Abstract
A comparison of national surveys on oral health suggested that the population of South Korea has a better periodontal health status than that of Japan, despite their similar inherent backgrounds. Here, we investigated differences in oral bacterial assemblages between individuals from those two countries. To exclude potential effects of oral health condition on the microbiota, we selected 52 Korean and 88 Japanese orally healthy adults (aged 40–79 years) from the participants of two cohort studies, the Yangpyeong study in South Korea and the Hisayama study in Japan and compared the salivary microbiomes. The microbiota of the Japanese individuals comprised a more diverse community, with greater proportions of 17 bacterial genera, including Veillonella, Prevotella and Fusobacterium, compared to the microbiota of the Korean individuals. Conversely, Neisseria and Haemophilus species were present in much lower proportions in the microbiota of the Japanese individuals than the Korean individuals. Because higher proportions of Prevotella and Veillonella and lower proportions of Neisseria and Haemophilus in the salivary microbiome were implicated in periodontitis, the results of this study suggest that the greater proportion of dysbiotic oral microbiota in the Japanese individuals is associated with their higher susceptibility to periodontitis compared to the Korean individuals.
Similar content being viewed by others
Introduction
South Korea and Japan are neighboring countries located in East Asia that are separated by a sea. The populations of both countries are composed primarily of the Asian mongoloid race and the genotypes of South Koreans are more similar to those of Japanese than other Asian people1. Both countries are industrialized and have a high standard of living and they share many similar demographic and socioeconomic metrics, such as high life expectancy2, low total fertility rate per female2, high national income per capita2 and a high percentage of people who have attained tertiary education3.
The number of dentists per 10,000 people is higher in Japan (7.4) than in South Korea (5.0)2. Universal health insurance has been instituted in both countries, but Japanese national insurance covers a wider array of oral health services than Korean national insurance in many cases (e.g., denture prosthesis). Given these circumstances, Japanese individuals likely have easier access to dental care than those in South Korea. Furthermore, a comparison of national surveys in South Korea4 and Japan5 found higher percentages of daily use of dental floss (11.1 and 16.7% of adults in South Korea and Japan, respectively) and interproximal brushes (9.5 and 28.8%) and a lower percentage of current smokers (39.1 and 20.1%) in Japan than South Korea, suggesting a high level of concern for oral health among Japanese adults. Nevertheless, interestingly, the prevalence of periodontitis in Japan (44.7 and 41.5% of people aged over 19 years in 2005 and 2011, respectively) is higher than in South Korea (32.9 and 23.9% of people aged over 18 years in 2008 and 2010, respectively), based on the results of national surveys4,6,7,8.
Periodontitis is a gingival inflammation that results in the destruction of tooth-supporting tissue and is a major cause of tooth loss in adults9. It is recognized as a chronic polymicrobial disease caused by accumulation of dental plaque in the gingival sulcus and several oral bacteria that have been identified frequently in diseased sites, such as Porphyromonas gingivalis, Tannerella forsythia and Treponema denticola, have received considerable attention as direct pathogens10. However, these bacteria could play an alternative role in the etiology of periodontal disease. When P. gingivalis was inoculated into the oral cavity of specific-pathogen-free (SPF) and germ-free (GF) mice, only SPF mice developed bone loss; moreover, a shift in the commensal oral microbiota was observed in SPF mice infected with P. gingivalis11. These results suggest that the entire microbiota may also be associated with bone loss. We showed that the relative abundances of dominant indigenous bacteria in saliva differed according to periodontal heath condition12. Here, we hypothesized that the lower susceptibility to periodontal disease in the South Korean than the Japanese population might reflect differences in the bacterial assemblage in the oral cavity between the two countries.
In the present investigation, we compared the salivary microbiome and oral health status of adult participants of the following cohort studies; the Yangpyeong cohort study in South Korea and the Hisayama cohort study in Japan. The bacterial community structure was assessed using a molecular approach based on the 16S rRNA gene and we investigated differences in the oral microbiome between the populations of South Korea and Japan.
