Abstract
Sugarcane (Saccharum spp. hybrids) is a world-wide cash crop for sugar and biofuel in tropical and subtropical regions and suffers serious losses in cane yield and sugar content under salinity and drought stresses. Although real-time quantitative PCR has a numerous advantage in the expression quantification of stress-related genes for the elaboration of the corresponding molecular mechanism in sugarcane, the variation happened across the process of gene expression quantification should be normalized and monitored by introducing one or several reference genes. To validate suitable reference genes or gene sets for sugarcane gene expression normalization, 13 candidate reference genes have been tested across 12 NaCl- and PEG-treated sugarcane samples for four sugarcane genotypes using four commonly used systematic statistical algorithms termed geNorm, BestKeeper, NormFinder and the deltaCt method. The results demonstrated that glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and eukaryotic elongation factor 1-alpha (eEF-1a) were identified as suitable reference genes for gene expression normalization under salinity/drought-treatment in sugarcane. Moreover, the expression analyses of SuSK and 6PGDH further validated that a combination of clathrin adaptor complex (CAC) and cullin (CUL) as reference should be better for gene expression normalization. These results can facilitate the future research on gene expression in sugarcane under salinity and drought stresses.
Similar content being viewed by others
Introduction
A large scale loss of plant production occurred when abiotic stresses happen in planting season under different agricultural production systems, which may result in 70% reduction of the potential yields of crop plants1. In the growth and developmental periods, crop would suffer seasonal floods and drought, temperature extremes or salinity all the year round. Globally, a data from FAO (2004) showed that about 22% of the agricultural land is saline and the increasingly damage caused by drought has been reported to limit plant growth and development following the loss of productivity and especially crop species2,3. Thus, salinity and drought stresses are two of the most seriously abiotic stresses that pose a threat on crop productivity worldwide.
Sugarcane (Saccharum spp. hybrids) is a world-wide cash crop for sugar, biofuel and other food industries such as rum in tropical and subtropical regions. It serves as the world's largest crop and was cultivated on about 26.1 million hectares in 2012 in 101 countries, estimated by FAO. As a highly productive C4 grass and a bio-fuel source, more interests have been paid on sugarcane recently4. Though sugarcane can overcome a short period of water deficit during the final developmental growth when sucrose accumulating, a long term of drought can cause significant cane yield and sugar content losses5. Traditional breeding strategy for resistance improvement is lagging behind the demand of commercial need for lack of knowledge about stress tolerance-related traits, lack of efficient selection techniques and low of genetic variance and fertility6. Recently, subtractive hybridization7,8, cDNA microarray9,10, transcript expression profile11, proteome12, transcriptome13 and microRNA-seq14, known as differential display and serial analysis of gene expression techniques and strategies, have been used to identify genes and molecular markers involved in stress responses and the following genetic modifications with these genes or application of these markers would be an enormous advantage for improvement of breeding15. The previously researches showed that such a large number of genes with minor gene effects regulate water deficiency-mediated stress tolerance, which referred to complex quantitative traits that vary in plants6,16. Thus, the regulation study of hundreds of stress-related genes is a key step to explain these stress-related molecular mechanisms. Real-time quantitative polymerase chain reaction (qRT-PCR), which is a high-throughput technique with wide application in living organisms, is powerful to target and quantify gene expression17. Although qRT-PCR has a numerous advantage in normalize gene expression, several factors, such as sample amount, RNA integrity, reverse transcription (RT) efficiency, cDNA quality, sample-to-sample variations in amplification efficiency, or even the tissue and cell activities for sampling, can significantly influence the normalization of gene expression when perform analysis18,19.
As a requirement for obtaining heavily reliable measurement of gene expression by qRT-PCR, introducing one or several reference genes, which stably express across various experimental samples and serve as the internal control, is an essential way to balance the variation between samples or reactions17,20,21. The expression of most housekeeping genes varied under different biotic or abiotic stresses and it means that no single gene is suitable for all kinds of normalization experiments20,22. So, it suggested that a stable-evaluating selection of reference genes should be performed before using these genes for normalization20,22. In the present study, several statistical algorithms, such as geNorm17, NormFinder18, BestKeeper23, deltaCt method24 and RefFinder WEB-based software25, were developed and employed to assess the expression stability of a series of candidate genes, including 18S ribosomal RNA (18S rRNA), eukaryotic elongation factor 1-alpha (eEF-1a), eukaryotic initiation factor 4-alpha (eIF-4α), ubiquitin (UBQ), cullin (CUL), clathrin adaptor complex (CAC), tonoplastic intrinsic protein (TIPS-41), 25S ribosomal RNA (25S rRNA), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), β-actin (ACT), β-tubulin (TUB), anthranilate phosphoribosyl transferase (APRT) and pseudo response regulator (PRR), which stably expressed in Oryza sativa26, Zea mays27 and Brassica juncea28 or had been evaluated in sugarcane29,30. Besides, two genes, sugarcane shaggy-like kinase gene8 (SuSK, NCBI GenBank EST database Acc: FG804674) and 6-phosphogluconate dehydrogenase31 (Sc6PGDH, GenBank Accession No. KF921299), were also used to further validate the reference accuracy of novel developed candidate gene(s), together with a reference gene ACT* (β-actin) employed by Patade et al.8. Here, we expected to select suitable reference genes that could express stably in the stage of mRNA transcript profile and serve as normalization factors for qRT-PCR experimental analysis in sugarcane under salinity and drought stresses.
Results
Selection of candidate reference genes, amplification specificity and PCR efficiency
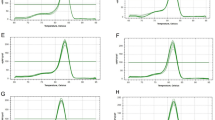
A set of 13 candidate reference genes, including four traditional housekeeping genes, 25S rRNA, GAPDH, ACT and TUB and nine new candidate reference genes, APRT, PRR, 18S rRNA, eEF-1a, eIF-4α, UBQ, CAC, TIPS-41 and CUL, were targeted in this study (Table 1). The seven of new candidates were achieved from O. sativa, Z. mays or B. juncea for expressing stably across different developmental stages or/and abiotic stresses. APRT and PRR have been validated as endogenous reference genes in sugarcane genome. Yet two genes, SuSK and 6PGDH, were used to further validate the reference accuracy of novel developed candidate controls, as well as a reference gene ACT*. The primers of SuSK and ACT* were achieved from previously study8 and the primers of 6PGDH from Yang et al.31. Moreover, the pair-primers of candidate genes that have been tested by melting curve and agarose gel electrophoresis were specific. The correlation coefficient (R2) values of all candidates changed from 0.9876 to 0.9999 across these cDNA diluted-points (Table 2). Concurrently, PCR efficiency (E) values of all pair-primers varied between 93.24%–113.56% (Table 2).
