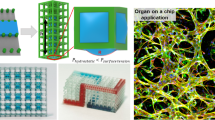
Abstract
We report a general cell surface molecular engineering strategy via liposome fusion delivery to create a dual photo-active and bio-orthogonal cell surface for remote controlled spatial and temporal manipulation of microtissue assembly and disassembly. Cell surface tailoring of chemoselective functional groups was achieved by a liposome fusion delivery method and quantified by flow cytometry and characterized by a new cell surface lipid pull down mass spectrometry strategy. Dynamic co-culture spheroid tissue assembly in solution and co-culture tissue multilayer assembly on materials was demonstrated by an intercellular photo-oxime ligation that could be remotely cleaved and disassembled on demand. Spatial and temporal control of microtissue structures containing multiple cell types was demonstrated by the generation of patterned multilayers for controlling stem cell differentiation. Remote control of cell interactions via cell surface engineering that allows for real-time manipulation of tissue dynamics may provide tools with the scope to answer fundamental questions of cell communication and initiate new biotechnologies ranging from imaging probes to drug delivery vehicles to regenerative medicine, inexpensive bioreactor technology and tissue engineering therapies.
Similar content being viewed by others
Introduction
The ability to direct cell behavior and tissue formation in vitro and in vivo is a central design feature for the development of a range of biomaterials, cell biotechnologies and tissue engineering based therapies for improving human health1,2,3,4,5. Traditional molecular biology methods have significantly advanced cell function understanding and provided a range of tools for manipulating cell behavior. Recently, cell surface engineering strategies that use bottom-up chemical approaches have gained increasing attention due to their ability to affect cell surface interactions but not require genomic manipulations6,7,8. Several chemical strategies have been used to tailor cell surfaces including metabolite analogues, cationic polymer adhesion and polymersome attachment9,10,11,12. An alternate chemical approach has used the addition of synthetic lipids delivered directly to cells in culture in order to add new functions to cell membranes13,14. New methods that rewire cell surfaces with the capability to control cell interconnectivity in space and time would allow for further exploration of a range of fundamental cell behavior studies and provide new ways to install imaging probes, advance cell based biotechnologies and accelerate regenerative medicine and tissue engineering based therapies.
Herein, we develop a general strategy that delivers photo-active and bio-orthogonal chemistry via liposome fusion to cell surfaces for subsequent in-situ tailoring for on-demand microtissue assembly and disassembly. We demonstrate this photo-active cell surface engineering system by conjugating and tracking cell surface ligands and applying a photo-cleavable click (oxime) type ligation between cells for the spatial and temporal control of multilayer tissue assembly and disassembly for generating multicellular tissues as well as manipulating stem cell differentiation. This methodology allows for real-time manipulation of tissue dynamics and may provide tools with the scope to answer fundamental questions of cell communication and initiate new biotechnologies ranging from imaging probes to drug delivery vehicles to regenerative medicine, inexpensive bioreactor technology and tissue engineering therapies.
Results and discussion
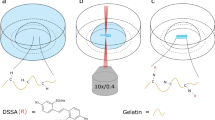
To generate tissue assemblies containing multiple cell types for a range of fundamental cell behavior, cell imaging and tissue engineering applications, we used a bottom-up synthetic approach to rewire cell surfaces. Cell surface tailoring was achieved by a straightforward new liposome fusion method to incorporate complementary bio-orthogonal molecules capable of an intercellular click chemical reaction upon physical cell-cell contact13,14. The external cell surface click conjugation between cells proceeds at physiological conditions in the presence of serum and allows for stable cell interconnectivity. The limited suite of bio-orthogonal click reactions is increasingly becoming important tools in chemical biology and cell biological research15,16. To access spatial and temporal control of cell-cell interactions, the synthetic ligation tether between cells was engineered to contain a photochemical cleavage site17. Remote controlled tissue disassembly proceeds by a programmed photo-initiated cleavage of the intercellular ligation tether (Fig. 1 top). The key features are the delivery of synthetic chemical groups to cell surfaces (via liposome fusion)13, the intercellular oxime click ligation bond16 (bio-orthogonal) and a photo-cleavage site contained within the oxyamine lipid tether (Fig. 1 bottom)17.
Schematic describing the molecular level control of tissue assembly and disassembly via a chemoselective, bio-orthogonal and photo-switchable cell surface engineering approach.
