Abstract
Steps and their associated adatoms extensively exist and play prominent roles in affecting surface properties of materials. Such impacts should be especially pronounced in two-dimensional, atomically-thin membranes like graphene. However, how single adatom behaves at monatomic steps of few-layer graphene is still illusive. Here, we report dynamics of individual adatom at monatomic steps of free-standing few-layer reduced graphene under the electron beam radiations and demonstrate the prevalent existence of monatomic steps even down to unexpectedly ultrasmall lateral size of a circular diameter of ~5 Å. Single adatom prefers to stay at the edges of the atomic steps of few-layer reduced graphene and evolve with the steps. Moreover, we also find that how the single adatom behaves at atomic step edges can be remarkably influenced by the type of adatoms and step edges. Such single adatoms at monatomic steps and ultrasmall atomic steps open up a new window for surface physics and chemistry for graphene-based as well as other two-dimensional materials.
Similar content being viewed by others
Introduction
Steps and adatoms on them extensively exist on the surfaces of materials and play prominent roles in surface physical and chemical properties, e.g. adsorption, diffusion, transport, catalysis and crystal growth1,2,3. Such effects should be especially significant in two-dimensional, atomically-thin graphene4,5,6,7,8,9,10. In general, single layer graphene has no conventional steps on its surface because of single-atom thickness11. However, steps may exist on the surface of few-layer graphene which has attracted equal attention to single-layer graphene for its unique tunable electronic properties12,13,14,15,16,17,18. To date, single adatom behavior at monatomic steps of few-layer graphene is still illusive, posing significant challenges in obtaining ultraclean samples with many atomic steps as well as atomic-resolution imaging of atomically-thin membranes. Here, we report the dynamics of single adatom at monatomic steps of free-standing few-layer reduced graphene under electron beam radiations. We demonstrate the existence of monatomic steps even down to unexpectedly ultrasmall lateral size of a circular diameter of ~5 Å on few-layer reduced graphene (FLRG). We also find that single adatom prefers to stay at the atomic step edges of few-layer reduced graphene and evolve with atomic steps. Moreover, single adatom behavior at atomic steps is found to be influenced by the type of adatoms and step edges. Such single adatoms and ultrasmall atomic steps open a new window of surface physics and chemistry for graphene-based and other two-dimensional materials.
One of the biggest challenges for imaging adatoms at atomic steps on few-layer graphene is to obtain ultraclean few-layer samples with many atomic steps. Since the polymer-based transfer of mechanically-exfoliated graphene and the graphene grown by chemical vapor deposition (CVD) often induce contamination on surfaces which can not be ruled out thoroughly even after high-temperature annealing19, in this study, we prepare few-layer reduced graphene by polymer-free and solution-based chemical reduction of exfoliated graphite oxide (see Methods)20,21,22,23. We find that the modified two-step chemical reduction enables to remove most oxygenous contents in exfoliated graphite oxide so as to obtain FLRG (Fig. 1a). The FLRG samples were further characterized with X-ray photoelectron spectra (XPS) and electron energy loss spectroscopy (EELS). The presence of C 1s XPS spectra indicates that the obtained FLRG has a high quality with few oxygenous contents (Supplementary Fig. S1a). The EELS spectra and the sharp σ* peak at the carbon K edge (Supplementary Fig. S1b) further confirm the existence of conjugation in FLRG (Fig. 1a)24, consistent with the C 1s XPS spectra.
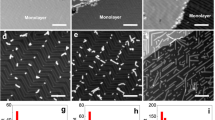
Mapping and atomic-resolution imaging of atomic steps on the free-standing few-layer reduced graphene.