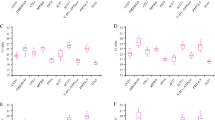
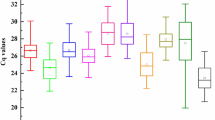
As showed in Table 2, the mean Ct values of 13 candidate reference genes, for all sample groups, changed at a wide range (14.12–28.67) and SD varied from 0.43 to 2.44 and CV from 3.04% to 8.88%. Comparing to all the candidates in all groups, 25S rRNA had a highest expression level (mean Ct ± SD = 14.12 ± 0.43), following 18S rRNA (mean Ct ± SD = 15.16 ± 0.89), whereas PRR (mean Ct ± SD = 28.67 ± 1.74) accumulated less than all the others. In terms of Ct value, 25S rRNA and 18S rRNA were closely abundant and the expression level of ACT, GAPDH and eEF-1a were similar and moderate (24.63, 24.78 and 24.47), as well as TUB and UBQ (26.85 and 26.77), while CUL, TIPS-41, APRT, eIF-4α, CAC and PRR accumulated much less equally. Additionally, 25S rRNA was the least variable candidate gene with a CV of 3.04% among 13 candidates and TIPS-41 showed the most variable expression across salinity and drought stresses. The rest of 11 genes varied at a small range (from 4.30% to 6.06%). The ranking of gene stability by CV was 25S rRNA>eIF-4α>ACT>UBQ>TUB>APRT>CAC>GAPDH>CUL>18S rRNA>eEF-1a>PRR>TIPS-41 (Table 2).
Analysis of gene expression stability
The stability values (SV) were used to rank genes from the most stable to the least stable. The relative expression stability values of candidate reference genes under salinity stress, drought stress and combination of salinity and drought stresses ranked by geNorm, NormFinder, deltaCt and BestKeeper were listed in Table 3, Table 4 and in Table 5, respectively. The mean Ct values of candidate reference genes in 12 sample groups were listed in Table S1.
In term of the top five ranked candidate genes from geNorm, NormFinder, deltaCt method or BestKeeper under salinity stress and drought stress separately, GAPDH, eEF-1a, eIF-4α, PRR, TUB and CUL had a stable expression at least in two systematic statistical algorithms when the seedlings treated with salinity stress and the same as GAPDH, eEF-1a, eIF-4α, UBQ, PRR and CUL when the seedlings treated with drought stress. But only GAPDH was ranked at the top five when evaluated by all four algorithms under salinity stress, while eEF-1a performed better than GAPDH on geNorm, NormFinder and deltaCt (Table 3). For the drought stress samples, none of candidate genes was identically ranked at the top five list generated by four algorithms, but GAPDH showed a stable performance when valued by NormFinder, deltaCt and BestKeeper and eEF-1a by geNorm, NormFinder and deltaCt (Table 4). For the salinity stress sample set, geNorm suggested GAPDH and eEF-1a were the most stable two according to the stability measure (M) and eIF-4α was the perfect one among the 13 candidates according to the identification of NormFinder and deltaCt (Table 4). The most stable gene ranked by geNorm was CAC together with APRT across four drought stress samples and Normfinder identified eIF-4α as the most stable one and deltaCt identified PRR as the best one within the same sample pool. It's interesting that 25S rRNA was identified as the better one at the two sample sets above by BestKeeper comparing to the other candidate genes (Table 4).
When the two data sets from salinity and drought stresses were analyzed together, some of GAPDH, eEF-1a, eIF-4α, CAC and CUL ranked top at least in two methods of geNorm, NormFinder, deltaCt and BestKeeper (Table 5). Four of the top five genes were CAC, CUL, eEF-1a and GAPDH at both geNorm and deltaCt and three of the top five genes were eIF-4α, eEF-1a and GAPDH at NormFinder. The least variable gene among these 13 candidate genes was 25S rRNA, following 18S rRNA, UBQ, eIF-4α and GAPDH (Table 5).
In the current study, the pearson correlation coefficient was employed to decide the consistencies of the rank of candidate reference genes that given by geNorm, NormFinder, deltaCt method and BestKeeper (Table 6). The stability values of three sample sets (NaCl, PEG and NaCl&PEG) were inputted into SAS S21.0 separately (Table 6). Though the pearson correlations achieved from the calculations were positive, the correlation between geNorm and BestKeeper were beyond the reach of significant, as well as NormFinder and BestKeeper under NaCl treatment (Table 6). Yet the most significant correlation of the rank of all candidate genes ranked by two methods was NormFinder and deltaCt in salinity and drought treatments (r = 0.910), following NormFinder VS deltaCt (r = 0.896) and NormFinder VS BestKeeper in drought treatment and geNorm in salinity and drought treatments.
Combination of reference genes for gene normalization under salinity and drought stresses
Introducing only one reference gene would cause deviation when performance gene quantification by qRT-PCR17. Thus, Vandesompele et al. suggested that at least a couple of genes were required for data normalization and recommended a method that identify the optimum number of reference genes by geNorm, through analyzing the pairwise variation (V = Vn/Vn+1) between normalization factors (NFn and NFn+1) and calculating Vn/Vn+1 ratio by adding gene one by one until the gene (n+1) makes no sense to count normalization factor17. By setting a threshold of V = 0.15, Vandesompele et al. suggested that the optimal genes set recommended by the program for a reliable normalization is not necessary to add one more gene. In this study, the data showed that V2/3 = 0.069 was less than 0.15, which suggested the combination of GAPDH and eEF-1a was good enough to normalize gene expression under salinity stress (Fig. 1). For drought stress in sugarcane, the combination of CAC and APRT, which had a V2/3 = 0.130, was the best one for accurate normalization and addition of another gene (CUL) or more had only little influence on the normalization (Fig. 1). In all salinity and drought stress samples, the same effect was found. In revealed that CAC together with CUL (V2/3 = 0.138) could provide a reliable reference for gene expression normalization. In spite of that, all V3/4 values from salinity stress samples set, drought stress samples set and all samples set were smaller than V2/3 (Fig. 1). And as the data showed above, the top three rank of candidate genes were GAPDH, eEF-1a and CUL under salinity stress, CAC, APRT and CUL under drought stress and CAC, CUL and eEF-1a under both salinity and drought stresses when evaluated by geNorm.
