Abstract
The stage of evolution is the population of reproducing individuals. The structure of the population is known to affect the dynamics and outcome of evolutionary processes, but analytical results for generic random structures have been lacking. The most general result so far, the isothermal theorem, assumes the propensity for change in each position is exactly the same, but realistic biological structures are always subject to variation and noise. We consider a finite population under constant selection whose structure is given by a variety of weighted, directed, random graphs; vertices represent individuals and edges interactions between individuals. By establishing a robustness result for the isothermal theorem and using large deviation estimates to understand the typical structure of random graphs, we prove that for a generalization of the Erdős-Rényi model, the fixation probability of an invading mutant is approximately the same as that of a mutant of equal fitness in a well-mixed population with high probability. Simulations of perturbed lattices, small-world networks and scale-free networks behave similarly. We conjecture that the fixation probability in a well-mixed population, (1 − r−1)/(1 − r−n), is universal: for many random graph models, the fixation probability approaches the above function uniformly as the graphs become large.
Similar content being viewed by others
In physics, a system exhibits universality when its macroscopic behavior is independent of the details of its microscopic interactions1. Many physical models are conjectured as universal and long programs have been carried out to establish this mathematically2,3. However such universality conjectures have been lacking in biological models.
It is well known that population structure can affect the behavior of evolutionary processes under both constant selection4,5,6,7,8,9,10, on which we focus here and frequency dependent selection11,12,13,14,15,16,17,18,19,20,21,22,23. However, so far deterministic and highly organized population structures have received the most attention24,25,26,27,28; while some populations are accurately modeled in this way29,30,31,32,33, often a random structure is far more appropriate to describe the irregularity of the real world34,35,36,37. Random population structures have been considered numerically, but analytical results have been lacking7,13,38.
The Moran process considers a population of n individuals, each of which is either wild-type or mutant with constant fitness 1 or r respectively, undergoing reproduction and death39. At each discrete time step an individual is chosen randomly for reproduction proportional to its fitness; another individual is chosen uniformly at random for death and is replaced by a new individual of the same phenotype as the reproducing individual. In the long run, the process has only two possible outcomes: the mutant fixes and the wild-type dies out or the reverse. When a single mutant is introduced randomly into a homogenous, wild-type population, we call the probability of the first eventuality the fixation probability.
Fixation probabilities are of fundamental interest in evolutionary dynamics40. For a well-mixed population as described above, the fixation probability, denoted

depends on r and n41,42. Fixation probabilities also depend on population structure43,44, which is modeled by running the process on a graph (a collection of n vertices with edges between them) where vertices represent individuals and edges competition between individuals. Population structure forces reproducing individuals to replace only individuals with whom they are in competition, as described by the graph and thus death is no longer uniformly at random but among only the reproducing individual's neighbors. See the SI for details.
With this enrichment of the model, the effects of population structure can be understood. Simple one-rooted population structures are able to suppress selection and reduce evolution to a standstill, while intricate, star-like structures can amplify the intensity of selection to all but guarantee the fixation of mutants with arbitrarily slight fitness advantages7. The former has been proposed as a model for understanding the necessity of hierarchical lineages of cells to reduce the likelihood of cancer initiation45. Some population structures have fixation probabilities which are given exactly by  and a fundamental result, called the isothermal theorem (stated precisely in Theorem si.1.1), gives conditions for this7. As a special case of these conditions are all symmetric population structures or graphs with undirected edges. More generally, a graph is called isothermal if the sums of the outgoing and ingoing edge weights are the same for all subsets of the graph's vertices (or equivalently if a graph's weighted adjacency matrix is doubly stochastic). This is our first hint of universality but it was not the first time certain quantities were observed as independent of population structure. Maruyama introduced geographical population structure by separating reproduction, which occurs within sub-populations and migration, which occurs between sub-populations and found that the fixation probability was the same as that of a well-mixed population structure46. In the framework of evolutionary graph theory, Maruyama's model would correspond to a symmetric graph. In this sense his finding is a special case of the isothermal theorem.