(a), Atomic step mapping on the BF STEM images. (b), Identification of layer numbers on the HAADF STEM image for (a). (c), Atomic-resolution imaging of one monatomic step between the bilayer and trilayer graphene indicated by white arrows in (a) and (b) using BF STEM. (d), The image from (c) with the atomic step area at top layer outlined. (e), HAADF STEM images of one monatomic step. Red arrow indicates adatoms at step edges. (f), The image from (e) with the atomic step area at top layer outlined. Scale bars in (a), (b), 1 nm; scale bars in (c–f), 0.5 nm. (a) and (c–f) are low-pass-filtered images to remove noise. Green line, atomic steps between single layer (I) and bilayer (II). Red line, atomic steps between bilayer (II) and three layer (III). Brown line, atomic steps between three layer (III) and four layer (IV). Blue line, atomic steps between bilayer (II) and four layer (IV). Purple line, atomic steps between single layer (I) and three layer (III). Black line, atomic steps between single layer (I) and four layer (IV).
Figure 1 shows bright-field (BF) and high angle annular dark-field (HAADF) scanning transmission electron microscopy (STEM) images of several typical areas of free-standing FLRG. The HAADF STEM images are very sensitive to layer numbers and the layers in the BF-STEM images can be identified with the corresponding HAADF STEM images (Fig. 1a,b, Supplementary Fig. S2)25. By quantifying HAADF intensity and its differences in areas, layer numbers for different layers (I–IV layers) on few-layer reduced graphene could be identified (Fig. 1b, Supplementary Fig. S2). The HAADF intensity is found to change linearly with layer numbers (Supplementary Fig. S2c). The atomic steps seen in the BF STEM images (Fig. 1a) can be determined under the assistance of their corresponding HAADF STEM images (Fig. 1b). The atomic steps on the FLRG can be monatomic (green, red and brown lines for I–II, II–III and III–IV steps, respectively), two-atom high (blue and purple lines for II–IV and I–III steps, respectively) and even three-atom high (black line for I–IV steps). The three-atom steps are much rougher than the monatomic ones, as demonstrated in both BF and HAADF STEM images.
Interestingly, there appear a large amount of atomic steps in the FLRG with the ultrasmall lateral size down to ~1 nm or even a few angstroms in diameter for the quasi-circular steps (Supplementary Fig. S3). Supplementary Figure S3b,c presents quasi-circular monatomic step edges with a diameter of ~5 Å. Moreover, monatomic steps represent the most common ones observed for both I–II, II–III and III–IV steps, albeit that several two-atom-high steps are also detected (Fig. 1a, Supplementary Fig. S3). However, the three-atom-high steps are rarely observed. Occasionally, some step edges can be defined by complex step edges with half monatomic steps and half two- or three-atom-high steps (area defined by green, purple and black lines in Fig. 1a and Supplementary Fig. S4). The complex atomic step-defined area can also be ultrasmall in lateral size even down to ~5 Å in circular diameter (Supplementary Fig. S4). In addition, the areas defined by ultrasmall quasi-circular step edges can reach up to about one quarter of the observed surfaces (~200 nm2), totally different from surfaces of the CVD graphene and mechanically-exfoliated graphene. The presence of a large amount of atomic steps on the FLRG should be attributed to the ultrasonic exfoliation processing where very local, inhomogeneous defects and stress in FLRG facilitate the formation of circular step edges with ultrasmall lateral size26,27,28.
Figure 1c–f shows atomic-resolution imaging of monatomic step edges. The atomic steps edges are often contaminated more severely by the adatoms or impurities than other areas (Fig. 1b,e,f and Supplementary Fig. S3). However, one can still find some ultraclean and highly ordered areas on the FLRG samples that allow us to study the local atomic structures at the atomic step edges. The atomic arrangements near the monatomic step edges are mostly hexagonal rings with a few pentagonal rings (Fig. 1d,f). No dominant zigzag or armchair edges can be observed. Instead, all observed atomic step edges are the mixture of zigzag and armchair edges and the reconstructed edges beyond zigzag and armchair (Fig. 1d,f), consistent with other experimental and theoretical studies of graphene edges29,30,31,32. The direct observation of atomic steps here can extract atomic-resolution information on the atomic step edge areas, which help understand the atomic and molecular behaviors of both atomic steps and adatoms.
