Abstract
Fatty acid degradation protein D32 (FadD32), an enzyme required for mycolic acid biosynthesis and essential for mycobacterial growth, has recently been identified as a valid and promising target for anti-tuberculosis drug development. Here we report the crystal structures of Mycobacterium smegmatis FadD32 in the apo and ATP-bound states at 2.4 Å and 2.25 Å resolution, respectively. FadD32 consists of two globular domains connected by a flexible linker. ATP binds in a cleft at the interface between the N- and C-terminal domains and its binding induces significant local conformational changes in FadD32. The binding sites of meromycolic acid and phosphopantetheine are identified by structural comparison with other members of the adenylating enzyme superfamily. These results will improve our understanding of the catalytic mechanism of FadD32 and help in the design of inhibitors of this essential enzyme.
Similar content being viewed by others
Introduction
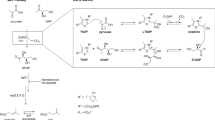
Tuberculosis (TB) is an infectious disease caused by the bacillus Mycobacterium tuberculosis. This ancient disease remains a major global health problem; one third of the world’s population is estimated to be latently infected with M. tuberculosis and approximately 9 million people developed TB and 1.5 million died from this disease in 20131. The continuing emergence of multidrug-resistant, extensively drug-resistant and totally drug-resistant M. tuberculosis severely threatens global TB control2,3 and development of new anti-TB drugs is therefore urgently required4. Targeting cell wall biogenesis in M. tuberculosis for drug development has proved to be very promising5. The mycobacterial cell wall is comprised of three covalently linked macromolecules: peptidoglycan, arabinogalactan and mycolic acids6. Recently, several proteins related to mycobacterial cell wall biogenesis have been validated as new drug targets, including the L,D-transpeptidase LdtMt2 involved in peptidoglycan biosynthesis7, the decaprenylphosphoryl-β-D-ribose 2′-epimerase DprE1 involved in arabinogalactan biosynthesis8 and fatty acid degradation protein D32 (FadD32), polyketide synthase PKS13 and trehalose monomycolate transporter Mmpl3, all of which are involved in mycolic acid biosynthesis9,10,11.
FadD32 is one of the 34 FadD proteins of M. tuberculosis annotated as putative fatty acyl-CoA synthetases12 and its role in mycolic acid biosynthesis is to activate C48–C64 meromycolic acid for condensation by PKS1313. Gene knockdown experiments have demonstrated that FadD32 inhibition can severely compromise the growth of M. tuberculosis both in vitro and inside macrophages14. FadD32 belongs to the adenylating enzyme superfamily that includes acyl-CoA synthetases, the adenylation domains of nonribosomal peptide synthetases and firefly luciferases. These enzymes consist of two domains and catalyze two half-reactions: an adenylate-forming reaction which leads to formation of acyl-AMP from a carboxylate substrate and ATP and a thioester-forming reaction which leads to formation of acyl-CoA from acyl-AMP and CoA or formation of acyl-ACP from acyl-AMP and ACP15. Although FadD32 was initially found to catalyze only the first half-reaction and was named fatty acyl-AMP ligase (FAAL)12, later evidence demonstrated that it is a bifunctional enzyme: during mycolic acid biosynthesis FadD32 catalyzes both the formation of meromycoloyl-AMP from meromycolic acid and ATP and the subsequent acyl chain transfer from meromycoloyl-AMP to the phosphopantetheinyl arm of the N-terminal ACP domain of PKS1316,17. Surprisingly, structural and biochemical studies on M. tuberculosis FAAL28 (or FadD28) suggest that FAALs (including FadD32) cannot catalyze the second half-reaction because of an insertion motif that hinders rotation of their C-terminal domains18. In addition, the FAAL28 structure only contains an N-terminal domain and so the catalytic mechanism of FAALs remains elusive. As FadD32 is a promising drug target, two groups have developed high-throughput screening methods to identify its inhibitors19,20,21. One set of inhibitors are 4,6-diaryl-5,7-dimethyl coumarin derivatives, which have activity comparable with that of isoniazid (a first-line anti-TB drug) in animal models of TB. Here, we have determined the structure of FadD32 in the apo and ATP-bound states in order to provide insights into FadD32 substrate recognition and catalysis and thus assist in the design of new inhibitors of FadD32.
