Abstract
Grey reef sharks (Carcharhinus amblyrhynchos) can be one of the numerically dominant high order predators on pristine coral reefs, yet their numbers have declined even in the highly regulated Australian Great Barrier Reef (GBR) Marine Park. Knowledge of both large scale and fine scale genetic connectivity of grey reef sharks is essential for their effective management, but no genetic data are yet available. We investigated grey reef shark genetic structure in the GBR across a 1200 km latitudinal gradient, comparing empirical data with models simulating different levels of migration. The empirical data did not reveal any genetic structuring along the entire latitudinal gradient sampled, suggesting regular widespread dispersal and gene flow of the species throughout most of the GBR. Our simulated datasets indicate that even with substantial migrations (up to 25% of individuals migrating between neighboring reefs) both large scale genetic structure and genotypic spatial autocorrelation at the reef scale were maintained. We suggest that present migration rates therefore exceed this level. These findings have important implications regarding the effectiveness of networks of spatially discontinuous Marine Protected Areas to protect reef sharks.
Similar content being viewed by others
Introduction
The conservation of higher order predators is important for the maintenance of marine biodiversity, due to their potentially pivotal role in shaping communities1. Predator removal may have far reaching consequences on lower trophic levels2, including cascading effects on both abundance2,3 and behavior4. This can reduce ecosystem resilience to natural and anthropogenic disturbances5 and in extreme cases lead to ecosystem phase-shifts6. The grey reef shark, Carcharhinus amblyrhynchos, the whitetip reef shark, Triaenodon obesus and the blacktip reef shark, C. melanopterus are abundant mesopredators that occupy the highest trophic level of permanent food webs in coral reefs7,8. Recently it has been shown that pristine coral reefs have high biomasses of higher order predators and reef sharks can make up the majority of that biomass9,10,11. Their potential loss is therefore a major threat to the long term resilience of coral reef ecosystems.
Reef sharks exhibit a combination of life history strategies and habitat specialization that make them particularly susceptible to anthropogenic pressure. They are slow growing, have delayed sexual maturity and low reproductive output12,13 and are therefore susceptible to even modest levels of fishing pressure14. As most species are strictly associated with coral reefs15,16, they are further threatened by the ongoing loss of coral reef habitat17 due to the synergistic effects of overfishing18, eutrophication19, ocean acidification20 and increased sea surface temperatures21. Not surprisingly, their numbers have been shrinking over recent decades14,22,23,24, particularly in areas open to fisheries, raising concerns about the long-term viability of reef shark populations, even in highly regulated marine parks such as the Great Barrier Reef Marine Park (GBR), Australia14,24.
The grey reef shark is the most abundant reef shark in the Indo-Pacific, accounting for up to 50% of higher order predator biomass on healthy reefs9. Within the GBR the grey reef shark is the most threatened by fishing pressure, having the fastest rate of decline of all reef shark species14,24. It has been suggested that grey reef sharks are protected when they inhabit very small but strictly enforced MPAs, where their density can be an order of magnitude higher than in neighboring areas open to fisheries23,24,25. However, acoustic telemetry studies show that grey reef sharks may exhibit limited site fidelity, with some individuals detected transiting across multiple reefs, repeatedly crossing the boundaries of different management zones26,27.
To be effective in increasing local biomass of target species, MPAs must meet two requirements: 1) they need to protect the target species by encompassing a large proportion of its home range; and 2) they need to be at least partially self-seeding (animals within an MPA need to contribute to the next generation)28,29. In this study we use empirical and simulated genetic data to investigate gene-flow for the grey reef shark within the GBR. We hypothesize that if grey reef shark reef fidelity is high, patterns of genetic structure will arise over large distances. Similarly, if self-recruitment is high, patterns of genotypic spatial autocorrelation should be detected i.e. genetic distance between individuals should increase with geographic distance.
Results
Diversity indices
Genetic data were obtained for a total of 121 grey reef sharks sampled along a latitudinal gradient spanning more than 1200 km and 9° of latitude (23° to 14° S, Fig. 1, Supplementary Table S1) within the GBR. We genotyped all individuals at 16 microsatellite loci. Overall diversity indices for the microsatellite dataset are given in the supplementary materials (Supplementary Table S2) and additional diversity indices for each region are given in Table 1. The number of alleles at each locus ranged from 3 to 34, observed heterozygosity for each locus ranged from 0.356 to 0.958 (Supplementary Table S2). None of the loci significantly deviated from Hardy-Weinberg Equilibrium (HWE) and there was no evidence of Linkage Disequilibrium (LD). Within each region, allelic richness (Ar) ranged from 5.6 to 6.93 and observed heterozygosity over all loci ranged from 0.79 to 0.8 (Table 1). Fixation indices (Fis) were small or negative (Table 1) and non-significant (95% confidence intervals overlapped with 0). We obtained sequence data for 813 bp of the ND4 gene from 109 individuals (Table 1, GenBank accession numbers KT326195-KT326303). A total of 10 polymorphic sites were present, of which 7 were parsimony informative (3 singletons). Ten distinct haplotypes were identified and haplotype diversity (h) ranged from 0.61 to 0.8 (Table 1).
