Abstract
Endogenous retroviruses (ERVs) are remnants of ancient retroviral infections of host germ-line cells. While most ERVs are defective, some are active and express viral proteins. The RD-114 virus is a replication-competent feline ERV and several feline cell lines produce infectious RD-114 viral particles. All domestic cats are considered to have an ERV locus encoding a replication-competent RD-114 virus in their genomes; however, the locus has not been identified. In this study, we investigated RD-114 virus-related proviral loci in genomes of domestic cats and found that none were capable of producing infectious viruses. We also found that all domestic cats have an RD-114 virus-related sequence on chromosome C2, termed RDRS C2a, but populations of the other RDRSs are different depending on the regions where cats live or breed. Our results indicate that RDRS C2a, the oldest RD-114-related provirus, entered the host genome before an ancestor of domestic cats started diverging and the other new RDRSs might have integrated into migrating cats in Europe. We also show that infectious RD-114 virus can be resurrected by the recombination between two non-infectious RDRSs. From these data, we conclude that cats do not harbor infectious RD-114 viral loci in their genomes and RD-114-related viruses invaded cat genomes multiple times.
Similar content being viewed by others
Introduction
Endogenous retroviruses (ERVs) are retroviruses which have been integrated into the host genome of germ-line cells, comprising approximately 10% of the host genome in mammals1. ERVs behave as host genes and are transmitted from parents to offspring in a Mendelian manner. Although most ERVs are inactivated through the accumulation of mutations and deletions, leading to an introduction of stop codons within coding sequences, some ERVs are active and have the potential to produce infectious viral particles.
The RD-114 virus is a replication-competent feline ERV2 and several feline cell lines produce infectious RD-114 viral particles3. The RD-114 virus was first isolated from human rhabdomyosarcoma (RD) cells during passages through fetal cat brains and was mistakenly regarded as the first human retrovirus4. Subsequently, it was revealed that cellular RNA from cats contain sequences homologous to RD-114 viral RNA5,6, leading to the discovery of RD-114 virus-related proviral sequences in the cat genome7,8,9,10. In addition, several groups reported that infectious RD-114 viral particles were produced, either spontaneously or after chemical induction, from cells originating from domestic cats11,12,13. From these data, it was concluded that the RD-114 virus was an active ERV of cats and not an exogenous human retrovirus.
Several research groups have searched for the active RD-114 proviral loci2,14,15. Spodick et al. identified thirteen RD-114 virus-related clones, consisting of conserved gag-pol and unrelated env regions from six domestic cat genomes14. At the same time, another group identified nine endogenous RD-114 virus-related sequence clones representing at least eight distinct loci from cat DNA libraries2. Although the gag-pol region of the clones was conserved, the env region displayed considerable divergence from the replication-competent RD-114 virus clone derived from human RD cells, both in size and sequence homology, coinciding with previous results. In their followup study, feline chromosomes which contain RD-114 virus-related segments were identified with variation of distinct viral segments among individual cats15. These two groups identified the restriction endonuclease-digested genome fragments corresponding to fragments of the infectious RD-114 virus clone in cat genomic DNA and concluded that the active infectious RD-114 viral locus may be a single copy. The former group detected a 4.8-kb PstI-digested pol-env fragment in all cats genomic DNAs examined14 and the latter group detected a 1.0-kb BamHI-digested gag fragment2,15. Although the latter fragment was found to be located on chromosome B315, the locus of the former fragment was not identified. Further it has not been confirmed whether both of them were present on the same single locus as the complete full-length provirus. After these studies, a new endogenous type C retrovirus named Felis catus endogenous retrovirus (FcEV) which consists of the RD-114 virus-related gag-pol gene and unrelated env gene was discovered in all domestic cats genomes examined16. They reported that there were fifteen to twenty FcEVs in domestic cat genomes and suggested that all or most RD-114 virus-related sequences (RDRSs) possessing unrelated env genes are more likely to be FcEV integrations16,17. Therefore, as mentioned above, the RD-114 proviral locus encoding an infectious RD-114 virus has not been identified.
In this study, we searched for loci encoding infectious RD-114 virus. Contrary to our expectations, we could not find any RDRSs capable of producing infectious RD-114 virus, but discovered that a RD-114 virus-related virus (RDRV) nearly identical to the RD-114 virus can be produced by recombination between two defective RD-114 virus-related proviruses. We also noticed that most RDRSs have not been fixed in domestic cat genomes, suggesting multiple invasions of cat genomes by the ancient RDRV. Our study sheds light on host-virus interactions and provides novel genetic markers of domestic cats available for tracing footsteps of cats.
