144–3154 (2012).
Vanommeslaeghe, K., Raman, E. P. & MacKerell, A. D. Jr. Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J. Chem. Inf. Model. 52, 3155–3168 (2012).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: Algorithms for Highly Efficient, Load-Balanced and Scalable Molecular Simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Suhre, K. & Sanejouand, Y. H. ElNemo: a normal mode web server for protein movement analysis and the generation of templates for molecular replacement. Nucleic Acids Res. 32, W610–614 (2004).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant 81273438), the Innovation Program of Shanghai Municipal Education Commission (Grant 13ZZ044), the Fundamental Research Funds for the Central Universities (Grant WY1113007), the 111 Project (Grant B07023) and the Swedish National Infrastructure for Computing (Grants SNIC2014-11-31 and SNIC2014-1-326).
Author information
Authors and Affiliations
Contributions
J.C. and X.S. carried out the molecular dynamics simulations. J.C. and Y.T.a designed the study and analyzed the data. Y.T.a was responsible for the project. J.C., X.S., W.L., G.L., Y.T.b and Y.T.a contributed to writing and commenting on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Cheng, J., Sun, X., Li, W. et al. Molecular switches of the κ opioid receptor triggered by 6′-GNTI and 5′-GNTI. Sci Rep 6, 18913 (2016). https://doi.org/10.1038/srep18913
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18913
This article is cited by
-
Bias-inducing allosteric binding site in mu-opioid receptor signaling
SN Applied Sciences (2021)
-
Computational insights into the G-protein-biased activation and inactivation mechanisms of the μ opioid receptor
Acta Pharmacologica Sinica (2018)
-
Allosteric inhibitor remotely modulates the conformation of the orthestric pockets in mutant IDH2/R140Q
Scientific Reports (2017)
-
Allosteric modulation model of the mu opioid receptor by herkinorin, a potent not alkaloidal agonist
Journal of Computer-Aided Molecular Design (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.