and a fundamental result, called the isothermal theorem (stated precisely in Theorem si.1.1), gives conditions for this7. As a special case of these conditions are all symmetric population structures or graphs with undirected edges. More generally, a graph is called isothermal if the sums of the outgoing and ingoing edge weights are the same for all subsets of the graph's vertices (or equivalently if a graph's weighted adjacency matrix is doubly stochastic). This is our first hint of universality but it was not the first time certain quantities were observed as independent of population structure. Maruyama introduced geographical population structure by separating reproduction, which occurs within sub-populations and migration, which occurs between sub-populations and found that the fixation probability was the same as that of a well-mixed population structure46. In the framework of evolutionary graph theory, Maruyama's model would correspond to a symmetric graph. In this sense his finding is a special case of the isothermal theorem.
However, the assumptions of the isothermal theorem sit on a knife edge—when any small perturbation is made to the graph, the assumptions no longer hold and the original isothermal theorem is silent. In particular, it cannot be applied to directed, random graphs. We address these shortcomings in Section si.1, where we strengthen the forward direction of the isothermal theorem by proving a deterministic statement: we weaken the theorem's assumptions to be only approximately true for a graph G and show that the conclusion is still approximately true, that is, the fixation probability of a general graph  is approximately equal to
is approximately equal to  . We call this the robust isothermal theorem (rit).
. We call this the robust isothermal theorem (rit).
Theorem (Robust isothermal theorem). Fix 0 ≤ ε < 1. Let Gn = (Vn, Wn ≡ [wij]) be a connected graph. If for all nonempty S ⊊ Vn we have

where and
and are the sums of the outgoing and ingoing edges respectively, then
are the sums of the outgoing and ingoing edges respectively, then

The proof begins by ignoring spacial structure and considering only the number of mutants. Since  depends only on the ratio of the probability of increasing to the probability of decreasing the number of mutants for each subset, a bound on these ratios and a coupling argument establish that
depends only on the ratio of the probability of increasing to the probability of decreasing the number of mutants for each subset, a bound on these ratios and a coupling argument establish that  is close to
is close to  . Finally, the mean value theorem and smoothness properties of
. Finally, the mean value theorem and smoothness properties of  simplify the bound and yield the result. We remark that assumption (2) is necessary in the sense that there are graphs whose fixation probability is far from
simplify the bound and yield the result. We remark that assumption (2) is necessary in the sense that there are graphs whose fixation probability is far from  but whose weighted adjacency matrix is arbitrarily close to being doubly stochastic (see (si.1.10) for an example).
but whose weighted adjacency matrix is arbitrarily close to being doubly stochastic (see (si.1.10) for an example).
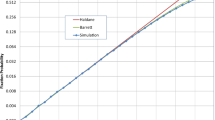
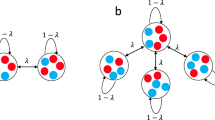
The proof verifies something essential for the process: as in physics, our laws should not depend on arbitrarily small quantities nor make disparate predictions for small perturbations of a system. The rit generalizes the isothermal theorem in this sense; if an isothermal graph is perturbed with strength ε such that the assumption (2) holds, then its fixation probability is close to that of the original graph (Figure 1). There are many ways of rigorously perturbing a graph, so we do not make a precise definition of perturbation here. All we claim is that any perturbation which changes the assumptions of the rit continuously can be controlled. The rit has many useful applications and is our first ingredient to universality.
The robust isothermal theorem guarantees that the fixation probability of each approximately isothermal graph lies in the green region.