Results and Discussion
Overall structure of FadD32
Full-length FadD32 from M. tuberculosis (MtFadD32) has been reported to be recalcitrant to crystallization22. As our own initial studies indicated that MtFad32 tended to aggregate, we decided to crystallize FadD32 from M. smegmatis (MsFadD32), which has 74% sequence identity to MtFadD32 (Supplementary Fig. 1). We expressed full-length MsFadD32 and its N-terminal domain in Escherichia coli and purified the protein to homogeneity (Supplementary Fig. 2). To detect the interaction between FadD32 and ATP, we performed isothermal titration calorimetry (ITC) experiments. ITC results showed that ATP bound to full-length MsFadD32 with a Kd value of approximately 36 μM, while AMP (control) did not bind to the protein (Supplementary Fig. 3a,b); Moreover, the N-terminal domain of MsFadD32 (N-MsFadD32) alone could not bind ATP (Supplementary Fig. 3c). Although N-MsFadD32 could be easily crystallized, we could only obtain crystals of full-length MsFadD32 by co-crystallizing the SUMO fusion protein and ATP. We determined the structures of apo N-MsFadD32 and ATP-bound MsFadD32 at 2.4 Å and 2.25 Å resolution by molecular replacement. We observed one molecule of MsFadD32 and one molecule of SUMO in the asymmetric unit of the crystal structure of ATP-bound MsFadD32, with a critical crystal contact mediated by SUMO (Supplementary Fig. 4). Data collection and refinement statistics are shown in Table 1.
The overall structure of MsFadD32 consists of two distinct domains, a large N-terminal domain comprising residues 1–483 and a smaller C-terminal domain comprising residues 484–630. The N-terminal domain can be further divided into three subdomains (A, B and C) and an insertion motif that is located between subdomains B and C. Subdomains A and B each have a central β-sheet that is flanked by α-helices on both sides. The two subdomains juxtapose to form a five-layered αβαβα structure. Subdomain C contains a distorted six-stranded β-barrel and a small α-helix. The C-terminal domain is composed of a three-stranded β-sheet surrounded by four α-helices and an additional small two-stranded β-sheet (Fig. 1a). Consistent with previous observations in E. coli FAAL (EcFAAL)23, the insertion motif interacts with both N- and C-terminal domains in MsFadD32. The interactions between the insertion motif and N-terminal subdomain C contain extensive hydrophobic interactions and three hydrogen bonds (Fig. 1b). These hydrophobic interactions were reported to be present in all FAALs and play an important role in anchoring the insertion motif to the N-terminal domain24. The insertion motif interacts with the C-terminal domain through two charge-stabilized hydrogen bonds between Arg368 and Asp499, hydrophobic interactions between Leu371 and Tyr496 and a hydrogen bond between the main-chain carbonyl oxygen of Asn372 and the side-chain nitrogen of Asn494 (Fig. 1c). These interactions likely exist in all FadD32 proteins of the Corynebacterineae as the residues involved are highly conserved (Supplementary Fig. 1).
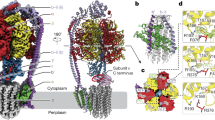
Overall structure of ATP-bound MsFadD32.
(A) Ribbon representation of the MsFadD32 structure. The three subdomains of the N-terminal domain are colored green, wheat and cyan, respectively. The insertion motif and C-terminal domain are colored orange and magenta, respectively. ATP is shown as yellow sticks. (B) The interactions between the insertion motif and N-terminal subdomain C. (C) The interactions between the insertion motif and C-terminal domain. (D) The ATP-bound MsFadD32 adopts an atypical adenylate-forming conformation. In (C–D), the residues involved are shown as sticks and the hydrogen bonds are shown as dashed lines.
