Abstract
The Malay people are an important ethnic composition in Southeast Asia, but their genetic make-up and population structure remain poorly studied. Here we conducted a genome-wide study of four geographical Malay populations: Peninsular Malaysian Malay (PMM), Singaporean Malay (SGM), Indonesian Malay (IDM) and Sri Lankan Malay (SLM). All the four Malay populations showed substantial admixture with multiple ancestries. We identified four major ancestral components in Malay populations: Austronesian (17%–62%), Proto-Malay (15%–31%), East Asian (4%–16%) and South Asian (3%–34%). Approximately 34% of the genetic makeup of SLM is of South Asian ancestry, resulting in its distinct genetic pattern compared with the other three Malay populations. Besides, substantial differentiation was observed between the Malay populations from the north and the south and between those from the west and the east. In summary, this study revealed that the genetic identity of the Malays comprises a mixed entity of multiple ancestries represented by Austronesian, Proto-Malay, East Asian and South Asian, with most of the admixture events estimated to have occurred 175 to 1,500 years ago, which in turn suggests that geographical isolation and independent admixture have significantly shaped the genetic architectures and the diversity of the Malay populations.
Similar content being viewed by others
Introduction
The Malay people, who generally inhabit the Malay Archipelago, particularly the Peninsular Malaysia, is a group of genetically diverse populations1. As a major ethnic group settling in such a strategic hub in Southeast Asia, the Malay population may provide evidence for the complex historical background and demographic information in this region, which has never been fully understood to date.
In Malaysia, the Malays consist of several sub-groups, e.g., Melayu Minang, Melayu Jawa, Melayu Bugis and Melayu Kelantan, based on their respective geographical origins. Malay sub-groups have various historical origins and genetic affinities1,2. Both ancient and recent human activities have introduced gene flows from other populations into the Malays3,4,5. The indigenous groups (Orang Asli) in Malaysia, including Proto-Malay, Senoi and Negrito, interacted with the Malays because their habitats were situated adjacent to one another. The Malays are genetically related to the Orang Asli despite differences in their physical features5. For example, Negritos have short stature and very dark skin, which are more similar to the phenotypes of African Pygmies than to those of other Southeast Asian populations6,7. In addition, populations from other regions of Asia, even from other continents, have cultural and genetic influences on the Malays to various extents. In particular, Chinese, Indians, Arabians and Europeans have substantially influenced the region since ancient times and have had a more considerable impact in recent centuries due to the colonization and globalization of the entire human society, thus leading to the mosaic genomic pattern in the Malays2,5,8. Besides Malaysia, the Malays are distributed in the surrounding islands, e.g. Peninsular Indo-China, Singapore, Java and Sri Lanka9. Malay populations share a common origin with other Austronesian populations9. During the 1400s and 1500s, the Malacca Sultanate was established and subsequently dominated the entire Malay Archipelago and the Malay language was established as the official language in that region, replacing Sanskrit. The Malays, especially those in the western and southern Malaysia, have had frequent interactions with other Austronesian populations through trading or other social activities10. These early contacts did not result in a large-scale of population admixture, but intermarriage did exist. Gradually, the Malays mixed with the local residents in the surrounding islands and have developed into various Malay ethnic groups2. These Malay ethnic groups affect not only the demography but also the culture, including religions and languages, outside Malaysia. However, the Malay people have been largely under-represented in studies on human genetic diversity and are not on the population list of large-scale international projects, such as the the International Haplotype Map Project11 (HapMap, http://www.hapmap.org), the Human Genome Diversity Project12 (HGDP, http://www.hagsc.org/hgdp) and the 1000 Genomes Project13 (http://www.1000genomes.org). Some recent studies on Asian populations have included a few Malay samples from Malaysia, Indonesia and Singapore8,14,15, but other Malay populations residing outside of Southeast Asia such as those living in Sri Lanka, have not been well studied.
