Abstract
Lactobacillus delbrueckii subsp. bulgaricus develops acid tolerance response when subjected to acid stress conditions, such as the induction of enzymes associated with carbohydrate metabolism. In this study, pyk gene encoding pyruvate kinase was over-expressed in heterologous host Lactococcus lactis NZ9000 and SDS-PAGE analysis revealed the successful expression of this gene in NZ9000. The survival rate of Pyk-overproducing strain was 45-fold higher than the control under acid stress condition (pH 4.0). In order to determine the transcription factor (TF) which regulates the expression of pyk by bacterial one-hybrid, we constructed a TF library including 65 TFs of L. bulgaricus. Western blotting indicated that TFs in this library could be successfully expressed in host strains. Subsequently, the promoter of pfk-pyk operon in L. bulgaricus was identified by 5′-RACE PCR. The bait plasmid pH3U3-p01 carrying the deletion fragment of pfk-pyk promoter captured catabolite control protein A (CcpA) which could regulate the expression of pyk by binding to a putative catabolite-responsive element (5′-TGTAAGCCCTAACA-3′) upstream the -35 region. Real-time qPCR analysis revealed the transcription of pyk was positively regulated by CcpA. This is the first report about identifying the TF of pyk in L. bulgaricus, which will provide new insight into the regulatory network.
Similar content being viewed by others
Introduction
Lactobacillus delbrueckii subsp. bulgaricus (L. bulgaricus) is a homofermentative facultative anaerobe, which has been commonly used in many fermented dairy products such as yoghurt and Italian cheese1. In addition, immune modulation and diarrhea-alleviating effects of this bacterium have been reported previously, suggesting its potential as a probiotic culture2. During its growth, L. bulgaricus gradually acidifies the environment through the conversion of pyruvate to lactate, which leads to acidification of the medium to approximately pH 3.83. Moreover, acidity in the stomach (pH 1.5–2) is also another acid stress encountered by L. bulgaricus during consumption3,4. Under these environmental conditions, acid tolerance response plays an important role in bacterial survival.
L. bulgaricus could develop acid tolerance response when subjected to moderate acid stress conditions, such as the induction of glycolysis-associated enzymes, the rerouting of pyruvate metabolism to fatty acid biosynthesis, as well as some modulations in protein synthesis3,4,5,6. In our previous study, enzymes involved in glycolysis, such as glucokinase (GlcK), enolase (Eno) and pyruvate kinase (Pyk), were found to be more abundant after acid stress treatment in L. bulgaricus CAUH16. The Pyk was up-regulated at both mRNA (10.78-fold) and protein (1.86-fold) levels after acid adaptation6. It is noteworthy that Pyk have also been observed to be more abundant in other Gram-positive bacteria under low pH conditions7,8. In Lactococcus lactis MG1363, the expression of pyk was increased 3.3-fold after acid treatment8. In Streptococcus mutans, Pyk was up-regulated more than 2.3-fold under acid stress7. Thus, it seems that Pyk probably contributes to acid stress tolerance in bacteria.
Pyruvate kinase (EC 2.7.1.40), a final-stage enzyme in glycolysis, catalyzes the transfer of a phosphoryl group from phosphoenolpyruvate (PEP) to adenosine diphosphate (ADP), generating adenosine triphosphate (ATP) and pyruvate9. This reaction is essentially irreversible in vivo and appears to be one of the major control points for the regulation of the glycolytic flux10. Moreover, the product pyruvate feeds into a number of metabolic pathways that places this enzyme at a primary metabolic intersection11. Therefore, Pyk plays an important role in both energy generation and control of intracellular metabolic flux distribution12,13,14,15. In L. bulgaricus, previous study revealed that the pyk gene is co-transcribed with the gene encoding phosphofructokinase (Pfk) and these two genes constitute the pfk-pyk operon16. However, the transcriptional regulation mechanism of this operon is still unclear. Therefore, determination of the transcription factor of pyk gene will provide new insight into the stress adaptive regulation of L. bulgaricus.
In this study, expression of pyk gene in L. lactis was carried out to investigate whether overproduction of Pyk would increase acid resistance in the heterologous host. The results indicated that the over-expression of pyk confers remarkable acid tolerance on the host L. lactis. Subsequently, we constructed a L. bulgaricus transcription factor library and employed the bacterial one-hybrid system to further determine the transcription factor that regulated the expression of pyk.
Results
Heterologous expression of pyk in L. lactis NZ9000 enhances acid tolerance
DNA sequencing results verified that DNA length of the amplified gene pyk was 1,770-bp-long, which predicted an open reading frame encoding 589 amino acids and a TAA stop codon. The nucleotide sequence of the amplified PCR product showed 99% identity with the pyk gene in L. bulgaricus ATCC11842 (Ldb0839) (GenBank Accession No. CR954253.1) and this sequence showed one mutation at position 33 (G to A). But, this mutation did not result in change in amino acid sequence. The SDS-PAGE analysis revealed the strong production of an expected 63 kDa protein in L. lactis NZPyk upon induction with 10 ng mL−1 nisin (Fig. 1A, lane 4), indicating the successful expression of pyk in L. lactis NZ9000. In the absence of nisin, there was no significant difference in the survival rate between L. lactis NZPyk and the control strain after the acid stress treatment (P > 0.05). However, the induced Pyk-overproducing strain showed markedly higher acid resistance than the controls, i.e. more than 45-fold increase in survival under low-pH condition (Fig. 1B). This indicates that the heterologous expression of pyk gene enhanced acid tolerance in the host strain L. lactis NZPyk.