Results
Characterization of RDRS loci in the genomes of domestic cats
Firstly, we searched for RD-114 proviral loci in the genome database of domestic cats (Felis_catus_6.2) using the SSEARCH program18 with the complete genome sequence of RD-114 strain CRT1 (GenBank accession number: AB559882.1)19 as a query; however, we could not identify any proviruses which were identical to the sequences of three previously reported infectious RD-114 virus clones20. Instead, we found one defective RDRS locus on feline chromosome C2 and designated it as RDRS C2a (proviruses were named after the chromosome numbers on which they are located). Quite recently, by using the sequences of RD-114 virus infectious clones as queries, Tamazian et al. reported that there are twelve RD-114 virus-like elements with more than 50% sequence identity, which cover at least 50% of query sequences in domestic cat genomes21. However, their sequences are not available for further analyses. Next, we investigated whether RDRS(s) capable of inducing RD-114 virus are present in all cat genomes by Southern blot analysis. By using HindIII and a gag probe, we detected a 2.6-kb fragment corresponding to the HindIII fragment of RD-114 virus molecular clones in RD-114 virus producer Crandell-Rees feline kidney (CRFK) cells and RD-114 virus non-producer cell lines such as G355-5 (a feline fetal astrocyte cell line), Felis catus whole fetus-4 (fcwf-4) (a feline macrophage-like cell line) and FeT-J (an interleukin-2-independent feline T-cell line) cells (Fig. 1B). By using HindIII and an env probe, we detected a 3.2-kb fragment corresponding to the HindIII fragment of RD-114 virus molecular clones only in CRFK cells but not RD-114 virus non-producer cell lines (Fig. 1D). In accordance with the previous studies2,14,15, there are polymorphisms present in RD-114 virus-related sequences with a conserved env region. By southern blotting hybridization using SacII, EcoRI or SphI and probes for gag, pol and env genes, we revealed that RD-114 virus-related env copy numbers were less than gag-pol copy numbers (Fig. 1B, C, D). Inverse nested PCR targeting RD-114 virus env identified three proviruses (designated as RDRS A2, C2a and C1, respectively) in CRFK cells (Fig. 2 and Fig. S1). We also identified three proviruses (RDRS C2a, E3 and D4) in primary feline peripheral blood mononuclear cells (PBMCs) and two proviruses (RDRS C2a and C2b) in FER cells (feline embryonic fibroblasts that produce RD-114-related virus) (Fig. 2 and Fig. S1). Again, these proviruses were not identical to the sequences of infectious RD-114 virus clones. Four of these proviruses (RDRS A2, C1, D4 and C2b) have the entire open reading frames (ORFs) for env and two of them (RDRS C1 and C2b) have intact ORFs for all viral genes (gag, pol and env) (Fig. 2). We found that the 3.2-kb HindIII fragment containing env could be generated by digestion of four RDRSs (RDRS A2, C1, D4 and C2b) which have the intact ORF for env gene. RDRS C2a of CRFK, G355-5 cells and feline PBMCs (n = 2) have stop codons within all viral genes in the same fashion. Phylogenetic analysis of RDRSs and RD-114 virus clones indicated that RDRS C2a belongs to a separated group from the other RDRSs and infectious RD-114 virus clones (Fig. 3). In feline cell lines, although all lines have RDRS C2a, RD-114 viral producer cell lines do not have any other loci in common (Table 1). These data suggest that a single RDRS is not responsible for the production of RD-114 viral particles. We further investigated cat populations for the possession of RDRSs in their genomes. Similar to the data obtained from cell lines, we also found that all cats have RDRS C2a in common, while the possession patterns of the other RDRS loci varied depending on the regions where cats live or breed (Fig. 4A).
Detection of RD-114 proviruses in the genomes of feline cell lines by Southern blotting analyses.
(A) Restriction endonuclease maps of an infectious molecular clone of RD-114 virus, termed pSc3c. Probes for gag, pol and env genes are shown as bars. (B–D) Southern blotting analyses of several feline cell lines. Genomic DNAs were digested with HindIII, SacII, EcoRI or SphI and then subjected to Southern blotting analyses using gag (B), pol (C) and env (D) probes. HindIII-digested fragments corresponding to pSc3c are shown by white arrowheads. Asterisks indicate RD-114 viral env locus detected in common in all cell lines examined.
Structures of RDRSs.
(A) Structures of the genomes of full length RDRSs. Open triangle indicates a deletion of nucleotides and filled triangle indicates an insertion of nucleotides. Green, orange and blue boxes indicate gag, pol and env ORFs, respectively. Numbers (%) above the diagrams indicate homologies with RD-114 infectious clones in each gene. (B) Schematic LTR structures of RDRSs and RD-114 virus clones, pSc3c and pCRT1. DR-A, direct repeat A; DR-B, direct repeat B; CAAT, CAAT box; TATA, TATA box; TSS, transcription start site; poly A, poly A signal. (C) Features of LTRs of RDRS proviruses and estimated integration time.
The maximum likelihood phylogenetic trees of RDRSs.
We used nucleotide sequences of gag, pol, env and full length excluding LTR. The general time-reversible (GTR) model of nucleotide substitution with the addition of invariant sites (I) and a gamma distribution of rates across sites (Γ) was used to infer the phylogenies. A2, RDRS A2; C2a, RDRS C2a; E3, RDRS E3; D4, RDRS D4; C2b, RDRS C2b; BaEV M7, Baboon endogenous virus strain M7.
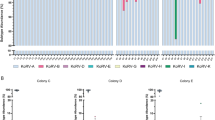
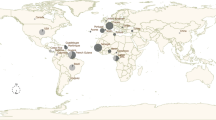
Possession of RD-114 viral infectious-type env and RDRSs.
(A) Population of RDRS proviruses in feline PBMC. Regions that the cats originally lived in were indicated based on a previous study40. (B) Position of reverse primer to detect infectious-type env indicated by blue characters and arrowheads. Red characters are the RDRS C2a-specific insertion. Numbers are defined as positions of pSc3c. (C) Cat's journey and process of RDRV integration. Virus particle with blue hexagonal shape and orange hexagonal indicate the oldest RDRV and the new RDRVs, respectively. The picture of cats and the map were drawn by S.S. using the Microsoft PowerPoint.