To shed light on the dynamics of both ultrasmall monatomic steps and adatoms at the step edges under electron beam radiations, we present HAADF STEM images of the dynamics of a monatomic step edge with adatoms (Fig. 2a–f and Supplementary Movie S1). The atomic step edge (edge 2) first combines with another atomic step edge below (edge 3) (Fig. 2a–c). At different time, one atomic step edge in the left (edge 1) joins the previously combined step edges (Fig. 2d–f). Figure 2a–f shows the adatoms, which are identified as sodium (Na) atoms by x-ray photoelectron spectra (XPS) together with the HAADF intensity analysis from experimental and simulated HAADF STEM images (Fig. 3 and Supplementary Figs. S5 and S6)33. The sodium adatoms at the edges show a high mobility at the edges and always move along the edges (Fig. 2a–f and Supplementary Movie S1). Moreover, we also show dynamics of another ultrasmall monatomic step edge with few adatoms at edges in Supplementary Fig. S7. Despite of their similar lateral size, the atomic step edges in Supplementary Fig. S7 show different behaviors from the step edges in Fig. 2a–f and are much more stable with their shapes changed only slightly during the observation. Although there are other step edges of ultrasmall lateral size near atomic step edges, no combination takes place in the whole imaging process (Supplementary Fig. S7). These differences may result from the local strain in the step area, from the adatoms at the step edges and from the step edge structures.
Dynamics of single adatoms and monatomic steps with ultrasmall lateral size on the free-standing few-layer reduced graphene.
(a–f), Snapshot of HAADF STEM images at different time: 3 s (a), 54 s (b), 126 s (c), 177 s (d), 222 s (e) and 252 s (f). The atomic steps are indicated by white arrows. Each image is produced from the overlapping of 10 continuous snapshots and low-pass filtered. (g–i), Snapshot of HAADF STEM images at different time: 3 s (g), 123 s (h) and 300 s (i). Each image is low-pass filtered. Individual adatoms are highlighted by white arrows. Scale bars, 1 nm. The dynamics is also shown in Supplementary Movies S1 and S2.
Apart from the presence of the Na adatoms, we also detect other species of single adatoms, sulfur (S) adatom, at step edges on the FLRG and monitor the dynamics of single S adatoms at step edges (Fig. 2g–i and Supplementary Movie S2). The adatoms at the atomic step edges (indicated by arrows in Supplementary Fig. S8) have similar dip in BF intensity and similar increase in HAADF intensity. The three marked single adatoms are all sulfur atoms which are identified by the XPS spectra and HAADF intensity analysis of experimental and simulated HAADF STEM images (Fig. 3 and Supplementary Fig. S6)33. Interestingly, the S adatoms at the step edges (indicated by white arrows) preferred to stay at the original place as opposed to the high mobility of Na adatoms. The S adatoms rarely move along the step edges and do not move at all at the edges (Fig. 2g–i and Supplementary Movie S2). The different atomic behaviors for different adatoms at step edges could provide new routes to tuning the physical and chemical properties of graphene surfaces.
The different atomic behaviors of Na and S adatoms at steps can be interpreted from the interactions of adatoms with step edges. First-principles calculations (details in Methods) indicate that the adatoms, whatever sulfur or sodium, prefer to stay at the step edges (Supplementary Figs. S9–S12). For example, the absorption energy for Na at the edge is calculated to be 1.0 eV for the armchair edges, 2.0 eV for the zigzag edges and 0.7 eV for the reconstructed, non-armchair and non-zigzag edges, which are lower than that far away from step edges. On the other hand, the absorption energy for S at the edge is 3.8 eV for the armchair, 5.7 eV for the zigzag and 3.1 eV for the reconstructed, non-armchair and non-zigzag edges, which are lower than that far away from step edges (Supplementary Figs. S9–S12). Importantly, energy barrier for the S and Na adatoms moving along the edges differs significantly (Fig. 4 and Supplementary Fig. S13): the energy barrier for S adatom moving along the armchair edges is ~1.4 eV, over 5 times higher than Na adatom (Fig. 4a,c). Similarly, the energy barrier for S adatom moving along the zigzag edges is calculated to be ~0.65 eV, twice that for Na adatoms at the zigzag edges (Fig. 4b,d). Similar situation is also seen for the reconstructed, non-zigzag and non-armchair edges. The energy barrier for the S adatom moving along the edges is over 15 times higher than that for the Na adatom (Supplementary Fig. S13).