The heterologous expression of pyk gene with nisin induction detected by SDS-PAGE and the survival of L. lactis NZ9000ck and L. lactis NZPyk after acid stress.
(A) Soluble extracts were analyzed on 12% denaturing SDS-PAGE. Lane 1, NZ9000ck without nisin induction; Lane 2, NZPyk without nisin induction; Lane 3, NZ9000ck with 10 ng mL–1 nisin induction; Lane 4, NZPyk with 10 ng mL–1 nisin induction. Red arrow indicates the over-produced Pyk. (B) Survival rate is calculated as the ratio of the number of colonies obtained on GM17 plates after and before acid treatment. Data are the averages from three independent experiments.
Lactic acid production was reduced by over-expression of Pyk in L. lactis NZ9000
L. lactis NZPyk were grown at 30 °C in GM17 broth medium with or without nisin induction, respectively. To determine whether addition of nisin affected the growth of L. lactis NZPyk, the cell counts of this strain were enumerated at 2 h and 4 h after nisin induction. As shown in Fig. 2, the cell counts were 9.18 ± 0.10 log (CFU mL−1) at 4 h in the absence of nisin and 9.10 ± 0.16 log (CFU mL−1) in the presence of nisin, respectively. However, the concentration of lactic acid produced by strain NZPyk with nisin induction was 49.73 ± 2.34 mM after 4 h incubation, whereas the concentration of lactic acid was 66.08 ± 2.43 mM in the absence of nisin (Fig. 2). These results indicated that over-expression of Pyk led to the lower lactic acid production in L. lactis NZ9000, which further verified the hypothesis that pyruvate metabolism would be rerouted to fatty acid biosynthesis under acid stress condition in L. bulgaricus, resulting in a possible modification of the cell membrane rigidity and impermeability to enhance acid tolerance.
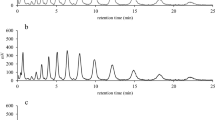
Growth of L. lactis NZPyk in GM17 broth with or without nisin induction and the effect of Pyk overproduction on the lactic acid monitored by gas chromatography.
Symbols: ■, cell viability of NZPyk without nisin induction; ○, cell viability of NZPyk with 10 ng mL−1 nisin induction; ▲, the concentration of lactic acid produced by strain NZPyk without nisin induction; □, the concentration of lactic acid produced by strain NZPyk with10 ng mL−1 nisin induction. Data are the averages from three independent experiments.
Construction of L. bulgaricus transcription factor library
For bacteria-one-hybrid analysis, a TF library containing 65 L. bulgaricus TFs was constructed using the vector pB1H2ω2-Prd as described previously17. Each recombinant plasmid in this TF library was verified by sequencing. To further confirm that the expression of TF as carboxy-terminal fusion to the omega-subunit of RNA polymerase, 7 TFs (TF06, TF17, TF27, TF35, TF54, TF59 and TF65) were randomly selected from the TF library and then whole-cell lysates were prepared and subjected to Western blotting using anti-FLAG antibody. As shown in Fig. 3, positive signal for each TF was observed on the Western blotting membrane, indicating that these omega-linked TFs were successfully expressed in the host E. coli US0. According to these results, we extrapolate that all TFs in this library could be successfully expressed in the host E. coli US0 and this L. bulgaricus TF library could be used for the subsequent bacterial one-hybrid analysis.
Detection of the expressed omega-TF fusion proteins by Western blotting analysis.
Lane 1: Dual color prestained broad molecular weight protein marker (10–170 kDa).Lane 2: Omega-Zif268 fusion protein17 was used as a positive control. Lane 3 to 9: TF06, TF17, TF27, TF35, TF54, TF59 and TF65, respectively. The theoretical molecular weights of the omega-TF constructs are: Zif268 = 23 kDa, TF06 = 40 kDa, TF17 = 49 kDa, TF27 = 31 k Da, TF35 = 42 kDa, TF54 = 36 kDa, TF59 = 22 kDa, TF65 = 30 kDa, marked with blue arrows respectively.
Mapping the transcription start site of pfk-pyk operon by 5′-RACE PCR
In order to construct the bait plasmids for bacterial one-hybrid analysis, 5′-RACE PCR was used to identify the promoter of pfk-pyk operon. A 335-bp DNA fragment was amplified from the 5′ end tailed cDNA using Nested Universal Primer A and gene-specific primer Internal-R. Only one DNA product could be obtained, suggesting that the pfk-pyk transcript was initiated at the single site. As shown in Fig. 4A, sequencing results indicated that the nucleotide immediately downstream of SMARTer IIA Oligonucleotide was transcription start site (TSS). This TSS (G) was located at position -43 relative to the start codon of pfk gene in L. bulgaricus. The potential -35 (AAGACT) and -10 (TATGAT) elements were present at position -72 and -49 relative to the ATG initiation codon of pfk, respectively (Fig. 4B).