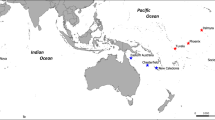
Sampling design. Genetic samples were collected from 121 grey reef sharks in three distinct regions of the GBR and the Coral Sea (Herald Cays).
In the North GBR, 32 individuals were sampled from 2 reefs; in the Central GBR, 54 individuals from 9 reefs and in the southern GBR we sampled 27 individuals from 3 reefs. Furthermore, 8 individuals were sampled from the Herald Cays in the Coral Sea. The map was produced using the package “maps” in the R statistical environment75.
Spatial structure analyses
Pairwise Fst estimates between regions for the mtDNA dataset were small (<0.015) and did not significantly differ from 0 (p > 0.27). Similarly, all measures of genetic differentiation obtained from the microsatellite dataset were not significantly different from 0 (Fig. 2). For the whole dataset Meirman’s F’st was very small (0.003) and non-significant (p > 0.3). The results from the PCA analysis (Fig. 3A) revealed no difference in overall genetic variance among regions; the 95% confidence ellipses of each region were clearly overlapping and densities along the first two principal components were very similar (data not shown). Even the Discriminant Analysis of Principal Components (DAPC), which is widely used to resolve weak genetic structure30,31, failed to reveal any clear partitioning between regions (Fig. 3B).Together these results are strong evidence of extensive gene flow over the entire sampling area.
Results from PCA (A) and DAPC (B) analyses.
Groups were defined as geographic regions within the sampling area (North GBR, Central GBR, South GBR and Coral Sea). Eigenvalues representing the variance explained by principal components (A) and discriminant factors (B) are shown in the scree plots. The x and y axes represent, respectively, the first and second principal components (A) and the first and second discriminant factors (B). Individuals are represented by dots, groups are color-coded and depicted by 95% inertia ellipses.
Spatial autocorrelation analyses did not reveal any spatial structure regardless of whether the dataset was analysed as a whole, or split according to sex (Supplementary Fig. S1). Autocorrelation coefficients (r) were very close to 0 and their 95% confidence interval always overlapped with 0 and fell well within the expectations of the null model. None of the correlograms were statistically significant according to the non-parametric heterogeneity test developed by Smouse, et al.32 (p > 0.1). This complete lack of spatial autocorrelation suggests that reef fidelity persisting for generations is unlikely and is consistent with the results from between region comparisons for both the mtDNA and microsatellite datasets.
Simulations
In contrast with the results from the empirical microsatellite and mitochondrial datasets, genetic differentiation between regions was clearly evident in the simulations, even when migration between neighbouring reefs was set at 25% (Fig. 4A). Estimates of overall Meirman’s F’st range from 0.49 (1% migration) to 0.08 (25% migration), were all significant (p < 0.05) and their confidence interval did not overlap with the F’st value for the observed data. Positive spatial autocorrelation was detected in all simulations and the extent of spatial autocorrelation was largely dependent on migration rates (Fig. 4B). Positive spatial-autocorrelation was detected in our simulations despite the fact that simulation parameters were overly conservative and that we used smaller sample sizes than in our empirical dataset (see Supplementary Materials). These results suggest that the lack of spatial structure observed in the empirical dataset is not the result of limited analytical power and likely reflects very high levels of connectivity.
Results from the simulated datasets under different migration scenarios.
Top graph (A) shows estimated global Meirman’ s F’st estimates between regions. Bottom graph (B) shows spatial autocorrelation estimates (r across different spatial scales: within reef, within region and across the entire simulated seascape). All scenarios resulted in positive and significant patterns of spatial autocorrelation. Error bars represent 95% confidence intervals obtained from 10 independent simulations. Grey lines in both (A,B) represent the estimates obtained from the empirical dataset.
Discussion
This first assessment of genetic connectivity in grey reef sharks was carried out across a latitudinal gradient spanning more than 1200 km and 9° of latitude i.e. 2/3 of the entire length of the Great Barrier Reef. Despite this spatial scale, we found no evidence of genetic differentiation across the study area and no evidence of fine-scale spatial structure. Our simulated data shows that we had the statistical power to detect structure at the regional geographical scale as well as fine scale spatial autocorrelation, even assuming high migration rates (~25% of individuals) between neighbouring reefs. With respect to temporal scales relevant to ecological processes, we found no genotypic spatial autocorrelation within reefs. Structuring of allele frequencies via genetic drift may be effectively prevented by a small number of migrants per generation in large populations and therefore may only become evident across large spatial and temporal scales33. It is possible therefore that large scale dispersal by a portion of the migrants resulted in an overestimate of genetic drift in our simulations. Genotypic spatial autocorrelation on the other hand reflects individual dispersal across a few generations34, is most pronounced at small spatial scales, measurable within less than 10 generations and reaches quasi-stationarity within 50–100 generations35. We can therefore conclude that the lack of spatial autocorrelation reflects sufficient dispersal over a few generations to prevent genotypic structure from accumulating within the GBR for grey reef sharks.