Results
We performed oral examinations on 543 adult residents of Yangpyeong county in South Korea in 2011 and 2,272 residents of Hisayama town in Japan in 2007 and collected their saliva as part of longitudinal cohort studies performed in each country (the Yangpyeong and Hisayama studies, respectively). The oral examinations were conducted by two examiners in the Korean study and nine examiners in the Japanese study. Although the examiners showed high kappa values (>0.75) within each study, no calibration was performed between the Korean and Japanese examiners, which may impose some limitations when comparing the oral clinical conditions of the two studies. The general and clinical parameters of each study population are given in Table S1. The composition of the salivary bacterial populations of all 2,815 individuals in both populations was compared by terminal restriction fragment length polymorphism (T-RFLP) analysis, which is a fingerprinting approach based on the 16S rRNA gene. The principal components analysis (PCA) diagram showed that Korean individuals were localized in the negative direction of principal component 1 (PC1) relative to Japanese individuals (Figure S1A). A significant difference in T-RFLP profiles between Korean and Japanese subjects was confirmed by PERMANOVA (P < 0.001). The loading plot of the first two principal components suggested that the microbiota of Korean individuals contained higher proportions of Neisseria, Haemophilus and Porphyromonas and lower proportions of Prevotella and Veillonella compared with Japanese individuals (Figure S1B and Table S2).
Although notable differences were observed in the T-RFLP profiles of the Korean and Japanese individuals, the possibility exists that they simply reflect differences in the oral health conditions of the two study populations described in Table S1. Therefore, we selected 140 orally healthy individuals from each study population (52 Korean and 88 Japanese individuals), according to the following inclusion criteria; ≥ 24 teeth, no use of dentures, no decayed teeth, ≤10 caries-experienced teeth, no sites with periodontal pockets (≥4 mm), no sites of severe clinical attachment loss (≥5 mm) and ≤20% of sites showing bleeding on probing. The individuals' microbiomes were compared in greater detail using barcoded pyrosequencing analysis of the 16S rRNA gene. We determined 1,006,247 bacterial 16S rRNA gene sequences (containing the V1–V2 region), of which 419,397 passed quality control tests (average length, 333 ± 7 bases; Table S3). The sequences were assigned to 2,703 species-level operational taxonomic units (OTUs) using a cutoff distance of 0.03. The principal coordinate analysis (PCoA) plot based on weighted UniFrac, which is a phylogeny-based distance metric, revealed that the overall composition of the microbiota of the Korean individuals was distinct from that of the Japanese individuals, even when limited to orally healthy individuals (partial η2 = 0.18, P < 0.001, PERMANOVA; Figure 1A). The analysis also suggested the geographic location had a greater effect on the microbiota than age, gender, obesity, or smoking status, although smoking status did demonstrate a significant effect on the microbiota (Figure S2 and Table S4). In addition, the discrimination was stronger with the weighted UniFrac (Figure 1A) than with the unweighted metric (partial η2 = 0.03, P < 0.001, PERMANOVA; Figure S3), suggesting that based on geographical location, the microbiota had greater differences in community structure than community membership. Other general health conditions such as diabetes, heart disease and antibiotic use were not considered in the present study. These are factors that will require further exploration in future studies.
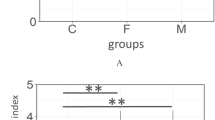
(A) Principal coordinate analysis (PCoA) plot showing similarity relationships among bacterial community samples from 140 orally healthy individuals (52 South Koreans and 88 Japanese) using the weighted UniFrac distance metric. To correct the unequal number of sequences, we evaluated it based on 1500 randomly selected sequences per sample. The two components explained 48.1 and 16.2% of the variance, respectively. Samples collected in the two countries are depicted using different colors and shapes. (B) Mean number of operational taxonomic unit (OTU), Shannon diversity index and phylogenetic diversity in the salivary microbiota of orally healthy individuals in South Korea (n = 52) and Japan (n = 88). To correct the unequal number of sequences, we evaluated each index of 1500 randomly selected sequences. The significance of differences was evaluated using Student's t-tests. The error bars indicate 95% confidence intervals. (C) Relative abundances of 21 bacterial genera that differed significantly between South Korean and Japanese orally healthy individuals (P < 0.05). The significance of differences was evaluated using Wilcoxon rank-sum test. The error bars indicate 95% confidence intervals.