(A) The 5′ end sequence of 5′-RACE PCR products. TSS, transcription start site. RBS, ribosome-binding site. (B) Sequence analysis of the promoter region upstream pfk-pyk operon. Putative −35 and −10 sequences are underlined. The putative catabolite-responsive element (cre) is enclosed in the box. The putative RBS site is shown in italics. (C) Linear map of pfk and pyk with the genomic DNA flanking these genes in L. bulgaricus. Two deletion fragments of the pfk-pyk promoter Ppfk, designated as p01 and p02, were used for constructing the bait plasmids. Numbers indicate positions relative to the start codon of pfk.
Expression of pyk gene was regulated by CcpA in L. bulgaricus
For the subsequent bacterial one-hybrid analysis, two deletion fragments of the promoter region (p01 and p02) were inserted respectively into the bait plasmid pH3U3-MCS (Fig. 4C). The transformants containing pH3U3-p01 or pH3U3-p02 could grow on the 5-FOA selective plates. These results indicated that deletion fragments did not self-activate the expression of reporter gene URA3. Thus, pH3U3-p01 and pH3U3-p02 can be used to capture the TF which binds to the regulatory element upstream pfk-pyk operon. These two bait plasmids were respectively co-transformed with TF library into the host strain, meanwhile a pB1H2-Prd derivative (pB1H2ck) lacking the TF gene was used as negative control. As shown in Fig. 5, only the strain co-transformed with pH3U3-p01 and TF library was able to grow on the selective NM plates (no histidine and 5 mM 3-AT). Plasmids isolated from these positive transformants were sequenced and sequence analysis revealed that the TF binding to p01 fragment was catabolite control protein A (CcpA). This transcription factor acts by binding to a consensus sequence called catabolite-responsive element (cre), usually found in the promoter region or the 5′ part of catabolite regulated genes18. In this study, a putative cre (5′-TGTAAGCCCTAACA-3′) upstream the -35 region was identified by using Target Explore (Fig. 4B), which was absent in the p02 fragment. These results suggested that the expression of pyk gene in L. bulgaricus was regulated by CcpA.
Identifying the transcription factor of L. bulgaricus pyk gene by bacterial one-hybrid.
The pH3U3-p01 and pH3U3-p02 were respectively co-transformed with TF library into E. coli US0 and transformants were grown on a selective NM medium plate. The plasmid pB1H2ck without TF gene was used as negative control. Each spot represents a 10-fold serial dilution of recovered cells from left to right (100 to 10−2).
Generally, CcpA and seryl-phosphorylated HPr are able to form a complex that enables CcpA to bind a cre sequence19. The binding capacity of CcpA was affected by the level of seryl-phosphorylated HPr, which varied in response to growth conditions (e.g. sugar utilization)20,21. Previous studies have shown that the transcription of pepQ encoding prolidase in L. delbrueckii is positively regulated by the binding of CcpA to a cre site located immediately upstream of the –35 region of its promoter22,23. Moreover, PepQ activity was 1.7 to 2.0-fold higher in cells grown in the presence of glucose compared to a culture with lactose, suggesting that the binding of CcpA to cre sites is dependent on the composition of the culture medium22. In this work, L. bulgaricus CAUH1 was respectively grown in MRSS (MRS broth devoid of beef extract) supplemented with 2% glucose or 2% lactose to investigate the role of CcpA in the regulation of pyk expression. Real-time quantitative PCR (RT-qPCR) analysis indicated that the expression level of pyk was 3.7 ± 0.2 fold higher in cells grown in MRSS supplemented with 2% glucose than that in cells grown in MRSS with 2% lactose. Thus, the transcription of pfk-pyk operon was positively regulated by CcpA in L. bulgaricus.
Discussion
In our previous study, proteomics approach complemented by transcriptional analysis revealed that Pyk might contribute to the acid tolerance of L. bulgaricus. However, the paucity of efficient transformation methods and effective molecular tools for gene inactivation severely limits directly functional identification in L. bulgaricus CAUH1. Therefore, heterologous expression of pyk gene was carried out using the L. lactis NICE system to investigate whether overproduction of Pyk would increase acid resistance in a heterologous host. In this study, improved acid resistance phenotype was observed in the host strain by overexpressing pyk, suggesting its contribution to acid tolerance response. In addition, Pyk-overproducing strain showed more than 10-fold increased viability than the control in GM17 liquid medium containing 1.25% w/v ox gall (Figure S1A). The overproduction of Pyk could also enhance cold resistance of the host strain, i.e. about 5-fold increase in survival under low temperature condition (10 °C) (Figure S1B). Previous studies indicated that Pyk was more abundant in some lactic acid bacteria and bifidobacteria after bile treatment24,25,26,27. This protein was also significantly induced in L. acidophilus RD758 during the cold adaptation28. Therefore, we supposed that the overexpression of pyk in the host strain played an important role in enhancing the resistance to multiple stresses.