Our results have important implications for the management of this species and are in sharp contrast with the findings of several previous studies on other reef associated sharks36,37,38,39. In the closely related Carcharhinus melanopterus there are clear patterns of genetic structure between reefs separated by as little as 50 km within French Polynesia36,37. Similarly, a recent study on gene flow of the other principal reef associated carcharhinid (Triaenodon obesus), also revealed very strong genetic structure (Φst = 0.25) between North and Central GBR38. Perhaps more surprising than the lack of spatial structure within the GBR is an absence of genetic partitioning between the GBR and the Coral Sea (Herald Cays). The Herald Cays are isolated coral reefs located approximately 200 km east of the edge of the continental shelf where the outer reefs of the GBR lie. They are separated from the GBR by waters deeper than 1000 m. Therefore, it is evident that grey reef sharks not only disperse between semi isolated reefs within the GBR, but have the capacity for long distance dispersal across oceanic waters. Heupel, et al.26 reported that in June 2008 an individual grey reef shark undertook a large scale movement (124 km) from Osprey Reef in the Coral Sea to the Ribbon Reefs in the North GRB. This same individual returned to Osprey reef in October of that year40. Our results concord with these observations and suggest that movements on the scale of 100 s of km across oceanic waters occur at a rate sufficient to prevent detectable genetic drift between isolated reefs in the Coral Sea and the GBR.
Grey reef sharks are often described as strict habitat specialists, associated exclusively with coral reef habitats15,17,25. Previous telemetry studies suggested that grey reef sharks exhibit a high degree of site fidelity in isolated seamounts, as most animals show year-round residency within the study area40,41. This has led to the presumption that even within the GBR grey reef sharks will exhibit a high degree of reef fidelity and therefore will be well protected by strictly enforced but small marine protected areas23,24. This interpretation has been supported by robust estimates of grey reef shark abundances across different management zones, showing that grey reef shark densities are up to an order of magnitude higher in strictly enforced no access areas compared to areas open to fisheries23,24. Robbins, et al.24 and Rizzari, et al.23 concluded that differences in fishing mortality are the main cause of the observed differences and that therefore even very small MPAs can be very effective in increasing grey reef shark biomass if they are well complied with. In support, Hisano, et al.14 argued that quantitative projections of abundances in fished versus unfished reefs calculated from differences in fishing mortality are also compatible with the observed differences.
This interpretation, however, relies on the assumption that grey reef sharks within each reef represent a closed population14. That is, it assumes: 1) that grey reef sharks are protected from fishing throughout most of their home range and 2) that reef sharks within each MPA contribute to the biomass of future generations within the same MPA. Telemetry studies carried out in the same regions do not support the first assumption. Heupel, et al.26 found that in the North GBR, grey reef sharks show little fidelity to the reef where they were initially tagged, consistently moving across different management zones. Even in the Central GBR, where reefs are isolated by much larger distances, 40% of the grey reef sharks (90% of adult males) were recorded moving between reefs, often crossing different management zones in a period of less than 2 years27. Furthermore we found no evidence of genetic spatial autocorrelation, suggesting that that migration rates between neighbouring reefs likely exceed 25%.
Heupel, et al.26 suggested that differences in fishing mortality may not be the only driver of the differences in abundances observed by Robbins, et al.24 and that other factors, such as differences in carrying capacity between areas open to fisheries and protected areas, might also be at play. It is possible that additional factors may affect grey reef shark abundances in strictly enforced protected areas. For example, strictly enforced MPAs have much larger fish biomasses42 compared to areas open to fisheries and therefore could potentially sustain higher predator abundances. Espinoza, et al.25 also found a positive effect of hard coral cover on grey reef shark abundance. Reefs with high hard coral cover provide higher structural complexity and this in turn may sustain a higher biomass43 of potential prey. It is well established that MPAs that have been enforced for extended periods of times are more effective at preventing losses in hard coral cover44. In the GBR, no-take areas have on average higher coral cover than areas open to fisheries45 and experience much lower frequencies of crown-of-thorns starfish (Acanthaster planci) outbreaks46, a leading cause of decline in coral cover in the region47.