The present study aimed to investigate the genetic relationship and population structure of the Malays from both Southeast Asia and South Asia to extend our knowledge about population history and the genetic landscape of human populations in Southeast Asia. Analyses were conducted by using 288,660 single nucleotide polymorphisms (SNPs) shared by 133 Malay individuals from Peninsular Malaysia, Singapore and Sri Lanka (abbreviated to PMM, SGM and SLM, respectively), as well as approximately 50,000 SNPs shared by 12 Indonesian Malays (IDM) located in Sumatra Island, 30 SGM and 38 PMM (20 Minangkabau, or locally known as Melayu Minang, a sub-tribe of Malay; 18 Kelantan Malay, or locally known as Melayu Kelantan) from the HUGO PanAsia SNP Consortium (PASNP)8. We also integrated 3,170 individuals from 90 worldwide populations with our data to obtain a higher resolution in deciphering the genetic diversity of the Malay populations. Genetic structure investigation was based on model-free methods that provided direct information on population relationships and the genetic makeup. Furthermore, we inferred the population admixture history of each Malay population to illustrate the population genetic structure patterns.
Results
Population genetic relationships
As shown in the population phylogenetic tree (Fig. 1, Supplementary Figure S1), populations from closer geographical areas showed greater genetic similarities and were thus clustered. Among the four Malay groups, SGM, PMM and IDM were closely clustered and shared a distinct clade with other Southeast Asians, whereas IDM showed a closer relationship to populations from Sumatra and Java than to other Southeast Asians. On the other hand, SLM was lying on a cline of Southeast Asian and South Asian. Based on the genetic distance measured by FST, SLM showed a closer relationship with South Asian populations (FST = 0.004–0.024) than with Southeast Asians (FST = 0.010–0.057). In addition, the genetic difference between SLM and the other Sri Lankan ethnic groups (FST = 0.007–0.012) was larger than that between any other Sri Lankan ethnic groups (FST = 0.001–0.003).
Phylogenetic tree showing genetic relatedness of four Malay populations.
A phylogenetic tree was constructed by using the neighbor-joining method, taking YRI as the outgroup. The pair-wise population distance was measured by global FST with 1,000 bootstrapping repeats. Bootstrap values are noted on the branches. Population IDs are shown in Supplementary Table S4. Geographical groups are indicated by colors. The four Malay populations are highlighted in bold font with red asterisks.
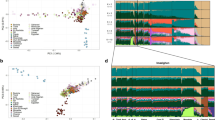
Similar population relationships were observed by using principal component analysis (PCA). At the global scale, the four Malay groups were considerably distinct from Africans and similar to other non-Africans (Fig. 2a). South Asians (Indians and Sri Lankans) constituted a bridge between the Europeans and other Asian populations (Fig. 2). SLM, as well as the Uyghurs that have been proven to be an admixed population16, were situated between other Sri Lankan populations and Southeast Asians (Fig. 2). In general, the geographical distribution of these populations was fully reflected by their genetic relationships, although exceptions existed. However, PMM showed closer affinity to the Southeast Asian populations located outside Malaysia despite that they inhabited the adjacent regions of Peninsular Malaysia, possibly due to the population isolation and local adaptation of the Orang Asli populations, which in turn resulted from their special living environment and demographic history5 (Fig. 2b). By gradually excluding the populations less closely related to the Malays (Supplementary Figure S2), we finally identified that the western Indonesians had the closest relationship to PMM, SGM and IDM, followed by Filipino, Thai, Proto-Malay and Bidayuh (a native population from southern Sarawak). Furthermore, population differentiation was also observed between Malay populations from the north (SLM and Kelantan Malay) and the south (Minangkabau, SGM and IDM) (Pearson product-moment correlation coefficient (PCC) = 0.781 between the PCA coordinates and the geographical latitude, p-value < 2.2 × 10−16) and between those from the west (SLM) and the east (PMM, SGM and IDM) (PCC = 0.9 between the PCA coordinates and the geographical longitude, p-value < 2.2 × 10−16) (Supplementary Figure S3).
Plots of the first two principal components for hierarchical analyses of worldwide populations.
(A) 2,713 individuals representing 83 populations from Africa, Europe and Asia. (B) 2,597 individuals representing 82 populations from Europe and Asia (excluding YRI). PMMK and PMMM denote Kelantan Malay and Minangkabau, respectively.