It is noteworthy why the overproduction of Pyk contributes to the resistance to multiple stresses. According to the previous studies, the over-expression of pyk gene in E. coli and L. lactis significantly enhanced the activity of Pyk12,14. Nuclear magnetic resonance (NMR) analysis revealed that the rate of fructose 1, 6-bisphosphate (FBP) consumption was notably accelerated during glucose catabolism, whereas phosphoenolpyruvate (PEP) decreased to undetectable levels14. It has been firmly established in lactic acid bacteria that concentrations of PEP are relatively low in rapidly metabolizing cells29. Furthermore, PEP could result in allosteric inhibition of phosphofructokinase (Pfk) which is another rate limiting enzyme in the upper part of glycolysis30,31. Thus, the lower PEP concentration might cause an increased Pfk activity in Pyk-overproducing strains. These metabolic modulations allowed a greater glycolytic flux and produced more energy-rich intermediates (e.g. ATP and NADH) for bacteria to confront with environmental stress12. Moreover, lactic acid was observed to be decreased in Pyk-overproducing strain in this study. This was consistent with the previous observation that the utilization of pyruvate appeared to be rerouted toward fatty acid biosynthesis instead of other pathways (e.g. butanoate metabolism and lactic acid synthesis) in L. lactis and L. bulgaricus5,6,14. Therefore, Pyk overproduction was extrapolated to result in a rerouting of pyruvate metabolism to fatty acid biosynthesis and thereby presumably enhance the rigidity and impermeability of cellular membrane which has been considered to play an important role for bacterial survival under acid, bile and cold stress5,28,32.
Bacterial one-hybrid system was employed to identify the transcription factor that regulated the expression of pyk gene. The results revealed that the transcription of pyk gene in L. bulgaricus was regulated by CcpA. CcpA is a DNA-binding protein belonging to the Lacl/GalR family of bacterial transcription factors. Previous studies have shown that the DNA-binding activity of CcpA is triggered by the effector HPr-Ser-P33 and the binding of CcpA to its regulatory sites is dependent on the transport and metabolism of carbon sources in L. bulgaricus22,23. L. bulgaricus is able to transport glucose via the phosphotransferase system (PTS), but prefers lactose over glucose and transports this disaccharide via the non-PTS transporter LacS34,35. This protein is highly homologous to the LacS in Streptococcus thermophiles and also has a C-terminal hydrophilic IIA-like domain which can be phosphorylated by HPr-His-P33,36. The phosphorylation of LacS was reported to stimulate its lactose transport activity and then enhance the lactose/galactose exchange reaction37,38. Therefore, when L. bulgaricus cells were grown on the PTS-transported glucose, HPr was phosphorylated on serine residue by HPr kinase/phosphorylase (HPrK) in an ATP-dependent reaction21 and triggered the DNA-binding activity of CcpA. By contrast, when L. bulgaricus cells were grown on lactose, dephosphorylation of HPr-Ser-P was catalyzed by HPrK to release the HPr, which was further phosphorylated on histidine-15 residue to increase the lactose uptake rate. Taken all together, when L. bulgaricus was grown in the presence of lactose, HPr-Ser-P/HPr-His-P ratio is lower than that grown in the presence of PTS-transported glucose, which led to the decreased binding capacity of CcpA to its regulatory sites.
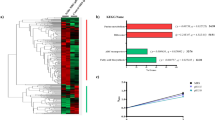
In Gram-positive bacteria, the CcpA/HPr-Ser-P complex can bind to a cis-acting catabolite response element (cre, with consensus sequence WTGNAARCGNWWW.CA, where W is A or T, R is A or G) that is commonly located in the proximity of promoters, thereby either repressing or activating the transcription of downstream genes or operons39. The binding of CcpA to cre sites upstream pfk-pyk operon has been studied in several lactic acid bacteria (Table 1)40,41,42,43. In L. lactis, L. plantarum and Streptococcus bovis, CcpA activates the transcription of pfk-pyk operon by binding to cre site upstream the -35 region and recruiting RNA polymerase to the promoter via direct protein-protein interaction44. However, the pfk-pyk operon was reported to be repressed by CcpA in L. casei, since there was another cre binding site between the -35 and -10 region43. Thus, CcpA could repress the transcription of pfk-pyk operon by looping DNA in the promoter region. In the present study, only one putative cre (5′-TGTAAGCCCTAACA-3′) is identified upstream the -35 region. In addition, RT-qPCR analysis revealed that the expression level of pyk was 3.7 ± 0.2 fold higher in cells grown in MRSS supplemented with 2% glucose than that in cells grown in MRSS with 2% lactose. These results indicated that the transcription of pyk in L. bulgaricus was positively regulated by CcpA. To our knowledge, this is the first report about the transcription factor of pyk gene in L. bulgaricus, which will provide new insight into the regulation network of L. bulgaricus CAUH1.
Methods
Bacterial strains and growth conditions
The bacterial strains and plasmids used in this study are listed in Table S1. L. bulgaricus CAUH1 was cultivated in de Man-Rogosa-Sharpe (MRS) broth medium and incubated statically at 37 °C. L. lactis NZ9000 was grown at 30 °C in GM17 (M17 broth supplemented with 0.5% w/v D-glucose). Escherichia coli were propagated aerobically at 37 °C in Luria-Bertani (LB) broth. For selection, media were supplemented with the relevant antibiotic at the following concentrations: 100 μg mL−1 ampicillin or 25 μg mL−1 kanamycin for E. coli; 10 μg mL−1 chloramphenicol for L. lactis NZ9000.