Structural comparisons with other adenylating enzymes reveal that ATP-bound MsFadD32 exists in the adenylate-forming conformation. However, the invariant Lys601 in motif A10 (Supplementary Fig. 1) located in the C-terminal domain does not point into the active site and interact with ATP but forms a charge-stabilized hydrogen bond with Glu315 located in the N-terminal subdomain B (Fig. 1d). We postulate that upon binding of its other substrate, meromycolic acid, Lys601 will be orientated into the active site and interact with both ATP and meromycolate, thereby forming the true adenylate-forming conformation observed in the phenylalanine-activating subunit of gramicidin synthetase 1 (PheA)25 and other adenylating enzymes15.
ATP-binding site
In the MsFadD32-ATP complex, ATP sits in a cleft at the interface between the N- and C-terminal domains (Fig. 2a) and the residues involved in ATP binding are all from the large N-terminal domain. The adenine moiety of ATP is sandwiched between the side chains of Tyr343 and Ile480 on one side and the main-chain atoms of Ser314–Pro316 on the other. The adenine ring stacks against Tyr343 and is stabilized by a hydrogen bond between its N6 amino group and the main-chain carbonyl oxygen of Ser342. The ribose moiety of ATP is anchored by two hydrogen bonds between its two hydroxyls and the carboxylate of Asp469, an invariant residue among all adenylating enzymes15. Mutation of the equivalent residue in FadD13 (D382A) has been shown to significantly affect enzyme activity26. The three phosphate groups of ATP are held in place mainly by the P-loop27 residues (Thr187–Arg192 of motif A3 (Supplementary Fig. 1)). In particular, the guanidinyl group of Arg192, a residue conserved in all FadD32 proteins in the Corynebacterineae (Supplementary Fig. 1), forms bidentate hydrogen bonds with the γ-phosphate. In addition, the main-chain amide nitrogen of Ala346 forms a hydrogen bond with the α-phosphate and the guanidinyl group of Arg483 located in the interdomain hinge region forms bidentate hydrogen bonds with the β-phosphate. The Mg2+ ion, which bridges the β- and γ-phosphates, is coordinated by two oxygen atoms from the β- and γ-phosphates and four water molecules. Two of the four water molecules are stabilized by forming hydrogen bonds with the carboxylate of the invariant Glu34715.
ATP binding and conformational changes in MsFadD32.
(A) The binding environment for ATP. Top panel: the 2Fo – Fc electron density (contoured at 1.5σ) is shown for ATP. Bottom panel: the Mg2+ ion between the β- and γ-phosphates of ATP is shown as a green sphere and the four surrounding water molecules are shown as red spheres. (B) ATP binding-induced conformational changes in MsFadD32. The fragments of apo and ATP-bound MsFadD32 involved in conformational changes are colored cyan and green, respectively.
ATP binding-induced conformational changes
Superposition of the structures of the apo and ATP-bound MsFadD32 revealed conformational changes within the N-terminal domain. Firstly, to accommodate the adenine moiety of ATP, the side-chain hydroxyl of Ser314, which forms a hydrogen bond with the main-chain carbonyl oxygen of Gly344 in the apo state, rotates 180°. Secondly, the side-chain carboxylate of Asp232, which forms a hydrogen bond with the side-chain hydroxyl of Thr187 in the apo state, rotates approximately 90° to make room for the α-phosphate of ATP. Thirdly, the P-loop moves a relatively large distance (5–7 Å) to stabilize the three phosphate groups of ATP. Of note, the conformational change in the P-loop is accompanied by the extension of a neighboring α-helix where Arg118, which in the apo state forms hydrogen bonds with Thr191 and Arg192, forms hydrogen bonds with Thr187 in the ATP-bound state (Fig. 2b).