The lack of genetic structure in this study may be the result of either high juvenile or adult dispersal, or both, albeit available movement data point to adults representing the dispersal stage. In two acoustic tagging studies conducted on the GBR, adult grey reef sharks were found to exhibit limited reef fidelity and move repeatedly between multiple reefs. Espinoza, et al.27 reported that approximately 90% of adult males were detected on multiple (up to five), semi-isolated reefs in the Central GBR, while only one out of 12 tracked juveniles was detected at more than one reef. Heupel, et al.26 also found that small juveniles exhibited high reef fidelity, while adults move continuously between reefs. Espinoza, et al.48 found that larger grey reef sharks and silvertip sharks (Carcharhinus albimarginatus) and especially large males, have wider home ranges that encompass multiple reefs. Given that larger sharks can traverse greater distances for comparable energetic expenditure49 and are less vulnerable to predation, increased dispersal by adults is not surprising. However, a closely related species (C. melanopterus) has been shown to exhibit ontogenetic dispersal between inshore nursery and offshore reefs50. While there is no empirical evidence of the existence of nursery areas for grey reef sharks (juveniles co-occur with adults on coral reefs), the hypothesis that ontogenetic dispersal occurs cannot be discarded. Future genetic studies could compare genetic structure in different life-stages (juveniles, sub adults and adults), to test ontogenetic dispersal differences.
Espinoza, et al.27 suggested that in grey reef sharks high adult male dispersal may provide benefits in terms of genetic and demographic connectivity and/or provide foraging opportunities while reducing competition with large resident females. Our data do not show any evidence of sex-bias in dispersal, however these findings do not necessarily refute the hypothesis of sex-bias dispersal if we assume female migration was sufficient to prevent any detectable effect of genetic drift. Espinoza, et al.27 found that >25% of mature females were detected on two distinct reefs within a period of 2 years. Since they investigated movement patterns between semi-isolated reefs (10 s of km apart) within the Central GBR, it is logical to assume that in other regions of the GBR where reefs are less isolated, movements between reefs would occur at higher frequencies, given that reefs that are separated by small distances may be perceived as a continuous habitat26,29. It therefore seems feasible, particularly given the results of our simulations, that female dispersal is large enough to mask any effect of sex bias dispersal at the spatial scale of this study.
The widespread gene flow documented in this study, along with the findings of recent acoustic telemetry studies on grey reef shark movement suggest that while MPAs play an important role in protecting grey reef sharks, they are no panacea for reef shark conservation. Complementary management strategies appear necessary. The shortcomings of the current model of discontinuous networks of MPAs for the protection of coral reef organisms with adult dispersal have recently been highlighted by Momigliano, et al.29 and Espinoza, et al.48. Potential solutions that have been proposed include the design of MPAs which include closely spaced (<20 km apart) reefs48 and the establishment of protected (i.e. no fishing) connectivity corridors that provide for movements between MPAs, thereby extending protection for animals with larger home ranges29. Furthermore, complementary management strategies based on fishery restrictions, such the ones implemented in eastern Australia in 2009, appear necessary.
Material and Methods
Sample collection
Fin clips from 121 grey reef sharks were obtained along a latitudinal gradient spanning more than 1200 km and 9° of latitude (23° to 14° S, Fig. 1) within the GBR. Specimens from the North GBR (2 reefs, N = 32), the Central GBR (9 reefs, N = 54), the Coral Sea (1 reef, N = 8) and the South GBR (1 reef, N=2) were collected between February 2001 and April 2005 and represent a subsample from Robbins12. The remaining individuals from the South GBR (2 reefs, N = 25) were collected in April 2013 by hook and line. Sharks were captured in shallow waters (<30 m) on the reef slopes of Wistari Reef and Heron Island reef using heavy handlines and 16/0 tuna circle hooks to reduce the risk of deep hooking. A fin clip of approximately 15 mm2 was taken before the animals were released. Sampling was conducted in accordance with Macquarie University’s approved guidelines and the sampling protocol was approved by Macquarie University’s Animal Ethics Committee (AEC reference No.: 2012/044). Samples were collected under permit G13/35796.1 issued by the Great Barrier Reef Marine Park Authority (GBRMPA). Samples were stored in 95% ethanol at −20 °C. Details of sampling, including geographic coordinates of all sampling locations, are given in the supplementary materials (Supplementary Table S1).
Laboratory procedures
DNA was extracted from fin clips of 121 grey reef sharks using a modified Chelex protocol51. A fragment of the NADH deydrogenase subunit 4 (ND4) gene approximately 850 bp long was amplified by Polymerase Chain Reaction (PCR) using the primers ND452 and H12293-Leu53. We amplified a suite of 15 microsatellite loci recently isolated for the grey reef sharks by Momigliano, et al.54 and an additional microsatellite locus (Cpl169) which was originally developed for C. plumbeus by Portnoy, et al.55. The 16 microsatellite loci used are listed in the supplementary materials (Supplementary Table S2).