Heterologous expression and acid stress survival experiments
Standard PCR was carried out using Ex Taq polymerase according to the manufacturer’s instructions (Takara, Dalian, China). The pyruvate kinase gene pyk was amplified by PCR from the chromosomal DNA of L. bulgaricus CAUH1 using the primer pair: forward 5′- CATGCCATGGGAACGAAGATTGTTAGTACTTTAG-3′ and reverse 5′- CCGAGCTCGCAATCCTAGATTACAGGTTTG-3′. Restriction sites used for subsequent cloning are underlined: NcoI and SacI for the forward and reverse primers, respectively. The PCR amplicon was digested and then inserted into expression vector pNZ8148. Subsequently, the ligation mixture was transformed into L. lactis NZ9000 according to previously described procedures45. The recombinant plasmid pNZPyk was sequenced and further analyzed with the DNAMAN software package (Lynnon Biosoftware, Vaudreuil, Quebec, Canada). The recombinant strain with pNZPyk was designated L. lactis NZPyk. Meanwhile, a control strain (L. lactis NZ9000ck) was constructed by introducing the empty vector pNZ8148 into L. lactis NZ9000. Overnight cultures of L. lactis NZ9000ck and L. lactis NZPyk were respectively inoculated into 10 mL of fresh GM17 supplemented with 10 μg mL−1 chloramphenicol (1% inoculums). When cell density reached an OD600 nm of 0.3, nisin was added (final concentration: 10 ng mL−1) and further incubated for 2 h at 30 °C. Sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) analysis was used to investigate the expression of pyk in L. lactis. To assay low-pH survival on nisin-induced cultures, aliquots of 1 mL were collected and cells were resuspended in the same volume of fresh medium adjusted to pH 4.0 with lactic acid. Samples were taken after 1 h incubation at 30 °C and 10-fold serial dilutions were spread on GM17 plates with chloramphenicol. Survival rates were calculated by dividing the number of colony-forming units (CFU) per mL after incubation at pH 4.0 by the number of CFU per mL immediately after resuspension.
Quantification of lactic acid production in Pyk-overproducing strain by gas chromatography
To investigate whether Pyk overproduction affects the lactic acid production, 5 mL L. lactis NZPyk cultures grown in the absence or presence of 10 ng mL−1 nisin were collected respectively. Cells were removed by centrifugation and the supernatants were analyzed by gas chromatography as described previously with slight modifications to quantify lactic acid production46. The standard solutions were prepared in five different concentrations (1 mM, 10 mM, 20 mM, 40 mM and 100 mM) by diluting a 1 M lactic acid stock solution (Sigma, St Louis, MO, USA) to obtain calibration curve. Subsequently, the standard solutions and samples were methylated by sulfuric acid-methanol method47. After extraction with chloroform, 2 μL organic phase was analyzed by an high resolution gas chromatography (GC; Agilent 6890 Series gas chromatography system; Agilent Technologies, PA, USA) equipped with a flame ionization detector (FID) using a HP-FFAP column (30 m × 0.53 mm × 1.0 μm, Agilent). The injection temperature and detector temperature were both 230 °C. The column temperature was held at 50 °C for 1 min after injection, increased at a rate of 10 °C/min to 140 °C, held at 140 °C for 1 min, increased at 30 °C/min to 240 °C and held at 240 °C for 1 min. The carrier gas was nitrogen and the flow was 1 ml/min. The concentration of lactic acid produced by strain NZPyk was determined using a calibration curve and this experiment was performed in triplicate.
Construction of L. bulgaricus transcription factor library
In order to identify the transcription factor (TF) which regulates the expression of pyk by bacterial one-hybrid, L. bulgaricus TF library was constructed using the vector pB1H2ω2-Prd17. All TFs were predicted from the genome of L. bulgaricus according to DNA-binding domain (DBD) database48 and RegPrecise49. Then 65 putative TF genes were amplified using their specific primers (Table S1) and inserted into the KpnI and XbaI restriction sites of pB1H2ω2-Prd, resulting in a series of pB1H2ω2-derived plasmids (from pB1H2ω2-TF01 to pB1H2ω2-TF65). Then, these recombinant plasmids were transformed into the E. coli DH5α, respectively. Each recombinant plasmid was sequenced and further analyzed with the DNAMAN software package. To further investigate whether TF has expressed as a carboxy-terminal fusion to the ω-subunit of RNA polymerase, 7 of the 65 TFs were randomly selected and Western blotting was carried out with monoclonal ANTI-FLAG M2 (Sigma, St Louis, MO, USA; cat.# F1804). A subgenomic library for L. bulgaricus transcription factors was produced by mixing these recombinant plasmids.
Determination of the transcription start site of pfk-pyk operon
To determine the transcription start site of pfk-pyk operon, 5′ rapid amplification of cDNA ends (5′-RACE) experiment was performed by using the SMARTer™ RACE cDNA amplification kit (Clontech Laboratories, Takara Bio Company, Mountain View, CA) (41). Total RNA was isolated using TRIzol reagent according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA). Extracted RNA was examined by 1.5% (w/v) agarose electrophoresis, then quantified by a Qubit fluorometer and a Qubit RNA assay kit (Invitrogen, Eugene, Oregon, US). The first strand cDNA was generated by reverse transcription PCR from 1 μg total RNA of L. bulgaricus using the Random primer (N-9) and tailed at the 5′ end by SMARTer IIA Oligonucleotide (5′-AAGCAGTGGTATCAACGCAGAGTACGCGGG-3′) according to the manufacturer’s protocol. Subsequently, the 5′-RACE fragment was amplified by PCR from the cDNA product using the primer pair: Nested Universal Primer A (5′-AAGCAGTGGTATCAACGCAGAGT-3′) and a gene-specific primer Internal-R (5′-GCTCAATACCAGCCAGCTGTCCTTC-3′). The 5′-RACE products were cloned into the pGM-T vector (Tiangen, Beijing, China) and ten clones were sequenced to identify the transcription start site of pfk-pyk operon. The promoter region of pfk-pyk operon was further analyzed using the online promoter prediction tools NNPP and BPROM50,51.
