Abstract
Recent studies show that targeting gene promoter or 3′ terminal regions of mRNA with siRNA induces target gene transcription. However, the ability of exon-targeting siRNA to affect transcription has yet to be demonstrated. We designed and synthesized siRNA against various exons in the gelsolin gene (GSN) to knockdown GSN transcript in KYSE150 and KYSE450 cells. Surprisingly, we found that siGSN-2, targeting the GSN twelfth exon, induced GSN gene transcription detected by real time RT-PCR. An siGSN-2 co-precipitation assay was performed and H3 histone, previously shown to correlate with gene transcription, was detected in the siGSN-2 pull-down pellet. However, H3 histone was not detected in an siGSN-1-precipitated pellet, which resulted in GSN knockdown. In addition, siGSN-2 decreased stress fibers, lamellipodia and filopodia, demonstrating that siGSN-2 induced GSN transcription activation and exerted biological function. In conclusion, our finds reveal siRNA, which is derived from target gene exon, can form the complex with H3 histone to be involved in the regulation of gene expression.
Similar content being viewed by others
Introduction
Small dsRNAs, including siRNA and microRNA, down-regulate homologous mRNA in cells to result in target gene post-transcriptional silencing1. The target gene mRNA template is cleaved and degraded via the Dicer endoribonuclease activated by the RNA-induced silencing complex (RISC) composed of siRNA and multiple proteins, including SND1, AEG-1, Ago and other proteins2,3. MicroRNA binding to RISC results in gene mRNA degradation if its seed region completely matches the mRNA template, or it inhibits gene mRNA translation4,5. However, some small dsRNAs, known as RNA activation (RNAa), up-regulate gene transcription by mechanisms distinct from RNAi. For example, an siRNA targeting CpG-rich regions or CpG islands in gene promoters activates gene expression6. Li et al. showed an siRNA targeting a region outside gene promoter CpG islands also adheres to RNAi rules activating gene transcription7 and an endogenous microRNA-373 induces expression of genes with complementary promoter sequences8. Yue et al. identified a non-coding transcript that overlaps the 3′ terminus of the progesterone receptor (PR) gene and found that small RNAs complementary to sequences beyond the 3′ terminus of PR mRNA modulate PR expression by recruiting Argonaute 2 to the non-coding transcript to further induce chromatin changes at the PR promoter9.
Gelsolin, encoded by the GSN gene, can sever, cap and nucleate actin filaments and thus plays a critical role in maintaining the normal cellular motility and morphology10,11,12. Gelsolin is a downstream effector of RAC that regulates organization of actin filaments and gelsolin knockdown induces RAC overexpression and increases amounts of F-actin in stress fibers and lamellipodia13,14. However, gesolin up-regulation inhibits RAC expression and reduces amounts of F-actin15. Gelsolin plays a key role as a tumor suppressor in human bladder cancer cells16. In the early stages of both human and animal tumor carcinogenesis, down-regulation of gelsolin has been widely observed17. Gelsolin down-regulation may be the mechanism through which tumor cells evade apoptotic signaling pathways18. Gelsolin may later become up-regulated to enhance metastasis. Therefore, gelsolin plays a key role in human cancer cells. However, the delineating biological function of gelsolin in human cancer cells is still unclear.
In the present study, during examination of the biological function of gelsolin, we designed several siRNAs targeting various exons in the GSN gene and found that an siRNA targeting the eleventh exon decreased GSN transcription. However, an siRNA targeting the twelfth exon caused GSN transcription up-regulation. We investigated the possibility that the twelfth exon by siRNA targeting activates gene transcription through mechanisms distinct from those of targeting gene promoters or 3′ terminal regions of mRNA.
Results
Up-regulation of GSN expression by siGSN-2
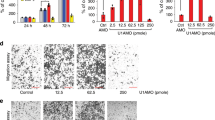
During the process of exploring endogenous GSN protein biological functions, we tempted to make GSN level knockdown by RNA interference. Therefore, we designed three siRNAs and the target sequence of siGSN-1, siGSN-2 or siGSN-3 against GSN exon was individually depicted (Figure 1a). 2 μg siGSN-1, siGSN-2 or siGSN-3 was individually transfected into KYSE150 cells cultured in 6-well plate and GSN level was detected by real time RT-PCR. Transfection of siGSN-1 resulted in GSN down-regulation. However, siGSN-2 transfection up-regulated GSN expression. siGSN-3 transfection did not alter GSN transcript expression. To confirm that siGSN-2 results in GSN up-regulation, we increased the siGSN-2 amount to 4 μg to treat KYSE150 cells cultured in 6-well plate. At this siRNA concentration, GSN was still up-regulated (Figure 1b). To further demonstrate that siGSN-2 induced GSN expression up-regulation, siGSN-2 was added to KYSE450 cells cultured in 6-well plate and siGSN-1 was selected as control. The result showed that GSN was down-regulated with 2 μg siGSN-1, whereas GSN was up-regulated with 2 μg siGSN-2. Similarly, 4 μg siGSN-2 up-regulated GSN compared to the scramble (Figure 1c).