(top) Two or more different cell types are engineered via liposome fusion to present complementary and bio-orthogonal molecules on their cell surfaces. Upon interaction, a stable and covalent click (oxime) type reaction occurs and tissue like assemblies are formed. After a defined period of time, tissue disassembly can be remotely controlled by photo-cleavage of the intercellular ligation tether. (bottom) A schematic showing the interfacial molecular lock and cleave sequence for conjugating and releasing cell associations. The lipid lock molecule contains both a photocleavable group and an aminooxy group for bio-orthogonal oxime ligation to ketone presenting cells. Disassociation of cell assemblies via photo-cleavage occurs upon UV illumination.
To demonstrate temporal control of tissue assembly and disassembly, we first delivered the key functional groups to different cell populations (Fig. 2). We generated three liposome populations containing the photo-oxyamine lipid (1), ketone lipid (2) and oxyamine lipid (11) respectively (Supporting scheme S1, Supporting Fig. S1–S4). By mixing these lipid-like molecules with background lipids (palmitoyl-oleoyl phosphatidylcholine (POPC) and positively charged, 1,2-dioleoyl-3-trimethylammonium-propane (DOTAP)), we generated liposomes that can easily fuse into a variety of cells to deliver the photo-oxyamine, ketone and oxyamine groups (Fig. 2A–C). As a demonstration, we applied this liposome fusion method to non-adherent Jurkat cells to generate different two cell populations; one presenting photo-oxyamine, ketone groups and the other presenting oxyamine groups. As controls, when regular Jurkat cells were mixed with ketone tailored Jurkat cells (Jurkat-ketone), no cell assembly was observed (Fig. 2D). When oxyamine tailored Jurkat cells (Jurkat-ONH2) or photo-oxyamine tailored Jurkat cells (Jurkat-photo-ONH2) were mixed with Jurkat-ketone, rapid multi-cell spheroid assemblies were generated via the intercellular oxime click ligation (Fig. 2E–F). When mild UV illumination was applied (365 nm, 10 mW/cm2, 5 min), the photolabile oxime linkage connecting cells was cleaved resulting in the complete disassembly of the microtissue into individual cells.
The design of liposomes to deliver chemoselective, bio-orthogonal and photo-active groups to cell surfaces and the subsequent application to assemble and disassemble microtissues on demand.
(A) A photolabile oxyamine molecule is mixed with POPC and positively charged DOTAP to generate photo-active liposomes that are delivered to cell surfaces via liposome fusion. (B) Dodecanone is mixed with POPC and DOTAP molecules to generate ketone presenting liposomes that are delivered to cell surfaces via liposome fusion. (C) O-dodecyloxyamine is mixed with POPC and DOTAP molecules to generate oxyamine presenting liposomes that are delivered to cell surfaces via liposome fusion. (D) No cell assembly occurred when control Jurkat cells were mixed with ketone presenting Jurkat cells (6). (E) Programmed microtissue assembly when Jurkat-ONH2 (10) cells were mixed with jurkat-ketone cells (6). No disassembly occurred upon uv light illumination. (F) Cell Assembly proceeded upon mixing of jurkat-photo-ONH2 (5) with jurkat-ketone (6) cells. Microtissue disassembly to single cells proceeded upon cleavage of the intercellular linkage through uv light illumination.
To generate and control the size of co-culture aggregate cell assemblies, the concentration of the mixed tailored cells in solution and duration of interaction were varied. As expected, higher concentrations of cells and longer durations resulted in larger co-culture spheroids (Supporting Fig. S5). As an important control, cell viability studies showed no difference between cell populations that were tailored with and without functional groups via liposome fusion (Supporting Fig. S6). Due to the photo-cleavable nature of the intercellular oxime ligation, spheroid disassembly proceeded upon illumination with UV light (365 nm, 10 mW/cm2, 5 min). The behaviors of newly disassembled cells were indistinguishable from control cells (viability, migration, proliferation). This strategy may allow for temporal control for a range of autocrine and paracrine signaling studies and provide new ways to study co-culture and multi-cell type associations.
As a further application, cells that were rewired with the photo-oxyamine were seeded onto patterned materials presenting aldehyde groups (Fig. 3). Cells specifically attached to the material through the interfacial oxime ligation bypassing non-specific associations used in regular tissue culture and native ligand-receptor based adhesion (integrin-extracellular matrix)18,19,20. Upon exposure to UV light (365 nm, 10 mW/cm2 5 min) the photo-oxime was cleaved and the cells detached from the substrate (Supporting Fig. S7). This strategy allows for non-adherent cells to become adhesive to tailored materials and in combination with microfluidic technology may allow for new cell sorting, cell patterning and tissue capture/release biotechnologies21.