All three indices of alpha diversity (the number of OTUs, Shannon diversity index and phylogenetic diversity) indicated that salivary bacterial populations in the Korean individuals contained less diverse communities compared to the Japanese individuals (Figure 1B). The vast majority of the sequences were assigned to five bacterial phyla (Firmicutes, Proteobacteria, Bacteroidetes, Actinobacteria and Fusobacteria); TM7, Spirochaetes, SR1, Tenericutes, Cyanobacteria and Planctomycetes were also identified, but at lower proportions in both study populations. The sequences obtained in this study represented 97 genera and 43 upper-level taxa; 73 of the 97 genera were present in both study populations. Ten genera, including Streptococcus, were found in all 140 subjects and constituted 86.0 ± 5.0% (mean ± SD) of the microbiota (Figure S4). Four bacterial genera, Neisseria and Haemophilus, in particular, were significantly more predominant in Korean subjects compared to Japanese individuals, while 17 genera, including Veillonella, Prevotella, Fusobacterium, Gemella and Granulicatella, were present in significantly higher proportions in the salivary bacterial populations of Japanese individuals (Figure 1C).
We also performed partial least-squares discriminant analysis (PLS-DA) with the OTU dataset to identify the key OTUs responsible for the differences in the microbiome of Korean and Japanese individuals. OTUs with variable importance in projection (VIP) values >2 were selected as important contributors to the separation (Figure 2). One OTU, which corresponded to Neisseria flavescens, was strongly associated with the microbiota of Korean individuals. In contrast, 12 OTUs that corresponded to bacterial species such as Veillonella rogosae, Fusobacterium nucleatum, Prevotella melaninogenica and Gemella sanguinis were included in the microbiomes of Japanese individuals.
Relative abundance distribution of operational taxonomic units (OTUs) that were important contributors to differences in the salivary microbiome between orally healthy individuals in Japan and South Korea that were identified using partial least-squares discriminant analysis (PLS-DA).
OTUs with variable importance in projection (VIP) values >2 were selected. The relative abundance of each OTU was normalized to have a mean of 0 and standard deviation of 1 (z-score normalization) and is represented by the color intensity of each grid (blue, low abundance; red, high abundance). The OTUs were ordered according to the results of a hierarchical cluster analysis using correlation distance with average linkage (shown as a dendrogram on the left).
Discussion
Distinctive salivary microbiomes in different geographic groups were reported by Nasidze et al.13 in a comparison of Batwa Pygmies from Uganda and African agriculturalists from Sierra Leone and the Democratic Republic of the Congo. They suggested that their distinct lifestyle and diet (hunter–gatherer versus agricultural styles) had an impact on the microbiota. The present study also demonstrated clear differences in the salivary microbiome between orally healthy individuals inhabiting Yangpyeong county in South Korea and Hisayama town in Japan. Both the Korean and Japanese individuals enrolled in this study lived in a suburb of a metropolitan area in an industrialized country in East Asia; therefore, their lifestyles and dietary habits, as well as sanitary and socioeconomic status, were presumably more similar than between the above-mentioned African peoples. Although a recent study indicated that ethnic affiliation influences oral microbial community14, the Korean and Japanese genotypes are very close, even among Asian peoples1. Nevertheless, notable differences exist in diet and environmental factors between Japan and Korea. For example, Koreans prefer their food spicier or saltier compared to Japanese15 and most Koreans consume kimchi, a spiced fermented cabbage, almost daily16. Such spicy or salty and fermented foods might affect oral bacterial species. Although the determinants of the differences in the microbiome in this study remain unclear, the geographical differences suggest that environmental factors that differ between South Korea and Japan influence the overall composition of the salivary microbiome.
