Abstract
H5N6 avian influenza viruses (AIVs) may pose a potential human risk as suggested by the first documented naturally-acquired human H5N6 virus infection in 2014. Here, we report the first cases of fatal H5N6 avian influenza virus (AIV) infection in a domestic cat and wild birds. These cases followed human H5N6 infections in China and preceded an H5N6 outbreak in chickens. The extensive migration routes of wild birds may contribute to the geographic spread of H5N6 AIVs and pose a risk to humans and susceptible domesticated animals and the H5N6 AIVs may spread from southern China to northern China by wild birds. Additional surveillance is required to better understand the threat of zoonotic transmission of AIVs.
Similar content being viewed by others
Introduction
On May 7, 2014, China formally confirmed the first human infection with an H5N6 avian influenza virus (AIV)1,2. The patient, a 49-year-old male in Sichuan province, was hospitalized with severe pneumonia and died. Notably, the identification of the first human case of H5N6 virus infection coincided with reports of human infections with H7N9 and H10N8 AIVs in China3,4,5. H7N9 and H10N8 AIVs have also been shown to infect some animals, including feral dogs6 and tree sparrows7, suggesting that mammals and wild birds may play a role in the ongoing circulation of novel AIVs that sporadically infect humans in China. Here, we report the first cases of fatal H5N6 AIV infection in a domestic cat and wild birds.
Results
Phylogenetic Analysis
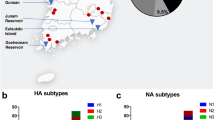
The lung specimen from the deceased cat and all lung specimens from three deceased swan geese were positive for H5N6 AIV. No other influenza virus subtypes were detected. The viral genomes of each isolate were sequenced to assess the molecular and epidemiological characteristics of these H5N6 strains. The PB2, PB1, PA, HA, NP, NA, M and NS segments of A/cat/Sichuan/SC18/2014 (H5N6) show 99.5%, 99.5%, 99.3%, 99.4%, 97.3%, 98.9%, 99.9% and 99.6% nucleotide sequence identity to the corresponding segments of A/swan goose/Jilin/JL01/2014 (H5N6). Phylogenetic analysis of the HA gene sequences indicated that the HA segments of each isolate were derived from the Asian H5N1 clade 2.3.4.6 HA lineage (Fig. 1). Collectively, the gene segments of A/cat/Sichuan/SC18/2014 and A/swan goose/Jilin/JL01/2014 were most closely related to a human H5N6 influenza isolate named A/Sichuan/26221/2014 (Table 1). Phylogenetic analyses suggest that the newly isolated H5N6 viruses possess high nucleotide sequence identity to H5N6 AIV subtypes previously isolated in southern China (Fig. 1 and Figure S1-S7). We speculate that the H5N6 virus may have emerged in Sichuan province in early 2014 and subsequently spread to Jilin provinces through the migration of wild birds.
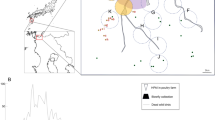
Phylogenetic tree of the hemagglutinin (HA) genes of influenza A (H5N6) viruses, China, 2014. Triangles indicate viruses characterized in this study. Other H5N6 viruses are indicated in circles. Horizontal branch lengths are proportional to genetic distances. Scale bar indicates nucleotide substitutions per site.
Analysis of molecular features associated with AIV virulence, transmissibility and antiviral resistance
We next analyzed whether these H5N6 strains possessed key molecular features associated with increased virulence in mammals, mammalian transmissibility and antiviral resistance (Table 2). The H5N6 viruses displayed some molecular markers associated with increased virulence and transmission in mammals. A/cat/Sichuan/SC18/2014 (H5N6) and A/swan goose/Jilin/JL01/2014 (H5N6) possessed an alanine at position 160 which is associated with H5 transmissibility in ferrets and an aspartic acid residue at M1 position 30, an alanine residue at M1 position 215 and a serine residue at NS1 position 42 which are associated with increased virulence in mice (Table 2). Several studies have suggested that NA stalk truncation enhances H5N1 virulence in mice and increases replication and virulence of AIVs in chicken eggs and poultry8,9,10. A deletion of amino acids 59-69 of the NA stalk has been reported in some H5N6 viruses, although no such stalk deletion was found in A/cat/Sichuan/SC18/2014 (H5N6) and A/swan goose/Jilin/JL01/2014 (H5N6). Molecular markers of oseltamivir- and amandatide-resistance were also not present in the NA and M1 protein sequences of the H5N6 viruses, indicating that they should be still sensitive to influenza antiviral drugs.