Determination of the optimal number of reference genes under salinity stress and drought stress.
The pairwise variation (Vn/Vn + 1) was analyzed between the normalization factors NFn and NFn + 1 by geNorm program to determine the optimal number of reference genes required for qRT-PCR data normalization. NFn, the Normalization Factor was based on n reference genes included in present study; The calculation of Vn/Vn + 1 value was based on two sequential normalization factors. We proposed 0.15 as a threshold value, which suggested that adding one more gene into the combination of reference genes is not required. As showed in Fig. 1, the optimal number of reference genes under salinity stress (a), drought stress (b) or salinity stress plus drought stress (c) were two. It suggested that the top two ranked by geNorm in this current study was the best combination.
Expression normalization of SuSK and 6PGDH genes based on different reference gene/genes
To validate the effectiveness of reference genes selected in the current study, the expressions of two gene, SuSK and 6PGDH, were detected with the samples treated by salinity or drought stress for 12 h in sugarcane varieties Liucheng03-182, "ROC"20 and YC05-179 and the samples treated by salinity or drought stress for 12 h, 24 h and 48 h in FN40 plantlets (Fig. 2) together with each control samples of these four cultivars. The data was normalized with the 2−ΔΔCt method. SuSK is a sugarcane gene significantly induced by short-term treatment of NaCl or PEG8. Based on the recommended gene/genes in this current study, GAPDH and eEF-1a together with ACT* and ACT, or reference gene sets, GAPDH+eEF-1a (served as the combination of the two best gene in drought treatment), GAPDH+eEF-1a+CUL, CAC+CUL and CAC+CUL+eEF-1a. The expression of SuSK showed a similar expression tendency under salinity stress treatment for all six sample groups but with different fold of up/down-regulation (Fig. 2, a). In Liucheng03-182, when normalized by a single reference gene ACT or ACT*, the most significantly up-regulated expression of SuSK was found (P<0.01), following closely up-regulated expression of SuSK by CAC+CUL, CAC+CUL+eEF-1a, GAPDH+eEF-1a+CUL, GAPDH, GAPDH+eEF-1a (Fig. 2, a). In "ROC"20 salinity-stress treated sample, the expression of SuSK exhibited a slight and similar up-regulation expression when the reference of GAPDH+eEF-1a and ACT* (p<0.01) were employed, comparing with the reference of GAPDH, CAC+CUL+eEF-1a and GAPDH+eEF-1a+CUL (p<0.01), in which a slight down regulation of SuSK was found. Following that, a more significant down-regulation of SuSK was found when normalized by gene(s) ACT and CAC+CUL (p<0.01) (Fig. 2, a). In the YC05-179 salinity stress treated sample, a slight up-regulation of SuSK expression was found with reference gene(s) GAPDH+eEF-1a, following by ACT, GAPDH, GAPDH+eEF-1a+CUL and CAC+CUL+eEF-1a, while a slight down-regulation of SuSK expression was found when normalized by ACT*, following by CAC+CUL, all of these were significantly different (p<0.01) (Fig. 2, a). In FN40 salinity-stress treated samples, the accumulation of SuSK decreased with the increasing of the exposure time and the fold up-regulation of SuSK was significantly different with different references (p<0.01) except CAC+CUL and CAC+CUL+eEF-1a at 12h and GAPDH+eEF-1a+CUL, CAC+CUL and CAC+CUL+eEF-1a at 48h (Fig. 2, a). Besides, the magnitude of up-regulation was relatively lower in the reference gene sets of CAC+CUL and CAC+CUL+eEF-1a than those of the others in three FN40 salinity treated samples (Fig. 2, a). When using the single gene ACT or the combinations of CAC+APRT, CAC+APRT+CUL, CAC+ CUL and CAC+CUL+eEF-1a as the internal control in drought stress samples, SuSK had a similar expression tendency (Fig. 2, b). Obviously, The accumulation of SuSK increased both in Liucheng03-182 and in three FN40 samples, but slightly decreased both in "ROC"20 and YC05-179 except when normalized by GAPDH and GAPDH+eEF-1a (GAPDH+eEF-1a was treated as the combination of the two best genes and the recommended gene set in salinity treatment in the current study) (Fig. 2, b). Exactly, the most up-regulation of SuSK in Liucheng03-182 was found with the reference of ACT (p<0.01), following the reference of CAC+APRT, CAC+APRT+CUL, ACT* secondly, CAC+ CUL and CAC+CUL+eEF-1a; thirdly, GAPDH and GAPDH+eEF-1a lastly (Fig. 2, b). In "ROC"20 and YC05-179, only the reference of GAPDH and GAPDH+eEF-1a separately showed that the expression of SuSK was up-regulated differently (P<0.01) comparing with the reference of ACT, CAC+CUL, CAC+APRT+CUL, CAC+APRT, CAC+CUL+eEF-1a and ACT*. In FN40 PEG-treated samples, the relative expression of SuSK increased continuingly when normalized by ACT*, GAPDH and GAPDH+eEF-1a, whereas a different expression pattern of SuSK was found when normalized by the remaining reference gene/genes, which firstly increased at 12 h and following a decrease at 24 h and a rising trend at 48 h (Fig. 2, b). The most obvious up-regulation of SuSK was found with reference of GAPDH at 12 h (p<0.01) while the lowest up-regulation was in FN40 48 h treated samples with reference of ACT (p<0.01) (Fig. 2, b). Actually, the variation of SuSK expression was only slight when normalized by CAC+CUL and CAC+CUL+eEF-1a across three FN40 PEG-treated samples (Fig. 2, b).
Normalized expression of SuSK and 6PGDH under salinity stress, drought stress or salinity stress plus drought stress in sugarcane.
SuSK (sugarcane Shagging-like kinase) and 6PGDH (6-phosphogluconate dehydrogenase) both were salinity and drought stress response genes in sugarcane. In this current study, the normalization of SuSK (a) and 6PGDH (b) employed a single refernce gene, ACT* ACT and GAPDH, or the reference gene set, GAPDH+eEF-1a, CAC+CUL, GAPDH+eEF-1a+CUL and CAC+CUL+eEF-1a as reference control in salinity stress treatment experiment, and. SuSK (c) and 6PGDH (d) ACT* ACT and GAPDH, or reference gene set, GAPDH+eEF-1a, CAC+APRT, CAC+CUL, CAC+APRT+CUL and CAC+CUL+eEF-1a as reference control in drought stress treatment experiment. Using 2−ΔΔCt to normalize the SuSK and 6PGDH, all the control samples from different cultivars were converted into 1.