In a previous study on the population of Japan using T-RFLP and clone library analyses, we found that salivary microbiomes with larger proportions of Prevotella and Veillonella were associated with periodontitis, whereas larger proportions of Neisseria, Haemophilus and Porphyromonas were associated with periodontal health12. Similarly, in this study, the T-RFLP profiles of orally healthy individuals involved in pyrosequencing analysis suggested that their microbiomes comprised higher proportions of Neisseria and Haemophilus and lower proportions of Prevotella and Veillonella compared to those of the remaining individuals of both cohorts (Figure S5 and Table S5). Therefore, the salivary microbiome composition of the Korean individuals could be healthier than that of the Japanese subjects, even though all of these individuals were orally healthy.
Imbalances in commensal microbiota of the intestinal mucosa, so-called dysbiosis, have been reported to drive mucosal inflammation17,18. In particular, Elinav et al.18 demonstrated that dysbiosis with expansion of Prevotellaceae and TM7 results in transmissible autoinflammation in the murine intestine. Also, oral dysbiosis is likely characterized by greater proportions of certain bacterial genera, including Prevotella, Veillonella and TM7, which elicit an inflammatory response in the gingival mucosa. Indeed, Said etal.19 reported that the salivary microbiome of patients with inflammatory bowel disease comprised higher proportions of Prevotella and Veillonella and lower proportions of Streptococcus, Neisseria, Haemophilus and Gemella compared with healthy controls and that higher levels of several inflammatory cytokines and IgA relative to healthy controls were detected in their saliva. However, further study, possibly using animal models, is required to clarify the role of dysbiotic oral microbiota in the onset of gingivitis and periodontitis.
Twelve OTUs were selected as important contributors (VIP values >2) to the salivary microbiomes of the Japanese subjects in the PLS-DA (Figure 2), but none corresponded to well-known periodontal pathogens such as P. gingivalis, T. denticola, or T. forsythia10. However, note that Fusobacterium nucleatum was present at high levels in the microbiomes of Japanese individuals. This species is regarded as an important “bridge” organism that coaggregates with early colonizers, such as Streptococcus species and late colonizers, including the above-mentioned periodontal pathogens, in dental plaque development20. The reduced proportion of F. nucleatum in the microbiome might also be associated with lower susceptibility to periodontitis in the Korean individuals through inhibition of pathogenic plaque development. On the other hand, one OTU, which corresponded to N. flavescens, was strongly associated with the microbiota of the Korean individuals. Although no previous report has implicated this species in periodontal health, its potential role as an oral health marker and probiotic species should be assessed in future studies.
Hisayama town is recognized to be demographically representative of Japan according to the national census21. The difference in periodontal health observed in this study was consistent with the results of national surveys. However, Yangpyeong county might not be socio-demographically representative of South Korea. Hence, this survey might not be representative of the oral health condition and the oral microbiome composition. Further studies in different regions of Japan and South Korea are needed to clarify cross-national differences.
This study revealed geographical differences in the salivary microbiomes of people living in South Korea and Japan. The greater proportion of presumed dysbiotic oral microbiota in the Japanese individuals is likely associated with their higher susceptibility to periodontitis compared with the Korean individuals. The oral microbiome is influenced by environmental factors such as dietary habits and lifestyle13,22. Improvements in environmental factors identified based on cross-national differences between South Korea and Japan might provide a novel and effective approach to maintaining periodontal health.
Methods
Ethics statement
All participants understood the nature of the study and provided informed consent. The Ethics Committee of Kyushu University in Japan (the reference number: 19B-1, 24-129) and Seoul National University in Korea (the reference number: S-D20100006) approved this study design and the procedure for obtaining informed consent. All experiments were performed in accordance with the approved guidelines.
Study population
Saliva samples were collected from participants of the Hisayama cohort study in Japan and the Yangpyeong cohort study in South Korea. Details of each study are described below.