Empirical Data Analysis
Mitochondrial DNA sequences were aligned using Bioedit56. Molecular indices, including haplotype diversity (h) and nucleotide diversity (π) were estimated in DNAsp57. Pairwise Fst between regions based on ND4 gene sequences were calculated in Arlequin 3.558.
Microsatellites were scored using GeneMapper (Version 4) and alleles were binned using the software TANDEM following the method outlined by Matschiner and Salzburger59. The dataset was tested for significant departures from HWE in Genepop 4.260 and for the presence of null alleles using the software MICROCHECKER61. Tests for LD were carried out using the software FSTAT version 2.9.3.262, using a sequential Holm-Bonferroni correction. Because there was no statistically significant departure from HWE and no evidence of linkage disequilibrium, all loci were used in the analyses. The microsatellite dataset is publicly accessible through the Dryad repository (doi:10.5061/dryad.362s5).
Allele frequencies and measures of genetic differentiation were estimated in the R statistical environment using the package diveRsity63. Different measures of genetic differentiation (Weir and Cockerham Fst64, Hedrick’s G’st65 and Jost’s D66) and their 95% confidence intervals were estimated using 1000 bootstraps using the same package. Furthermore, Meirman’s F’st67 were calculated in the software GenAlEx 6.568.
A Principal Component Analysis (PCA), using region as the grouping factor, was performed using the package Adegenet69. The principal components obtained from PCA were used as input synthetic variables to perform a Discriminant Analysis of Principal Components (DAPC) (for a full description of the method see Jombart, et al.31 and Horne, et al.70). The number of principal components to be retained in the discriminant analysis (N) was determined using alpha-score optimization, a method that finds a trade-off between discriminative power and model over-fitting.
To assess for the presence of spatial structure, we estimated genotypic spatial autocorrelation across three distance classes; 1) within single reefs 2) within the same geographic region 3) across the entire sampling area. We calculated individual based genetic distances in GenAlEx 6.5, using the codom-genotipic genetic distance option (for a detailed description of the method see Smouse and Peakall71). Autocorrelation coefficients (r)71,72 were calculated for each distance class in GenAlEx 6.5. We used 999 random permutations to generate 95% confidence intervals for the null model of no spatial autocorrelation and 999 bootstraps to generate 95% confidence intervals of r for each distance class. The analysis was performed first for all sampled individuals and then separately for males and females to detect possible sex-bias in spatial autocorrelation. The overall significance of each correlogram was tested using the non-parametric heterogeneity test outlined in Smouse, et al.32.
Simulations
To determine whether the statistical analyses performed were capable of detecting genetic structuring at different levels of between-reef migration, we carried out a range of simulations using the software EASYPOP73. We created an 8 × 8 network of 64 reefs, with each reef encompassing an area of 20 km2. The census size of each reef was approximated based on the densities provided in the literature from pristine GBR reefs (2 sharks per hectare24). We assumed an effective population size (Ne) to census size (Nc) ratio of 1:1074 and created a dataset with the same number of loci and the locus diversity of the empirical dataset. Genetic drift was simulated for 100 generations with a 2-dimensional stepping stone model and levels of between reef migrations were set at 1%, 10% and 25%. Ten simulations were carried out for each migration level. We randomly sampled 10 individuals from three simulated reefs in each region of the simulated seascape and carried out the same repertoire of analyses that were carried out for the empirical dataset. The value of each of the simulation parameters used was chosen to provide a conservative estimate of genetic drift: that is, the effect of genetic drift and the subsequent genetic structuring within the simulated dataset would be likely less than in our real data set (see Supplementary Material for details).
Additional Information
How to cite this article: Momigliano, P. et al. Connectivity in grey reef sharks (Carcharhinus amblyrhynchos) determined using empirical and simulated genetic data. Sci. Rep. 5, 13229; doi: 10.1038/srep13229 (2015).
References
Heithaus, M. R., Frid, A., Wirsing, A. J. & Worm, B. Predicting ecological consequences of marine top predator declines. Trends Ecol. Evol. 23, 202–210 (2008).
Myers, R. A., Baum, J. K., Shepherd, T. D., Powers, S. P. & Peterson, C. H. Cascading effects of the loss of apex predatory sharks from a coastal ocean. Science 315, 1846–1850 (2007).
Bascompte, J., Melián, C. J. & Sala, E. Interaction strength combinations and the overfishing of a marine food web. Proc. Natl. Acad. Sci. USA 102, 5443–5447, 10.1073/pnas.0501562102 (2005).