The 6PGDH is a sugarcane gene that involves in the positive response to the salt stress31. Comparing with the reference of single gene ACT* and ACT or the reference of gene(s) GAPDH, GAPDH+eEF-1a, CAC+CUL, GAPDH+eEF-1a+CUL and CAC+CUL+eEF-1a, the expression of 6PGDH showed a significantly different fold up-regulation (p<0.01) in Liucheng03-182, while the reference of ACT and CAC+CUL showed a significant down-regulation of 6PGDH comparing with the reference of the remaining reference(s), which showed that the expression of 6PGDH kept at a low up-regulation level in "ROC"20 12 h NaCl-treated sample (Fig. 2, c). However, in YC05-179 the accumulation of 6PGDH increased as treated with NaCl (Fig. 2, c). The highest accumulation of 6PGDH was found when normalized by the combination of GAPDH+eEF-1a or ACT (p<0.01), following a moderate accumulation by GAPDH, GAPDH+eEF-1a+CUL or CAC+CUL+eEF-1a and a lower accumulation by CAC+CUL or ACT in order. In FN40 NaCl-treated samples, when normalized with the reference of ACT*, GAPDH, GAPDH+eEF-1a and GAPDH+eEF-1a+CUL, the accumulation of 6PGDH increased firstly at 12 h, then following a down-regulation at 24 h and a rising at 48 h (Fig. 2, c). Conversely, 6PGDH showed a transcript decrease at 12 h and 24 h and then a increasing trend occurred at 48 h when used ACT, CAC+CUL or CAC+CUL+eEF-1a as the reference (Fig. 2, c). In PEG-treated samples, the similar tendencies of 6PGDH expression were found when employed GAPDH and GAPDH+eEF-1a or ACT*, ACT, CAC+APRT, CAC+CUL, CAC+APRT+CUL and CAC+CUL+eEF-1a as reference control (Fig. 2, d). In Liucheng03-182, the references of ACT*, ACT, GAPDH+eEF-1a, CAC+APRT and CAC+CUL showed obviously different fold up/down-regulation of 6PGDH (p<0.01). In "ROC"20, the reference of GAPDH, GAPDH+eEF-1a, CAC+APRT and CAC+CUL (or ACT*) indicated that 6PGDH has a significantly different level of 6PGDH expression (p<0.01) (Fig. 2, d). Yet the expression of 6PGDH was closely up-regulated in YC05-179 except the normalization of GAPDH and GAPDH+eEF-1a (Fig. 2, d). In FN40 PEG-treated samples, all of the reference controls employed in this normalization gave a first decreasing accumulation of 6PGDH expression at 12 h comparing to the control sample and following a slight increasing accumulation at 24 h (Fig. 2, d). However, in 48 h PEG-treated sample the expression of 6PGDH exhibited significantly different accumulation levels when normalized with different references (Fig. 2, d). The results showed that the 6PGDH expression with normalization of GAPDH had a straight increasing to the highest level at 48 h (p<0.01) comparing with a slight increasing with normalization of ACT*, ACT and GAPDH+eEF-1a, whereas the normalization using CAC+APRT or CAC+APRT+CUL as the reference control, the 6PGDH transcript exhibited a decreasing accumulation at 48 h, as well as using CAC+CUL or CAC+CUL+eEF-1a (Fig. 2, d).
Across six samples from four cultivars, the expression patterns of SuSK and 6PGDH, which were normalized by different reference controls under discussion in this current study, were analyzed with significant difference test and pearson correlation test. The results showed that the single reference gene (ACT*, ACT and GAPDH) exhibited a significant difference and the highest relative level of SuSK and 6PGDH expressions, which indicated that the expressions of SuSK and 6PGDH were induced by PEG and NaCl treatment obviously (Table 7). Yet comparing with the single gene reference, the combination of GAPDH+eEF-1a under salinity treatment, CAC+APRT and CAC+APRT+CUL under drought treatment, or CAC+CUL and CAC+CUL+eEF-1a under both two treatments for normalization suggested that the expression of SuSK and 6PGDH were significantly different but at a lower relative expression level (Table 7). Though significantly differences would be found between GAPDH and GAPDH+eEF-1a, GAPDH+eEF-1a and GAPDH+eEF-1a+CUL or CAC+APRT and CAC+APRT+CUL, the correlation analysis based on the fold up/down-regulation of SuSK and 6PGDH showed that the correlation coefficients between these pair sets (except employed GAPDH and GAPDH+eEF-1a for 6PGDH normalization under NaCl treatment) together with CAC+CUL and CAC+CUL+eEF-1a were positive significantly. It also indicated that the expression patterns of SuSK and 6PGDH were nearly identical when normalized with either of two reference gene sets (Table 8).
Discussion
Major environmental stresses, such as drought and salinity, contribute to the gap between actual and potential crop yields. To guarantee a sustainable crop yield, it is imperative to design and develop better crop varieties with in-built tolerance to the harmful effects of constantly changing environmental factors5,35,36. To characterize the function of the target genes, some drought- and salty-treated experiments should be implemented under artificial and controllable conditions to study the gene expression variation. qRT-PCR has been commonly recognized as a reliable and accurate method for detecting expression of gene and has a huge advantage in the study of molecular mechanism. However, to minimize the variation caused by RNA integrity, cDNA synthesis, PCR reaction, the tissue or cell activities, reference gene selection should be required18. Sugarcane is the most important sugar crop which accounts for 80% sugar production in the world and 92% of that in China and it is also an important energy crop which can be used effectively for biofuel production. The development of sugarcane cultivars tolerant to osmotic and drought stresses could allow for the expansion of plantations to sub-prime regions. Having a good knowledge of the molecular mechanisms related to drought-responses and the relationship between water-utilized and carbon-fixed in plants would greatly help to increase yield37 and even breeding. A number of novel drought- or salt-related genes/microRNAs have been identified in sugarcane5,6,7,35,36,37,38. Though Iskandar et al.29 had identified GAPDH as a reliable reference for normalizing the transcriptional profile of target genes in two sugarcane cultivars and three Saccharum species and Ling et al.30 had identified GAPDH, eEF-1a and eIF-4a as the most stable and suitable references genes for normalizing target gene related to various experimental samples, several genes, such as β-actin, tubulin, 25S rRNA, GAPDH and poly-ubiquitin, were still frequently used as the reference genes for the normalization of gene expression in sugarcane.