Liposome delivery of a photolabile group to cell surfaces for controllable cell capture and release on a patterned material surface.
(A) Fibroblasts presenting the photo-active oxyamine chemoselectively adhere to materials presenting aldehyde groups (B, C) through an interfacial oxime linkage. The rewired cells could then be selectively released from the material upon UV illumination (D).
Fig. 4 shows the construction of a multi-layered tissue co-culture system based on rewiring cell surfaces via liposome fusion. Two different types of cells (hMSCs and fibroblasts), tailored with ketone and photo-oxyamine respectively, were peeled from tissue culture plates and assembled via the oxime ligation to form a stable multi-layered tissue co-culture system. Images of the co-culture tissue showed regions of multilayer and monolayer. The cell layers only adhered if the complementary chemistry was present on their cell surfaces (Supporting Fig. S8). These results show that large tissues can be easily assembled via specific cell surface ligation and may provide new efficient and inexpensive bio-reactor routes to generate complex tissues and organs22,23. Furthermore, red fluorescent protein (RFP) fibroblasts and green fluorescent protein (GFP) fibroblasts were used in combination with the bio-orthogonal liposome fusion method to demonstrate that multilayer co-cultures can be assembled with control of thickness and orientation (Fig. 4 F–G).
Construction of multi-tissue co-culture systems.
Two different types of cells (fibroblasts and hMSCs), tailored with photo-oxyamine (A) and ketone (B) respectively, were peeled from tissue culture plates and assembled via the oxime ligation (C, D) to form a stable multi-tissue co-culture system. (E) Phase contrast microscopy of the tissue showing regions of multilayer and monolayer. (F) Monolayer of mixed RFP and GFP cells with no ketone or oxyamine chemistry. (G) Multilayer of mixed RFP and GFP cells via oxime ligation. (H) Oriented RFP and GFP cells via sequential addition of cells presenting ketone and oxyamine groups. The cell layers only adhered if the complementary chemistry was present on their cell surfaces.
Since the linkage between cell layers contained a photo-cleavage site, the co-culture tissue could be separated upon UV exposure (Supporting Fig. S9). These results show that the oxime ligation between cells can be scaled from initial liposome fusion (nanometer scale) to small clusters of cells (micrometer scale), to large tissue patches (centimeter scale). Furthermore, upon addition of induction media, the click ligated tissue patches containing hMSCs could differentiate to adipocytes, fibroblasts and osteoblasts24,25 (Supporting Fig. S10). Because a range of cell lines may be integrated with stem cells to generate co-culture multi-layers, new stem cell differentiation studies and higher order 3-dimensional tissues may be possible26,27. This strategy is general and may be used to produce complex multi-cell type structures for a range of regenerative medical applications (stem cell co-cultures, tissue grafts, etc…) and as a high-throughput tissue chip screening technology.
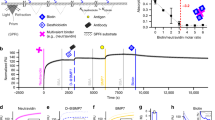
We used flow cytometry to quantify and characterize the amount of photo-oxyamine lipid delivered to cells via liposome fusion, for subsequent photo-oxime ligation induced microtissue assembly (Fig. 5). The photo-oxyamine was designed to have 3 components that are essential for spatial and temporal control of cell interconnectivity (Fig. 5A). A lipid hydrophobic component to insert into membranes, a photo-cleavable center and a bio-orthogonal oxyamine group for cell surface ligation of a range of ligands or other cells presenting ketone groups.
Characterization of the photo-oxime reaction at cell surfaces.
(A) NMR characterization of the photo-oxyamine molecule. Each part of the molecule is designed to optimize membrane insertion, photo-cleavability and availability of the oxyamine group for intercellular ligation. (B) Flow cytometry analysis to determine the amount of photo-oxyamine molecules per cell surface. Fibroblast cells were cultured with or without (control) photo-oxyamine containing liposomes for different time periods. The cells were then reacted with a ketone containing fluorescent calcein dye. The cells were then tested against a standard bead (~107 beads/mL) with known fluorescein molecule density. Approximately 25 × 103 cells were counted for all samples. Samples were run in triplicate and the mean fluorescence intensity values are displayed. (C, D) Histograms relating the number of cells counted as a function of fluorescence intensity are shown and labeled as control (without photo-oxyamine) and 12 h, 36 h, 72 h and 96 h (with photo-oxyamine). (E) Micrograph of cells presenting photo-oxyamine reacted with calcein. When no photo-oxyamine is present in the cell surface, no fluorescence is observed. (F) A cartoon describing the interfacial reaction between the surface-engineered cells and a self-assembled monolayer (SAM) on a gold substrate. Cells tailored with the photo-oxyamine group are seeded onto a SAM presenting aldehyde groups to form an interfacial oxime ligation. After washing and removing the cells, the photo-oxyamine lipid is pulled out of the cell membrane and remains ligated to the SAM via the covalent oxime bond. (G) MALDI-MS characterization of the resulting substrate shows an interfacial reaction occurs between the rewired cell surfaces and the aldehyde substrate via oxime ligation.