Wirsing, A. J., Heithaus, M. R., Frid, A. & Dill, L. M. Seascapes of fear: evaluating sublethal predator effects experienced and generated by marine mammals. Mar. Mamm. Sci. 24, 1–15 (2008).
Ruppert, J. L., Travers, M. J., Smith, L. L., Fortin, M.-J. & Meekan, M. G. Caught in the middle: combined impacts of shark removal and coral loss on the fish communities of coral reefs. PloS one 8, e74648 (2013).
Estes, J. A. & Duggins, D. O. Sea otters and kelp forests in Alaska: generality and variation in a community ecological paradigm. Ecol. Monogr. 65, 75–100, 10.2307/2937159 (1995).
Mourier, J., Planes, S. & Buray, N. Trophic interactions at the top of the coral reef food chain. Coral Reefs 32, 285–285 (2013).
Heupel, M. R., Knip, D. M., Simpfendorfer, C. A. & Dulvy, N. K. O. Mar. Ecol. Prog. Ser. 495, 291–298 (2014).
Friedlander, A. M. et al. The real bounty: marine biodiversity in the Pitcairn Islands. PloS one 9, e100142 (2014).
Sandin, S. A. et al. Baselines and degradation of coral reefs in the northern Line Islands. PloS one 3, e1548 (2008).
Stevenson, C. et al. High apex predator biomass on remote Pacific islands. Coral Reefs 26, 47–51, 10.1007/s00338-006-0158-x (2007).
Robbins, W. D. Abundance, demography and population structure of the grey reef shark (Carcharhinus amblyrhynchos) and the whitetip reef shark (Triaenodon obesus)(Fam. Charcharhinidae) Doctoral dissertation thesis. James Cook University, (2006).
Chin, A., Simpfendorfer, C., Tobin, A. & Heupel, M. Validated age, growth and reproductive biology of Carcharhinus melanopterus, a widely distributed and exploited reef shark. Mar. Freshw. Res. 64, 965–975 (2013).
Hisano, M., Connolly, S. R. & Robbins, W. D. Population growth rates of reef sharks with and without fishing on the Great Barrier Reef: robust estimation with multiple models. PloS one 6, e25028 (2011).
Chin, A., Tobin, A., Simpfendorfer, C. & Heupel, M. Reef sharks and inshore habitats: patterns of occurrence and implications for vulnerability. Mar. Ecol. Prog. Ser. 460, 115–125 (2012).
Rizzari, J. R., Frisch, A. J. & Magnenat, K. A. Diversity, abundance and distribution of reef sharks on outer-shelf reefs of the Great Barrier Reef, Australia. Mar. Biol. 161, 2847–2855 (2014).
Chin, A., Kyne, P. M., Walker, T. I. & McAuley, R. An integrated risk assessment for climate change: analysing the vulnerability of sharks and rays on Australia’s Great Barrier Reef. Global Change Biology 16, 1936–1953 (2010).
Hughes, T. P. Catastrophes, phase shifts and large-scale degradation of a Caribbean coral reef. Science 265, 1547–1547 (1994).
Fabricius, K. E. Effects of terrestrial runoff on the ecology of corals and coral reefs: review and synthesis. Mar. Pollut. Bull. 50, 125–146 (2005).
Fabricius, K. E. et al. Losers and winners in coral reefs acclimatized to elevated carbon dioxide concentrations. Nature Climate Change 1, 165–169 (2011).
Hoegh-Guldberg, O. et al. Coral reefs under rapid climate change and ocean acidification. Science 318, 1737–1742 (2007).
Graham, N. A., Spalding, M. D. & Sheppard, C. R. Reef shark declines in remote atolls highlight the need for multi‐faceted conservation action. Aquat. Conserv. 20, 543–548 (2010).
Rizzari, J. R., Frisch, A. J. & Connolly, S. R. How robust are estimates of coral reef shark depletion? Biol. Conserv. 176, 39–47 (2014).
Robbins, W. D., Hisano, M., Connolly, S. R. & Choat, J. H. Ongoing collapse of coral-reef shark populations. Curr. Biol. 16, 2314–2319 (2006).
Espinoza, M., Cappo, M., Heupel, M. R., Tobin, A. J. & Simpfendorfer, C. A. Quantifying shark distribution patterns and species-habitat associations: implications of Marine Park Zoning. PloS one 9, e106885 (2014).
Heupel, M. R., Simpfendorfer, C. A. & Fitzpatrick, R. Large–scale movement and reef fidelity of grey reef sharks. PloS one 5, e9650 (2010).
Espinoza, M., Heupel, M. R., Tobin, A. & Simpfendorfer, C. Residency patterns and movements of grey reef sharks (Carcharhinus amblyrhynchos) in semi-isolated coral reef habitats. Mar. Biol. 162, 343–358, 10.1007/s00227-014-2572-x (2015).