siGSN-2-mediated up-regulation of GSN expression involves transcription
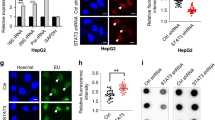
Since siGSN-2 induced GSN up-regulation, we first speculated that siGSN-2 was involved in GSN transcription. Therefore, we transfected cells with biotin-labeled siGSN-2 and pulled down the potential siGSN-2-containing protein complexes (Figure 2a). To determine if transfection with biotin-labeled siGSN-1 still resulted in GSN knockdown and biotin-labeled siGSN-2 resulted in GSN expression up-regulation, GSN mRNA was detected by real time RT-PCR at 48 h post-transfection. The results showed that GSN was down-regulated in siGSN-1-1- or siGSN-1-2- treated KYSE450 cells, respectively. However, GSN was up-regulated in siGSN-2-1 or siGSN-2-2 treated KSYE450 cells, respectively (Figure 2b). The results demonstrated that biotin labeling did not significantly alter the effects of our siRNAs on GSN expression. A previous study showed that histone H3 played an important role during gene transcription20. Therefore, we first speculated that histone H3 might exist in the biotin pull-down complexes from nuclear and cytoplasmic extracts. Histone H3 was co-precipitated with siGSN-2-1 and siGSN-2-2 in nuclear extracts. However, histone H3 was not detected in the cytoplasmic extract. Following transfection and pull down of complexes after siGSN-1-biotin or scramble-biotin transfection, histone H3 was not detected from either nuclear or cytoplasmic extracts (Figure 2c).
Redistribution of the actin cytoskeleton is inhibited by siGSN-2
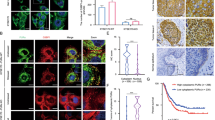
Since GSN up-regulation results in a reduction of F-actin filaments15, to further demonstrate siGSN-2 involvement in up-regulation of GSN expression, siGSN-2 was transfected into cells and then F-actin was observed. We observed that siGSN-2 inhibited F-actin redistribution and made stress fibers formation impeded and meanwhile, made lamellipodia and filopodia formation become less (Figure 3a). The result of western blot analysis further demonstrated that GSN protein was slightly up-regulated with siGSN-2 transfection (Figure 3b).
Discussion
Although promoter-targeting and 3′UTR-targeting siRNAs have been shown to be capable of transcriptional activation7,9, there are still no reports of exon-targeting siRNAs with similar capability. In this present study, we report that siGSN-2 targeting the twelfth exon of gene GSN up-regulates GSN transcript.
The RNA-induced silencing complex (RISC), which is a multiprotein complex composed of a bound siRNA and the RNA endoribonuclease Dicer complex, induces target gene transcript degradation21. In the present study, siGSN-1 induces GSN transcript knockdown and forms a RISC complex distinct from the complex composed of siGSN-2 and accessory proteins that induce GSN transcription up-regulation. Lysine methylation of H3K4, H3K36 and H3K79 correlates with transcription activation22,23,24. Our detection of histone H3 in the siGSN-2-biotin pull-down complexes indicates that siGSN-2 stimulates H3K4, H3K36 and H3K79 lysine residue methylation. However, the exact nature of the lysine residue modification is still not clear. In fact, we reasoned that siGSN-2 was involved in the transcription activation through the interaction between siGSN-2 and H3 protein (Figure 2c). H3 protein was expressed in the nuclei and the interaction between siRNA and H3 protein had been reported. Li L.C et al. reported that siRNA targeting gene promoter caused target gene expression up-regulation by inducing H3 histone demethylation in the promoter region7.
Gelsolin, a well-known actin-binding protein, can sever, cap and nucleate actin filaments and plays a critical role in maintaining the normal cellular motility and morphology and gelsolin up-regulation inhibits actin filament elongation. The fact that siGSN-2 up-regulates GSN expression is further verified by siGSN-2 inducing the reduction of stress fibers, lamellipodia and filopodia formation in KYSE450 cells. Also, the result of western blot showed that siGSN-2 induced GSN protein expression up-regulation, which caused the reduction of stress fibers, lamelipodia and filopodia formation of cells (Figure 3b).