Timelines for acquisitions of RDRSs in feline species
We next estimated the integration time of RDRSs. The divergence accumulated between 5′ and 3′ LTRs over time was previously used as a molecular clock to investigate the integration time of ERV22. Thus, we tried to estimate the integration time of the RDRSs by assessing the sequence differences between 5′ and 3′ LTRs. The proposed rate of 1.2 × 10−8 substitutions/site/year in cats was adopted for calculating proviral insertion time23. Four of the six proviruses (RDRS A2, E3, D4 and C2b) have identical 5′ and 3′ LTRs, which are suggestive of relatively recent integrations into the host genome, less than 0.2 million years ago (MYA), possibly by RDRV infection (Fig. 2C). On the other hand, RDRS C1 and C2a have one and eight nucleotide differences, resulting in an estimated time of integration of RDRS C1 and C2a around 0.2 and 1.6 MYA, respectively (Fig. 2C). These periods are after domestic cats separated from the Sand cat lineage24. The time of ERV integration can be also estimated by determining whether phylogenetically closely related animal species share these proviruses in the same genome location. The oldest RDRV provirus C2a was not present in the genome of the Leopard cat (Prionailurus bengalensis) (n = 4) which inhabits Vietnam. Instead, we found a 54-bp sequence within the same locus. Intriguingly, the 54-bp sequence is part of a SINE element (Repbase ID: SINEC_Fc2), predicted by the Repeatmasker program (http://www.repeatmasker.org/) (Fig. 5). The SINE-like sequence is defined by a tRNA-related region, harboring RNA polymerase III-specific internal promoter with its conserved regulatory elements, A-box and B-box, followed by a microsatellite region (TC)n and by an A/T-rich tail with the polyadenylation signal AATAAA25,26. The 54-bp sequence in the leopard cat genome contains the A-box, but the domestic cat's genome lacks this part of the SINE element. In other Felidae, including the Serval cat (Leptailurus serval), Snow leopard (Panthera uncia) and Tiger (Panthera tigris) (whole genome shotgun sequences, accession number: NW_006711997.1), the full length SINE elements were maintained in their genomes at the same location as the leopard cat, suggesting that SINEC_Fc2 integrated into the genome of the common ancestor of Felidae species and then RDRS C2a integration occurred, leading to the disruption of a part of the SINE sequence (Fig. 5). Anai et al. also reported the absence of the RD-114 viral env gene in the Tsushima cat (Prionailurus bengalensis euptilurus)27,28. The domestic cat is estimated to have separated from the leopard cat lineage approximately 6.2 MYA28. We hypothesize that the oldest RDRV C2a entered the ancestor of domestic cats from 6.2 to 1.6 MYA before the diversification of cat breeds.
Comparison of sequences of RDRS C2a integration sites in domestic cat's (Felis catus) and other Felidae species' genomes.
(A) Alignment of sequences of 5′ and 3′flanking regions of RDRS C2a in domestic cat (Felis catus) and their corresponding sites' sequences in leopard cat (P. bengalensis), Serval cat (L. serval), Snow leopard (P. uncia) and Tiger (P. tigris). Asterisks indicate conserved nucleotides between four species. Target site duplications (TSDs) for SINEC_Fc2-like sequence are shaded in black. (B) Schematic view of RDRS C2a integration site and adjacent sequences of Felidae genomes. Lime green arrow indicates RDRS C2a. Pink and pink-red arrows indicate SINEC_Fc2 and C SINEC_Fc2-like sequence, respectively and aqua arrow indicates LINE-like sequences. Short orange lines at both sides indicate primer positions used for PCR.
Integration of RDRS coding for infectious-type env
Because RDRS sequences vary depending on where cats live or breed, it is possible that unidentified RDRS loci are present in domestic cat genomes of breeds investigated above. We developed a PCR method to differentiate infectious-type env on any loci from the defective C2a-type env (Fig. 4B). RDRS C2a has a 15-bp insertion in the env coding region, whereas the other RDRSs do not. We designed a reverse primer to step over the region of the 15-bp insertion present in RDRS C2a (Fig. 4B). As a result, we revealed that 40 and 56% of domestic cats in Europe and North America have infectious-type env, respectively, whereas almost all domestic cats in Asia do not (Fig. 4A). Asian domestic cats containing infectious-type env were mongrel cats and their origins were not identified. Interestingly, none of the Asian cats have any RDRSs harboring infectious-type env identified in this study. Similarly, most Scottish fold cats have RDRS harboring infectious-type env; however, they are unknown RDRSs. These data may indicate that unidentified RDRS(s) with infectious-type env are present in Scottish fold cats. Domestic cats (Felis silvestris catus) originate from the African wildcat (Felis silvestris lybica), a wild cat living in the Middle East and then spread over the world to Europe or a different route to Asia24 (Fig. 4C). Our results suggested that the newer RDRVs (i.e. RDRSs possessing infectious-type env) might have infected domestic cats after cats started moving from the Middle East. There is a possibility that some additional loci with the infectious-type env were acquired in cats through introgression of newer RDRSs into their genomes by crossbreeding between different feline species or cat breeds21,29,30,31. Although the original cat breeds of CRFK, 3201 (feline thymic lymphoma) and FER cells possessing the newer RDRSs are unclear, these cell lines were established in the United States or the United Kingdom32,33,34, suggesting that they originated from cats living in these countries.
Infectivity of RDRSs
Among the RDRS loci, RDRS C1 and C2b have intact ORFs for gag, pol and env genes and RDRS C1 was most similar to the sequences of infectious RD-114 virus clones20. RDRS C1 has five differences, when compared with an infectious RD-114 virus clone, termed pSc3c; LTR has one additional direct repeat (termedΔDR-A), one point mutation at primer binding site (the “a” to “g” change at position 491 of pSc3c) and three point mutations in pol gene (“a” to “g” at 3748, “t” to “c” at 5394 and “a” to “g” at 5719) resulting in amino acid substitutions (Fig. 6A). To examine if RDRS C1 encodes infectious RD-114-like virus, we transfected human embryonic kidney (HEK) 293T cells with a plasmid encoding RDRS C1. We then collected culture supernatants at 48 hours post transfection and inoculated them onto HEK293T cells transduced with the LacZ marker gene [HEK293T(LacZ) cells]. Virus titers produced in the culture supernatants were measured by the LacZ marker rescue assay as described previously35. As a result, we found that RDRS C1 was not infectious. We further evaluated the effects of the three point mutations in the pol region on the infectivity of RDRS C1. We constructed single mutants, which were made by site-directed mutagenesis36. Two mutants were not infectious, whereas one mutant (5394t mutant) with a mutation introduced in the integrase core domain (Pfam accession number: PF00665), predicted by Pfam, (http://pfam.sanger.ac.uk/) had infectivity as an RD-114 virus clone (Fig. 6B and C). From these results, we conclude that the “t” at 5394 is crucial for infectivity for RDRS C1 infectivity. The same mutation in RDRS C1 has also been observed in the sequences of two unrelated cats.
Infectivity of RDRS C1.