1) Hisayama cohort study
A prospective population-based follow-up study of cardiovascular disease has been performed since 1961 in the town of Hisayama, which is a suburb of the Fukuoka metropolitan area in western Japan21. The population of the town is approximately 8,000 and it has been shown to be demographically representative of Japan based on the national census21. As part of the follow-up survey, we performed a health examination, including a dental examination, of Hisayama residents aged 40 years and older in 2007. Among all residents aged 40–79 years, 2,861 residents consented to participate in the study. Dental and medical examinations were performed on 2,669 subjects. Of these, 397 subjects who had missing data or, from whom sufficient saliva was not collected were excluded. Ultimately, 2,272 individuals (1,011 males, 1,261 females) were used in this study.
2) Yangpyeong cohort study
As part of the Korean Genomic Study for cardiovascular disease (KoGES-CVD), the Yangpyeong cohort study has been undertaken since 2004 in Yangpyeong county, a suburb of the Seoul metropolitan area, located 45 km east of Seoul, in South Korea. The population of Yangpyeong county is 78,490 and the number of residents who are aged 40–79 years is 45,298 according to the national population census in 2010. Among them, 3,398 residents participated in the baseline during the first phase, 2004–2006. Dental examinations were added to the KoGES-CVD study during the third phase, 2010–2013 and saliva samples were collected as part of the study in 2011. A total of 676 males and females aged 40–79 years living in Yangpyeong county were voluntarily recruited and received dental and medical examinations. Among these, 133 subjects who had missing data or, from whom sufficient saliva was not collected were excluded. Finally, 543 individuals (198 males, 345 females) were included in the analysis.
Dental examination
The number of teeth, dental caries (the number of decayed, missing and filled teeth) and denture wearing were examined in both the Hisayama and Yangpyeong studies. The number of decayed, missing and filled teeth value signifies teeth with caries experience and represents the caries status of the individual. Periodontal condition was evaluated based on periodontal pocket depth (PD), clinical attachment loss (CAL) and bleeding on probing (BOP). The periodontal pocket is the pathologic space between the gingiva and the tooth root and its depth is used for the clinical diagnosis of periodontal disease. The depth of these spaces is normally 1–3 mm in periodontally healthy individuals, but deepens as supporting connective tissue and alveolar bone are destroyed by persistent gingival inflammation9. A deep pocket usually indicates existing periodontal inflammation, whereas severe attachment loss is usually indicative of a history of periodontal destruction, which does not always suggest periodontal inflammation. BOP is bleeding caused by picking the inside of the periodontal pocket with a periodontal pocket probe and is a visible symptom of gingival inflammation that is suggestive of an active phase of periodontal disease. In the Hisayama study, periodontal status was assessed using PD, CAL and BOP at two sites for all teeth (the mesio-buccal and mid-buccal sites), except for the third molars since these teeth when partially impacted frequently have pseudopockets, based on the third National Health and Nutrition Examination Survey (US) III method. Details of the periodontal examination have been described elsewhere23. In the Yangpyeong study, PD, CAL and BOP were measured at six sites per tooth (mesio-buccal, mid-buccal, disto-buccal, mesio-lingual, mid-lingual and disto-lingual) on the following index teeth: 12, 15, 17, 21, 24, 26, 32, 35, 37, 41, 44 and 46. To compare periodontal status between the Hisayama and Yangpyeong studies, we selected data for PD, CAL and BOP on mesio-buccal and mid-buccal sites for the 12 above-mentioned index teeth.
Saliva collection and DNA extraction
After the dental examination, the subjects were asked to chew gum for 2 min and stimulated saliva samples were collected in sterile plastic tubes. The samples were stored at −30°C until further analysis. DNA extraction from saliva samples was performed according to a protocol described previously24.