Multiple population admixture in Malays
An unsupervised ADMIXTURE analysis, without prior information of individual ancestry, was performed to investigate patterns of admixture in Asia. Nine ancestral clades were assumed to reflect the major genetic components in Asia (Fig. 3, Supplementary Figure S4). Collectively, South Asians and East Asians had their distinct genetic patterns. European, Central Asian and South Asian components predominated the South Asian genomes, whereas a large proportion of the East Asian component and a small proportion of the Southeast Asian component comprised the East Asian genomes. However, the Southeast Asian populations had more complex and diverse genetic makeup than the South Asians and the East Asians. Malay populations were admixed with multiple ancestries with no representative component identified in them, but most Malays had a genetic pattern distinct from other Southeast Asian populations.
Locations and genetic makeup of the Malays and other populations.
The averaged genetic makeup across individuals of each population are indicated by the bars. Each color represents an independent cluster at K = 9. Southeast Asian 1 and Southeast Asian 2 represent the aboriginal Southeast Asian component and Austronesian component, respectively. All the Malay populations are arranged in the dashed box. Population IDs are shown in Supplementary Table S4. The map is generated using the packages of R v2.11.1, including mapdata v2.2-3, mapplots v1.5 and maps 2.3-9 (http://cran.r-project.org/web/packages/).
Malay populations shared four major components, e.g., East Asian, South Asian, Austronesian and aboriginal Southeast Asian. East Asians, which contribute 4%–16% of the Malay genomes, had interactions with Malays at very recent time (100–200 years ago, assuming a single generation time of 25 years was applied throughout this study) (Table 1, Supplementary Table S1). We observed a substantial proportion of South Asian ancestry in the Malay populations as well, ranging from 3% to 34%, with the greatest proportion observed in the SLM and the most ancient admixture date in the PMM (625–1,400 years ago) and IDM (1,350–2,250 years ago). In addition, the proportions of Austronesian admixture (represented by the Taiwanese aborigines; labeled as Southeast Asian 1 in Fig. 3) and aboriginal Southeast Asian admixture (represented by Proto-Malays; labeled as Southeast Asian 2 in Fig. 3) were within the ranges of 15%–31% and 17%–62%, respectively, in the Malay populations, with a greater proportion in PMM, SGM and IDM. We observed nearly constant proportions (<5%) of components from African, European, hunter-gatherer (represented by Malaysian Negrito) and Oceanian (represented by Papuan) in most of the Southeast Asians examined in this study. However, these were disregarded in our interpretation because these could have resulted from the ascertainment bias. The time of admixture was estimated by using ALDER throughout the present study (see Materials and Methods).
While the genetic makeup of the four Malay populations shared similar ancestral components, the proportion of these components in each population substantially differed. In PMM, SGM and IDM, the Austronesian and Southeast Asian aboriginal components had larger proportions than the East Asian and South Asian components. Specifically, very little genetic difference was found between PMM and SGM, in which the two major components were of nearly equal proportions and together comprised 60%–70% of the entire genome. IDM also showed a very similar genetic pattern, but was more marginally affected by the Austronesian population (Austronesian component: 62% in IDM, 31%–45% in SGM and PPM). However, the genetic admixture pattern of SLM was unique compared to the other Malay populations, exhibiting admixture between South Asians and Malays. The proportions of different components were relatively more comparable in SLM. For instance, the proportions of Central Asian and South Asian components in SLM were intermediate between those observed in Southeast Asians and South Asians and a similar pattern was detected for the Southeast Asian component (Austronesians and Proto-Malays) and East Asian component.
To quantify the contribution of each ancestral component to the differentiation of Malay populations, we calculated the correlation between the proportions of each ancestral component and the values along PC1 of the 4 Malay populations in Supplementary Figure S2a. Supplementary Table S2 showed that all the four major components in the Malays were correlated with PC1 to different extents (p < 0.01). For example, the South Asian component was most highly correlated with the Malay differentiation (PCC = 0.927, p-value < 2.2 × 10−16), whereas the East Asian component showed a much less significant correlation (PCC = 0.268, p-value = 1.03 × 10−4). Furthermore, the European and Central Asian components, although they were not major components in the Malays, also largely contributed to the Malay differentiation (PCC = 0.807 and PCC = 0.579, respectively).
Discussion
In this study, we conducted a genome-wide analysis of four geographical Malay populations from Malaysia, Singapore, Indonesia and Sri Lanka. We have analyzed the genetic diversity of the Malay populations and revealed substantial population differentiation and gene flow among these different geographical populations.