(A) Differences between RDRS C1 and an infectious molecular clone, pSc3c. Numbers indicate nucleotide locations at pSc3c. Shaded regions are functional domains of pol region predicted by Pfam (PR, aspartyl protease; RT, reverse transcriptase; RH, RNaseH; IN, integrase core domain [Pfam accession number: PF00077, PF00078, PF00075 and PF00665, respectively]). Differences between RDRS C1 and pSc3c are indicated by vertical lines (point mutations) and filled triangle (insertion). RDRS C1 mutants have mutations marked by crosses. (B, C) LacZ marker rescue assay performed using TE671 cells as target cells. Infection with LacZ pseudotype viruses (RD-114 virus, RDRS C1 wild-type and RDRS C1 mutants) were visualized by X-Gal staining (B) and the virus titers were expressed as f.f.u./ml (C). Assays were performed in triplicate and the data are shown as the mean viral titers ± standard errors.
In addition to CRFK cells, FER cells produce infectious RD-114-related viral particles3. Because FER cells have RDRS C2b which has intact ORFs for gag, pol and env genes (Fig. 2), we suspected that RDRS C2b may encode an infectious RD-114-like virus. RDRS C2b has a 9-bp insertion at the pol coding region, resulting in the insertion of three amino acids in comparison with pSc3c (Fig. S2). We examined the infectivity of RDRS C2b in the same way as described above. Consistent with results of the other RDRSs identified in this study, we found that RDRS C2b was not infectious (Fig. S2).
Recombination of RDRSs
Because we could not find any loci which encode infectious RD-114 virus, we hypothesized that recombination between RDRSs resulted in the production of infectious RD-114 viral particles. We attempted to reproduce the recombination event by transfection of two RDRS plasmids. Because CRFK cells, from which infectious RD-114 virus is produced, have RDRS A2, C2a and C1 loci, we suspected that an infectious RDRV might be generated by recombination between RDRS A2 and C1. We transfected two plasmids encoding RDRS A2 and C1 and succeeded in recovering replication-competent viral particles (Fig. 7A). By comparing sequences of RDRS A2 and C1 and the recombinant RDRV obtained by transfection of RDRS A2 and C1 (hereinafter referred to as RDRV AC), it was suggested that three changes had occurred (two in the middle of pol and one in pol-env region) (Fig. 7B). Of the two single nucleotide differences between RDRV AC and pSc3c at positions 3224 and 4357, only the “c” to “a” at 3224 results in an amino acid substitution. The amino acid difference was not present in the predicted functional domains of Pol. The length of LTR of RDRV AC was 27 bp shorter than that of the recombination origin RDRS C1 at the repeat region (Fig. 7C). The 27-bp sequence was repeated in the U3 region of LTR in pSc3c, RDRS A2 and C1 and termed direct repeat A1 (DR-A1). Altogether, infectious RD-114 virus-like particles can be generated by recombination between defective RDRS A2 and C1.
Recombination between RDRS A2 and C1.
(A) Growth of RDRV AC in HEK293T cells. The RDRS plasmids were transfected into HEK293T cells and supernatants were inoculated into HEK293T(LacZ) cells and then virus production was monitored by the LacZ marker rescue assay. Assays were performed in triplicate and the data are shown as the mean viral titers ± standard errors. (B) Comparison of nucleotide sequences of pSc3c, RDRS A2, C1 and RDRV AC virus. The nucleotide positions of pSc3c are shown. Dark grey arrowheads indicate positions of primers used for PCR. Small letters indicate nucleotides. Capital letters in parentheses indicate amino acids. Dots indicate differences between RDRV AC and the other clones. Colors of dots are dependent on where are mutations from (Red, derived from A2; Blue, from C1; Gray, from A2 or C1; Black, undetermined). (C) LTR structures of RD-114 virus clones (pSc3c and pCRT1), RDRS A2, C1 and RDRV AC.
Discussion
The RD-114 virus was originally isolated from human RD cells after passages in fetal cat brains4. Because the parent cell line (RD cells) does not contain the RD-114 virus genome, there has been speculation about the origin of the RD-114 virus. In previous studies, restriction endonuclease map analysis and Southern blot hybridization of cat cellular DNA revealed that there are approximately twenty RDRSs in domestic cat genomes but most have a deleted-env region2,14,15. The 4.8 kb PstI-digested fragment corresponding to a complete env region was detected in all cat DNAs examined as a single copy in the previous study; however, they did not isolate a clone containing a complete RD-114 provirus14. We confirmed that the PstI-digested env fragments derived from RDRS C2a is similar to infectious RD-114 virus clones, pSc3c and pCRT1, in size (4.7 kb) with high sequence homology (95.7%, data not shown). These data suggest that the single locus in all cat genomes which Spodick et al. identified was in fact RDRS C2a and not the complete infectious RD-114 provirus. Another group identified eight RDRSs (Ren 18c, 20a, 8a, 7c, 10a and 6c) in domestic cat genomes2 and found that the 1.0-kb BamHI gag fragment specific to the RD-114 virus clones was present in cat genomes as a single copy on chromosome B3; however, domestic cats are polymorphic with respect to the presence or absence of this RD-114 virus-related segment2,15. Although it has been proposed that all domestic cats harbor infectious RD-114 provirus in their genomes, the proviral locus has not been identified even after extensive mining of feline genome databases.
In this study, we identified six RDRSs in domestic cats genomes. These proviruses are not identical to the sequences of infectious molecular clones of RD-114 virus reported previously20. RDRS C1, whose sequence is most similar to those of RD-114 infectious clones, could not produce infectious viral particles. The defect was ascribed to a single nucleotide mutation resulting in an amino acid substitution in the integrase core domain which plays a key role in catalysis37. The sequences of RDRS LTRs are rich in diversity, suggesting that each RDRS entered the host genome at different time points. Five of the RDRSs (excluding RDRS C2a) have identical 6-bp target site duplications (TSDs) (Table 2) and there were no differences between 5′ and 3′ LTRs in RDRSs A2, C2b and E3. These data suggest that LTR-LTR recombination did not occur and RDRS invasions into cat genomes occurred relatively recently. Except for RDRS C2a, RDRSs have diverse TSDs, indicating that multiplication of RDRSs occurred through viral re-infection but not genome rearrangements. The mechanism of multiple invasions of RDRVs is unknown at present. Generally, it is considered that integration of ERVs into host genomes confers resistance to the infection of exogenous counterparts38. However, it is possible that the expression of newly integrated RDRS is tightly regulated in cats, especially in germ-line cells, to allow multiple integrations of RDRV in cats.