T-RFLP analysis
From each DNA sample, internal regions of 16S rRNA genes were amplified using the universal forward primer 8F (5′-AGA GTT TGA TYM TGG CTC AG-3′) labeled at the 5′ end with 6-carboxyfluorescein (6-FAM) and the universal primer 806R (5′-GGA CTA CCR GGG TAT CTA A-3′) labeled at the 5′ end with hexachlorofluorescien (HEX). PCR amplification, purification, digestion by the restriction enzyme HaeIII and electrophoresis were performed as described previously12. Prior to alignment of T-RFLP profiles, the peaks representing less than 1% of the total peak area were excluded and the percentages of the remaining peaks recalculated. T-RFLP profiles containing electrophoresis data based on each 5′-terminal restriction fragment (TRF) labeled with different fluorescent dyes (6-FAM and HEX) per subject were aligned TRFs with molecular weights that differed 40 or less were considered identical. The aligned profiles were subjected to principal-component analysis (PCA), after excluding TRFs that were detected in fewer than 10% of the subjects. PCA was performed using library ade4 in R version 3.0.1 (available from URL: http://www.r-project.org/). Candidate bacterial species that corresponded to combinations of a 6-FAM labeled TRF and a HEX labeled TRF were selected from 831 oral bacterial 16S rRNA gene sequences (HOMD 16S rRNA RefSeq Version 13.2) deposited in the Human Oral Microbiome Database25. The matching window was set to a molecular weight of ± 600.
Barcoded pyrosequencing analysis
Barcoded pyrosequencing analysis of the 16S rRNA gene was performed for 52 Korean and 88 Japanese individuals that were in a healthy oral condition. The V1–V2 regions of 16S rRNA genes from each sample were amplified using the following primers: 338R with the 454 Life Sciences (Roche, Basel, Switzerland) adaptor B sequence (5′-CCT ATC CCC TGT GTG CCT TGG CAG TCT CAG TGC TGC CTC CCG TAG GAG T -3′) and 27F with the 454 Life Sciences adaptor A and sample-specific 10-base barcodes (5′-CCA TCT CAT CCC TGC GTG TCT CCG ACT CAG XXX XXX XXX XAG AGT TTG ATC MTG GCT CAG-3′). PCR amplification was performed as previously described26. The amplicons were gel-purified using a Wizard SV Gel and PCR Clean-Up System (Promega, Madison, WI) according to the manufacturer's instructions. DNA concentration and quality were assessed using a spectrophotometer (NanoDrop Technologies, Wilmington, DE) and equal amounts of DNA were pooled together. Pyrosequencing was conducted using a 454 Life Sciences Genome sequencer FLX instrument (Roche) at Hokkaido System Science Co., Ltd. (Sapporo, Japan) in three runs as described in Table S5.
Data analysis and taxonomy assignment
Sequences were excluded from the analysis using a script written in PHP if they were shorter than 240 bases or had an average quality score <25 and they were subsequently removed using a script written in R if they did not include the correct forward and reverse primer sequences, had a homopolymer run >6 nt or contained ambiguous characters. The remaining sequences were assigned to the appropriate sample by examining the 10-base barcode sequence. Similar sequences were clustered into operational taxonomic units (OTUs) using UCLUST27 in QIIME28, with a minimum pairwise identity of 97%. The most abundant sequence in each OTU was chosen to represent that OTU. The representative sequences were aligned using PyNAST29 and the Greengenes database30 using a minimum identity of 75%. Chimeras were removed from the representative set after being identified using Chimera Slayer31 and after verifying that the putative chimera appeared in only one sample. After chimera elimination, a relaxed neighbor-joining tree was built using FastTree32. The UniFrac metric33 was used to determine the dissimilarity between any pair of bacterial communities. The similarity relationship, assessed using the UniFrac metric, was presented in a PCoA plot, drawn using R 3.0.1. OTU number, Shannon diversity index and phylogenetic diversity34—the sum of all branch lengths in a 16S rRNA gene phylogenetic tree of each sample—were also calculated using R. The taxonomy of representative sequences was determined using the RDP classifier with a minimum support threshold of 60% and the RDP taxonomic nomenclature (to the genus level). For each representative sequence of OTUs that were identified as key phylotypes responsible for differences between Korean and Japanese subjects, nearest-neighbor species with over 98% identity were selected as candidates using BLAST searches against 831 oral bacterial 16S rRNA gene sequences (HOMD 16S rRNA RefSeq version 13.2) in the Human Oral Microbiome Database25.