In general, all the four Malay populations were inferred to be substantially admixed with multiple ancestries from East Asian, South Asian, Austronesian and Southeast Asian aboriginal people around 175–1,500 years ago (Fig. 3, Table 1), which could be attributable to their complex origins and frequent interactions with surrounding ethnic groups, as reported in historical records. As early as 2,000 years ago, several ancient Malay states emerged in the coastal areas of the Malay Peninsula, Sumatra, western Java and western Borneo. During the 7th–13th centuries, the Malay culture reached its golden age and influenced the entire Malay Archipelago2,17. In the 15th century, the Malacca Sultanate initiated a major revolution in Malay history by promulgating its languages, religions and traditions, which has also recently influenced the immigrants from other countries2. Since the year 1299, the advent of the Kingdom of Singapura, a small Malay kingdom, in the modern-day island nation Singapore started the inhabitation of Malays in Singapore18. Although Sri Lanka and Southeast Asian countries have had very early interactions on politics, religion and culture since the 11th century19, the earliest Malay inhabitants of Sri Lanka were brought in laborers from Indonesia and Malaysia by the Dutch and British colonizers during the 16th–17th century20,21. The present-day SLM, which comprise 0.3% of the Sri Lankan population, are the descendants of the mixture of the local Sri Lankan people with those Malays20. Meanwhile, the first millennium of the Common Era also witnessed the trade migration of South Asians into the Malay kingdoms, as well as the arrival of Chinese, Arabian and Persian merchants in Island Southeast Asia2,22. From 1500s towards the end of the 20th century, European countries, including Portugal, Netherlands and Britain, colonized the majority of Southeast and South Asian countries, bringing about mating and gene flow across continents. More recently, due to the development of transportation and the globalization of the entire human society, the Southeast and South Asian populations have thus had more frequent gene flow from each other and from their surrounding neighborhoods, e.g., East Asians. Although previous genetic studies investigated less comprehensive Malay samples covering a smaller geographical range of Malay, the results indeed concurred with our findings of extensive genetic diversity among the Malays1,8.
The present study provides the first identification of the distinct genetic makeup of SLM compared to the other three Malay populations. SLM inherited much more South Asian ancestry, whereas PMM, SGM and IDM are genetically similar to most of the other Southeast Asian populations, with minor fluctuations in the proportions of each genetic component across populations. Two factors have facilitated to shaping SLM to be a genetically intermediate population between South Asians and Southeast Asians (Figs 1 and 2). First, SLM originated from the admixture of Sri Lankan populations and the Malays. Assuming the Sri Lankan ethnic groups and Malay to be the ancestral populations, we identified 55%–61% of Sri Lankan component and 39%–45% of Malay component in SLM (Supplementary Table S3). Second, Sri Lanka is located in the strategic region bridging South Asia and Southeast Asia and thus has been an important trading center as well as a melting pot of various ethnicities.
Unlike those isolated or ancient populations in Southeast Asia (e.g., Negritos), the Malays do not have a representative genetic component. Instead, four major ancestral components constitute the Malay genomes, making the Malays distinguishable from most of the other Southeast Asian populations (Fig. 3). The Austronesian and Southeast Asian aborigines have been determined to be the predominant components. As indicated by our results, aboriginal Taiwanese (Ami and Atayal) and Proto-Malays have the largest proportion of these two components; however, we were not able to determine the origin of the components because these could have been derived from any other source populations that have yet to be identified by our analyses. The genetic pattern of the Austronesian component across Southeast Asians follows the Austronesian migration route supported in the “Out of Taiwan” hypothesis23,24,25, which influenced the eastern part of Southeast Asian more than the western part of it. However, the Proto-Malay component, which has been reported to be an ancient Southeast Asian component26, is centered in the west. Considering that the Proto-Malays are an isolated aboriginal population in this region, it is likely that the Proto-Malay genetic component in other Southeast Asian populations, especially those located outside Peninsular Malaysia, was obtained from the broadly migrated Malay population, which had very close interactions with Proto-Malays. This observation also has important implications to the human migration history in Southeast Asia. Apart from the four major components, we observed small amount of other components in the Malays. These components were not included in our interpretation because they could be resulted from ascertainment bias. In ADMIXTURE analyses, we only used SNPs shared by global populations, leading to the reduction of the number of rare SNPs in the SNP panel of each population, thus providing less population-specific information. In this case, some genomic regions could be origin-ambiguous and have been assigned to multiple continental populations. The major human migration events that occurred in Southeast Asia may have geographical limitations, which means that different populations obtained the major genetic contributions from their respective close neighbors, thus leading to a correlation between geographical coordinates (both latitude and longitude) and population differentiation. This indicates the effect of geographical independency in human genetics. Even so, we did not rule out other migration events that would have had a smaller genetic impact. These findings may provide crucial clues to the peopling in Southeast Asia.