Each edge of the 4 × 4, 2-dimensional torus is perturbed by a uniform random value from [−δ/2, δ/2] where the total of the perturbations for one vertex are conditioned to sum to 0. As the perturbation strength decreases through δ ∈ [0.4, 0] and the graph approaches isothermality, the bound improves and converges uniformly to the solid black line,  . The figures of square lattices show how random perturbations shift the graphs from isothermality, as the perturbation strength decreases from left to right; we draw each graph with the directed edges' thickness proportional to their weight and the vertices' color given by the sum of the weights of edges pointing to them. In the bottom row, empirical estimates of the fixation probabilities (small circles) are plotted against the values predicted by
. The figures of square lattices show how random perturbations shift the graphs from isothermality, as the perturbation strength decreases from left to right; we draw each graph with the directed edges' thickness proportional to their weight and the vertices' color given by the sum of the weights of edges pointing to them. In the bottom row, empirical estimates of the fixation probabilities (small circles) are plotted against the values predicted by  (solid lines) and, despite the perturbations to the graphs, their fixation probabilities lie close to
(solid lines) and, despite the perturbations to the graphs, their fixation probabilities lie close to  .
.
Robustness is essential for the analysis of random graphs. We say a random graph model exhibits universal Moran-type behavior if its fixation probability behaves like  as the graph becomes large. That is, as the graphs become large their macroscopic properties, fixation probabilities, are independent of their microscopic structures, the distributions of individual edges. Mathematically, we ask that the random variable
as the graph becomes large. That is, as the graphs become large their macroscopic properties, fixation probabilities, are independent of their microscopic structures, the distributions of individual edges. Mathematically, we ask that the random variable  converges in probability to 0, as n goes to infinity. For finite values of n, we can require finer control over this convergence such that
converges in probability to 0, as n goes to infinity. For finite values of n, we can require finer control over this convergence such that

where the functions δ(n) = o(1) and ε(n) = o(1) can be specified. For the generalized Erdős-Rényi model34 where edges are produced independently with fixed probability p (see Definitions si.2.4 and si.4.1) we prove universality. In Sections si.2 and si.4 we analyze the typical behavior of random graphs and show that with very high probability they satisfy the assumptions of the rit, giving us the paper's main result:
Theorem.Let be a family of random graphs where the directed edge weights are chosen independently according to some suitable distribution (the outgoing edges may be normalized to sum to 1 or not). Then there is a constant C > 0, not dependent on n, such that the fixation probability of a randomly placed mutant of fitness r > 0 satisfies
be a family of random graphs where the directed edge weights are chosen independently according to some suitable distribution (the outgoing edges may be normalized to sum to 1 or not). Then there is a constant C > 0, not dependent on n, such that the fixation probability of a randomly placed mutant of fitness r > 0 satisfies

uniformly in r with probability greater than 1 − exp(−ν(log n)1+ξ), for some positive constants ξ and ν.
The proof applies the rit to random graphs, showing that with high probability they satisfy assumption (2) with ε approximately order  . This relies on two main results. Using large deviation estimates, we show that the sum of the ingoing edge weights to each vertex (its temperature) are within approximately order
. This relies on two main results. Using large deviation estimates, we show that the sum of the ingoing edge weights to each vertex (its temperature) are within approximately order  of 1 with high probability (Lemma si.2.7) and that sum of the outgoing (and ingoing) edge weights of each subset are at least the same order as the size of the subset or its complement for some uniform constant with high probability (Lemma si.2.8).
of 1 with high probability (Lemma si.2.7) and that sum of the outgoing (and ingoing) edge weights of each subset are at least the same order as the size of the subset or its complement for some uniform constant with high probability (Lemma si.2.8).
This theorem isolates the typical behavior of the Moran process on these random structures. It can be interpreted as stating that random processes generating population structures where vertices and edges are treated independently and interchangeably will almost always produce graphs with Moran-type behavior. While such processes can generate graphs which do not have Moran-type behavior (for example one-rooted or disconnected graphs), these graphs are generated with very low probability as the size of the graphs becomes large. Moreover, it improves upon diffusion approximation methods by explicitly controlling the error rates47.