Remarkably, number of RDRS proviral copies vary considerably with type of cell lines and cat breeds. All cats have RDRS C2a in common, but all proviral genes (gag, pol and env) are disrupted in RDRS C2a. The other RDRSs have not yet been fixed in domestic cat genomes. Furthermore, we found that most Asian cats are free from new RDRSs harboring infectious-type env.
Because domestic cats originated from the African wildcat living in the Middle East (Felis silvestris lybica)24, we propose the following scenario (Fig. 4C) for RDRV integration into domestic cat genomes in Europe and Asia. (i) Firstly the ancestral virus of RDRS C2a (RDRV C2a) infected the domestic cat progenitor. (ii) The African wildcats were domesticated in the Middle East where agriculture started around 10,000 years ago. (iii) Then, some domesticated cats moved to Europe and America as ship's cats with Vikings or traders and became common throughout Europe39. Ship's cats could travel to different harbors for chance encounters with wildcats for interbreeding39, diversifying the genomes of their offspring but at the same time putting themselves at risk for new RDRV infection and subsequent integration by infectious-type RDRSs. (iv) The other cats went to Asia as guardians of Buddhist shariha along well-established trade routes, termed the Silk Road39. With no native wildcats that lived near this road to interbreed with, Oriental domestic cats soon began evolving in their own way39 without the risk of infection by new RDRVs.
British Shorthair (BSH) was established in England from European Shorthair which first appeared in Sweden and later American Shorthair (ASH) and American Curl (a variant of ASH) originated from BSH40. Some of these cat breeds have RDRS E3 but others do not. These data indicate that RDRS E3 would be a useful genetic marker to trace the footsteps of these breeds from Europe to America. In future studies, we will investigate the presence of RDRSs in domestic cats living in various regions as well as closely related wild cats to further map cat migration.
Several feline cell lines including CRFK and FER cells produce RD-114 or RD-114-related viruses, but there are no proviruses identical to previously reported RD-114 viral sequences. Several novel retroviruses have been reported to be generated by the recombination of ERVs41,42,43. Therefore, we hypothesized that RD-114 virus-like particles were generated by the recombination between two defective RDRS proviruses. By co-transfection of two defective RDRS A2 and C1, we demonstrated that infectious RD-114-like viral particles were generated through recombination. The resultant recombinant, RDRV AC and infectious clones of RD-114 virus were nearly identical in coding sequences; however, the LTR of RDRV AC was not identical to those of the infectious clones, pCRT1 and pSc3c. The changes in LTR length frequently occurred in repeat sequences during cell passages in vitro as observed in other gammaretroviruses such as porcine endogenous retrovirus44,45. FER cells also produce RD-114-related viral particles3, despite the lack of RDRS A2 and C1 in their genomes (Table 1). Similar to the case of CRFK cells, we speculated that an RD-114-related virus could have been generated by recombination between RDRS C2b which has the infectious-type env and some other unidentified RDRSs or FcEV.
Retroviral recombination events occur by template switching during reverse transcription46,47. Xenotropic murine leukemia virus-related virus (XMRV) is one such virus which has arisen through the recombination of two defective murine ERVs during passages of human prostate cancer xenografts in immunocompromised nude mice48. The defective viruses recombined in human cells but not in mouse cells because the XMRV is xenotropic. Similarly, the first RD-114 virus may have arisen by recombination in human RD xenografts in cat brains4,8. RD-114-related viruses have also been induced by chemicals in feline cells without xenotransplantation11,12,13, where a RD-114-related virus may have arisen in feline cells without contact with cells from other species. In these cases, defective viral particles co-packaged with RDRS genome infected feline cells and recombined during reverse transcription. Resultant replication-competent viruses may have expanded in the feline cells, because feline cells are permissive to RD-114 virus3,49. Additionally, we should consider a possibility that RDRS C1 produces infectious viral particles by spontaneous mutation like Emv2 in C57BL mice50,51,52. Emv2 has a defect in the pol gene, compromising the ability of the Emv2 to produce infectious virus particles53. However, Emv2 can produce an infectious endogenous ecotropic murine leukemia virus (E-MLV) through the acquisition of a spontaneous mutation in the pol gene in vivo in aging C57BL mice50 or by backcrossing between C57BL and E-MLV-negative mice51. Therefore, we cannot exclude the possibility that RDRS C1 produces infectious viral particles through spontaneous mutations in vivo.
In the previous study, we revealed ubiquitous expression of the RD-114 viral receptor, termed ASCT2, in domestic cat tissues49, suggesting that RD-114 viral recombination events may occur in cats and the infectious RD-114 virus may be generated in some groups of cats having RDRSs harboring infectious-type env. Recently, it was reported that in immune-compromised mice, ERVs in the host genome were resurrected and induced lymphoma54,55. In these cases, receptors for infectious ERVs are expressed in tissues of mice and infectious ERVs re-infected the host cells and induced lymphoma possibly via insertion of the ERVs in the vicinity of cellular proto-oncogenes54,55. In this study, most RDRSs are not yet fixed in cat genomes, but some of them have the potential to generate a recombinant virus which could re-infect the host cells. This recombinant virus may re-infect cat cells which do not express RDRS Env, leading to new integrations. Some of the new integrations may induce diseases in cats, if the integration occurred in the vicinity of proto-oncogenes. While it is still unknown whether RD-114 virus causes any diseases in cats, our report that RD-114 virus can be generated by recombination will be useful for further investigation of the pathogenicity of this virus.
Methods
Ethics statement
All animal care and procedures that were performed in this study conformed to the guidelines for animal experiments at Institute for Virus Research in Kyoto University (IVRKU) and all experimental protocols were approved by the Committee on the Ethics of Animal Experiments of IVRKU. Blood samples of domestic cats were collected for diagnostic purpose in veterinary clinics. Genomic DNAs of Leopard cats, serval and snow leopard were stored samples which had been used in our previous studies56,57.