In a previous report, siRNA-mediated target gene transcription activation was complementary to the promoter or 3′UTR sequence7,9. However, in the present study, siGSN-2, derived from the twelfth GSN exon, is not complementary to the GSN promoter sequence. One possible mechanism could be that siGSN-2 is just used as a nuclear acid scaffold for an siGSN-2 complex which up-regulates GSN transcription activation, making it unnecessary for siGSN-2 to directly bind to the GSN promoter to activate GSN transcription. In fact, Small RNAs serving as the scaffold have been previously reported25.
To our knowledge, this is the first report that siRNA, derived from target gene exon sequence, involves in target gene transcription activation by mechanisms distinct from those siRNAs from promoter or 3′UTR in regulation of gene expression.
Method
Cell culture
KYSE150 and KYSE450 human esophageal cancer cell lines were kindly provided by Dr. Ming-Zhou Guo (Department of Gastroenterology & Hepatology, Chinese PLA General Hospital, Beijing, China). KYSE150 and KYSE450 cells were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum and grown under a 5% CO2 atmosphere at 37°C.
siRNA transfection
siRNA against GSN were individually transfected into KYSE150 cells or KYSE450 cells by X-tremeGene siRNA transfection reagent (Roche, USA) and GSN mRNA transcripts were detected by real time RT-PCR at 48 h post-transfection. Primers and siRNA oligoes were individually synthesized by BGI and GenePharma (China) and the sequence for the primers and siRNA as follows: siGSN-1: sense 5′-GACAGUGCAGCUGAGGAACGGAAAU-3′; antisense 5′-AUUUCCGUUCCUCAGC-UGCACUGUC-3′. siGSN-2: sense 5′-GAAAGGAGGU-GUGGCAUCAGGAUUC-3′; antisense 5′-GAAUCCUGAUGCCACACCUCCUUUC-3′.siGSN-3:sense5′-AUCUUUGUCUGGAAAGGCAAGCAGG-3′; antisense 5′-CCUGCUUGCCUUUCCAGACAAAGAU-3′. Scramble siRNA: sense 5′-UUCUCCGAACGUGUCACGU-3′; antisense 5′-ACGUGACACGUUCGGAGAA-3′. Real time RT-PCR primers for GSN were: forward 5′-CGTGCCCACCAACCTTTA-3′; reverse 5′-TCATTGCCCAGCCAGTAG-3′. Real time RT-PCR primers for β-Actin were: forward 5′-AGCGAGCATCCCCCAAAGTT-3′; reverse: 5′-GGGCACGAAGGCTCATCATT-3′.
Biotin labeling of siGSN
Biotin labeling of siGSN was performed by GenePhama (GenePharma, Shanghai, China). The 5′ end of the siGSN-1 sense strand, labeled with biotin, was designated as siGSN-1-1 and that of the siGSN-1 antisense strand, labeled with biotin, was designated siGSN-1-2. Similarly, the 5′ ends of the siGSN-2 sense and antisense strands labeled by biotin were denoted siGSN-2-1 and siGSN-2-2, respectively. The 5′end of scrambled RNA labeled with biotin was used as negative control.
RT-PCR, real-time PCR and western blot
RNA was extracted with Trizol and RT-PCR and real-time PCR were performed using PrimeScript® RT reagent Kit with gDNA Eraser and SYBR® Premix Ex Taq™ (DRR047A and DRR081A; Takara, Dalian, China) according to the manufacturer's instructions. Briefly, the contaminated DNA in 1 μg total RNA was firstly removed at 42°C for 2 min as following conditions: 10 μl reaction volume including 1 μg total RNA, 1 μl gDNA Eraser, 2 μl buffer and then cDNA was synthesized as 37°C, 15 min; 85°C, 5 sec. 1 μl cDNA was used as real-time PCR template and real-time PCR was performed by 7300 real-time PCR system (Life technology, American) as following conditions: 95°C, 30 sec; 95°C, 5 sec; 60°C, 31 sec; total 40 cycles. β-actin was used as internal control and water as PCR negative control (NTC). The fold changes of GSN mRNA level were calculated with 2−ΔΔCt method. Total proteins prepared in RIPA buffer, were separated in SDS-polyacrylamide gelelectrophoresis (SDS-PAGE) and transferred onto PVDF membrane (Millipore, USA) and rabbit anti-human GSN primary antibody and HRP-conjugated secondary antibody were from Sigma-Aldrich Company and the bands of GSN protein was analyzed by Image J software. GAPDH was used as internal control and GAPDH primary antibody was from Santa Cruz Biotechnology Company.