The result holds with high probability but sometimes this probability becomes close to 1 only as the graphs become large. The necessary graph size depends on the distribution from which the random graph's edge weights are drawn. In particular, it depends inversely on the parameter p from the generalized Erdős-Rényi model, which is the probability that there is an edge of some weight between two directed vertices. The smaller this parameter the more disordered and sparse the random graphs and the less uniform their vertices' temperatures, which all tend to decrease the control over the graph's closeness to isothermality, (2). Regardless, our choice of the parameters ξ and ν guarantees that the bound (si.2.54) decays to 0 and that it holds with probability approaching 1 as n becomes large.
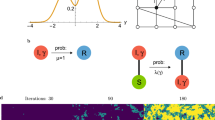
We investigated the issues of convergence for small values of n numerically to illustrate our analytical result (Figure 2). For Erdős-Rényi random graphs (see Section si.2 with the distribution chosen as Bernoulli), we generated 10 random graphs according to the procedure outlined in Definition si.2.4 for fixed values of 0 < p < 1. On each graph the Moran process was simulated 104 times for various values of 0 ≤ r ≤ 10 to give the empirical fixation probability, that is, the proportion of times that the mutant fixed in the simulation. Degenerate graphs were not excluded from the simulations but rather than estimating their fixation probabilities, we calculated them exactly, so that 1-rooted graphs were given fixation probability 1/n and many-rooted and disconnected graphs were given fixation probability 0. Trivially, such 1-rooted graphs are suppressors—that is, the fixation probability of a mutant of fitness 0 < r < 1 (and a mutant of fitness r > 1) is greater than (and less than respectively) the mutant's fixation probability in a well-mixed population—but suppressing graphs without these degenerate properties were also observed. As the graphs become larger their fixation probabilities match  closely and degeneracy becomes highly improbable as predicted by our result.
closely and degeneracy becomes highly improbable as predicted by our result.
The fixation probability of the generalized Erdős-Rényi random graphs converge uniformly to  .
.
The three columns from left to right correspond to Erdős-Rényi random graphs with decreasing connection probabilities p = 1, p = 0.6 and p = 0.3. The representative random graphs in the top row show both the increasing sparsity and disorder as p decreases and the elimination of degeneracy (rootedness and disconnectedness) and the increasing uniformity of temperature as the graph sizes increase. In the middle row, empirical estimates of the fixation probabilities (small circles) are plotted against the values predicted by  (solid lines). When p = 1 the graphs are isothermal and thus correspond exactly to their predicted values which can be seen even more clearly in the bottom row, where the difference of the empirical fixation probabilities and their predicted values display as stochastic fluctuations about 0. For p = 0.6 and p = 0.3, the convergence of the empirical values to
(solid lines). When p = 1 the graphs are isothermal and thus correspond exactly to their predicted values which can be seen even more clearly in the bottom row, where the difference of the empirical fixation probabilities and their predicted values display as stochastic fluctuations about 0. For p = 0.6 and p = 0.3, the convergence of the empirical values to  as the graphs increase in size is apparent. Smaller graphs are typically suppressors as illustrated by the clear sign change at r = 1 in the difference of empirical and predicted values, whereas larger graphs fluctuate about 0. This phenomenon is due not only to the higher probability of obtaining degenerate graphs—simulations produced strongly connected, small suppressors. Moreover, the convergence is patently slower in n for smaller values of p.
as the graphs increase in size is apparent. Smaller graphs are typically suppressors as illustrated by the clear sign change at r = 1 in the difference of empirical and predicted values, whereas larger graphs fluctuate about 0. This phenomenon is due not only to the higher probability of obtaining degenerate graphs—simulations produced strongly connected, small suppressors. Moreover, the convergence is patently slower in n for smaller values of p.