Liposomes containing the photo-oxyamine lipid were synthesized and then delivered to fibroblasts in culture. To measure the amount of photo-oxyamine incorporated, the cells chemoselectively reacted via oxime formation with a fluorescent calcein dye containing a ketone group. FACS analysis determined the amount of photo-oxyamine molecule present at the cell surface after various time points after liposome fusion (Fig. 5B–E). As expected, the cells initially had the highest amounts of photo-oxyamine molecule, which gradually decreased over time as the cells grew and divided. This decrease is due to multiple cell divisions and therefore dilution of the photo-oxyamine lipid molecule over subsequent generations of cells. Depending on the cell line, this dilution took weeks and many rounds of cell divisions. After dilution, the cell lines were indistinguishable from untreated cells and could again be used in normal cell culture or for future liposome fusion enhancement or new cell surface tailoring. These results and observations are analogous to classical cDNA transient transfections but in this case with photo-active bio-orthogonal lipids being the transfected biomolecule. Furthermore, FACS analysis is able to quantitate the amount of lipid at different time points and shows the amount of lipid can be controlled by adjusting the liposome fusion conditions (duration, concentration) and allows for multiple fusions of different lipid like molecules (for. eg. different bio-orthogonal lipids for hydrazone, huisgen, diels-alder, thiol-ene type conjugations or lipids with fluorescent, spin label, radiolabel etc. properties) or serial fusions at different time points.
As further characterization of cell surface presenting photo-oxyamine groups, we developed a novel mass spectrometry method (Fig. 5F). Lipid transfected cells were added to SAM surfaces presenting aldehyde groups, which allow cells to attach to the surface via the interfacial oxime ligation16,21. Cells were then removed from the surface (lysed) with vigorous agitation through a stream of PBS solution and H2O. Due to the covalent nature of the interfacial oxime bond between the cells and SAM surface, the lipid was essentially pulled out of the membrane and remained conjugated to the substrate. Because the substrate is conductive, MALDI mass spectrometry could be performed directly on the substrate to characterize the interfacial reaction. Fig. 5G shows MALDI analysis and clearly shows the presence of the oxime molecule on the substrate. This strategy may be used to characterize a range of lipid presenting molecules on cell surfaces and as a new pull-down method of proteins if the lipid molecule spans the plasma membrane and covalently associates with cytoplasmic proteins.
To demonstrate the spatial and temporal control of tissue assembly using the intercellular photo-oxime strategy, we generated multilayers of adipocytes and hMSC cells (Fig. 6). We first added a dodecanone liposome to hMSCs, which readily presented the ketone from the cell surface. Addition of adipocytes presenting photo-oxyamine to the hMSCs rapidly generated a two layer structure. In comparison, no layers are formed without the complementary chemistry presented from either cell type. Fig. 6A–C shows images and confocal representations of the layers. Upon UV illumination through a mask, the adipogenic cells were released from the hMSCs due to the cleavage of the intercellular photo-oxime ligation tether.
General schematic, optical and confocal images of chemoselective and photo-switchable tissue assembly and disassembly.