Comparing to the results showed in the present study, geNorm, NormFinder and deltaCt mostly showed consistency of the rank of candidate reference genes. Bestkeeper showed the least correlation of the rank of all the candidate genes with geNorm. Depending on the pearson correlations achieved from the calculations, we prefer to select the reference genes base on the rank gave by geNorm NormFinder and deltaCt. In the current study, we collected 12 sample groups from four sugarcane varieties and attempted to select the best-suited reference genes from 13 candidates, 25S rRNA, GAPDH, ACT, TUB, APRT, PRR, 18S rRNA, eEF-1a, eIF-4α, CAC, TIPS-41, CUL and UBQ for testing across salinity (NaCl) and drought (PEG) treatment. Then the expression profiles of SuSK and 6-PGDH were normalized with reference gene(s) recommended by geNorm, NormFinder, deltaCt and BestKeeper synthetically and reference gene sets recommended by geNorm above. The results showed that none of gene ranked at the first by four different statistical algorithms simultaneously, but some consistency could be found among geNorm, NormFinder and deltaCt. When analyzed the data from salinity-treated samples, only GAPDH and eEF-1a were identified for their stable expression across four sugarcane cultivars by three of the four programs employed in this study. It was in consistent with the results reported that GAPDH expressed stable across different tissues in two sugarcane cultivars and three other Saccharum species29. An overview from Kozera et al. emphasized that though the use of GAPDH in many studies brings variable normalization results when exhibited to some experimental conditions, GAPDH is one of the most commonly used reference genes due to its different sequence and validation in each case separately39. In this study, eEF-1a expression was identified at a stable level with respect to salinity and drought treatments when evaluated by geNorm, NromFinder and deltaCt and some previous studies have proved that eEF-1a was suitable for gene transcript profile normalization in O. sativa, Cucumis sativus, Carica papaya, Gossypium hirsutum L. and Vigna mungo under abiotic stress26,40,41. For these reasons, GAPDH and eEF-1a recommended by at least three systematic statistical algorithms in this study could be accepted as normalization factor in sugarcane when exposed to salinity and drought. eIF-4α exhibited high stability across two methods viz., NormFinder and deltaCt, but poor performance across geNorm and BestKeeper. According to previous reports, though eIF (Eukaryotic initiation factor) performed good across different tissues in Musa paradisiaca, Lycoris longituba, Hevea brasiliensis and Coffea spp., it was less stable across multiple stresses except in C. papaya42. It meant that eIF-4α was not as suitable as GAPDH and eEF-1a for gene normalization in the current study, as well as the previously used reference gene ACT that ranked at the bottom of 13 candidates. The data also showed that 25S rRNA and TUB were ranked at mid-position of the list from four algorithms or even worse. These results agreed with Iskandar et al. discussion about 25S rRNA, TUB, ACT and GAPDH29. In the previous study, Jain et al. showed that the expression of UBQ5 was more stable than that of UBQ10 in tested tissue samples26, but performed poorly stable in this current study. So does 18S rRNA and 25S rRNA, which was highly expressed. CAC and TIPS-41 was the best recommended combination in Brassica juncea28. However, TIPS-41 was not suitable to serve as a reference gene for its unstable expression across treated samples according to all four algorithms. CAC together with CUL, which exhibited a more stable expression pattern according to geNorm, NormFinder and Bestkeeper27, was recommended as the reference gene combination in the present study. Due to most of the candidate genes were member of multigene family, different sequences from one family mostly probably have different stability performance in each case39 and the current study aimed to select suitable reference gene normalization salinity and drought response. It may be the main reason that these candidates exhibited different stability pattern in sugarcane although some were referenced from sugarcane closely related species (Rice and Maize). APRT and PRR with low copies number in sugarcane genome were few detected at the transcript level.
Using GAPDH, eEF-1a, ACT*, ACT, GAPDH+eEF-1a, GAPDH+eEF-1a+CUL, CAC+CUL and CAC+CUL+eEF-1a as internal controls, the expression of SuSK and 6PGDH were detected by qRT-PCR in this study. In salinity or drought treatment, the result showed that the expression trends of SuSK and 6PGDH across different groups were closely consistency under different reference gene/genes, but some different would be found in several groups separately. The expression profiles or transcript abundance were normalized by the less stable reference gene (ACT), the most stable reference gene (GAPDH) together with ACT*, the combination of the two most stable genes (GAPDH and eEF-1a), reference gene sets recommended by geNorm or combinations of the top three rank reference genes by geNorm. The above six strategies were employed in the present study to validate the suitability of the candidate reference gene/gene set under investigation. Thus, the results suggested that the expressions level of SuSK and 6PGDH with different reference gene sets can be ranked in a nearly identical order from high to low under PEG or NaCl treatment in either of six sample groups. Though the reference of single gene, ACT*, ACT or GAPDH, showed similar fold of regulations of SuSK and 6PGDH with gene set(s) under one or more of six samples, it exhibited a more variable expression of SuSK and 6PGDH across six sample groups. Actually, the expression of SuSK was less variable when normalized by GAPDH+eEF-1a, CAC+CUL, CAC+APRT, GAPDH+eEF-1a+CUL, CAC+CUL+eEF-1a and CAC+APRT+CUL across all six salinity- or drought-treated sample groups. When normalized by CAC+CUL and CAC+CUL+eEF-1a, the expression of SuSK was moderately induced across all salinity or drought-treated sample groups, as well as the expression of 6PGDH. For salinity-treated sample groups, the normalization with reference of the best recommended gene set, GAPDH+eEF-1a, showed that the expression level of SuSK and 6PGDH were lower than most of the remaining references employed in Liucheng03-182 and "ROC"20 but higher in YC05-179 and FN40 in this test. However, when this two genes, GAPDH and eEF-1a, served as combination of two best genes under drought treatment, the transcript level of SuSK and 6PGDH were more significantly higher than most of the other references in "ROC"20, YC05-179 and FN40. For the reference of CAC+APRT, the best combination under drought treatment, the data showed that the accumulation of SuSK and 6PGDH changed easily. Using a single reference gene for gene normalization was considered as less reliable than that of gene set17,19. It means that the normalizations with the reference of CAC+APRT, GAPDH+eEF-1a or CAC+CUL were more reliable than those with GAPDH, ACT* or ACT.