Bacterial one-hybrid analysis
Bacterial one-hybrid system was carried out to determine the transcription factor of pyk as described previously with slight modifications52. Two deletion fragments of the pfk-pyk promoter Ppfk, designated as p01 and p02 respectively, were obtained by PCR with specific primers (Table S2). The p01 fragment lacking the −10 region was located at nucleotides −206 to −57 relative to the start codon of pfk and the p02 fragment lacking the −35 region was located at nucleotides −72 to 0 (Fig. 3C). The PCR products were digested with NotI and EcoRI and then ligated with pH3U3-MCS to generate the bait plasmid pH3U3-p01 and pH3U3-p02. These recombination plasmids were transformed into E. coli US0 for self-activation assays as described previously52. In order to screen the TF which could bind to the regulatory element upstream the pfk-pyk operon, pH3U3-p01 and pH3U3-p02 were respectively co-transformed with TF library into the host US0 and transformants were grown on a selective NM medium plate containing 5 mM 3-amino-1, 2, 4-triazole (3-AT), 100 μg mL−1 ampicillin and 25 μg mL−1 kanamycin. The plates were incubated at 37 °C for 36–48 h. Subsequently, plasmids isolated from the positive transformants were sequenced and further analyzed with DNAMAN. The TF binding site was predicted using an online database RegPrecise (http://regprecise.lbl.gov/RegPrecise/index.jsp) and Target Explorer (http://te.cryst.bbk.ac.uk/).
Real-time quantitative PCR
In order to further investigate the role of CcpA in the regulation of pyk expression, L. bulgaricus CAUH1 was respectively grown in MRSS (MRS broth devoid of beef extract) supplemented with 2% glucose or 2% lactose and then real-time quantitative PCR (RT-qPCR) was employed to analyze the expression level of pyk in L. bulgaricus CAUH1. Total RNA was isolated using TRIzol Reagent (Invitrogen) according to the manufacturer’s instructions and digested with RNase-free DNase I. Purified RNA was then applied to synthesize the first-strand cDNA, which was used as the template in RT-qPCR. Specific primers (Table S3) for pyk and the reference gene 16S rRNA were designed using PRIMER V5 software (PREMIER Biosoft International, Palo Alto, CA) and their specificity was checked before the quantitative analysis. Gene expressions were normalized by the ΔΔCT method53 and this experiment was performed in triplicate and the average results were reported.
Statistical analysis
All experimental data are shown as the mean ± S.D. Data were analyzed using SPSS (PASW) Statistics 19.0 (Version 19). Student’s unpaired t test was employed. A P value < 0.05 was considered to be statistically significant.
Additional Information
How to cite this article: Zhai, Z. et al. Functional role of pyruvate kinase from Lactobacillus bulgaricus in acid tolerance and identification of its transcription factor by bacterial one-hybrid. Sci. Rep. 5, 17024; doi: 10.1038/srep17024 (2015).
References
Hao, P. et al. Complete sequencing and pan-genomic analysis of Lactobacillus delbrueckii subsp. bulgaricus reveal its genetic basis for industrial yogurt production. Plos One 6, e15964 (2011).
Mercenier, A., Pavan, S. & Pot, B. Probiotics as Biotherapeutic Agents: Present Knowledge and Future Prospects. Curr. Pharm. Design 9, 175–191 (2003).
Lim, E. M., Ehrlich, S. D. & Maguin, E. Identification of stress-inducible proteins in Lactobacillus delbrueckii subsp. bulgaricus. Electrophoresis 21, 2557–2561 (2000).
Silva, J., Carvalho, A. S., Teixeira, P. & Gibbs, P. A. Effect of stress on cells of Lactobacillus delbrueckii sp. bulgaricus. J. Food Technol. 3, 479–490 (2005).
Fernandez, A. et al. Rerouting of pyruvate metabolism during acid adaptation in Lactobacillus bulgaricus. Proteomics 8, 3154–3163 (2008).
Zhai, Z. et al. Proteomic characterization of the acid tolerance response in Lactobacillus delbrueckii subsp. bulgaricus CAUH1 and functional identification of a novel acid stress-related transcriptional regulator Ldb0677. Environ. Microbiol. 16, 1524–1537 (2014).
Len, A. C., Harty, D. W. & Jacques, N. A. Proteome analysis of Streptococcus mutans metabolic phenotype during acid tolerance. Microbiology 150, 1353–1366 (2004).
Budin-Verneuil, A., Pichereau, V., Auffray, Y., Ehrlich, D. S. & Maguin, E. Proteomic characterization of the acid tolerance response in Lactococcus lactis MG1363. Proteomics 5, 4794–4807 (2005).
Valentini, G. et al. The allosteric regulation of pyruvate kinase: a site-directed mutagenesis study. J. Biol. Chem. 275, 18145–18152 (2000).
Zoraghi, R. et al. Functional analysis, overexpression and kinetic characterization of pyruvate kinase from methicillin-resistant Staphylococcus aureus. Biochemistry 49, 7733–7747 (2010).
Muñoz, M. E. & Ponce, E. Pyruvate kinase: current status of regulatory and functional properties. Comp. Biochem. Physiol. B 135, 197–218 (2003).