(A) Top row. Two different cell populations (hMSCs and adipocytes) tailored with ketones and photo-oxyamine, respectively, were conjugated together via the chemoselective oxime ligation to form a multilayer co-culture system (B, middle row). (C) Bottom row. Select photo-disassembly of the tissue multilayer to a monolayer occurred upon uv exposure through a mask. After partially exposing to UV, the co-culture system was partially disassembled to form specially controlled monolayer again. (D) Schematic and corresponding phase contrast and optical images displaying the formation, differentiation and release of 3D dynamic tissues using a Fibroblast/hMSC co-culture. Ketone tailored hMSCs (8) were cultured on a substrate (105 cells/mL, 3 d) and formed a 2D monolayer as shown by the image in (E). Fibroblasts presenting photo-oxyamine were then added (105 cells/mL, 2 d), producing a 3D multi-layered, interconnected co-culture tissue (F). When the appropriate induction media is used, the hMSCs differentiated into adipocytes, resulting in a 3D multi-layered co-culture of Fibroblasts and adipocytes (G). Photo-release of Fibroblasts from the co-culture left only the adhered adipocytes on the surface as a 2D monolayer shown by lower (H) and higher (I) magnified images. Adipocytes were stained for lipid vacuoles (red, Oil Red O) and nucleus (blue, Harris Hemotoxylin).
In order to extend this strategy for potential stem cell differentiation and tissue engineering applications, we generated cell layers through the photo-oxime ligation between hMSCs and fibroblasts (Fig. 6D–I). After incubating in stem cell induction media, the hMSCs differentiated to adipocytes and subsequent UV illumination and media exchange resulted in the release of the fibroblasts generating relatively pure adipogenic cells. It should be noted that the liposome fusion method is general and can be introduced to many cell lines.
Conclusion
We have developed a novel liposome fusion system to deliver bio-orthogonal photo-active lipid and ketone-lipid like molecules to cell membranes. Upon mixing these cells in different formats, co-culture spheroids and multilayers could be generated due to an intercellular oxime (click) ligation. Furthermore, we also showed the feasibility of producing macroscopic-scale tissue and achieving multiple tissue-tissue assembly, which could lead to a series of tissue engineering, especially organ printing technologies. Since the ligation tether contains a photo-cleavable group, remote control of disassembly could be achieved upon UV light illumination. We demonstrated this system in several cell lines to generate switchable co-culture spheroids and multi-layers. Flow cytometry and mass spectrometry analysis quantified and characterized the interfacial cell surface reaction. The ability to engineer cell surfaces with a straightforward and inexpensive liposome fusion strategy will find wide use in fundamental studies of membrane biophysics, paracrine signaling and adapt to generate new biomaterials and as a biotechnology platform for screening complex cell behaviors in tissue microarrays. Several other bio-orthogonal chemical ligation strategies including diels-alder, huisgen, oxime, hydrazone, thiol-ene, etc. may be used to tailor cell surfaces with nanoparticles, redox groups and a range of other molecules for targeted delivery and as cell tracking and imaging beacons. The spatial and temporal control of cell interactions between multiple different cell types will lead to new studies of dynamic cellular communication. Furthermore, the combination of bioreactor technologies with intercellular ligation methods may provide new ways to generate large-scale complex multi-cell type tissues. When combined with traditional polymer scaffolds, molds or printing technologies, a range of complex 3D tissues and organs may be possible for an array of biomedical diagnostic and transplantation applications28,29,30.
Methods
Structure and list of molecules, liposomes and cells used in this study are shown in Supplementary Information (scheme S1). Synthesis and other experimental procedures and characterizations are detailed in Methods and Supplementary Information. O-Dodecyloxyamine was synthesized as previously described14. Egg palmitoyl-oleoyl phosphatidylcholine (egg-POPC) were purchased from Avanti Polar Lipids (Alabaster, AL) and all other chemicals were obtained from Sigma-Aldrich or Fisher. Swiss 3T3 albino mouse fibroblasts were obtained from ATCC. Human mesenchymal stem cells (hMSCs), basic medium, growth medium and differentiation medium were obtained from Lonza.
Liposome preparation
Liposomes were prepared as previously reported13,14. To generate photo-oxyamine (1) or ketone (2) liposome, photo-oxyamine or dodecanone (60 μL, 10 mM solution in CHCl3) were dissolved with egg-POPC (450 μL, 10 mg/mL in CHCl3) and 1,2-dioleoyl-3-trimethylammonium-propane (DOTAP, 10 μL, 10 mg/mL in CHCl3) in chloroform followed by concentration under high vacuum for 4 h. The dried lipid samples were then reconstituted and brought to a final volume of 3 mL in PBS buffer, pH 7.4. The contents of the vial were warmed to 50°C and sonicated for 20 min, in a tip sonicator, until the solution became clear and liposomes containing photo-oxyamine or ketone groups were formed.