The major limitation of the present study lies in the possible bias of the estimated admixture time and fraction. For some populations, the sample size and SNP density were not highly appropriate for use in the analyses. On one hand, most of the Southeast Asian genotypes, including IDM, from PASNP8, only harbored 50,000 SNPs and comprised less than 20 individuals. On the other hand, we have integrated data from various international projects and genotyping platforms hence the number of SNPs in the combined datasets that were then applied to different analyses is very small. In this case, although PCA, genetic component analysis and the construction of population phylogenetic tree are sufficiently robust to fluctuations in sample size or SNP density, the estimation of admixture time and fraction, which were based on the calculation of linkage disequilibrium (LD) decay, might be highly accurate. Therefore, a more comprehensive collection of larger numbers of samples and SNP markers will be necessary in the future studies. Besides, the existing methods for admixture analysis are not powerful enough to handle the various Southeast Asian ethnic groups with complex historical and genetic backgrounds. This can be explained by two factors. First, our estimation was based on the hybrid isolation (HI) model27, which might not be representative of the real cases. Second, the populations referred to as the ancestries also recently obtained genetic materials from other populations, which could bias the estimation of the impact of ancestral populations on their admixed descendants. Therefore, more sophisticated methods may better facilitate the construction of human migration history in Southeast Asia.
In summary, the present study provides the first demonstration of the genetic relationship and genetic makeup of various Malay populations and indicated a complex human population admixture history in Southeast Asia. We propose that geographical isolation and independent admixture history have significantly shaped the genetic architectures and the diversity of the Malay populations. An important direction for future work is to extend to a wider coverage of Malay groups and increase the SNP density, to allow a more detailed investigation of the population genetic diversity in Southeast Asia.
Methods
Populations and samples
The present study focused on 213 unrelated Malay individuals from 4 Malay groups that were collected from different datasets. All the 27 SLM samples, as well as 538 Sri Lankan samples from other ethnic groups (303 Tamil, of which 200 were of Indian ancestry, 35 Burger and 200 Sinhalese) were genotyped on an Illumina 2.5 M array. The other 186 Malay samples were previously reported, including 89 SGMs from the Singapore Genome Variation Project14 (SGVP, http://www.statgen.nus.edu.sg/~SGVP/), 30 SGMs, 12 IDMs and 38 PMMs from PASNP8 and 17 PMMs as reported in Deng et al. (2014)5. Notably, the PMM population in the present study consisted of two subgroups: 35 Kelantan Malays and 20 Minangkabau.
To characterize the genetic variation of the 4 Malay groups on a global scale, we also included 3,170 unrelated individuals from 90 worldwide populations reported in HapMap Phase 311, HGDP12, SGVP14, PASNP8 and our previous study5. The details of sampled populations are summarized in Supplementary Table S4.
Sample and SNP quality control
Our analyses excluded samples with missingness greater than 5%. Then, we used PLINK v1.0728 (http://pngu.mgh.harvard.edu/~purcell/plink/) to control the data quality. Stringent sample filtering was conducted within each of the examined populations and in total, 113 samples with missing rates of >5% were excluded from the subsequent analyses (Supplementary Table S4). We then generated several combined datasets for different purposes of analysis and discarded SNPs with missing rates of >5% and minor allele frequency of <5%. The summary of all datasets can be found in Supplementary Table S5.