Meromycolate-binding site
There is a hydrophobic tunnel in the ATP-bound MsFadD32 structure at the opposite side of the phosphate tail of ATP. We found that, upon superposition of the structures of ATP-bound MsFadD32 and dodecanoyl-AMP-bound EcFAAL, the AMP components of ATP and dodecanoyl-AMP nearly overlap and the C12 tail of dodecanoyl-AMP points into the hydrophobic tunnel of MsFadD32 (Fig. 3a). However, unlike the huge hydrophobic channel present in the human Ag-presenting molecule CD1b28, the hydrophobic tunnel in MsFadD32 is clearly too small to accommodate the very long (C48–C64) meromycolic acid (Fig. 3b). We suggest that the hydrocarbon tail of meromycolate may protrude from the bottom of the hydrophobic tunnel and enter into another hydrophobic tunnel present in a partner protein of FadD32, such as PKS13. The direct interaction between FadD32 and PKS13 has been implicated in previous biochemical studies16. Consistent with this hypothesis, we found that MsFadD32 and MsPKS13 coeluted from a gel filtration column (Supplementary Fig. 5).
Phosphopantetheine-binding site
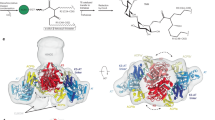
According to previous biochemical and structural studies on other adenylating enzymes15, the phosphopantetheine-binding tunnel of MsFadD32 is expected to be present at the interface between the N- and C-terminal domains and to be created when the C-terminal domain rotates approximately 140° to form the thioester-forming conformation. Of note, the glycine residue in the A8 motif that interacts with the β-alanine moiety of the pantetheine in other adenylating enzymes15, i.e. Gly492, is conserved in FadD32 (Supplementary Fig. 1). In the current adenylate-forming conformation, one half of the phosphopantetheine-binding tunnel in MsFadD32 can be identified by superposing the structures of ATP-bound MsFadD32 and the adenylate analog- and phosphopantetheine-bound PA1221 protein29. Upon superposition, the AMP components of ATP and the adenylate analog nearly overlap and the pantetheine-binding tunnel of PA1221 corresponds well with a tunnel located at the interdomain interface of MsFadD32. Phe284 is located at the entrance of the putative pantetheine-binding tunnel of MsFadD32 (Fig. 4). A previous study has shown that mutation of the equivalent residue of MtFadD32 (F291L) results in resistance to 4,6-diaryl-5,7-dimethyl coumarin derivative inhibitors in M. tuberculosis19. We therefore suggest that these potential drugs inhibit FadD32 activity by binding to the entrance of the phosphopantetheine-binding tunnel, thus preventing pantetheine binding, consistent with a previous finding that 4,6-diaryl-5,7-dimethyl coumarin derivatives inhibit acyl chain transfer from acyl-AMP to the phosphopantetheinyl arm of PKS13 or the mycobacterial acyl carrier protein (AcpM)19.
Phosphopantetheine-binding site.
(A) Structural superposition of ATP-bound MsFadD32 and adenylate analog- and phosphopantetheine-bound PA1221 protein. The N- and C-terminal domains of MsFadD32 are colored green and wheat, respectively. The N- and C-terminal subdomains and the peptidyl carrier protein domain of PA1221 are colored pale cyan, white and pink, respectively. The covalently-connected adenylate analog and phosphopantetheine are shown as magenta sticks. Phe284 of MsFadD32 is shown as a red stick. (B) Surface representation of the two domains of MsFadD32 showing the phosphopantetheine-binding tunnel.