Blood and tissue samples
Domestic cat blood samples for isolation of PBMCs were provided from Fujimura Animal Hospital (Osaka, Japan), Kyoritsu Seiyaku Corporation (Tokyo, Japan) and Veterinary Clinics in Japan. The Snow leopard and Serval cat tissue samples were provided from Sapporo Maruyama Zoo (Hokkaido, Japan). These animals died from feline parvovirus infection57.
Cell cultures
FeT-J and 3201 cells were cultured in RPMI 1640 medium (Sigma-Aldrich, Tokyo, Japan) supplemented with 10% heat-inactivated fetal calf serum (FCS), penicillin (100 IU/ml) and streptomycin (100 μg/ml) (Invitrogen, Carlsbad, CA). CRFK, G355-5, fcwf-4, FER, AH927 (feline embryonic fibroblast), HEK293T and TE671 (human rhabdomyosarcoma) cells were cultured in Dulbecco's modified Eagle's medium (DMEM) (Sigma-Aldrich) supplemented with 10% FCS, penicillin (100 IU/ml) and streptomycin (100 μg/ml) (Invitrogen).
Genomic DNA isolation and genomic PCR
Genomic DNA was extracted from cell lines and PBMCs using a QIAamp DNA Blood Mini Kit (QIAGEN, Valencia, CA). Genomic PCR was performed by using PrimeSTAR GXL polymerase (TaKaRa, Shiga, Japan) or Ex Taq polymerase (TaKaRa). The former was used for cloning RDRSs and the recombinant RDRV and the later was used for screening for the presence of RDRSs. Amplicons were analyzed by electrophoresis in 0.8% or 2% agarose gels. Primers used in this study are listed in Table S1.
Southern blotting
A total of 5 μg (or 3 μg) genomic DNA was isolated from PBMCs or cell lines, digested with HindIII for detection of gag and env genes, SacII for detection of gag and pol genes, EcoRI for detection of env genes, or SphI for detection of gag and env genes. Digested fragments were separated by a 1.0% agarose gel and transferred to positive charged-nylon membrane (Roche Diagnostics, Indianapolis, IN). Hybridization was performed using digoxigenin (DIG)-labeled PCR probes and DIG easy Hyb (Roche) at appropriate temperatures (50°C for gag and pol, 65°C for env detection) for 16 hours. After hybridization, the membrane was washed with a low stringency buffer (0.1% SDS and 2 × SSC) at room temperature followed by a high stringency buffer (0.1% SDS and 0.1 × SSC) at 68°C. The hybridized bands were visualized using CDP-Star reagent (Roche) following the manufacturer's instructions. Probes were prepared from an infectious molecular clone of RD-114 virus, termed pCRT119, using DIG-labeled PCR probe synthesis kit (Roche) according to the manufacturer's instructions with primers used listed in Table S1.
Inverse PCR
Genomic DNA was digested with EcoRI or SphI and 100 ng of digested genomes were ligated by T4 ligase (TOYOBO, Osaka, Japan) at 16°C overnight. Ligated DNA was amplified with specific primers listed in Table S1 using PrimeSTAR GXL polymerase (TaKaRa). Second PCR products were cloned into a pCR4 Blunt TOPO vector (Invitrogen) and sequenced. Loci of RDRSs were determined by comparing flanking sequences with the genome database of domestic cats (Felis_catus_6.2).
Cloning of RDRSs
Full-length RDRSs were amplified using specific primers (listed in Table S1) on flanking regions and amplicons were cloned into pCR BluntII TOPO vector (Invitrogen).
Infection assay
HEK293T cells were transfected with 4 μg of RD-114 virus molecular clones or plasmids containing RDRSs using Lipofectamine2000 (Invitrogen). The pcDNA3.1 vector was used for mock transfection. Infectivities of RDRSs were examined by the LacZ marker rescue assay28. Culture supernatants were collected 48 hours post-transfection and filtrated through 0.45 μm-pore-size filters and then inoculated onto HEK293T(LacZ) cells with 8 μg/ml polybrene. After inoculation, supernatants were collected every four days. TE671 cells were seeded in 96-well plates at 1 × 104 cells per well one day before infection and diluted supernatants of inoculated HEK293T(LacZ) cells were inoculated onto TE671 cells in the presence of 8 μg/ml polybrene. Two days after inoculation, the inoculated cells were stained with X-Gal and virus titers, expressed as focus forming units (f.f.u.)/ml, were determined as described previously35,47. Assays were conducted in triplicate for each individual sample.
Preparation of RDRS C1 mutants
RDRS C1 mutants were made by DpnI method36. PCR was performed using PrimeSTAR GXL Polymerase (TaKaRa) with primers listed in Table S1. RDRS C1 plasmid DNA was used as template. Amplicons were digested with DpnI to eliminate the parental plasmid DNA. Digested DNA fragments were purified and transformed into Escherichia coli DH5 alpha.
RDRV AC isolation
HEK293T cells were transfected with 4 μg of pCRT1 or plasmids containing RDRSs using Lipofectamine2000 (Invitrogen). pcDNA3.1 vector (Invitrogen) was used for mock transfection. Culture supernatants were collected 48 hours post-transfection and filtrated through 0.45 μm-pore-size filters and then inoculated into HEK293T cells with 8 μg/ml polybrene (Sigma-Aldrich). After inoculation, supernatants were collected and filtrated every four days. TE671 cells were seeded in 6-well plates at 1 × 106 cells per well one day before infection and diluted supernatants of the inoculated HEK293T cells (36 d.p.i.) were inoculated onto TE671 cells in the presence of 8 μg/ml polybrene. Six hours after inoculation, genomic DNAs were extracted and proviruses were amplified by PCR. PCR was performed with PrimeSTAR GXL polymerase (TaKaRa) and primers listed in Table S1, according to the manufacturer's instructions. The forward primer was designed in the R region of 5′LTR and reverse primer was designed in the U5 region of 3′LTR (positions of primers were indicated in Fig. 7).