Statistical methods
Wright’s fixation index FST as a measure of population differentiation was calculated according to Weir and Hill29, which is an unbiased estimate that considers standardized sample sizes. Two independently integrated datasets with different numbers of SNPs were used to calculate the FST. Population phylogenetic trees were constructed based on pairwise population FST metrics using a neighbor-joining method provided by PHYLIP v3.6930 (http://evolution.genetics.washington.edu/phylip.html) and were visualized by MEGA v6.0631 (http://www.megasoftware.net). The bootstrap value was based on the 1,000 repeats of FST calculation by randomly sampling SNPs. PCA was conducted by using EIGENSOFT v3.032 (http://www.hsph.harvard.edu/alkes-price/software/). Prior to analysis, 24 SNPs were removed by LD pruning with a threshold of r2 = 0.8 for pairwise SNPs and 14 individuals (4 PMM, 6 SGM, 1 IDM and 3 SLM) were filtered out with a PC1 value >2-fold standard deviation in the PCA for each independent population. Population genetic clustering was conducted by using ADMIXTURE33 (https://www.genetics.ucla.edu/software/admixture/) specifying K = 2 to K = 12. Cross validation (CV) was used to assess the number of clusters best-fitting the data (Supplementary Figure S4). To quantify the gene flow, we applied an LD-based method ALDER34 (http://groups.csail.mit.edu/cb/alder/). We used the one-reference model in ALDER, assuming only a single reference population. To circumvent the uncertainties about the exact ancestral source, we have performed the same admixture test using all the populations from each ancestral group. Because various numbers of SNPs have been identified in the populations that are used in this analysis, which ranged from ~50,000 to ~2,000,000, in the independent analysis of each donor-receptor pair, we used the SNPs shared by the pair of populations to maximize the SNP density. Similarly, when estimating the ancestral contributions of the Sri Lankan ethnic groups to Sri Lankan Malays, we examined all possible combination of the two sources (one Sri Lankan population and one Malay population). Ten individuals were randomly sampled from each population to generate a fair sample size. The supervised STRUCTURE analysis is based on the admixture model with pre-assigned ancestral individuals and two predefined clusters (K = 2). We obtained locus-specific ancestry estimations by using the linkage model implemented in STRUCTURE version 2.335, with replicate runs of 20,000 burn-in iterations and 10,000 parameter-estimating iterations. The ancestral contributions to the Sri Lankan Malays were averaged across the whole genome. The PCC between the geographical coordinates and the PCA coordinates and between the proportions of genetic components and the PCA coordinates were calculated by using R v2.11.136 (http://www.r-project.org/) based on the Pearson’s correlation model.
Additional Information
How to cite this article: Deng, L. et al. Dissecting the genetic structure and admixture of four geographical Malay populations. Sci. Rep. 5, 14375; doi: 10.1038/srep14375 (2015).
References
Hatin, W. I. et al. Population genetic structure of Peninsular Malaysia Malay sub-ethnic groups. PLoS One 6, e18312 (2011).
Cavendish, M. Malaysia, Singapore, Brunei and the Philippines. 1584 (Marshall Cavendish Corporation, 2008).
Wheatley, P. The Golden Khersonese: Studies in the Historical Geography of the Malay Peninsula before AD 1500. 388 (University of Malaya Press, 1961).
Syukri, I. Sejarah Kerajaan Melayu Patani. 260 (Universiti Kebangsaan Malaysia, 2002).
Deng, L. et al. The population genomic landscape of human genetic structure, admixture history and local adaptation in Peninsular Malaysia. Hum. Genet. 133, 1169–1185 (2014).
Migliano, A. B. et al. Evolution of the pygmy phenotype: evidence of positive selection from genome-wide scans in African, Asian and Melanesian pygmies. Hum. Biol. 85, 251–284 (2013).
Stock, J. T. The skeletal phenotype of “Negritos” from the Andaman Islands and Philippines relative to global variation among hunter-gatherers. Hum. Biol. 85, 67–94 (2013).
The HUGO Pan-Asian SNP Consortium. Mapping human genetic diversity in Asia. Science 326, 1541–1545 (2009).
Bellwood, P. Prehistory of the Indo-Malaysian Archipelago. 384 (University of Hawai’i Press, 1997).
Sarji, A. & Hamid, A. The Encyclopedia of Malaysia: The Rulers of Malaysia. (Editions Didier Millet, 2011).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Li, J. Z. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Teo, Y.-Y. et al. Singapore Genome Variation Project: a haplotype map of three Southeast Asian populations. Genome Res. 19, 2154–2162 (2009).