In conclusion, we have presented the first crystal structures of FadD32, an enzyme essential for mycolic acid biosynthesis in mycobacteria and identified the binding sites of its substrates, including ATP, meromycolic acid and phosphopantetheine. The full-length structure of FadD32 shows that although the insertion loop has extensive interactions with the N-terminal domain, its interaction with the C-terminal domain is not strong. We therefore suggest that upon PKS13 binding, FadD32 may undergo a conformational change, leading to weakening of the interactions between the insertion loop and the C-terminal domain and rotation of the C-terminal domain to the thioester-forming conformation. Further mutational studies on the residues mediating the interactions between the insertion loop and the C-terminal domain will be required to study the catalytic mechanism of FadD32. In addition, the structures shed light on the mechanism of inhibition of FadD32 by 4,6-diaryl-5,7-dimethyl coumarin derivatives and lay the foundation for designing new inhibitors of this essential enzyme. We suggest that a small molecule which could bind to the interface between the insertion loop and the C-terminal domain and enhance their interactions would make a good inhibitor of FadD32 activity.
Methods
Protein expression and purification
The pMCSG7-N-MsFadD32 vector30 containing the N-terminal domain (residues 1–484) of M. smegmatis FadD32, pET28-SMT3-MsFadD32 vector31 containing SUMO-fused full-length M. smegmatis FadD32 and pMCSG7-MsPKS13ΔC vector containing residues 1–1042 of M. smegmatis PKS13 were constructed and transformed into E. coli BL21 (DE3) for overexpression. After IPTG induction at 289 K for 18 h, the cells were collected and resuspended in lysis buffer containing 20 mM Tris, pH 8.0, 500 mM NaCl and 20 mM imidazole. After sonication, the cell lysate was centrifuged at 16000 rpm, 277 K and the supernatant containing the tagged protein was applied to a nickel-affinity column pre-equilibrated with the lysis buffer. The recombinant protein was eluted with 250 mM imidazole and further purified by size-exclusion chromatography (SEC). The N-MsFadD32 protein sample was treated with TEV protease to remove the N-terminal His-tag before SEC. The resulting protein sample was >95% pure, as determined by SDS-PAGE.
Crystallization and structure determination
Crystallization trials were performed using the sitting-drop vapor-diffusion method. The protein was concentrated to approximately 10 mg ml-1 and mixed in a 1:1 ratio with reservoir solution. N-MsFadD32 crystallized in 0.1 M magnesium formate, 15% PEG 3350 at 289 K. For crystallization of SUMO-fused full-length MsFadD32 at 281 K, the protein was pre-incubated with 10 mM ATP and 20 mM MgCl2 at 277 K for 2 h and then mixed with 0.2 M sodium malonate, 20% PEG 3350. Diffraction data were collected from flash-frozen crystals at 100 K using a Rigaku MM007-HF CCD (Saturn 944HG) diffractometer and the beamline BL17U of the Shanghai Synchrotron Radiation Facility.
All data sets were processed using HKL200032. The structures of apo N-MsFadD32 and ATP-bound MsFadD32 were determined by molecular replacement using PHASER33 and search models derived from the N-terminal domain of MtFAAL28 (PDB ID: 3E53) and the C-terminal domain of EcFAAL (PDB ID: 3PBK). Models were built manually with COOT34 and refined with PHENIX35. The stereochemical quality of the refined structures was verified with PROCHECK36. Most of the residues have good geometry and no residues were present in the disallowed regions of the Ramachandran plots. Data collection and refinement statistics are presented in Table 1. All structural representations in this paper were prepared with PyMol (DeLano Scientific).
Isothermal titration calorimetry assay
The binding affinities between MsFadD32 and ligands were measured with an ITC200 micro-calorimeter (MicroCal). Full-length MsFadD32 and its N-terminal domain were prepared in a buffer containing 20 mM Tris, pH 8.0, 150 mM NaCl, 3 mM MgCl2. ATP or AMP was dissolved in the same buffer. Protein concentration in the measurement cell was 50 μM, while the ligand concentration in the titration procedure was adjusted to 600 μM. Titrations were performed at 298 K and data were fitted to a one-binding-site model using Origin 7.0 (MicroCal).