Emmerling, M., Bailey, J. E. & Sauer, U. Glucose catabolism of Escherichia coli strains with increased activity and altered regulation of key glycolytic enzymes. Metab. Eng. 1, 117–127 (1999).
Fry, B. et al. Characterization of growth and acid formation in a Bacillus subtilis pyruvate kinase mutant. Appl. Environ. Microbiol. 66, 4045–4049 (2000).
Ramos, A. et al. Effect of pyruvate kinase overproduction on glucose metabolism of Lactococcus lactis. Microbiology 150, 1103–1111 (2004).
Siddiquee, K. A. Z., Arauzo-Bravo, M. & Shimizu, K. Metabolic flux analysis of pykF gene knockout Escherichia coli based on 13C-labeling experiments together with measurements of enzyme activities and intracellular metabolite concentrations. Appl. Microbiol. Biot. 63, 407–417 (2004).
Branny, P., De La Torre, F. & Garel, J. R. The genes for phosphofructokinase and pyruvate kinase of Lactobacillus delbrueckii subsp. bulgaricus constitute an operon. J. Bacteriol. 178, 4727–4730 (1996).
Noyes, M. B. et al. A systematic characterization of factors that regulate Drosophila segmentation via a bacterial one-hybrid system. Nucleic Acids Res. 36, 2547–2560 (2008).
Fujita, Y., Miwa, Y., Galinier, A. & Deutscher, J. Specific recognition of the Bacillus subtilis gnt cis-acting catabolite-responsive element by a protein complex formed between CcpA and seryl-phosphorylated HPr. Mol. Microbiol. 17, 953–960 (1995).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 6, 613–624 (2008).
Rechinger, K. B., Siegumfeldt, H., Svendsen, I. & Jakobsen, M. “Early” protein synthesis of Lactobacillus delbrueckii ssp. bulgaricus in milk revealed by [35S] methionine labeling and two-dimensional gel electrophoresis. Electrophoresis 21, 2660–2669 (2000).
Titgemeyer, F. & Hillen, W. Global control of sugar metabolism: a gram-positive solution. Antonie Van Leeuwenhoek 82, 59–71 (2002).
Schick, J., Weber, B., Klein, J. R. & Henrich, B. PepR1, a CcpA-like transcription regulator of Lactobacillus delbrueckii subsp. lactis. Microbiology 145, 3147–3154 (1999).
Morel, F., Lamarque, M., Bissardon, I., Atlan, D. & Galinier, A. Autoregulation of the biosynthesis of the CcpA-like protein, PepR1, in Lactobacillus delbrueckii subsp bulgaricus. J. Mol. Microbiol. Biotechnol. 3, 63–66 (2001).
Sánchez, B. et al. Proteomic analysis of global changes in protein expression during bile salt exposure of Bifidobacterium longum NCIMB 8809. J. Bacteriol. 187, 5799–5808 (2005).
Bøhle, L. A. et al. Identification of proteins related to the stress response in Enterococcus faecalis V583 caused by bovine bile. Proteome Sci. 8, 1–12 (2010).
Burns, P. et al. Inside the adaptation process of Lactobacillus delbrueckii subsp. lactis to bile. Int. J. Food Microbiol. 142, 132–141 (2010).
Hamon, E. et al. Investigation of biomarkers of bile tolerance in Lactobacillus casei using comparative proteomics. J. Proteome Res. 11, 109–118 (2012).
Wang, Y., Delettre, J., Guillot, A., Corrieu, G. & Béal, C. Influence of cooling temperature and duration on cold adaptation of Lactobacillus acidophilus RD758. Cryobiology 50, 294–307 (2005).
Gunnewijk, M. G. & Poolman, B. Phosphorylation state of HPr determines the level of expression and the extent of phosphorylation of the lactose transport protein of Streptococcus thermophilus. J. Biol. Chem. 275, 34073–34079 (2000).
Auzat, I., Le Bras, G. & Garel, J.-R. The cooperativity and allosteric inhibition of Escherichia coli phosphofructokinase depend on the interaction between threonine-125 and ATP. Proc. Natl. Acad. Sci. USA 91, 5242–5246 (1994).
Kimmel, J. L. & Reinhart, G. D. Reevaluation of the accepted allosteric mechanism of phosphofructokinase from Bacillus stearothermophilus. Proc. Natl. Acad. Sci. USA 97, 3844–3849 (2000).
Taranto, M. P., Fernandez Murga, M. L., Lorca, G. & de Valdez, G. F. Bile salts and cholesterol induce changes in the lipid cell membrane of Lactobacillus reuteri. J. Appl. Microbiol. 95, 86–91 (2003).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 6, 613–624 (2008).
Chervaux, C., Ehrlich, S. D. & Maguin, E. Physiological study of Lactobacillus delbrueckii subsp. bulgaricus strains in a novel chemically defined medium. Appl. Environ. Microbiol. 66, 5306–5311 (2000).
Van de Guchte, M. et al. The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution. Proc. Nat. Acad. Sci. USA 103, 9274–9279 (2006).
Leong-Morgenthaler, P., Zwahlen, M. C. & Hottinger, H. Lactose metabolism in Lactobacillus bulgaricus: analysis of the primary structure and expression of the genes involved. J. Bacteriol. 173, 1951–1957 (1991).