Discussion
In this study, we provide the first evidence of fatal H5N6 AIV infection in domestic cats and wild birds. The extensive migration routes of wild birds may contribute to the geographic spread of H5N6 AIVs and pose a risk to humans and susceptible domesticated animals. Based on our phylogenetic analysis, we speculated that the H5N6 AIVs may spread from southern China to northern China by wild birds. This hypothesis is supported by the recent outbreak of H5N6 AIV among chickens in Harbin city, Heilongjiang province, China, which affected 20,550 chickens of which 17,790 died11. The H5N6-infected swan geese were identified in Baicheng, China which is adjacent to inner Mongolia. The middle migratory flyway of China includes inner Mongolia to the north and Sichuan province to the south12. Wild birds in or near Baicheng could migrate to Sichuan province via this path. Additionally, documentation of H5N6 infection of domestic cats raises the possibility that cats could play a role in the mammalian adaptation or perhaps transmission of novel avian influenza viruses. Therefore, continuous epidemiological monitoring of different animal species such as cats, dogs and wild birds should be performed to effectively reduce the threat of zoonotic transmission of influenza strains like the newly emergent H5N6 avian influenza virus.
Methods
Ethics statement
The protocol of the study was conducted in accordance with guidelines of animal welfare of World Organization for Animal Health. All experimental protocol were approved by the Review Board Military Veterinary Research Institute of the Academy of Military Medical Sciences.
Facilities
Studies with H5N6 AIVs were conducted in a biosecurity level 3 laboratory approved by the Military Veterinary Research Institute of the Academy of Military Medical Sciences.
Sample Collection
During May–June 2014, seventy feces specimens from wild birds and one lung specimen from a dead domestic cat were collected in areas with close-proximity to the residence of the patient infected with H5N6 AIV in Nanchong city, Sichuan province, southwest China. The domestic cat is suspected of having succumbed to AIV infection based on flu-like clinical signs and necropsy findings. Additionally, lung specimens were collected following the sudden and unexplained deaths of three swan geese in Baicheng city, Jilin province, northeast China. No additional information was available. We screened all specimens by RT-PCR for evidence of influenza virus infection.
Virus Isolation
We inoculated the allantoic cavities of 10-day-old specific pathogen-free embryonated chicken eggs with material from the H5N6 AIV-positive lung specimen from the domestic cat and one swan goose. After 60 hours incubation at 37 °C, we recovered two virus isolates, named A/cat/Sichuan/SC18/2014 (H5N6) and A/swan goose/Jilin/JL01/2014 (H5N6).
RNA isolation, PCR amplification and Sequencing
Viral RNA was isolated from the allantoic fluid of inoculated eggs using the RNeasy Mini kit (QIAGEN, Germantown, MD) according to the manufacturer’s protocol. Reverse transcription of viral RNA and subsequent PCR were performed using primers specific for each gene segment (sequences available upon request). PCR products were purified using the QIAquick PCR purification kit (QIAGEN, Germantown, MD) according to the manufacturer’s protocol. Viral gene segments were sequenced by the Beijing Genomics Institute (Beijing, China). The GenBank accession numbers of A/cat/Sichuan/SC18/2014 (H5N6) and A/swan goose/Jilin/JL01/2014 (H5N6) are KM873635 to KM873650, respectively.
Phylogenetic Analysis
To investigate the molecular and epidemiological characteristics and to determine the profile of genetic diversity, phylogenetic trees were constructed using molecular evolutionary genetics analysis MEGA 6 (http://www.megasoftware.net/mega.php) with the neighbor-joining (NJ) method to calculate distance. Bootstrap values were estimated for 1,000 replicates.
Additional Information
How to cite this article: Yu, Z. et al. Fatal H5N6 Avian Influenza Virus Infection in a Domestic Cat and Wild Birds in China. Sci. Rep. 5, 10704; doi: 10.1038/srep10704 (2015).
References
World Health Organization (WHO). WHO China statement on H5N6. (2014). (http://www.wpro.who.int/china/mediacentre/releases/2014/20140507/en/)(Accessed 7 May 2014).