Gel filtration assay
A Superdex 200 column was used for gel filtration assays and the column was pre-equilibrated with a buffer containing 20 mM Tris, pH 8.0, 150 mM NaCl. To examine the interaction between MsFadD32 and MsPKS13, SUMO-fused full-length MsFadD32 was treated with Ulp1 protease to remove the N-terminal SUMO protein and the resultant MsFadD32 was used in subsequent experiments. MsFadD32 pre-incubated with MsPKS13ΔC and individual MsFadD32 and MsPKS13ΔC proteins, were injected into the column and eluted at a flow rate of 0.4 ml min-1. Peak fractions were collected and analyzed by SDS-PAGE.
Additional Information
Accession codes: Coordinates and structure factors for apo N-MsFadD32 and ATP-bound MsFadD32 have been deposited in the Protein Data Bank under accession codes 5D6N and 5D6J, respectively.
How to cite this article: Li, W. et al. Crystal structure of FadD32, an enzyme essential for mycolic acid biosynthesis in mycobacteria. Sci. Rep. 5, 15493; doi: 10.1038/srep15493 (2015).
References
World Health Organization. Global Tuberculosis Report 2013 (WHO, 2013).
Gandhi, N. R. et al. Multidrug-resistant and extensively drug-resistant tuberculosis: a threat to global control of tuberculosis. Lancet 375, 1830–1843 (2010).
Dheda, K. et al. Global control of tuberculosis: from extensively drug-resistant to untreatable tuberculosis. Lancet Respir. Med. 2, 321–338 (2014).
Zumla, A., Nahid, P. & Cole, S. T. Advances in the development of new tuberculosis drugs and treatment regimens. Nat. Rev. Drug Discov. 12, 388–404 (2013).
Jackson, M., McNeil, M. R. & Brennan, P. J. Progress in targeting cell envelope biogenesis in Mycobacterium tuberculosis. Future Microbiol. 8, 855–875 (2013).
Brennan, P. J. Structure, function and biogenesis of the cell wall of Mycobacterium tuberculosis. Tuberculosis 83, 91–97 (2003).
Gupta, R. et al. The Mycobacterium tuberculosis protein LdtMt2 is a nonclassical transpeptidase required for virulence and resistance to amoxicillin. Nat. Med. 16, 466–469 (2010).
Makarov, V. et al. Benzothiazinones kill Mycobacterium tuberculosis by blocking arabinan synthesis. Science 324, 801–804 (2009).
Grzegorzewicz, A. E. et al. Inhibition of mycolic acid transport across the Mycobacterium tuberculosis plasma membrane. Nat Chem Biol 8, 334–341 (2012).
Stanley, S. A. et al. Diarylcoumarins inhibit mycolic acid biosynthesis and kill Mycobacterium tuberculosis by targeting FadD32. Proc Natl Acad Sci USA 110, 11565–11570 (2013).
Wilson, R. et al. Antituberculosis thiophenes define a requirement for Pks13 in mycolic acid biosynthesis. Nat Chem Biol 9, 499–506 (2013).
Trivedi, O. A. et al. Enzymic activation and transfer of fatty acids as acyl-adenylates in mycobacteria. Nature 428, 441–445 (2004).
Portevin, D. et al. The acyl-AMP ligase FadD32 and AccD4-containing acyl-CoA carboxylase are required for the synthesis of mycolic acids and essential for mycobacterial growth: identification of the carboxylation product and determination of the acyl-CoA carboxylase components. J. Biol. Chem. 280, 8862–8874 (2005).
Carroll, P., Faray-Kele, M. C. & Parish, T. Identifying vulnerable pathways in Mycobacterium tuberculosis by using a knockdown approach. Appl. Environ. Microbiol. 77, 5040–5043 (2011).
Gulick, A. M. Conformational dynamics in the acyl-CoA synthetases, adenylation domains of non-ribosomal peptide synthetases and firefly luciferase. ACS Chem. Biol. 4, 811–827 (2009).
Gavalda, S. et al. The Pks13/FadD32 crosstalk for the biosynthesis of mycolic acids in Mycobacterium tuberculosis. J. Biol. Chem. 284, 19255–19264 (2009).