Gunnewijk, M. G. W. et al. Hierarchical control versus autoregulation of carbohydrate utilization in bacteria. J. Mol. Microbiol. Biotechnol. 3, 401–413 (2001).
Gunnewijk, M. G. & Poolman, B. HPr (His∼P)-mediated phosphorylation differently affects counterflow and proton motive force-driven uptake via the lactose transport protein of Streptococcus thermophilus. J. Biol. Chem. 275, 34080–34085 (2000).
Fujita, Y. Carbon catabolite control of the metabolic network in Bacillus subtilis. Biosci. Biotechnol. Biochem. 73, 245–259 (2009).
Luesink, E. J., Van Herpen, R. E. M. A., Grossiord, B. P., Kuipers, O. P. & De Vos, W. M. Transcriptional activation of the glycolytic las operon and catabolite repression of the gal operon in Lactococcus lactis are mediated by the catabolite control protein CcpA. Mol. Microbiol. 30, 789–798 (1998).
Zotta, T. et al. Inactivation of ccpA and aeration affect growth, metabolite production and stress tolerance in Lactobacillus plantarum WCFS1. Int. J. Food Microbiol. 155, 51–59 (2012).
Asanuma, N., Kanada, K. & Hino, T. Molecular properties and transcriptional control of the phosphofructokinase and pyruvate kinase genes in a ruminal bacterium, Streptococcus bovis. Anaerobe 14, 237–241 (2008).
Viana, R., Pérez-Martínez, G., Deutscher, J. & Monedero, V. The glycolytic genes pfk and pyk from Lactobacillus casei are induced by sugars transported by the phosphoenolpyruvate:sugar phosphotransferase system and repressed by CcpA. Arch. Microbiol. 183, 385–393 (2005).
Rodionov, D. A. Comparative genomic reconstruction of transcriptional regulatory networks in bacteria. Chem. Rev. 107, 3467–3497 (2007).
de Ruyter, P. G., Kuipers, O. P. & de Vos, W. M. Controlled gene expression systems for Lactococcus lactis with the food-grade inducer nisin. Appl. Environ. Microbiol. 62, 3662–3667 (1996).
Yu, H. & Fang, H. Thermophilic acidification of dairy wastewater. Appl. Microbiol. Biotechnol. 54, 439–444 (2000).
Bricknell, K., Brook, I. & Finegold, S. Optimizing methylation conditions for gas liquid chromatography assay of lactic and succinic acid in biological samples. Chromatographia 12, 22–24 (1979).
Wilson, D., Charoensawan, V., Kummerfeld, S. K. & Teichmann, S. A. DBD–taxonomically broad transcription factor predictions: new content and functionality. Nucleic Acids Res. 36, D88–D92 (2008).
Novichkov, P. S. et al. RegPrecise: a database of curated genomic inferences of transcriptional regulatory interactions in prokaryotes. Nucleic Acids Res. 38, D111–D118 (2009).
Reese, M. G. Application of a time-delay neural network to promoter annotation in the Drosophila melanogaster genome. Comp. Chem. 26, 51–56 (2001).
Solovyev, V. & Salamov, A. Automatic annotation of microbial genomes and metagenomic sequences. Metagenomics and its applications in agriculture, biomedicine and environmental studies, 61–78 (2011).
Meng, X. & Wolfe, S. A. Identifying DNA sequences recognized by a transcription factor using a bacterial one-hybrid system. Nat. Protoc. 1, 30–45 (2006).
Schmittgen, T.D. & Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
Acknowledgements
This work was supported by the Program for New Century Excellent Talents in University [No. 2015QC086], the National Natural Science Foundation of China [No. 31171740], the Chinese Universities Scientific Fund [No. 2014JD011]. We thank Professor Willem M. de Vos (Wageningen University) for the gift of Lactococcus lactis NZ9000 and plasmid pNZ8148.
Author information
Authors and Affiliations
Contributions
Z.Z., Y.L. and Y.H. conceived of the experiment and participated in its design. Z.Z. carried out the heterologous expression, GC analysis, 5′-RACE and B1H. G.W. carried out the Western bloting. H.A. and Z.Z. were responsible for the L. bulgaricus transcription factor library. Z.Z. and Y.H. drafted the manuscript together. All authors read, commented and approved of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhai, Z., An, H., Wang, G. et al. Functional role of pyruvate kinase from Lactobacillus bulgaricus in acid tolerance and identification of its transcription factor by bacterial one-hybrid. Sci Rep 5, 17024 (2015). https://doi.org/10.1038/srep17024
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep17024
This article is cited by
-
Harnessing valorization potential of whey permeate for D-lactic acid production using lactic acid bacteria
Biomass Conversion and Biorefinery (2023)
-
Synthesis, characterization, antibacterial evaluation, 2D-QSAR modeling and molecular docking studies for benzocaine derivatives
Molecular Diversity (2021)
-
Whole-genome sequencing and genomic-based acid tolerance mechanisms of Lactobacillus delbrueckii subsp. bulgaricus LJJ
Applied Microbiology and Biotechnology (2020)
-
Engineered Zymomonas mobilis tolerant to acetic acid and low pH via multiplex atmospheric and room temperature plasma mutagenesis
Biotechnology for Biofuels (2019)
-
Improved acid-stress tolerance of Lactococcus lactis NZ9000 and Escherichia coli BL21 by overexpression of the anti-acid component recT
Journal of Industrial Microbiology and Biotechnology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.