In addition to the generalized Erdős-Rényi random graphs, we also considered the Watts-Strogatz model and the Barabási-Albert model. The Watts-Strogatz model35 produces random graphs with small-world properties, that is, high clustering and short average path length. The model has three inputs: a parameter 0 ≤ β ≤ 1, the graph size n and the mean degree 2k. Typically, the model produces random, undirected graphs, thus, to escape isothermality, it was modified slightly to produce weighted, directed graphs. We do this in the most natural way: we start with a directed 2k-regular graph where each node is connected to its 2k nearest neighbors if the graph is arranged on a cycle (Figure 3) and then we rewire each edge to a new vertex chosen uniformly at random with probability β independently. Since the number of edges leaving each vertex is fixed at 2k, the weight of each edge is exactly 1/(2k). Potentially, there can be multiple edges for one vertex to another, which we account for by summing the edge weights. The model may be viewed as an interpolation between an isothermal, 2k-regular graph and an Erdős-Rényi graph by the parameter β.
Small-world networks also show universal behavior.
Representative Watts-Strogatz random graphs display increasing disorder as the rewiring probability β increases from 0 to 1, which may be viewed as an interpolation between an isothermal graph and an Erdős-Rényi random graph. For all values of β the correspondence to  is striking but mathematical proof is lacking.
is striking but mathematical proof is lacking.
Moran-type behavior was observed in the Watts-Strogatz model for all values of the input parameters we simulated (Figure 3). While mathematical proof of universality in the Watts-Strogatz model is still needed, there is hope that the techniques of this paper may be applied in this situation as the in-degrees of the vertices are concentrated around 1 for graphs with large degree 2k.
Unlike the Erdős-Rényi and Watts-Strogatz models, scale-free networks are random graphs where the in-degrees of the vertices follow a power law. Normally, scale-free networks are undirected and unweighted. To produce weighted, directed scale-free networks, we modified the preferential attachment algorithm of Barabási-Albert48: we start with a connected cycle and then add directed edges of equal weight in sequence to a randomly selected vertex where the destination of each edge is selected proportional to the in-degree of the current vertices.
Surprisingly, even though there is a sense in which vertices are not treated interchangeably in the preferential attachment algorithm, Moran-type behavior was observed in all simulations (Figure 4). This is in contrast with the results in Lieberman et al. where they observed some amplification in scale-free networks7. The scale-free property is emergent and only becomes apparent as the graph becomes large, thus this increases the running time of the Monte Carlo method for estimating the fixation probability. More simulations are required here for conclusive findings and again, there are currently no mathematical results.
Simulations on graphs generated by preferential attachment yield fixation probabilities close to  .
.
Several scale-free networks with varying out degrees, m = 10, m = 20, m = 40 and m = 80, were generated using a preferential attachment algorithm. Histograms of the sum of the weights of edges pointing to each vertex are plotted next to each graph; however, the small graphs size limits the resemblance to a power law. Given the comparatively large size of the graphs, only a restricted number of simulations were performed, but the simulations corresponded to  without a tendency to amplify or suppress. More extensive work is required.
without a tendency to amplify or suppress. More extensive work is required.
In summary, we have generalized the isothermal theorem to make it biologically realistic and to increase its technical applicability. The conclusion of the robust isothermal theorem now depends continuously on its assumptions. With this new tool, we have proved analytically that fixation probabilities in a generalized Erdős-Rényi model converge uniformly to the fixation probability of a well-mixed population. In our proof, we identify the reason for this convergence and bound its rate. Thus, we confirm observations from many simulations and give a method of approximation with a specified error. Furthermore, we conjecture that many random graph models exhibit this universal behavior. However, it is easy to construct simple examples of random graphs which do not, thus it still remains to determine the necessary assumptions on the random graph model for it to exhibit universal behavior.
References
Deift, P. Universality for mathematical and physical systems. In: Proc. Int. Congr. Math., vol. I, 125–152 (European Mathematical Society, Zürich, 2007).
Erdős, L. & Yau, H.-T. Universality of local spectral statistics of random matrices. Bull. Am. Math. Soc. 49, 377–414 (2012).
Borodin, A. & Corwin, I. Macdonald processes. Probab. Theory Relat. Fields 158, 225–400 (2014).
Levin, S. A. & Paine, R. T. Disturbance, patch formation and community structure. Proc. Natl. Acad. Sci. 71, 2744–2747 (1974).