Energy diagram for single adatom diffusion at monatomic step edges.
(a), Na adatom at the armchair edges. (b), Na adatom at the zigzag edges. (c), S adatom at the armchair edges. (d), S adatom at the zigzag edges. The arrows indicate the moving direction of single adatom at step edges. The bottom layer graphene, top layer graphene, Na atom and S atom are indicated in pink, gray, purple and red, respectively. X axis shows the positions of adatom along the direction indicated by arrows in the insets.
These calculations indicate that the energy barrier for S adatoms is much larger than that for Na adatoms when moving along the step edges, independent of the types of the edges, providing thereby an energetical explanation to their observed different atomic behaviors at step edges. Moreover, our calculations also indicate that the effect of atomic arrangements of step edges on single adatom behaviors may depend on the adatoms. The energy barrier for the movement of Na adatoms along the step edge is very close in the cases of armchair, zigzag and reconstructed edges, while the barrier for S adatoms at the armchair and reconstructed edges are almost twice and four times larger than that at the zigzag edges, respectively (Fig. 4 and Supplementary Fig. S13). To gain more insights into the dynamics, the dynamic process of Na and S adatoms at the monatomic steps is calculated (details in Methods). The results indicate that the Na adatom is highly mobile in both cases, while the S adatom is relatively much more stable, consistent with the energy barrier calculations, which implies the important role of interactions between adatom and steps. The findings here therefore raise the likelihood that surface physical and chemical property of graphene can be modulated by adatoms and atomic arrangement at step edges.
Direct observation of dynamics of both atomic steps and single adatoms at atomic step edges on free-standing two-dimensional membranes under the electron radiations should have many implications. First, two-dimensional free-standing membranes with a few atom thickness represent a prototype for surface studies using TEM, paving a way to unravel the secrets of interactions between surfaces and impurity adatoms down to single atom level, which is very important for many physical processes and chemical reactions1,34. Secondly, monitoring the behaviors of single adatom and steps helps understand the roles of step edges on graphite surfaces or impurities in graphene on electrochemical activity35,36. Thirdly, the presence of a large amount of ultrasmall atomic steps on the surfaces of few-layer graphene also inspires new opportunities for graphene. In addition, adatoms may alter the surface properties of few-layer graphene by affecting dynamics of atomic steps and such change relies on the physical nature of adatoms and the interactions between adatoms and atomic step edges. Moreover, the ultrasmall atomic steps mean that they can appear in graphene nanoribbons with a diameter of a few nanometers37,38, implying that atomic steps and adatoms at the step edges can affect electronic properties of few-layer graphene nanoribbons. Further, the atomic step and adatoms can impact the molecular dynamics behavior of surfaces of two-dimensional atomic membranes39,40. The complete understanding of adatoms and atomic steps may therefore lead to a significant step forward for potential applications of two-dimensional membranes in nanoscale, molecular and atomic systems.
Methods
Preparation of few-layer reduced graphene
Few-layer reduced graphene was fabricated by chemical reduction of exfoliated graphite oxide20,21,22,23. In the modified process, a two-step chemical reduction was adopted to achieve a thorough reduction. Hydrazine was firstly applied in the first-step reduction, followed by the hydrothermal reduction at 150°C in a sealed autoclave with a maximum working pressure of 20 MPa. The obtained few-layer reduced graphene was then washed completely using distilled water and ethanol. To image the few-layer reduced graphene sample, the samples were re-dispersed in water/ethanol mixture (1:1) by sonication. The obtained solution was dropped on carbon grids, followed by drying at room temperature. XPS spectra of graphene were collected by AXIS ultra DLD (Shimadzu Kratos) with an X-ray source of monochromatic Al Ka and X-ray power of 15 kV and 10 mA.