Sequence Analyses
Sequencing was performed by a commercial DNA sequencing service (FASMAC, Kanagawa, Japan). Alignment of nucleotide sequences was computed using L-INS-i program in MAFFT version 758 and their phylogeny was inferred by RAxML version 7.2.6 that is based on the maximum likelihood method59.
GenBank accession number
RDRS A2, C2a, C1, E3, D4 and C2b are deposited in GenBank as LC005744- LC005749.
References
Gifford, R. & Tristem, M. The evolution, distribution and diversity of endogenous retroviruses. Virus Genes 26, 291–315 (2003).
Reeves, R. H. & O'Brien, S. J. Molecular genetic characterization of the RD-114 gene family of endogenous feline retroviral sequences. J Virol 52, 164–171 (1984).
Okada, M., Yoshikawa, R., Shojima, T., Baba, K. & Miyazawa, T. Susceptibility and production of a feline endogenous retrovirus (RD-114 virus) in various feline cell lines. Virus Res 155, 268–273 (2011).
McAllister, R. M. et al. C-type virus released from cultured human rhabdomyosarcoma cells. Nat New Biol 235, 3–6 (1972).
Okabe, H., Gilden, R. V. & Hatanaka, M. Extensive homology of RD114 virus DNA with RNA of feline cell origin. Nat New Biol 244, 54–56 (1973).
Okabe, H., Gilden, R. V. & Hatanaka, M. RD 114 virus-specific sequences in feline cellular RNA: detection and characterization. J Virol 12, 984–994 (1973).
Baluda, M. A. & Roy-Burman, P. Partial characterization of RD114 virus by DNA-RNA hybridization studies. Nat New Biol 244, 59–62 (1973).
Neiman, P. E. Measurement of RD114 virus nucleotide sequences in feline cellular DNA. Nat New Biol 244, 62–64 (1973).
Gillespie, D., Gillespie, S., Gallo, R. C., East, J. L. & Dmochowski, L. Genetic origin of RD114 and other RNA tumour viruses assayed by molecular hybridization. Nat New Biol 244, 51–54 (1973).
Benveniste, R. E. & Todaro, G. J. Segregation of RD-114 and FeL-V-related sequences in crosses between domestic cat and leopard cat. Nature 257, 506–508 (1975).
Livingston, D. M. & Todaro, G. J. Endogenous type C virus from a cat cell clone with properties distinct from previously described feline type C virus. Virology 53, 142–151 (1973).
Fischinger, P. J., Peebles, P. T., Nomura, S. & Haapala, D. K. Isolation of RD-114-like oncornavirus from a cat cell line. J Virol 11, 978–985 (1973).
Todaro, G. J., Benveniste, R. E., Lieber, M. M. & Livingston, D. M. Infectious type C viruses released by normal cat embryo cells. Virology 55, 506–515 (1973).
Spodick, D. A., Soe, L. H. & Roy-Burman, P. Genetic analysis of the feline RD-114 retrovirus-related endogenous elements. Virus Res 1, 543–555 (1984).
Reeves, R. H., Nash, W. G. & O'Brien, S. J. Genetic mapping of endogenous RD-114 retroviral sequences of domestic cats. J Virol 56, 303–306 (1985).
van der Kuyl, A. C., Dekker, J. T. & Goudsmit, J. Discovery of a new endogenous type C retrovirus (FcEV) in cats: Evidence for RD-114 being an FcEVGag-Pol/baboon endogenous virus BaEVEnv recombinant. J Virol 73, 7994–8002 (1999).
Kim, F. J., Battini, J. L., Manel, N. & Sitbon, M. Emergence of vertebrate retroviruses and envelope capture. Virology 318, 183–191 (2004).
Pearson, W. R. Searching protein sequence libraries: comparison of the sensitivity and selectivity of the Smith-Waterman and FASTA algorithms. Genomics 11, 635–650 (1991).
Yoshikawa, R., Sato, E., Igarashi, T. & Miyazawa, T. Characterization of RD-114 virus isolated from a commercial canine vaccine manufactured using CRFK cells. J Clin Microbiol 48, 3366–3369 (2010).
Shimode, S. et al. Sequence comparison of three infectious molecular clones of RD-114 virus. Virus Genes 45, 393–397 (2012).
Tamazian, G. et al. Annotated features of domestic cat - Felis catus genome. Gigascience 3, 13 (2014).
Martins, H. & Villesen, P. Improved integration time estimation of endogenous retroviruses with phylogenetic data. PLoS One 6, e14745 (2011).
Lopez, J. V., Culver, M., Stephens, J. C., Johnson, W. E. & O'Brien, S. J. Rates of nuclear and cytoplasmic mitochondrial DNA sequence divergence in mammals. Mol Biol Evol 14, 277–286 (1997).
Driscoll, C. A. et al. The Near Eastern origin of cat domestication. Science 317, 519–523 (2007).
Okada, N. SINEs: Short interspersed repeated elements of the eukaryotic genome. Trends Ecol Evol 6, 358–361 (1991).
Okada, N., Hamada, M., Ogiwara, I. & Ohshima, K. SINEs and LINEs share common 3′ sequences: a review. Gene 205, 229–243 (1997).
Anai, Y. et al. Infectious endogenous retroviruses in cats and emergence of recombinant viruses. J Virol 86, 8634–8644 (2012).
Johnson, W. E. et al. The late Miocene radiation of modern Felidae: a genetic assessment. Science 311, 73–77 (2006).
Beaumont, M. et al. Genetic diversity and introgression in the Scottish wildcat. Mol Ecol 10, 319–336 (2001).
Randi, E., Pierpaoloni, M., Beaumont, M., Ragni, B. & Sforzi, A. Genetic identification of wild and domestic cats (Felis silvestris) and their hybrids using Bayesian clustering methods. Mol Biol Evol 18, 1679–1693 (2001).
Oliveira, R., Godinho, R., Randi, E., Ferrand, N. & Alves, P. C. Molecular analysis of hybridisation between wild and domestic cats (Felis silvestris) in Portugal: implications for conservation. Conservation Genetics 9, 1–11 (2008).
Crandell, R. A., Fabricant, C. G. & Nelson-Ress, W. A. Development, characterization and viral susceptibility of a feline (Felis catus) renal cell line (CRFK). In Vitro 9, 176–185 (1973).