Levin, S. A. Population dynamic models in heterogeneous environments. Annu. Rev. Ecol. Syst. 7, 287–310 (1976).
Durrett, R. & Levin, S. A. Stochastic spatial models: a user's guide to ecological applications. Philos. Trans. R. Soc. London. Ser. B Biol. Sci. 343, 329–350 (1994).
Lieberman, E., Hauert, C. & Nowak, M. A. Evolutionary dynamics on graphs. Nature 312–316 (2005).
Broom, M. & Rychtá[rbreve], J. An analysis of the fixation probability of a mutant on special classes of non-directed graphs. Proc. R. Soc. A Math. Phys. Eng. Sci. 464, 2609–2627 (2008).
Frean, M., Rainey, P. B. & Traulsen, A. The effect of population structure on the rate of evolution. Proc. R. Soc. B Biol. Sci. 280, 20130211 (2013).
Antal, T., Redner, S. & Sood, V. Evolutionary dynamics on degree-heterogeneous graphs. Phys. Rev. Lett. 96, 188104 (2006).
Sinervo, B. & Lively, C. The rock–paper–scissors game and the evolution of alternative male strategies. Nature 380, 240–243 (1996).
Kerr, B., Riley, M. a., Feldman, M. W. & Bohannan, B. J. M. Local dispersal promotes biodiversity in a real-life game of rock-paper-scissors. Nature 418, 171–4 (2002).
Ohtsuki, H., Hauert, C., Lieberman, E. & Nowak, M. A. A simple rule for the evolution of cooperation on graphs and social networks. Nature 441, 502–505 (2006).
Chen, Y.-T. Sharp benefit-to-cost rules for the evolution of cooperation on regular graphs. Ann. Appl. Probab. 23, 637–664 (2013).
Broom, M. & Rychtá[rbreve], J. Game-theoretical models in biology. CRC mathematical and computational biology series (Chapman and Hall/CRC Press, Taylor and Francis Group, Boca Raton, FL, 2013).
Nowak, M. A. & May, R. M. Evolutionary games and spatial chaos. Nature 359, 826–829 (1992).
Tarnita, C. E., Wage, N. & Nowak, M. A. Multiple strategies in structured populations. Proc. Natl. Acad. Sci. 108, 2334–2337 (2011).
Szabó, G. & Fath, G. Evolutionary games on graphs. Phys. Rep. 446, 97–216 (2007).
Perc, M. Coherence resonance in a spatial prisoner's dilemma game. New J. Phys. 8, 22 (2006).
Perc, M. & Szolnoki, A. Social diversity and promotion of cooperation in the spatial prisoner's dilemma game. Phys. Rev. E 77, 11904 (2008).
Fu, F., Nowak, M. A. & Hauert, C. Invasion and expansion of cooperators in lattice populations: Prisoner's dilemma vs. Snowdrift games. J. Theor. Biol. 266, 358–366 (2010).
Pacheco, J. M., Traulsen, A. & Nowak, M. A. Coevolution of strategy and structure in complex networks with dynamical linking. Phys. Rev. Lett. 97, 258103 (2006).
Tarnita, C. E., Ohtsuki, H., Antal, T., Fu, F. & Nowak, M. A. Strategy selection in structured populations. J. Theor. Biol. 259, 570–581 (2009).
Johst, K., Doebeli, M. & Brandl, R. Evolution of complex dynamics in spatially structured populations. Proc. R. Soc. B Biol. Sci. 266, 1147–1154 (1999).
Broom, M., Rychtá[rbreve], J. & Stadler, B. Evolutionary dynamics on small-order graphs. J. Interdiscip. Math. 12, 129–140 (2009).
Ispolatov, J. & Doebeli, M. Diversification along environmental gradients in spatially structured populations. Evol. Ecol. Res. 11, 295–304 (2009).
Díaz, J. et al. On the fixation probability of superstars. Proc. R. Soc. A Math. Phys. Eng. Sci. 469, 20130193 (2013).