Cell culture
Human mesenchymal stem cells (hMSCs) were cultured as instructed by the vendor. After cells were washed with PBS and trypsinized for 3–5 minutes, they were centrifuged in serum containing medium and followed with gentle resuspending in serum free medium. The cells were then seeded onto transparent glass substrates and then incubated at 37°C in a humidified atmosphere of 5% CO2 overnight. Adipogenic differentiation was induced by adipogenic induction medium and kept by induction/maintenance cycles as described in the Lonza protocol. Osteogenic differentiation was induced by osteogenic induction medium provided by Lonza. 3T3 Swiss Abino Fibroblasts, RFP Expressing Human Neonatal Dermal Fibroblasts and C3H/10T1/2 cells were cultured in Dulbecco's modified Eagle medium (DMEM) with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. NIH3T3/GFP cells were cultured in DMEM containing 10% FBS, 0.1 mM MEM Non-Essential Amino Acids, 2 mM L-glutamine, 10 μg/mL Blasticidin and 1% penicillin/streptomycin. These cells were incubated at 37°C in a humidified atmosphere of 5% CO2 and released from tissue culture plates using 0.05% trypsin in 0.53 mM EDTA.
Immunohistochemistry
The substrates for confocal imaging were fixed with formaldehyde (3.2% in PBS) and permeated (PBS containing 0.1% Triton X–100). A fluorescent dye mixture, containing phalloidin-TRITC (actin) and DAPI (nucleus) was then made in PBS containing 5% normal goat serum and 0.1% Triton X –100. Cells were incubated with the dye solution for 2 h. The substrates were then secured in fluorescence mounting medium (Dako, Carpinteria, CA, USA), which enhances the visualization of cells when viewed under a fluorescent microscope with a glass cover slip. The substrates for adipogenic differentiation were washed by PBS and fixed in 3.2% formaldehyde for 30 minutes, followed with sterile water and 60% isopropanol for 5 minutes. Samples were then stained by Oil Red O for 5 minutes followed by Harris Hematoxylin for 1 minute. The substrates for collagen differentiation were fixed with formaldehyde and permeated with 0.1% Triton X–100. Monoclonal antibody of collagen I was applied for 1 h, then incubated with secondary antibody anti-mouse IgG (FITC conjugate) for 30 min and followed with DAPI for 30 min for nucleus staining (reagents from Fisher Scientific). The substrates for osteogenic differentiation were stained with sigmal Alkaline Phosphatase (ALP) kit (sigmal kit 85).
Confocal microscopy
Cell clusters (spheroids) and tissue formation were visualized with a Nikon Eclipse TE2000-E inverted microscope (Nikon USA, Inc., Melville, NY). Data was analyzed by Metamorph software and a spectral confocal microscope (LeicaMicrosystems, Bannockburn, IL). Three-dimensional reconstructions of fluorescent images were generated using Volocity software.
Flow cytometry
Fluorescence-activated cell sorting (FACS) analysis was performed to quantify the approximate number of photo-oxyamine lipids at the cell surface after membrane fusion. Liposomes were cultured with Fbs (3 mM in tris buffer, 400 μL added to 4 mL) to present photo-active group on cell surface (9). Ketone-conjugated fluorescein was then reacted with the engineered cells (0.15 mM in tris buffer, 2 h). This time course assay was conducted to determine whether the chemistry was being carried on after cell growth and division. A control cell population (not displaying photolabile lipids) was incubated with the ketone-fluorescein (0.15 mM in tris buffer, 2 h). After culturing for the appropriated time, the different cell populations were washed with PBS (3 × 5 mL), trypsinized (1 mL, 5 min, 37°C, 5% CO2), centrifuged (5 min, 1000 rpm), resuspended in RPMI (without phenol red), centrifuged (5 min, 1000 rpm) and resuspended in RPMI (~107 cells/mL). Fluorescence measurements were calibrated using RCP-5-30 beads (~107 beads/mL, Spherotech, Inc., Lake Forest, IL) of known fluorescein equivalent molecule density. Fluorescent intensities based on number of cells counted were compared to the standard bead and control cells lacking fluorescent molecule conjugation and approximate numbers of fluorescent compound bound to the surface was calculated. Flow cytometry was performed using a Dako CyAn ADP (Beckman-Coulter, Brea, CA) and data was analyzed with Summit 4.3 software. The error bars are represented as the mean fluorescence intensity SD of 3 trials.