Almany, G. et al. Connectivity, biodiversity conservation and the design of marine reserve networks for coral reefs. Coral Reefs 28, 339–351 (2009).
Momigliano, P., Harcourt, R. & Stow, A. Conserving coral reef organisms that lack larval dispersal:are networks of Marine Protected Areas good enough? Frontiers in Marine Science 2:16, 10.3389/fmars.2015.00016 (2015).
Horne, J. B., Momigliano, P., van Herwerden, L. & Newman, S. J. Murky waters: Searching for structure in genetically depauperate blue threadfin populations of Western Australia. Fisheries Research 146, 1–6 (2013).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet. 11, 94 (2010).
Smouse, P. E., Peakall, R. & Gonzales, E. A heterogeneity test for fine‐scale genetic structure. Mol. Ecol. 17, 3389–3400 (2008).
Ovenden, J. R. Crinkles in connectivity: combining genetics and other types of biological data to estimate movement and interbreeding between populations. Mar. Freshw. Res. 64, 201–207 (2013).
Stow, A., Sunnucks, P., Briscoe, D. & Gardner, M. The impact of habitat fragmentation on dispersal of Cunningham’s skink (Egernia cunninghami): evidence from allelic and genotypic analyses of microsatellites. Mol. Ecol. 10, 867–878 (2001).
Epperson, B. Estimating dispersal from short distance spatial autocorrelation. Heredity 95, 7–15 (2005).
Vignaud, T., Clua, E., Mourier, J., Maynard, J. & Planes, S. Microsatellite analyses of blacktip reef sharks (Carcharhinus melanopterus) in a fragmented environment show structured clusters. PloS one 8, e61067 (2013).
Vignaud, T. M. et al. Blacktip reef sharks, Carcharhinus melanopterus, have high genetic structure and varying demographic histories in their Indo‐Pacific range. Mol. Ecol. 23, 5193–5207 (2014).
Whitney, N. M., Robbins, W. D., Schultz, J. K., Bowen, B. W. & Holland, K. N. Oceanic dispersal in a sedentary reef shark (Triaenodon obesus): genetic evidence for extensive connectivity without a pelagic larval stage. J. Biogeogr. 39, 1144–1156 (2012).
Mourier, J. & Planes, S. Direct genetic evidence for reproductive philopatry and associated fine‐scale migrations in female blacktip reef sharks (Carcharhinus melanopterus) in French Polynesia. Mol. Ecol. 22, 201–214 (2013).
Barnett, A., Abrantes, K. G., Seymour, J. & Fitzpatrick, R. Residency and spatial use by reef sharks of an isolated seamount and its implications for conservation. PloS one 7, e36574 (2012).
Field, I., Meekan, M., Speed, C., White, W. & Bradshaw, C. Quantifying movement patterns for shark conservation at remote coral atolls in the Indian Ocean. Coral Reefs 30, 61–71 (2011).
Edgar, G. J. et al. Global conservation outcomes depend on marine protected areas with five key features. Nature (2014).
Gratwicke, B. & Speight, M. R. Effects of habitat complexity on Caribbean marine fish assemblages. Mar. Ecol. Prog. Ser. 292, 301–310 (2005).
Selig, E. R. & Bruno, J. F. A global analysis of the effectiveness of marine protected areas in preventing coral loss. PLoS ONE 5, e9278, 10.1371/journal.pone.0009278 (2010).
McCook, L. J. et al. Adaptive management of the Great Barrier Reef: A globally significant demonstration of the benefits of networks of marine reserves. Proc. Natl. Acad. Sci. 107, 18278–18285 (2010).
Sweatman, H. No-take reserves protect coral reefs from predatory starfish. Curr. Biol. 18, R598–R599 (2008).
De’ath, G., Fabricius, K. E., Sweatman, H. & Puotinen, M. The 27–year decline of coral cover on the Great Barrier Reef and its causes. Proc. Natl. Acad. Sci. 109, 17995–17999, 10.1073/pnas.1208909109 (2012).
Espinoza, M., Lédée, E. J., Simpfendorfer, C. A., Tobin, A. J. & Heupel, M. R. Contrasting movements and connectivity of reef-associated sharks using acoustic telemetry: implications for management. Ecol. Appl. (2015).
Parsons, G. Metabolism and swimming efficiency of the bonnethead shark Sphyrna tiburo. Mar. Biol. 104, 363–367 (1990).
Chin, A., Heupel, M. R., Simpfendorfer, C. A. & Tobin, A. J. Ontogenetic movements of juvenile blacktip reef sharks: evidence of dispersal and connectivity between coastal habitats and coral reefs. Aquat. Conserv. 23, 468–474, 10.1002/aqc.2349 (2013).
Walsh, P. S., Metzger, D. A. & Higuchi, R. Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. BioTechniques 10, 506–513 (1991).