Jamieson-Lane, A. & Hauert, C. Fixation probabilities on superstars, revisited and revised. arXiv Prepr. arXiv1312.6333 (2013).
Durrett, R. & Levin, S. The Importance of Being Discrete (and Spatial). Theor. Popul. Biol. 46, 363–394 (1994).
Hassell, M. P., Comins, H. N. & May, R. M. Species coexistence and self-organizing spatial dynamics. Nature 370, 290–292 (1994).
Rainey, P. B. & Rainey, K. Evolution of cooperation and conflict in experimental bacterial populations. Nature 425, 72–74 (2003).
Le Gac, M. & Doebeli, M. Environmental viscosity does not affect the evolution of cooperation during experimental evolution of colicigenic bacteria. Evolution 64, 522–33 (2010).
Allen, B., Gore, J. & Nowak, M. A. Spatial dilemmas of diffusible public goods. Elife 2, e01169 (2013).
Erdős, P. & Rényi, A. On the evolution of random graphs. Publ. Math. Inst. Hungarian Acad. Sci. 5 (1960).
Watts, D. J. & Strogatz, S. H. Collective dynamics of ‘small-world’ networks. Nature 393, 440–442 (1998).
Barrat, A. & Weigt, M. On the properties of small-world network models. Eur. Phys. J. B 13, 547–560 (2000).
Nakamaru, M. & Levin, S. A. Spread of two linked social norms on complex interaction networks. J. Theor. Biol. 230, 57–64 (2004).
Barbosa, V. C., Donangelo, R. & Souza, S. R. Early appraisal of the fixation probability in directed networks. Phys. Rev. E 82, 046114 (2010).
Moran, P. A. P. The statistical processes of evolutionary theory (Oxford University Press, Oxford, England, 1962).
Patwa, Z. & Wahl, L. M. The fixation probability of beneficial mutations. J. R. Soc. Interface 5, 1279–1289 (2008).
Maruyama, T. A Markov process of gene frequency change in a geographically structured population. Genetics 76, 367–377 (1974).
Nowak, M. A. Evolutionary dynamics: exploring the equations of life (Belknap Press of Harvard University Press, Cambridge, Massachusetts, 2006).
Barton, N. The probability of fixation of a favoured allele in a subdivided population. Genet. Res. 62, 149–157 (1993).
Whitlock, M. Fixation probability and time in subdivided populations. Genetics 779, 767–779 (2003).
Nowak, M. A., Michor, F. & Iwasa, Y. The linear process of somatic evolution. Proc. Natl. Acad. Sci. 100, 14966–14969 (2003).
Maruyama, T. A simple proof that certain quantities are independent of the geographical structure of population. Theor. Popul. Biol. 5, 148–154 (1974).
Hadjichrysanthou, C., Broom, M. & Kiss, I. Z. Approximating evolutionary dynamics on networks using a Neighbourhood Configuration model. J. Theor. Biol. 312, 13–21 (2012).
Barabási, A.-L. & Albert, R. Emergence of scaling in random networks. Science. 286, 509–512 (1999).
Acknowledgements
Financial support from the John Templeton Foundation is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
B.A. and M.A.N. performed the research and wrote and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Adlam, B., Nowak, M. Universality of fixation probabilities in randomly structured populations. Sci Rep 4, 6692 (2014). https://doi.org/10.1038/srep06692
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06692
This article is cited by
-
Fixation probabilities in network structured meta-populations
Scientific Reports (2021)
-
Modelling Evolution in Structured Populations Involving Multiplayer Interactions
Dynamic Games and Applications (2021)
-
Exploring and mapping the universe of evolutionary graphs identifies structural properties affecting fixation probability and time
Communications Biology (2019)
-
Moran-type bounds for the fixation probability in a frequency-dependent Wright–Fisher model
Journal of Mathematical Biology (2018)
-
Amplification on Undirected Population Structures: Comets Beat Stars
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.