Additionally, the results of significant difference test and pearson correlation test showed that the single reference gene (ACT*, ACT and GAPDH) exhibited the significant difference and the highest relative expression levels of SuSK and 6PGDH, which indicated that the expressions of SuSK and 6PGDH were induced by PEG and NaCl treatment obviously. Yet though significantly different between the reference of GAPDH and GAPDH+eEF-1a, GAPDH+eEF-1a and GAPDH+eEF-1a+CUL or CAC+APRT and CAC+APRT+CUL, the correlation analysis based on the fold up/down-regulation of SuSK and 6PGDH indicated that the correlation coefficients between the recommended two and the top three gene ranked by geNorm were positive significantly, which indicated that the expression patterns of SuSK and 6PGDH were nearly identical when normalized with either of two. This result indicated that the best reference gene set recommended by geNorm plus one more gene from the rank (the top three gene ranked by geNorm) failed to give any more precise normalization.
Methods
Plant materials and stress treatments
To achieve disease-free materials, the stem sections (nodes) from four cultivars, "ROC"20, FN40, Liucheng03-182 and YC05-179, were firstly treated with 50°C -water thermotherapy contained 100 mg·L-1 fungicide carbendazim (Friend, Zhengzhou, China) for 40 min and then planted in autoclaved soil (16 h light/8 h dark, 28°C in incubator) before harvested for meristem excision, callus induction, shoot differentiation and rooting32 and then the seedlings were placed in ddH2O and cultured for ten days. Salinity stress was induced by sodium chloride (NaCl) and drought stress by polyethylene glycol (PEG) 8000. The greatly consistent seedlings were divided into three groups. Two of them were placed into NaCl (250 mM)- or PEG 8000 (25% w/v)-water solutions for 12 h in flat-bottomed glass tubes separately. The third group without treatment (kept in distilled water) was harvested as control at 0 h. Three biological replications were contained in each sample group. All samples had been employed in the experiments that evaluating the stabilities of candidate reference genes in sugarcane under salinity and drought stresses. Additionally, the samples that treated with NaCl (250 mM)- and PEG 8000 (25% w/v)-water solutions for 12 h, 24 h and 48 h in FN40 seedlings together with previously FN40 untreated sample group (control) were collected and used in experimental validation together with the previous stress treated sample groups and control group from the remaining three cultivars. Three biological replications were included in each sample group collected in this current experiment.
In our study, each sample collected from each cultivar under different treatments was considered an experimental group as well as control sample, thus 12 groups (4 different cultivars and 2 different treatments for 12 h together with controls) were used for analyzing candidate reference stability and this 12 groups plus 4 groups, which contained PEG or NaCl 24 h and 48 h treated samples fromFN40), were used for experimental validation.
RNA isolation and cDNA synthesis
All samples were snap–frozen and stored at −80°C before RNA extraction. RNAprep Pure Plant Kit (polysaccharides & polyphenolics-rich, TIANGEN, Beijing, China) was used for RNA extraction, following RNA integrity analysis by agarose gel electrophoresis and RNA samples quality tests with synergy H1 Microplate Reader Multi-Mode (Bio-Tek, Vermont, USA). Finally, the samples with 260/280 ratio from 1.9 to 2.1, 260/230 ratio from 2.0 to 2.5 were chosen.
Subsequently, the first-strand cDNA was synthesized with a 10 μL reaction system following the instruction of TAKARA PrimeScrit RT reagent Kit (Perfect for Real Time) (TAKARA Biotechnology, Dalian, China). Then the quality and integrity of cDNA were checked by agarose gel electrophoresis and synergy H1 Microplate Reader Multi-Mode and stored at −20°C until use.
Screening of candidate reference genes and primer design
The primers of four traditional housekeeping genes, 25S rRNA, GAPDH, ACT and TUB, had been reported by Iskandar et al.29 and Que et al.33 and the pair-primers of two new candidate reference genes, APRT and PRR, were from Casu et al.34. The probe sequences from O. sativa (18S rRNA, AK059783; eEF-1a, AK061464; eIF-4α, AK073620; UBQ5, AK061988), Z. mays (CUL, GRMZM2G166694_T04) and Arabidopsis thaliana (CAC, AK317765.1 and TIPS-41, NM_119592.4) were used to search within sugarcane expressed sequence tags (ESTs) database (http://www.ncbi.nlm.nih.gov/). Subsequently, the target ESTs were identified by querying homologous sugarcane sequences together with A. thaliana genes mRNA complete CDS and probe-sequences. Using the tool Primer-BLAST (http://www.ncbi. nlm.nih.gov/tools/primer-blast/), the pair-primers were designed on the homolog regions that verified by DNAMAN with the reference sequences from O. sativa, Z. mays, B. juncea, T. aestivum or A. thaliana and the homologous sequences from sugarcane.
Two-step qRT-PCR
The 2.0 μL diluted-cDNA (10 ng total RNA) was used as template in a 20 μL-mixture qPCR reaction together with 10.0 μL SYBR Green Master Mix (Roche, New York City, USA), 10 pM of each primer and 6.4 μL ddH2O. The ABI 7500 real time PCR amplification regime (Applied Biosystems, Foster City, CA, USA) comprised 2 min denaturation at 50°C, 10 min at 95°C and then followed by 40 cycles (94°C for 15 s and 60°C for 60 s). To check the specificity of the amplicon, the qRT-PCR products of each gene were used for a electrophoresis with a 1.0% w/v agarose gel and the melting curves were evaluated in each reaction (data not shown). The Ct value from three biological replicates, which three technical replicates were included and resulted from three different RNA extractions, RT and qRT-PCR reactions, were collected for stabilities analysis. Using series dilution of cDNA from YC05-179 control sample as the templates, the standard curves were generated for each candidate reference gene prior to the qRT-PCR evaluation of these genes in sugarcane treated sample groups, then estimating the PCR amplification efficiencies of all genes through the slope of the standard curve with the formula as follow:

To calculate PCR efficiency and the correlation coefficient (R2) and generate standard curve, Ct values of each gene, which obtained from a range of ten-, five- or three-fold dilutions of YC05-179 control sample, were inputted into MicroSoft Excel 2003. Among the investigated genes, GAPDH and TUB were ten-fold, 25S rRNA, ACT, 18S rRNA, eEF-1a, CUL and eIF-4α were five-fold and CAC, TIPS-41, UBQ, APRT and PRR were three-fold of decrement on the YC05-179 control sample.