STEM imaging
STEM characterizations were conducted with the FEI TITAN Cubed 80–300 electron microscope with the CEOS spherical aberration corrector and Gatan imaging filter operated at 80 kV. BF and HAADF STEM images were simultaneously recorded. The dwell time on each shot was 3.9 μs. EELS measurements were carried out with a monochrometer. For movies, the total acquisition time including the live drift correction was typically about 5 min for 100 images with 512 × 512 pixels. The STEM images were simulated using WinHREM + STEM package (HREM Research).
First-principles calculations
Density-functional-theory (DFT) calculations were carried out by using Vienna ab initio simulation package (VASP)41,42,43,44. Local density approximation (LDA) was applied to address the exchange-correlation functional. The cutoff energy was set to be 500 eV. We applied the projector-augmented wave (PAW) method with 2 × 2 × 1 Monkhorst-Pack grids45,46. The energy was converged to an accuracy of 1.0−6 eV. All atoms were relaxed until the maximum atomic force was less than 1.0−2 eV/Å. To evaluate the diffusion barrier of Na/S atoms along the monatomic step edges, the climbing image nudged elastic band (CI-NEB) was adopted47,48. We considered two layers of graphene in the simulations. The bottom layer represented the pristine graphene and the top layer was the graphene ribbons with different edges (i.e. armchair, zigzag, or reconstruction edges). The adsorption energy of adatom (for example, Na atoms) is defined as:

where E(C + Na) is the total energy of Na atoms adsorbed on the graphene, E(Na) and E(C) are the energies of Na atoms and graphene system, respectively. To understand further the adatom dynamic processes observed under incident electron beams, DFT molecular dynamics calculations (DFTMD) are also performed for Na and S adatoms at zigzag and armchair step edges. The energy of electron beam in first calculation is 80 KeV, which is the same as the measured result. The energies transferred from the electron to target adatoms can be determined by T(Θ) = Tmaxcos2Θ, where Tmax, the maximum energies transferred to S and Na atoms, are ~6 eV and ~8 eV, respectively and Θ represents scattering angle from incident direction of electron. We simulated the dynamics of adatoms after being knocked by electrons at different incident angles within 0.6 ps and found that Na adatoms move along the step edges easily, while it is not the case for the S adatoms, consistent with the STEM observations. We also set up different initial knocking electron energies for the Na to make the transferred energy from electrons and found the same value of ~6 eV for both Na and S adatoms. The Na adatom remained much more mobile along the step edges than the S adatom, which is consistent with the energy barrier calculations, implying that the interactions between adatoms and atomic steps on surfaces of few-layer reduced graphene are critical to the dynamics of adatoms.
References
Ibach, H. Physics of surfaces and interfaces. (Springer, Berlin, 2006).
Fujita, T. et al. Atomic origins of the high catalytic activity of nanoporous gold. Nat. Mater. 11, 775–780 (2012).
Helveg, S. et al. Atomic-scale imaging of carbon nanofibre growth. Nature 427, 426–429 (2004).
Zhao, L. et al. Visualizing individual nitrogen dopants in monolayer graphene. Science 333, 999–1003 (2011).
Meyer, J. C., Girit, C. O., Crommie, M. & Zettl, A. Imaging and dynamics of light atoms and molecules on graphene. Nature 454, 319–322 (2008).
Suenaga, K. & Koshino, M. Atom-by-atom spectroscopy at graphene edge. Nature 468, 1088–1090 (2010).
Huang, P. Y. et al. Grains and grain boundaries in single-layer graphene atomic patchwork quilts. Nature 469, 389–392 (2011).
Meyer, J. C. et al. Experimental analysis of charge redistribution due to chemical bonding by high-resolution transmission electron microscopy. Nat. Mater. 10, 209–215 (2011).