siRNA co-precipitation assay
The equivalent number of KYSE450 cells was inoculated in four 6-well plates and siGSN-1-1, siGSN-1-2, siGSN-2-1, siGSN-2-2 or scramble siRNA, were individually transfected into KYSE450 cells. All the cells from each transfection were individually collected together at post-transfection 48 h and cytoplasm and nuclei were isolated according to http://www.lifetechnologies.com/cn/zh/home/references/protocols/cell-and-tissue-analysis/elisa-protocol/elisa-sample-preparation-protocols/nuclear-extraction-method-.html. Briefly, the cell pellet was suspended in 500 μl 1× hypotonic buffer solution (20 mM Tris-HCl, pH 7.4, 10 mM NaCl, 3 mM MgCl2) and placed on ice for 15 min; The cell suspension was centrifuged at 3000 g for 10 min, after which 25 μl NP-40 was added to the mixture. The supernatant (cytoplasmic extract) was used for siGSN-1-1, siGSN-2-1 and scramble RNA co-precipitation. The nuclear pellet was resuspended in 100 μl ice-cold nuclear extraction buffer mix (Biovision, San Francisco, USA) and then vortexed every 10 min for 1 h at 4°C, then nuclear extract was collected by centrifugation at 16000 g for 10 min. The nuclear extract was used for siGSN-1-2, siGSN-2-2 and scramble RNA co-precipitation. siGSN-1-1, siGSN-1-2, siGSN-2-1 and siGSN-2-2 in the cytoplasm or nuclear extract co-precipitation were performed by the binding of biotin to streptavidine coupled on the magnetic beads according to Lehrbach19.
Immunofluorescence
KYSE 450 cells were seeded on cover slides in the 6-well plate and 2 μg siGSN-2 was transfected into KYSE450 cells after 24 h and F-actin was stained with Alexa594-conjugated mouse anti-actin antibody (Cytoskeleton, California, USA) at 48 h post-transfection and stress fibers, lamellipodia and filopodia formation were observed with a laser confocal microscope (Olympus, Tokyo, Japan).
Statistical analysis
Statistical analyses were performed using SPSS 13.0 for windows (IBM, Chicago, IL, USA). A p value of less than 0.05 was considered statistically significant and each value was two-tailed.
References
Elbashir, S. M. et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001).
Yoo, B. K. et al. Increased RNA-induced silencing complex (RISC) activity contributes to hepatocellular carcinoma. Hepatology 53, 1538–1548 (2011).
Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343–349 (2004).
Guo, H. S., Xie, Q., Fei, J. F. & Chua, N. H. MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for arabidopsis lateral root development. Plant Cell 17, 1376–1386 (2005).
Cimmino, A. et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci U. S. A. 102, 13944–13949 (2005).
Reynolds, A. et al. Rational siRNA design for RNA interference. Nat Biotechnol. 22, 326–330 (2004).
Li, L. C. et al. Small dsRNAs induce transcriptional activation in human cells. Proc Natl Acad Sci U S A. 103, 17337–17342 (2006).
Place, R. F., Li, L. C., Pookot, D., Noonan, E. J. & Dahiya, R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc Natl Acad Sci U. S. A. 105, 1608–1613 (2008).
Yue, X. et al. Transcriptional regulation by small RNAs at sequences downstream from 3′ gene termini. Nat Chem Biol. 66, 21–629 (2010).
Selden, L. A. et al. Severing of F-actin by the amino-terminal half of gelsolin suggests internal cooperativity in gelsolin. Biophys J. 75, 3092–3100 (1998).
Gremm, D. & Wegner, A. Gelsolin as a calcium-regulated actin filament-capping protein. Eur J Biochem. 267, 4339–4345 (2000).
Ditsch, A. & Wegner, A. Nucleation of actin polymerization by gelsolin. Eur J Biochem. 224, 223–227(1994).
Small, J. V. Lamellipodia architecture: actin filament turnover and the lateral flow of actin filaments during motility. Semin Cell Biol. 5, 157–163(1994).
Tanaka, H. et al. siRNA gelsolin knockdown induces epithelial-mesenchymal transition with a cadherin switch in human mammary epithelial cells. Int J Cancer 118, 1680–1691 (2006).
Azuma, T., Witke, W., Stossel, T. P., Hartwig, J. H. & Kwiatkowski, D. J. Gelsolin is a downstream effector of rac for fibroblast motility. EMBO J 17, 1362–1370 (1998).
Tanaka, M. et al. Gelsolin: a candidate for suppressor of human bladder cancer. Cancer Res 55, 3228–3232 (1995).