Matrix-assissted laser-desorption/ionization mass spectrometry (MALDI-MS)
Gold-coated MALDI sample plates (123 × 81 mm) (Applied Biosystems, Foster City, CA) were prepared by electron-beam deposition (Thermionics Laboratory Inc, Hayward, CA) of titanium (5 nm) and then gold (12 nm). In order to form self-assembled monolayers (SAM) of alkanethiolates on the plates, the slides were immersed in a 1 mM solution of 1∶1 ratio mixture of 11-mercaptoundecanol and tetra(ethylene glycol)-terminated undecanethiol in EtOH for 12 h, rinsed with EtOH and dried and then partially oxidized to aldehyde by mild oxidant pyridinium chlorochromate (PCC), as previously reported31. Once removed from solution, the surfaces were rinsed with EtOH and dried before use. Cells tailored with photolabile oxyamine group are seeded onto the SAM presenting aldehyde group to form oxime ligation between the cell membrane and gold substrate. After washing and removing the cells, the bonded residue on the gold substrate was traced by MALDI-MS. MALDI analysis was carried out using an AB SCIEX TOF/TOF™ 5800 System (Applied Biosystems, Foster City, CA).
Cell patterning
Self-assembled monolayers (SAMs) presenting aldehyde and tetra(ethylene glycol) (EG4) groups were patterned on gold substrate at a ratio of 1∶9 using microfluidic oxidation to ensure that fbs were only adhering to the patterned surface regions that presented 10% aldehyde groups32. Fbs were cultured with photo-liposomes (4 h) and then seeded (~102 cells/mL, 2 h) to the patterned aldehyde surfaces. Media containing 10% calf bovine serum (CBS) and 1% penicillin/streptomycin was then added and the substrates were incubated at 37°C in 5% CO2 for 4 d. Cells cultured with liposomes, not containing the key functional groups, did not attach to the patterned surfaces. Substrates were then imaged by fluorescence microscopy.
References
Friedl, P. & Zallen, J. A. Dynamics of cell–cell and cell–matrix interactions in morphogenesis, regeneration and cancer. Curr. Opin. Cell Biol. 22, 557–559 (2010).
Lutolf, M. P. & Hubbell, J. A. Synthetic biomaterials as instructive extracellular microenvironments for morphogenesis in tissue engineering. Nat. Biotechnol. 23, 47–55 (2005).
Cukierman, E., Pankov, R., Stevens, D. R. & Yamada, K. M. Taking cell-matrix adhesions to the third dimension. Science 294, 1708–1712 (2001).
Abbott, A. Cell culture: biology's new dimension. Nature 424, 870–872 (2003).
Anderson, D. G., Burdick, J. A. & Langer, R. Smart biomaterials. Science 305, 1923–1924 (2004).
Saxon, E. & Bertozzi, C. R. Cell surface engineering by a modified staudinger reaction. Science 287, 2007–2010 (2000).
Gordon, E. J., Sanders, W. J. & Kiessling, L. L. Synthetic ligands point to cell surface strategies. Nature 392, 30–31 (1998).
Chan, Y. H. M. & Boxer, S. G. Model membrane systems and their applications. Curr. Opin. Chem. Biol. 11, 581–587 (2007).
Gartner, Z. J. & Bertozzi, C. R. Programmed assembly of 3-dimensional microtissues with defined cellular connectivity. Proc. Natl. Acad. Sci. USA. 106, 4606–4610 (2009).
Kellam, B., De Bank, P. A. & Shakesheff, K. M. Chemical modification of mammalian cell surfaces. Chem. Soc. Rev. 32, 327–337 (2003).
Martin, S. E. & Peterson, B. R. Non-natural cell surface receptors: synthetic peptides capped with N-cholesterylglycine efficiently deliver proteins into mammalian cells. Bioconj. Chem. 14, 67–74 (2003).
Sampathkumar, S.-G., Li, A. V., Jones, M. B., Sun, Z. & Yarema, K. J. Nat. Chem. Biol. 2, 149–152 (2006).
Dutta, D., Pulsipher, A., Luo, W. & Yousaf, M. N. Synthetic chemoselective rewiring of cell surfaces: generation of three-dimensional tissue structures. J. Am. Chem. Soc. 133, 8704–8713 (2011).
Dutta, D., Pulsipher, A., Luo, W., Mak, H. & Yousaf, M. N. Engineering cell surfaces via liposome fusion. Bioconj. Chem. 22, 2423–2433 (2011).
Sletten, S. M. & Bertozzi, C. R. From mechanism to mouse: a tale of two bioorthogonal reactions. Acc. Chem. Res. 44, 666–676 (2011).