Robertson, A. W. et al. Dynamics of Single Fe Atoms in Graphene Vacancies. Nano Lett. 13, 1468–1475 (2013).
Wang, H. et al. Doping monolayer graphene with single atom substitutions. Nano Lett. 12, 141–144 (2011).
Meyer, J. C. et al. The structure of suspended graphene sheets. Nature 446, 60–63 (2007).
Ohta, T., Bostwick, A., Seyller, T., Horn, K. & Rotenberg, E. Controlling the electronic structure of bilayer graphene. Science 313, 951–954 (2006).
Castro, E. V. et al. Biased bilayer graphene: semiconductor with a gap tunable by the electric field effect. Phys. Rev. Lett. 99, 216802 (2007).
Zhang, Y. et al. Direct observation of a widely tunable bandgap in bilayer graphene. Nature 459, 820–823 (2009).
Oostinga, J. B., Heersche, H. B., Liu, X., Morpurgo, A. F. & Vandersypen, L. M. Gate-induced insulating state in bilayer graphene devices. Nat. Mater. 7, 151–157 (2008).
Craciun, M. et al. Trilayer graphene is a semimetal with a gate-tunable band overlap. Nat. Nanotechnol. 4, 383–388 (2009).
Reina, A. et al. Large area, few-layer graphene films on arbitrary substrates by chemical vapor deposition. Nano Lett. 9, 30–35 (2008).
Lui, C. H., Li, Z., Mak, K. F., Cappelluti, E. & Heinz, T. F. Observation of an electrically tunable band gap in trilayer graphene. Nat. Phys. 7, 944–947 (2011).
Lin, Y.-C. et al. Graphene annealing: how clean can it be? Nano Lett. 12, 414–419 (2011).
Stankovich, S. et al. Graphene-based composite materials. Nature 442, 282–286 (2006).
Chang, H. et al. A Transparent, Flexible, Low-Temperature and Solution-Processible Graphene Composite Electrode. Adv. Funct. Mater. 20, 2893–2902 (2010).
Chang, H. et al. Thin Film Field-Effect Phototransistors from Bandgap-Tunable, Solution-Processed, Few-Layer Reduced Graphene Oxide Films. Adv. Mater. 22, 4872–4876 (2010).
Chang, H. & Wu, H. Graphene-Based Nanomaterials: Synthesis, Properties and Optical and Optoelectronic Applications. Adv. Funct. Mater. 23, 1984–1997 (2013).
Mkhoyan, K. A. et al. Atomic and electronic structure of graphene-oxide. Nano Lett. 9, 1058–1063 (2009).
Gass, M. H. et al. Free-standing graphene at atomic resolution. Nat. Nanotechnol. 3, 676–681 (2008).
Gómez-Navarro, C. et al. Atomic structure of reduced graphene oxide. Nano Lett. 10, 1144–1148 (2010).
Dreyer, D. R., Park, S., Bielawski, C. W. & Ruoff, R. S. The chemistry of graphene oxide. Chem. Soc. Rev. 39, 228–240 (2010).
Erickson, K. et al. Determination of the local chemical structure of graphene oxide and reduced graphene oxide. Adv. Mater. 22, 4467–4472 (2010).
Koskinen, P., Malola, S. & Häkkinen, H. Evidence for graphene edges beyond zigzag and armchair. Phys. Rev. B 80, 073401 (2009).
Kobayashi, Y., Fukui, K.-i., Enoki, T. & Kusakabe, K. Edge state on hydrogen-terminated graphite edges investigated by scanning tunneling microscopy. Phys. Rev. B 73, 125415 (2006).
Jia, X. et al. Controlled formation of sharp zigzag and armchair edges in graphitic nanoribbons. Science 323, 1701–1705 (2009).
Koskinen, P., Malola, S. & Häkkinen, H. Self-Passivating Edge Reconstructions of Graphene. Phys. Rev. Lett. 101, 115502 (2008).