Arèvalo, E., Davis, S. K. & Sites, J. W. Mitochondrial DNA sequence divergence and phylogenetic relationships among eight chromosome races of the Sceloporus grammicus complex (Phrynosomatidae) in central Mexico. Syst. Biol. 43, 387–418 (1994).
Inoue, J. G., Miya, M., Tsukamoto, K. & Nishida, M. Evolution of the Deep-Sea Gulper Eel Mitochondrial Genomes: Large-Scale Gene Rearrangements Originated Within the Eels. Mol. Biol. Evol. 20, 1917–1924, 10.1093/molbev/msg206 (2003).
Momigliano, P., Robbins, W. D., Gardner, M. & Stow, A. Characterisation of 15 novel microsatellite loci for the grey reef shark (Carcharhinus amblyrhynchos). Conservation Genetics Resources, 6, 661–663, 10.1007/s12686-014-0174-z (2014).
Portnoy, D., Mcdowell, J., Thompson, K., Musick, J. & Graves, J. Isolation and characterization of five dinucleotide microsatellite loci in the sandbar shark, Carcharhinus plumbeus. Mol. Ecol. Notes 6, 431–433 (2006).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41, 95–98 (1999).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular ecology resources 10, 564–567 (2010).
Matschiner, M. & Salzburger, W. TANDEM: integrating automated allele binning into genetics and genomics workflows. Bioinformatics 25, 1982–1983 (2009).
Raymond, M. & Rousset, F. GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J. Hered. 86, 248–249 (1995).
Van Oosterhout, C., Hutchinson, W. F., Wills, D. P. & Shipley, P. MICRO‐CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 4, 535–538 (2004).
FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9. 3) (Available from www.unil.ch/izea/softwares/fstat.html, 2001).
Keenan, K., McGinnity, P., Cross, T. F., Crozier, W. W. & Prodöhl, P. A. diveRsity: An R package for the estimation and exploration of population genetics parameters and their associated errors. Methods in Ecology and Evolution 4, 782–788 (2013).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution, 1358–1370 (1984).
Hedrick, P. W. A standardized genetic differentiation measure. Evolution 59, 1633–1638 (2005).
Jost, L. GST and its relatives do not measure differentiation. Mol. Ecol. 17, 4015–4026 (2008).
Meirmans, P. G. Using the AMOVA framework to estimate a standardized genetic differentiation measure. Evolution 60, 2399–2402 (2006).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28, 2537–2539 (2012).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Horne, J. B., Momigliano, P., Welch, D. J., Newman, S. J. & van Herwerden, L. Limited ecological population connectivity suggests low demands on self‐recruitment in a tropical inshore marine fish (Eleutheronema tetradactylum: Polynemidae). Mol. Ecol. 20, 2291–2306 (2011).
Smouse, P. E. & Peakall, R. Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82, 561–573 (1999).
Peakall, R., Ruibal, M. & Lindenmayer, D. B. Spatial autocorrelation analysis offers new insights into gene flow in the Australian bush rat, Rattus fuscipes. Evolution 57, 1182–1195 (2003).
Balloux, F. EASYPOP (version 1.7): a computer program for population genetics simulations. J. Hered. 92, 301–302 (2001).
Frankham, R. Effective population size/adult population size ratios in wildlife: a review. Genet. Res. 66, 95–107 (1995).
R Core Team. R: A language and environment for statistical computing. http://www.R-project.org (2014).
Acknowledgements
This study was funded by the Department of Biological Sciences at Macquarie University and the Sea World Research and Rescue Foundation (project SWR/7/2013)
Author information
Authors and Affiliations
Contributions
P.M. designed and conducted the study, analyzed the data and wrote the manuscript. A.S. and R.H. contributed to study design, data analysis, data interpretation and to the writing of the manuscript. W.R. contributed to sample collection, data interpretation and to the writing of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Momigliano, P., Harcourt, R., Robbins, W. et al. Connectivity in grey reef sharks (Carcharhinus amblyrhynchos) determined using empirical and simulated genetic data. Sci Rep 5, 13229 (2015). https://doi.org/10.1038/srep13229
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep13229
This article is cited by
-
Characterization of 35 new microsatellite markers for the blacktip reef shark (Carcharhinus melanopterus) and cross-species amplification in eight other shark species
Molecular Biology Reports (2023)
-
Genomic insights into the historical and contemporary demographics of the grey reef shark
Heredity (2022)
-
Recent expansion of marine protected areas matches with home range of grey reef sharks
Scientific Reports (2021)
-
Contrasting global, regional and local patterns of genetic structure in gray reef shark populations from the Indo-Pacific region
Scientific Reports (2019)
-
Repeated long-range migrations of adult males in a common Indo-Pacific reef shark
Coral Reefs (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.