Wong, L.-P. et al. Deep whole-genome sequencing of 100 Southeast Asian Malays. Am. J. Hum. Genet. 92, 52–66 (2013).
Xu, S. & Jin, L. A genome-wide analysis of admixture in Uyghurs and a high-density admixture map for disease-gene discovery. Am. J. Hum. Genet. 83, 322–336 (2008).
Milner, A. The Malays. (Wiley-Blackwell, 2010).
Abshire, J. E. The History of Singapore. 179 (Greenwood publishing group, 2011).
Sirisena, W. M. Sri Lanka and South-east Asia: Political, Religious and Cultural Relations from A.D. c. 1000 to c. 1500. 186 (Sri Lanka University Press, 1978).
De Silva Jayasuriya, S. Malay contacts with Sri Lanka. IIAS Newsl. 31, 28–29 (2003).
Crawford, J. History of Indian Archipelago. 481 (The Long Now Foundation, 1820).
Andaya, L. Y. Orang Asli and the Melayu in the History of the Malay Peninsula. J. Malayan Branch R. Asiat. Soc. 75, 23–48 (2002).
Ko, A. M.-S. et al. Early Austronesians: into and out of Taiwan. Am. J. Hum. Genet. 94, 426–436 (2014).
Bellwood, P. & Eusebio, D. The batanes archaeological project and the “Out of Taiwan” hypothesis for Austronesian dispersal. J. Austronesian Stud. 1, 1–32 (2005).
Gray, R. D., Drummond, A. J. & Greenhill, S. J. Language phylogenies reveal expansion pulses and pauses in Pacific settlement. Science 323, 479–483 (2009).
Hill, C. et al. Phylogeography and ethnogenesis of aboriginal Southeast Asians. Mol. Biol. Evol. 23, 2480–2491 (2006).
Jin, W., Wang, S., Wang, H., Jin, L. & Xu, S. Exploring population admixture dynamics via empirical and simulated genome-wide distribution of ancestral chromosomal segments. Am. J. Hum. Genet. 91, 1–14 (2012).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Weir, B. S. & Hill, W. G. Estimating F-statistics. Annu. Rev. Genet. 36, 721–750 (2002).
Felsenstein, J. PHYLIP-phylogeny inference package (Version 3.2). Cladistics 5, 164–166 (1989).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Loh, P.-R. et al. Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193, 1233–1254 (2013).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Ihaka, R. & Gentleman, R. A language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Acknowledgements
These studies were supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (CAS) (XDB13040100), by the National Science Foundation of China (NSFC) grants (91331204; 31171218). S.X. is Max-Planck Independent Research Group Leader and member of CAS Youth Innovation Promotion Association. S.X. also gratefully acknowledges the support of the National Program for Top-notch Young Innovative Talents of the “Wanren Jihua” Project.
Author information
Authors and Affiliations
Contributions
S.X. conceived and designed the study. A.K., J.E.S., B.A.Z., N.K. and A.R.W. contributed genotype data. L.D. analyzed data, with contribution from D.L., W.Y.S. and R.T.H.O. L.D. and S.X. wrote the paper. B.P.H. and Y.Y.T. contributed to revision of the manuscript. All authors reviewed and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Deng, L., Hoh, BP., Lu, D. et al. Dissecting the genetic structure and admixture of four geographical Malay populations. Sci Rep 5, 14375 (2015). https://doi.org/10.1038/srep14375
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14375
This article is cited by
-
Ancestry-informative marker (AIM) SNP panel for the Malay population
International Journal of Legal Medicine (2020)
-
Analysis of five deep-sequenced trio-genomes of the Peninsular Malaysia Orang Asli and North Borneo populations
BMC Genomics (2019)
-
SNP variants associated with non-Hodgkin lymphoma (NHL) correlate with human leukocyte antigen (HLA) class II expression
Scientific Reports (2017)
-
Contrasting Linguistic and Genetic Origins of the Asian Source Populations of Malagasy
Scientific Reports (2016)
-
Ethnic admixture affects diabetes risk in native Hawaiians: the Multiethnic Cohort
European Journal of Clinical Nutrition (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.