Krivanek, O. L. et al. Atom-by-atom structural and chemical analysis by annular dark-field electron microscopy. Nature 464, 571–574 (2010).
Horch, S. et al. Enhancement of surface self-diffusion of platinum atoms by adsorbed hydrogen. Nature 398, 134–136 (1999).
Davies, T. J., Hyde, M. E. & Compton, R. G. Nanotrench arrays reveal insight into graphite electrochemistry. Angew. Chem. Int. Ed. 44, 5121–5126 (2005).
Ambrosi, A. et al. Metallic impurities in graphenes prepared from graphite can dramatically influence their properties. Angew. Chem. Int. Ed. 51, 500–503 (2012).
Li, X., Wang, X., Zhang, L., Lee, S. & Dai, H. Chemically derived, ultrasmooth graphene nanoribbon semiconductors. Science 319, 1229–1232 (2008).
Bai, J. et al. Very large magnetoresistance in graphene nanoribbons. Nat. Nanotechnol. 5, 655–659 (2010).
Colson, J. W. et al. Oriented 2D covalent organic framework thin films on single-layer graphene. Science 332, 228–231 (2011).
Xu, K., Cao, P. & Heath, J. R. Graphene visualizes the first water adlayers on mica at ambient conditions. Science 329, 1188–1191 (2010).
Kresse, G. & Hafner, J. Ab initio molecular dynamics for liquid metals. Phys. Rev. B 47, 558 (1993).
Kresse, G. & Hafner, J. Ab initio molecular-dynamics simulation of the liquid-metal–amorphous-semiconductor transition in germanium. Phys. Rev. B 49, 14251 (1994).
Kresse, G. & Furthmüller, J. Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set. Comp. Mater. Sci. 6, 15–50 (1996).
Kresse, G. & Furthmüller, J. Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys. Rev. B 54, 11169 (1996).
Kresse, G. & Joubert, D. From ultrasoft pseudopotentials to the projector augmented-wave method. Phys. Rev. B 59, 1758 (1999).
Monkhorst, H. J. & Pack, J. D. Special points for Brillouin-zone integrations. Phys. Rev. B 13, 5188–5192 (1976).
Mills, G., Jónsson, H. & Schenter, G. K. Reversible work transition state theory: application to dissociative adsorption of hydrogen. Surf. Sci. 324, 305–337 (1995).
Henkelman, G., Uberuaga, B. P. & Jónsson, H. A climbing image nudged elastic band method for finding saddle points and minimum energy paths. J. Chem. Phys. 113, 9901–9904 (2000).
Acknowledgements
This work was supported by World Premier International Research Center Initiative (WPI), MEXT of Japan. H.X.C. acknowledges the funding from HUST, WPI-AIMR fusion research funding from MEXT and a Grant-in-Aid for Young Scientists B from JSPS (No. 25870057). Z.C.W. thanks financial supports from Grant-in-Aid for Young Scientists (A) (grant no. 24686069), NSFC (grant no. 11332013), JGC-S Foundation and Kurata Memorial Hitachi Science and Technology Foundation. We acknowledge Kazuyo Ohmura at the Institute for Materials Research, Tohoku University for XPS measurements and the support from NIMS-Nanotechnology Platform Project.
Author information
Authors and Affiliations
Contributions
H.X.C. designed the project and prepared the graphene. Y.I., K.K., M.S., T.N. and Z.C.W. conducted STEM measurements and simulations. Y.L. and Y.K. calculated the first-principles theoretical models. H.K.W. helped to analyze the results. All the authors commented on the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Supplementary Movie S1
Supplementary Information
Supplementary Movie S2
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Chang, H., Saito, M., Nagai, T. et al. Single adatom dynamics at monatomic steps of free-standing few-layer reduced graphene. Sci Rep 4, 6037 (2014). https://doi.org/10.1038/srep06037
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06037
This article is cited by
-
Single-Atom Catalysts: From Design to Application
Electrochemical Energy Reviews (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.