Leger, M. et al. The dual function of the Mycobacterium tuberculosis FadD32 required for mycolic acid biosynthesis. Chem. Biol. 16, 510–519 (2009).
Arora, P. et al. Mechanistic and functional insights into fatty acid activation in Mycobacterium tuberculosis. Nat. Chem. Biol. 5, 166–173 (2009).
Stanley, S. A. et al. Diarylcoumarins inhibit mycolic acid biosynthesis and kill Mycobacterium tuberculosis by targeting FadD32. Proc. Natl. Acad. Sci. USA 110, 11565–11570 (2013).
Galandrin, S. et al. Assay development for identifying inhibitors of the mycobacterial FadD32 activity. J Biomol Screen 18, 576–587 (2013).
Kawate, T. et al. Synthesis and structure-activity relationships of phenyl-substituted coumarins with anti-tubercular activity that target FadD32. Bioorg Med Chem Lett 23, 6052–6059 (2013).
Goyal, A. et al. Crystallization and preliminary X-ray crystallographic studies of the N-terminal domain of FadD28, a fatty-acyl AMP ligase from Mycobacterium tuberculosis. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 62, 350–352 (2006).
Zhang, Z. N. et al. Structural and functional studies of fatty acyl adenylate ligases from E. coil and L. pneumophila. J. Mol. Biol. 406, 313–324 (2011).
Goyal, A., Verma, P., Anandhakrishnan, M., Gokhale, R. S. & Sankaranarayanan, R. Molecular basis of the functional divergence of fatty acyl-AMP ligase biosynthetic enzymes of Mycobacterium tuberculosis. J. Mol. Biol. 416, 221–238 (2012).
Conti, E., Stachelhaus, T., Marahiel, M. A. & Brick, P. Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S. EMBO J. 16, 4174–4183 (1997).
Khare, G. et al. Dissecting the role of critical residues and substrate preference of a fatty acyl-CoA synthetase (FadD13) of Mycobacterium tuberculosis. Plos One 4, e8387 (2009).
May, J. J., Kessler, N., Marahiel, M. A. & Stubbs, M. T. Crystal structure of DhbE, an archetype for aryl acid activating domains of modular nonribosomal peptide synthetases. Proc. Natl. Acad. Sci. USA 99, 12120–12125 (2002).
Batuwangala, T. et al. The crystal structure of human CD1b with a bound bacterial glycolipid. J. Immunol. 172, 2382–2388 (2004).
Mitchell, C. A., Shi, C., Aldrich, C. C. & Gulick, A. M. Structure of PA1221, a nonribosomal peptide synthetase containing adenylation and peptidyl carrier protein domains. Biochemistry 51, 3252–3263 (2012).
Stols, L. et al. A new vector for high-throughput, ligation-independent cloning encoding a tobacco etch virus protease cleavage site. Protein Expr. Purif. 25, 8–15 (2002).
Bernier-Villamor, V., Sampson, D. A., Matunis, M. J. & Lima, C. D. Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108, 345–356 (2002).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Mccoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Laskowski, R. A., Macarthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
We would like to thank Dr. M. Fan for his help with structure determination and Yi Han, Yonglin Hu, Defeng Li and the staff at beamline 17U of Shanghai Synchrotron Radiation Facility for their help in data collection. We thank Yuanyuan Chen at the Core Facility for Protein Research, Institute of Biophysics, for assistance with ITC experiments. This work was supported by grants 2011CB910300, 2011CB911103 and 2013CB911500 from the National Basic Research Program of China and grant KSCX2-EW-J-3 from the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
W.L. and S.G. expressed, purified and crystallized FadD32. W.L. performed data collection and structure determination. W.L., S.G. and J.F. wrote and revised the manuscript. L.B. supervised the project and revised the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, W., Gu, S., Fleming, J. et al. Crystal structure of FadD32, an enzyme essential for mycolic acid biosynthesis in mycobacteria. Sci Rep 5, 15493 (2015). https://doi.org/10.1038/srep15493
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep15493
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.