Luo, W., Chan, E. W. L. & Yousaf, M. N. Tailored electroactive and quantitative ligand density microarrays applied to stem cell differentiation. J. Am. Chem. Soc. 132, 2614–2621 (2010).
Kloxin, A. M., Kasko, A. M., Salinas, C. N. & Anseth, K. S. Photodegradable hydrogels for dynamic tuning of physical and chemical properties. Science 324, 59–63 (2009).
Chen, C. S., Mrksich, M., Huang, S., Whitesides, G. M. & Ingber, D. E. Geometric control of cell life and death. Science 276, 1425–1428 (1997).
Grinnell, F. Fibroblast biology in three-dimensional collagen matrices. Trends Cell Biol. 13, 264–269 (2003).
Luo, Y. & Shoichet, M. S. A photolabile hydrogel for guided three-dimensional cell growth and migration. Nat. Mater. 3, 249–253 (2004).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235 (2007).
Nuttelman, C. R., Henry, S. M. & Anseth, K. S. Synthesis and characterization of photocrosslinkable, degradable poly(vinyl alcohol)-based tissue engineering scaffolds. Biomaterials 23, 3617–3626 (2002).
Albrecht, D. R., Underhill, G. H., Wassermann, T. B., Sah, R. L. & Bhatia, S. N. Probing the role of multicellular organization in three-dimensional microenvironments. Nat. Methods. 3, 369–375 (2006).
Dazzi, F. & Horwood, N. J. Potential of mesenchymal stem cell therapy. Curr. Opin. Oncol. 19, 650–655 (2007).
Pittenger, M. F. et al. Multilineage potential of adult human mesenchymal stem cells. Science 284, 143–147 (1999).
Keller, G. M. In vitro differentiation of embryonic stem cells. Curr. Opin. Cell Biol. 7, 862–869 (1995).
Schuldiner, M., Yanuka, O., Itskovitz-Eldor, J., Melton, D. A. & Benvenisty, N. Effects of eight growth factors on the differentiation of cells derived from human embryonic stem cells. Proc. Natl. Acad. Sci. USA. 97, 11307–11312 (2000).
Stevens, M. M. & George, J. H. Exploring and engineering the cell surface interface. Science 310, 1135–1138 (2005).
Petersen, T. H. et al. Tissue-engineered lungs for in vivo implantation. Science 329, 538–541 (2010).
Mironov, V. et al. Organ printing: tissue spheroids as building blocks. Biomaterials 30, 2164–2174 (2009).
Pulsipher, A., Westcott, N. P., Luo, W. & Yousaf, M. N. Rapid in situ generation of two patterned chemoselective surface chemistries from a single hydroxy-terminated surface using controlled microfluidic oxidation. J. Am. Chem. Soc. 131, 7626–7632 (2009).
Pulsipher, A. & Yousaf, M. N. Tandem surface microfluidic lithography and activation to generate patch pattern biospecific ligand and cell arrays. Langmuir 26, 4130–4135 (2010).
Acknowledgements
This work was supported by the Carolina Center for Cancer Nanotechnology Excellence (NCI), The Burroughs Wellcome Foundation (Interface Career Award), The National Science Foundation (Career Award), National Science and Engineering Research Council of Canada (NSERC) and the Canadian Foundation for Innovation.
Author information
Authors and Affiliations
Contributions
M.N.Y. designed the study. W.L., A.P. and D.D. performed the experiments. M.N.Y., W.L., D.D., A.P. and B.M.L. analyzed the data. M.N.Y. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reprints and permissions information is available online at www.nature.com/reprints.
Electronic supplementary material
Supplementary Information
Supporting Information Figures and Methods
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Luo, W., Pulsipher, A., Dutta, D. et al. Remote Control of Tissue Interactions via Engineered Photo-switchable Cell Surfaces. Sci Rep 4, 6313 (2014). https://doi.org/10.1038/srep06313
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06313
This article is cited by
-
Engineering organoids
Nature Reviews Materials (2021)
-
Coatings on mammalian cells: interfacing cells with their environment
Journal of Biological Engineering (2019)
-
Oncotically Driven Control over Glycocalyx Dimension for Cell Surface Engineering and Protein Binding in the Longitudinal Direction
Scientific Reports (2018)
-
Scaffold Free Bio-orthogonal Assembly of 3-Dimensional Cardiac Tissue via Cell Surface Engineering
Scientific Reports (2016)
-
Spatiotemporal control of cell–cell reversible interactions using molecular engineering
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.