Rojko, J. L. et al. Feline lymphomas: immunological and cytochemical characterization. Nature 19, 656–658 (1978).
Jarrett, O., Laid, H. M. & Hay, D. Determinants of the host range of feline leukemia viruses. J Gen Virol 20, 169–175 (1973).
Sakaguchi, S., Okada, M., Shojima, T., Baba, K. & Miyazawa, T. Establishment of a LacZ marker rescue assay to detect infectious RD114 virus. J Vet Med Sci 70, 785–790 (2008).
Li, S. & Wilkinson, M. F. Site-directed mutagenesis: a two-step method using PCR and DpnI. Biotechniques 23, 588–590 (1997).
Craigie, R. HIV integrase, a brief overview from chemistry to therapeutics. J Biol Chem 276, 23213–23216 (2001).
Palmarini, M., Mura, M. & Spencer, T. E. Endogenous betaretroviruses of sheep: teaching new lessons in retroviral interference and adaptation. J Gen Virol 85, 1–13 (2004).
Driscoll, C. A., Clutton-Brock, J., Kitchener, A. C. & O'Brien, S. J. The Taming of the cat. Genetic and archaeological findings hint that wildcats became housecats earlier--and in a different place--than previously thought. Sci Am 300, 68–75 (2009).
Menotti-Raymond, M. et al. Patterns of molecular genetic variation among cat breeds. Genomics 91, 1–11 (2008).
Henzy, J. E., Gifford, R. J., Johnson, W. E. & Coffin, J. M. A novel recombinant retrovirus in the genomes of modern birds combines features of avian and mammalian retroviruses. J Virol 88, 2398–2405 (2014).
Delviks-Frankenberry, K. et al. Generation of multiple replication-competent retroviruses through recombination between PreXMRV-1 and PreXMRV-2. J Virol 87, 11525–11537 (2013).
Paprotka, T. et al. Recombinant origin of the retrovirus XMRV. Science 333, 97–101 (2011).
Denner, J., Specke, V., Thiesen, U., Karlas, A. & Kurth, R. Genetic alterations of the long terminal repeat of an ecotropic porcine endogenous retrovirus during passage in human cells. Virology 314, 125–133 (2003).
Denner, J. Recombinant porcine endogenous retroviruses (PERV-A/C): a new risk for xenotransplantation? Arch Virol 153, 1421–1426 (2008).
Delviks-Frankenberry, K. A., Nikolenko, G. N. & Pathak, V. K. The "Connection" Between HIV Drug Resistance and RNase H. Viruses 2, 1476–1503 (2010).
Takeuchi, Y. et al. Host range and interference studies of three classes of pig endogenous retrovirus. J Virol 72, 9986–9991 (1998).
Arias, M. & Fan, H. The saga of XMRV: a virus that infects human cells but is not a human virus. Emerg Microbes Infect 3, e25 (2014).
Shimode, S., Nakaoka, R., Shogen, H. & Miyazawa, T. Characterization of feline ASCT1 and ASCT2 as RD-114 virus receptor. J Gen Virol 94, 1608–1612 (2013).
Odaka, T. Genetic transmission of endogenous N- and B-tropic murine leukemia viruses in low-leukemic strain C57BL/6. J Virol 15, 332–337 (1975).
Kozak, C. A. & Rowe, W. P. Genetic mapping of ecotropic murine leukemia virus-inducing loci in six inbred strains. J Exp Med 155, 524–534 (1982).
McCubrey, J. & Risser, R. Genetic interactions in the spontaneous production of endogenous murine leukemia virus in low leukemic mouse strains. J Exp Med 156, 337–349 (1982).
King, S. R., Berson, B. J. & Risser, R. Mechanism of interaction between endogenous ecotropic murine leukemia viruses in (BALB/c X C57BL/6) hybrid cells. Virology 162, 1–11 (1988).
Young, G. R. et al. Resurrection of endogenous retroviruses in antibody-deficient mice. Nature 491, 774–778 (2012).
Yu, P. et al. Nucleic acid-sensing Toll-like receptors are essential for the control of endogenous retrovirus viremia and ERV-induced tumors. Immunity 37, 867–879 (2012).
Nakamura, K. et al. Comparison of prevalence of feline herpesvirus type 1, calicivirus and parvovirus infections in domestic and leopard cats in Vietnam. J Vet Med Sci 61, 1313–1315 (1999).
Sassa, Y. et al. Successive deaths of a captive snow leopard (Uncia uncia) and a serval (Leptailurus serval) by infection with feline panleukopenia virus at Sapporo Maruyama Zoo. J Vet Med Sci 73, 491–494 (2011).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30, 772–780. (2013).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Acknowledgements
We are grateful to Dr. Masato Fujimura (Fujimura Animal Hospital) and Dr. Kazuo Kawakami (Kyoritsu Seiyaku Corporation) for providing domestic cat blood samples, Dr. Yasuhiro Takeuchi (University College London, London, U.K.) for providing TELCeB6/pRDF cells and TE671 cells chronically infected with RD-114 virus and Dr. Kenji Baba (Yamaguchi University, Yamaguchi, Japan) for helpful advice. We thank Peter Gee (Kyoto University, Kyoto, Japan) for his generous help in the preparation of the manuscript. This work was supported by JSPS KAKENHI Grant Numbers 265589 and 25891023 (to S.S. and S.N., respectively).
Author information
Authors and Affiliations
Contributions
S.S. and T.M. designed the experiments. S.S. performed the experiments. S.S., S.N. and T.M. analyzed data and wrote the manuscript. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Shimode, S., Nakagawa, S. & Miyazawa, T. Multiple invasions of an infectious retrovirus in cat genomes. Sci Rep 5, 8164 (2015). https://doi.org/10.1038/srep08164
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08164
This article is cited by
-
Establishment of CRFK cells for vaccine production by inactivating endogenous retrovirus with TALEN technology
Scientific Reports (2022)
-
The origin of porcine endogenous retroviruses (PERVs)
Archives of Virology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.