Gay, F. et al. In colon carcinogenesis, the cytoskeletal protein gelsolin is down-regulated during the transition from adenoma to carcinoma. Hum Pathol 39, 1420–1430 (2008).
Kwiatkowski, D. J. Functions of gelsolin: motility, signaling, apoptosis, cancer. Curr Opin Cell Biol. 11, 103–108 (1999).
Lehrbach, N. J. et al. LIN-28 and the poly (U) polymerase PUP-2 regulate let-7 microRNA processing in Caenorhabditis elegans. Nat Struct Mol Biol. 16, 1016–1020 (2009).
Yamamoto, Y., Verma, U. N., Prajapati, S., Kwak, Y. T. & Gaynor, R. B. Histone H3 phosphorylation by IKK-alpha is critical for cytokine-induced gene expression. Nature 423, 655–659 (2003).
Orban, T. I. & Izaurralde, E. Decay of mRNAs targeted by RISC requires XRN1, the Ski complex and the exosome. RNA. 11, 459–469(2005).
Santos-Rosa, H. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002).
Xu, L., Zhao, Z., Dong, A., Soubigou-Taconnat, L., Renou, J. P., Steinmetz, A., & Shen, W. H. Di- and tri- but not monomethylation on histone H3 lysine 36 marks active transcription of genes involved in flowering time regulation and other processes in Arabidopsis thaliana. Mol Cell Biol. 28, 1348–1360 (2008).
Steger, D. J. et al. DOT1L/KMT4 recruitment and H3K79 methylation are ubiquitously coupled with gene transcription in mammalian cells. Mol Cell Biol. 28, 2825–2839 (2008).
Ponchon, L. & Dardel, F. Recombinant RNA technology: the tRNA scaffold. Nat Methods 4, 571–576 (2007).
Acknowledgements
The authors wish to thank Professor Yun Wang from the Molecular pathology Lab in Shantou University Medical College for technical help with the siGSN co-precipitation assay. This work was supported by grants from the National Basic Research Program [2012CB526600 to L.Y. Xu], the National High Technology Research and Development Program of China [2012AA02A503 and 2012AA02A209 to E.M. Li] and the Natural Science Foundation of China-GuangDong Joint Fund [U0932001 to E.M. Li, U1301227 to L.Y. Xu].
Author information
Authors and Affiliations
Contributions
G.W.H. and L.D.L. performed the experiments and prepared the figures 1,2,3. E.M.L. and L.Y.X. conceived and designed the experiments and involved in the writing of the manuscript and the decision to submit the manuscript for publication.
GSN transcript was up-regulated with siGSN-2 transfection.
(a) Schematic diagram of siGSNs against target sequence of GSN gene exons. (b) GSN transcripts individually detected by real-time RT-PCR at 48 h post-transfection with 2 μg or 4 μg siGSN(−1, −2, −3) into KYSE150 cells cultured in 6-well plate. (c) GSN transcripts were individually detected by real time RT-PCR at 48 h post-transfection with 2 μg and 4 μg siGSN (−1, −2, −3) into KYSE450 cells cultured in 6-well plate. Results are mean ± standard error (SE). *p ≤ 0.05.
Histone H3 copreciptates with siGSN.
(a) Schematic diagram of the siGSN co-precipitation assay. (b) siGSN (−1–1, −1–2, −2–1, −2–2) labeled with biotin was individually transfected into KYSE450 cells and GSN transcripts were detected by real time RT-PCR at post-transfection 48 h. Results are expressed as mean ± SE. *p ≤ 0.05. (c) Histone H3 was detected in the siGSN (−1–1, −1–2, −2–1, −2–2) co-precipitated complexes from cytoplasmic or nuclear extracts probed by mouse anti-H3 antibody by western blot.
KYSE450 cell morphological alteration was observed by immunofluorescence staining and GSN protein was slightly up-regulated following 2 μg siGSN-2 or control siRNA (scramble) post-transfection 48 h.
(a) F-actin was labeled with Alexa594-conjugated mouse anti-actin antibody. Stress fiber, lamellipodia and filopodia were observed by laser confocal microscopy. Photos were observed at 40× optical magnification with a 2× digital zoom. Scale bar = 20 μm. (b) GSN protein was detected by western blot after 2 μg siGSN-2 or control siRNA (scrambled) transfection into KYSE 450 cells cultured in 6-well plate for 48 h (Left) and the bands of GSN protein was analyzed by Image J software (Right).
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Huang, GW., Liao, LD., Li, EM. et al. siRNA induces gelsolin gene transcription activation in human esophageal cancer cell. Sci Rep 5, 7901 (2015). https://doi.org/10.1038/srep07901
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07901
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.