Statistical analysis
All statistical analyses were conducted using R version 3.0.1. Number of OTU and Shannon index was calculated using vegan library. Phylogenetic diversity was calculated using pd function in picante library. Wilcoxon rank sum tests were used to compare microbial richness, microbial diversity and the relative abundance of each genus using wilcox_test function in coin library. We adjusted the obtained P-values by a Benjamini–Hochberg false discovery rate correction to evaluate the difference in the relative abundance of each genus. Permutational multivariate analysis of variance (PERMANOVA) was used to assess the effects of geographical differences between the South Korean and Japanese populations or host factors (age, gender, obesity, or smoking status) on the salivary microbiome using adonis function in vegan library based on 9,999 permutations. Partial least-squares discriminant analysis (PLS-DA) was used to distinguish Korean and Japanese subjects using the OTU data set and to identify the key OTUs responsible for the differences in the microbiome using mixOmics library. OTUs with high (>2) variable importance in projection (VIP) values35 were selected as important contributors for separation.
Additional Information
Accession codes: The obtained sequence data were deposited in DDBJ Sequence Read Archive under accession number DRA002249, DRA002250 and DRA002251.
References
Hugo Pan-Asian SNP Consortium. Mapping human genetic diversity in Asia. Science 326, 1541–1545 (2009).
World Health Organization. World Health Statistics. World Health Organization, Geneva, Switzerland (2013).
Organisation of Economic Co-operation and Development [OECD]. Education at a glance 2013: OECD Indicators. OECD Publishing. http://dx.doi.org/10.1787/eag-2013-en (2013) Data of access: 25/11/2013.
Korea Center for Disease Control and Prevention & Ministry of Health and Welfare. Korean national health and examination suveys: the 4th surveys. Available at http://knhanes.cdc.go.kr/ (2008) Data of access: 25/11/2013.
Ministry of Health, Labour and Welfare of Japan. The National Health and Nutrition Survey in Japan [in Japanese]. Available from URL: http://www.mhlw.go.jp/bunya/kenkou/eiyou/h21-houkoku.html (2009) Data of access: 25/11/2013.
Dental Health Division of Health Policy Bureau, Ministry of Health, Labour and Welfare Japan. Report on the Survey of Dental Diseases 2005 by Health Policy Bureau, Ministry of Health, Labour and Welfare Japan [in Japanese]. Available from URL: http://www.mhlw.go.jp/toukei/list/62-17c.html (2006) Data of access: 25/11/2013.
Dental Health Division of Health Policy Bureau, Ministry of Health, Labour and Welfare Japan. Report on the Survey of Dental Diseases 2011 by Health Policy Bureau, Ministry of Health, Labour and Welfare Japan [in Japanese] Available from URL: http://www.mhlw.go.jp/toukei/list/62-17c.html (2012) Data of access: 25/11/2013.
Korea Center for Disease Control and Prevention & Ministry of Health and Welfare. Korean national health and examination suveys: the 5th surveys. Available at http://knhanes.cdc.go.kr/ (2010) Data of access: 25/11/2013.
Pihlstrom, B. L., Michalowicz, B. S. & Johnson, N. W. Periodontal diseases. Lancet 366, 1809–1820 (2005).
Socransky, S. S., Haffajee, A. D., Cugini, M. A., Smith, C. & Kent, R. L., Jr Microbial complexes in subgingival plaque. J Clin Periodontol 25, 134–144 (1998).
Darveau, R. P., Hajishengallis, G. & Curtis, M. A. Porphyromonas gingivalis as a potential community activist for disease. J Dent Res 91, 816–820 (2012).
Takeshita, T. et al. The ecological proportion of indigenous bacterial populations in saliva is correlated with oral health status. ISME J 3, 65–78 (2009).
Nasidze, I. et al. High diversity of the saliva microbiome in Batwa Pygmies. PLoS One 6, e23352 (2011).
Mason, M. R., Nagaraja, H. N., Camerlengo, T., Joshi, V. & Kumar, P. S. Deep sequencing identifies ethnicity-specific bacterial signatures in the oral microbiome. PLoS One 8, e77287 (2013).
Lee, W. C., Lee, M. J., Kim, J. S. & Park, S. Y. Foodborne illness outbreaks in Korea and Japan studied retrospectively. J Food Prot 64, 899–902 (2001).