Data analysis
The Ct values over 40 would be quitted to avoid non-specific detection. According to the manual provided by the authors, the expression data from the mean Ct values of three biological replications 12 sample groups described previously were evaluated by freely available Visual Basic Applications for Microsoft Excel, namely geNorm17 and NormFinder18 and by RefFinder25, a WEB-based software (http://www.leonxie.com/referencegene.php). The Ct values were transformed into relative quantities (Q value) via the delta-Ct method, as following formula:

E, the PCR efficiency of candidate gene; ΔCt, the difference of Ct values between the treated-sample and the control-sample. Then the Q values were loaded into geNorm and NormFinder to achieve a stability measure (M) and a stability ranking. Optimal numbers of reference genes for gene expression normalization were required to calculate by geNorm. In addition, the measure results of BestKeeper and deltaCt method from RefFinder were employed, which required Ct input directly. Besides, the mean Ct values of 13 candidate reference genes were used to calculate standard deviation (SD) and co-variance (CV) on MicroSoft Excel 2003. Following that, pearson correlation values (r value) were calculated by inputting the stability values of 13 candidate genes that obtained from geNorm, NormFinder, deltaCt and BestKeeper into SAS S21.0. These values reflect the level of correlation between the results from geNorm, NormFinder and deltaCt. To valid the reference gene/genes selected in the present study, the relative expression level of SuSK and 6PGDH were normalized with the 2−ΔΔCt method after collected the Ct mean value of each biological replication from the samples treated by salinity or drought stresses for 12 h in sugarcane varieties Liucheng03-182, "ROC"20, YC05-179 and the samples treated by salinity or drought stress for 12 h, 24 h and 48 h in FN40 plantlets together with the control samples. Finally, the relative folds of SuSK and 6PGDH expression in experimental validation were used to calculate the difference of normalization with different reference gene or genes by inputting the values into DPS (v7.05) and the pearson correlation between gene(s) and gene set on SAS S21.0.
Conclusion
To validate suitable reference genes or gene sets for gene expression normalization in sugarcane under salinity and drought stresses, 13 candidate reference genes have been tested across four different sugarcane genotypes using four commonly used systematic statistical algorithms, which termed as geNorm, BestKeeper, NormFinder and deltaCt methods. The results showed that geNorm, NormFinder and deltaCt would give a more consistent rank of gene stability. Based on gene stability analysis, the current study confirmed that GAPDH or eEF-1a was identified as a suitable reference gene for gene expression normalization under salinity/drought treatments in sugarcane. Moreover, the expression analyses of SuSK and 6PGDH further validated that a combination of CAC and CUL as reference would be better in the current study. These results can facilitate the future research on gene expression in sugarcane under salinity- and drought-stresses.
References
Agarwal, P. K., Agarwal, P., Reddy, M. & Sopory, S. K. Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep. 25, 1263–1274 (2006).
Burke, E. J., Brown, S. J. & Christidis, N. Modeling the recent evolution of global drought and projections for the twenty-first century with the Hadley Centre climate model. J. Hydrometeor. 7, 1113–1125 (2006).
Shao, H.-B., Chu, L.-Y., Jaleel, C. A. & Zhao, C.-X. Water-deficit stress-induced anatomical changes in higher plants. C. R. Biol. 331, 215–225 (2008).
Cheavegatti-Gianotto, A. et al. Sugarcane (Saccharum X officinarum): a reference study for the regulation of genetically modified cultivars in Brazil. Trop. Plant Biol. 4, 62–89 (2011).
Begcy, K. et al. A novel stress-induced sugarcane gene confers tolerance to drought, salt and oxidative stress in transgenic tobacco plants. PLoS One 7, e44697, 10.1371/journal.pone.0044697 (2012).
Rodríguez, M., Canales, E. & Borrás-Hidalgo, O. Molecular aspects of abiotic stress in plants. Biotecnol. Apl. 22, 1–10 (2005).
Prabu, G., Kawar, P. G., Pagariya, M. C. & Prasad, D. T. Identification of water deficit stress upregulated genes in sugarcane. Plant Mol. Biol. Rep. 29, 291–304 (2011).
Patade, V. Y., Rai, A. N. & Suprasanna, P. Expression analysis of sugarcane shaggy-like kinase (SuSK) gene identified through cDNA subtractive hybridization in sugarcane (Saccharum officinarum L.). Protoplasma 248, 613–621 (2011).
Casu, R. E. et al. Identification of a novel sugar transporter homologue strongly expressed in maturing stem vascular tissues of sugarcane by expressed sequence tag and microarray analysis. Plant Mol. Biol. 52, 371–386 (2003).
Casu, R. E. et al. Identification of differentially expressed transcripts from maturing stem of sugarcane by in silico analysis of stem expressed sequence tags and gene expression profiling. Plant Mol. Biol. 54, 503–517 (2004).
Patade, V. Y., Bhargava, S. & Suprasanna, P. Transcript expression profiling of stress responsive genes in response to short-term salt or PEG stress in sugarcane leaves. Mol Biology Rep. 39, 3311–3318 (2012).
Zhou, G. et al. Proteomic analysis of osmotic stress-responsive proteins in sugarcane leaves. Plant Mol. Bio. Rep. 30, 349–359 (2012).
Manners, J. M. & Casu, R. E. Transcriptome analysis and functional genomics of sugarcane. Trop. Plant Bio. 4, 9–21, 10.1007/s12042-011-9066-5 (2011).
Gentile, A. et al. Effects of drought on the microtranscriptome of field-grown sugarcane plants. Planta 237, 783–798 (2013).
Sreenivasulu, N., Sopory, S. K. & Kavi Kishor, P. B. Deciphering the regulatory mechanisms of abiotic stress tolerance in plants by genomic approaches. Gene 388, 1–13, 10.1016/j.gene.2006.10.009 (2007).
Simon-Sarkadi, L., Kocsy, G., Sebestyén, Z. & Galiba, G. Deletions of chromosome 5A affect free amino acid and polyamine levels in wheat subjected to salt stress. Environ. exp. Bot. 60, 193–201 (2007).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Bio. 3, research0034 (2002).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250 (2004).