Report of Health and Family Planning Commission of Sichuan Province. The first human case of H5N6 bird flu was confirmed. (2014). (http://www.scwst.gov.cn/index.php/2012-07-24-12-47-36/9425-1h5n6)(Accessed 4 May 2014).
Chen, H. et al. Clinical and epidemiological characteristics of a fatal case of avian influenza A H10N8 virus infection: a descriptive study. Lancet 383, 714–721, 10.1016/S0140-6736(14)60111-2 (2014).
Gao, R. et al. Human infection with a novel avian-origin influenza A (H7N9) virus. N Engl J Med 368, 1888–1897, 10.1056/NEJMoa1304459 (2013).
Report of Health and Family Planning Commission of Jiangxi Province. The third human case of H10N8 bird flu was confirmed. (2014). (http://www.jxwst.gov.cn/gzdt/201402/t20140213_308109.htm)(Accessed 13 February 2014)
Su, S. et al. First Evidence of H10N8 Avian Influenza Virus Infections among Feral Dogs in Live Poultry Markets in Guangdong Province, China. Clin Infect Dis 59, 748–750, 10.1093/cid/ciu345 (2014).
Zhao, B. et al. Novel avian influenza A(H7N9) virus in tree sparrow, Shanghai, China, 2013. Emerg Infect Dis 20, 850–853, 10.3201/eid2005.131707 (2014).
Matsuoka, Y. et al. Neuraminidase stalk length and additional glycosylation of the hemagglutinin influence the virulence of influenza H5N1 viruses for mice. J. Virol. 83, 4704–4708, 10.1128/JVI.01987-08 (2009).
Zhou, H. et al. The special neuraminidase stalk-motif responsible for increased virulence and pathogenesis of H5N1 influenza A virus. PLoS one 4, e6277, 10.1371/journal.pone.0006277 (2009).
Munier, S. et al. A genetically engineered waterfowl influenza virus with a deletion in the stalk of the neuraminidase has increased virulence for chickens. J. Virol. 84, 940–952, 10.1128/JVI.01581-09 (2010).
Office of the Ministry of Agriculture. The outbreak of highly pathogenic avian influenza in poultry in Shuangcheng district of Harbin city. (2014). (http://www.moa.gov.cn/zwllm/yjgl/yqfb/201409/t20140901_4043080.htm).(Accessed 1 September 2014).
Fang, L. Q. et al. Environmental factors contributing to the spread of H5N1 avian influenza in mainland China. PloS one 3, e2268, 10.1371/journal.pone.0002268 (2008).
Acknowledgements
We thank Peter Wilker for editing the manuscript. This work was supported by the National Key Technologies R&D Program (no. 2013BAD12B04, KJYJ-2013-01-01 and 2012AA022006) and by the Department of Wildlife Conservation and Nature Reserve Management of the State Forestry Administration (SFA), People’s Republic of China.
Author information
Authors and Affiliations
Contributions
Y.G., J.Q., H.C. and X.X. designed the experiments. T.W., Y.L., Y.X., D.C., H.S., C.W., Y.L., Z.X. and W.L. performed samples collected. Z.Y., X.G. and S.L., performed virus isolation and RT-PCR assay. Y.L., S.L. and H.W. performed database searches and sequence analyses. Z.Y. and Y.G. wrote the main manuscript. All authors reviewed the manuscript.”
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0
About this article
Cite this article
Yu, Z., Gao, X., Wang, T. et al. Fatal H5N6 Avian Influenza Virus Infection in a Domestic Cat and Wild Birds in China. Sci Rep 5, 10704 (2015). https://doi.org/10.1038/srep10704
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep10704
This article is cited by
-
H5N1 infection impairs the alveolar epithelial barrier through intercellular junction proteins via Itch-mediated proteasomal degradation
Communications Biology (2022)
-
N-linked glycosylation at site 158 of the HA protein of H5N6 highly pathogenic avian influenza virus is important for viral biological properties and host immune responses
Veterinary Research (2021)
-
Reassortant Highly Pathogenic H5N6 Avian Influenza Virus Containing Low Pathogenic Viral Genes in a Local Live Poultry Market, Vietnam
Current Microbiology (2021)
-
A study of the relationship between human infection with avian influenza a (H5N6) and environmental avian influenza viruses in Fujian, China
BMC Infectious Diseases (2019)
-
Genetic characterization of the first detected human case of avian influenza A (H5N6) in Anhui Province, East China
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.