Lee, S., Sung, J., Lee, J. & Ko, G. Comparison of the gut microbiotas of healthy adult twins living in South Korea and the United States. Appl Environ Microbiol 77, 7433–7437 (2011).
Garrett, W. S. et al. Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell Host Microbe 8, 292–300 (2010).
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011).
Said, H. S. et al. Dysbiosis of salivary microbiota in inflammatory bowel disease and its association with oral immunological biomarkers. DNA Res 21, 15–25 (2014).
Kolenbrander, P. E., Palmer, R. J., Jr, Periasamy, S. & Jakubovics, N. S. Oral multispecies biofilm development and the key role of cell-cell distance. Nat Rev Microbiol 8, 471–480 (2010).
Hata, J. et al. Secular trends in cardiovascular disease and its risk factors in Japanese: half-century data from the Hisayama Study (1961–2009). Circulation 128, 1198–1205 (2013).
Stahringer, S. S. et al. Nurture trumps nature in a longitudinal survey of salivary bacterial communities in twins from early adolescence to early adulthood. Genome Res 22, 2146–2152 (2012).
Shimazaki, Y. et al. Effectiveness of the salivary occult blood test as a screening method for periodontal status. J Periodontol 82, 581–587 (2011).
Takeshita, T., Nakano, Y. & Yamashita, Y. Improved accuracy in terminal restriction fragment length polymorphism phylogenetic analysis using a novel internal size standard definition. Oral Microbiol Immunol 22, 419–428 (2007).
Chen, T. et al. The Human Oral Microbiome Database: a web accessible resource for investigating oral microbe taxonomic and genomic information. Database (Oxford) 2010, baq013 (2010).
Takeshita, T. et al. Enteral tube feeding alters the oral indigenous microbiota in elderly adults. Appl Environ Microbiol 77, 6739–6745 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336 (2010).
Caporaso, J. G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26, 266–267 (2010).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72, 5069–5072 (2006).
Haas, B. J. et al. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21, 494–504 (2011).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26, 1641–1650 (2009).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71, 8228–8235 (2005).
Faith, D. P. Conservation Evaluation and Phylogenetic Diversity. Biological Conservation 61, 1–10 (1992).
Perez-Enciso, M. & Tenenhaus, M. Prediction of clinical outcome with microarray data: a partial least squares discriminant analysis (PLS-DA) approach. Hum Genet 112, 581–592 (2003).
Acknowledgements
This study was supported in part by Grants-in Aid for Scientific Research 25463249 (T. T.), 22406034 (Y. S.) and 25293428 (Y. Y.) from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Contributions
T.T., Y.S., Y.Y. designed and supervised the research project. T.T. wrote the paper. H.-D.K., T.Y. and Y.Y. edited the paper. T.N. and Y.K. supervised the Hisayama cohort study. Y.S. supervised oral examination and saliva collection in Hisayama cohort study. T.T., K.M., M.F., Y.S, Y.S and S.A. conducted oral examination and saliva collection. D.-H.H., H.-D.K supervised oral examination and saliva collection in the Yangpyeong cohort study and K.M., D.-H.H. collected saliva. K.M, K.F. and T.T. conducted molecular analysis and data processing. M.F. analyzed clinical data. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplemental information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Takeshita, T., Matsuo, K., Furuta, M. et al. Distinct composition of the oral indigenous microbiota in South Korean and Japanese adults. Sci Rep 4, 6990 (2014). https://doi.org/10.1038/srep06990
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06990
This article is cited by
-
Microbiome composition comparison in oral and atherosclerotic plaque from patients with and without periodontitis
Odontology (2021)
-
Salivary microbiome in patients undergoing hemodialysis and its associations with the duration of the dialysis
BMC Nephrology (2020)
-
Response of the human gut and saliva microbiome to urbanization in Cameroon
Scientific Reports (2020)
-
Alteration of salivary microbiome in periodontitis with or without type-2 diabetes mellitus and metformin treatment
Scientific Reports (2020)
-
Impact of a vegan diet on the human salivary microbiota
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.