Janská, A. et al. The choice of reference gene set for assessing gene expression in barley (Hordeum vulgare L.) under low temperature and drought stress. Mol. Genet. Genomics 288, 639–649 (2013).
Nicot, N., Hausman, J.-F., Hoffmann, L. & Evers, D. Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J. Exp. Bot. 56, 2907–2914 (2005).
Ovesna, J. et al. Validation of the β-amy1 transcription profiling assay and selection of reference genes suited for a RT-qPCR assay in developing barley caryopsis. PloS one 7, e41886 (2012).
Lilly, S., Drummond, R., Pearson, M. & MacDiarmid, R. Identification and validation of reference genes for normalization of transcripts from virus-infected Arabidopsis thaliana. Mol. Plant Microbe In. 24, 294–304 (2011).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515 (2004).
Silver, N., Best, S., Jiang, J. & Thein, S. L. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol. Bio. 7, 33 (2006).
Xie, F., Xiao, P., Chen, D., Xu, L. & Zhang, B. miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol. Bio. 80, 75–84 (2012).
Jain, M., Nijhawan, A., Tyagi, A. K. & Khurana, J. P. Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem. Bioph. Res. Co. 345, 646–651 (2006).
Manoli, A., Sturaro, A., Trevisan, S., Quaggiotti, S. & Nonis, A. Evaluation of candidate reference genes for qPCR in maize. J Plant Physiol 169, 807–815, 10.1016/j.jplph.2012.01.019 (2012).
Chandna, R., Augustine, R. & Bisht, N. C. Evaluation of candidate reference genes for gene expression normalization in Brassica juncea using real time quantitative RT-PCR. PLoS One 7, e36918, 10.1371/journal.pone.0036918 (2012).
Iskandar, H. M. et al. Comparison of reference genes for quantitative real-time polymerase chain reaction analysis of gene expression in sugarcane. Plant Mol. Bio. Rep. 22, 325–337, 10.1007/BF02772676 (2004).
Ling, H., Wu, Q., Guo, J., Xu, L. & Que, Y. Comprehensive selection of reference genes for gene expression normalization in sugarcane by real time quantitative rt-PCR. PLoS One 9, e97469, 10.1371/journal.pone.0097469 (2014).
Yang, Y., Li, G., Su, Y., Guo, J. & Xu, L. Cloning and characterization of sugarcane 6-phosphgluconte dehydrogenase gene, positive response to salt stress. J. Fujian agriculture and forest university (Nature science edition) 43 (2014).
Ramgareeb, S., Snyman, S., Van Antwerpen, T. & Rutherford, R. Elimination of virus and rapid propagation of disease-free sugarcane (Saccharum spp. cultivar NCo376) using apical meristem culture. Plant Cell Tiss. Org. (PCTOC) 100, 175–181 (2010).
Que, Y. et al. Selection of control genes in real-time qPCR analysis of gene expression in sugarcane. Chinese J Trop Crop 30, 276–278 (2009).
Casu, R. E., Selivanova, A. & Perroux, J. M. High-throughput assessment of transgene copy number in sugarcane using real-time quantitative PCR. Plant cell Rep. 31, 167–177 (2012).
Rocha, F. R. et al. Signal transduction-related responses to phytohormones and environmental challenges in sugarcane. BMC genomics 8, 71 (2007).
Rodrigues, F. et al. Sugarcane genes differentially expressed during water deficit. Biologia Plantarum 55, 43–53 (2011).
Lembke, C. G., Nishiyama Jr, M. Y., Sato, P. M., de Andrade, R. F. & Souza, G. M. Identification of sense and antisense transcripts regulated by drought in sugarcane. Plant Mol. Bio. 79, 461–477 (2012).
Muthiah, M., Ramadass, A., Amalraj, R. S., Palaniyandi, M. & Rasappa, V. Expression profiling of transcription factors (TFs) in sugarcane X Colletotrichum falcatum interaction. J. Plant Biochem. Biot. 22, 286–294 (2013).
Kozera, B. & Rapacz, M. Reference genes in real-time PCR. J. App. Genet. 54, 391–406 (2013).
Wan, H. et al. Selection of appropriate reference genes for gene expression studies by quantitative real-time polymerase chain reaction in cucumber. Anal. Biochem. 399, 257–261 (2010).
Kundu, A., Patel, A. & Pal, A. Defining reference genes for qPCR normalization to study biotic and abiotic stress responses in Vigna mungo. Plant cell Rep. 32, 1647–1658 (2013).
Zhu, X. et al. Evaluation of new reference genes in papaya for accurate transcript normalization under different experimental conditions. PLoS One 7, e44405 (2012).
Acknowledgements
This work was funded by the National Natural Science Foundation of China (31271782), Natural Science Foundation of Fujian province, China (2012J01089), National High Technology Research and Development Program of China 863 Program Project (2013AA102604) and the Program for New Century Excellent Talents in Fujian Province University, China (JA14095).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: J.G., L.X., Y.Q. Performed the experiments: H.L., J.G., Q.W. Analyzed the data: J.G., H.L., L.X., Y.Q. Wrote the paper: J.G., H.L., L.X. Revised and approved the final version of the paper: L.X., Y.Q.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Table S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Guo, J., Ling, H., Wu, Q. et al. The choice of reference genes for assessing gene expression in sugarcane under salinity and drought stresses. Sci Rep 4, 7042 (2014). https://doi.org/10.1038/srep07042
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07042
This article is cited by
-
Selection and validation of reliable reference genes for quantitative real-time PCR in Barnyard millet (Echinochloa spp.) under varied abiotic stress conditions
Scientific Reports (2023)
-
Selection and validation of reference genes by RT-qPCR under photoperiodic induction of flowering in sugarcane (Saccharum spp.)
Scientific Reports (2021)
-
Selection of reference genes for normalization of microRNA expression in sugarcane stalks during its interaction with Colletotrichum falcatum
3 Biotech (2021)
-
Screening of stable internal reference gene of Quinoa under hormone treatment and abiotic stress
Physiology and Molecular Biology of Plants (2021)
-
Identification and validation of new reference genes for accurate quantitative reverse transcriptase-PCR normalization in the Antarctic plant Colobanthus quitensis under abiotic stress conditions
Polar Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.