Abstract
The fungus Harpophora oryzae is a close relative of the pathogen Magnaporthe oryzae and a beneficial endosymbiont of wild rice. Here, we show that H. oryzae evolved from a pathogenic ancestor. The overall genomic structures of H. and M. oryzae were found to be similar. However, during interactions with rice, the expression of 11.7% of all genes showed opposing trends in the two fungi, suggesting differences in gene regulation. Moreover, infection patterns, triggering of host defense responses, signal transduction and nutritional preferences exhibited remarkable differentiation between the two fungi. In addition, the H. oryzae genome was found to contain thousands of loci of transposon-like elements, which led to the disruption of 929 genes. Our results indicate that the gain or loss of orphan genes, DNA duplications, gene family expansions and the frequent translocation of transposon-like elements have been important factors in the evolution of this endosymbiont from a pathogenic ancestor.
Similar content being viewed by others
Introduction
Rice has been a major food source for people in Asia and Africa for centuries. However, a large proportion of the rice yield is annually lost to agricultural diseases and pests1. Rice blast, caused by the fungal pathogen Magnaporthe oryzae, is one of the most severe rice diseases and has been found almost everywhere rice is grown2,3.
Beneficial relationships between plants and microorganisms often occur in the rhizosphere and play important roles in ecosystems by improving plant growth or by helping plants to overcome biotic or abiotic stress4. Whereas mycorrhizal symbioses have long attracted significant interest5, root endophytes have only recently been recognized to also be fundamental components of ecosystems6,7. The fungus Harpophora oryzae was first described among endophytes residing in domestic Chinese wild rice (Oryza granulata), where H. oryzae can strongly promote rice growth and biomass accumulation8. In addition, H. oryzae can protect rice roots from invasion by M. oryzae and can induce systemic resistance to rice blast, which makes it an attractive candidate for biocontrol9. Phylogenetic analyses have shown that Harpophora has a close relationship to other members of the Magnaporthaceae, such as Gaeumannomyces and Magnaporthe8, most of which are plant pathogens10,11. H. oryzae, similar to M. poae and G. graminis, but unlike M. oryzae, only infects plant roots10. Some morphological aspects of penetration into the plant, such as the formation of appressoria or hyphopodia, are also similar between M. oryzae and H. oryzae9,12. However, H. oryzae only infects roots from the epidermis to the cortex without penetrating the stele9, whereas M. oryzae can invade vascular tissue, from which it systemically spreads to the aerial parts of the plant12.
The morphologic and phylogenetic relationships between H. oryzae and its close plant pathogenic relatives render it an attractive model for examining the evolutionary mechanisms that lead to a beneficial endophyte in an ancestral neighborhood of pathogens. Here, we describe the genome sequence of H. oryzae and present a comparative analysis of the transcriptomes of H. oryzae and M. oryzae during rice root infection. We show that H. oryzae arose from a pathogenic ancestor and that this change was accompanied by a significant expansion of transposable elements, the development of a retrotransposon surveillance pathway and the concomitant expansion of gene families related to a biotrophic lifestyle.
Results and Discussion
Genome sequencing and general features
The genome of H. oryzae R5-6-1 was sequenced using a combination of 454 pyrosequencing and the Illumina HiSeq 2000 sequencing platform. The total reads were 6562 Mb in length, representing an approximately 129-fold genome sequence coverage (Supplementary Table S1). A 50.78-Mb draft genome sequence of H. oryzae was assembled and contained 247 scaffolds (size: >200 bp; Table 1). The genome of H. oryzae was approximately 8% larger than those of M. oryzae, M. poae and G. graminis (Supplementary Table S2). A total of 14,575 protein-coding genes have been predicted for H. oryzae, 90.1% of which were validated using mRNA sequences. By the Core Eukaryotic Genes (CEGs) Mapping Approach pipeline (CEGMA v 2.0)13, the completeness of the H. oryzae genome was assessed to be 99.2%. The details of the sequence analysis are provided in the supplementary material (Supplementary Note 1).
Syntenic analysis and phylogenetic relationships
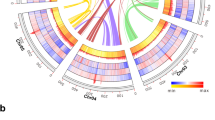
Pairwise sequence comparisons of the genome sequence of H. oryzae and those of its nearest phylogenetic neighbors revealed high degrees of macrosynteny between H. oryzae and M. poae or G. graminis. By contrast, only mesosynteny (i.e., chromosomes with similar gene contents but with different orders and orientations of genes14) was observed between H. oryzae and M. oryzae (Supplementary Fig. S1). A phylogenetic tree derived from a combined analysis of 280 single-copy orthologs [i.e., genes in the same Markov cluster (MCL); see below] from H. oryzae and 16 other fungi was consistent with known species phylogeny (Fig. 1a). However, species sharing the same nutritional lifestyle (i.e., pathogens, symbionts or saprophytes) were located randomly on the branches rather than clustering together, indicating that these lifestyles have been gained and lost several times during evolution. Ancestral state reconstruction (ASR) further showed that Harpophora, Magnaporthiopsis and Gaeumannomyces formed a monophyletic clade together with Nakataea and Magnaporthe. Bayesian inference (BI), maximum likelihood (ML) and maximum parsimony (MP) analyses all suggested that the ancestral state of H. oryzae and related species in clade A was that of a plant pathogen (Fig. 1b). Inferring a time scale from the phylogenetic analysis further revealed that M. oryzae diverged from G. graminis, H. oryzae and M. poae approximately 67 million years (MY) ago (Fig. 1a), which corresponds well with the time of origin of the first grass families (55–77 MY ago15,16,17). G. graminis then diverged from H. oryzae and M. poae 19 MY ago and H. oryzae and M. poae diverged approximately 15 MY ago (Fig. 1a). These divergence times among G. graminis, M. poae and H. oryzae correlate with the divergence times between the Triticeae (barley, wheat and oats, 13–25 MY17). Therefore, these results suggest that differentiation among M. oryzae, M. poae, G. graminis and H. oryzae occurred in response to the divergence of their hosts.
Phylogenomic relationships and the ancestral state reconstruction analysis between H. oryzae and related species.
(a) Maximum likelihood (ML) phylogenetic tree was constructed using the LG + I + G amino acid substitution model showing the evolutionary relationships of 5 symbiotic (red) species, 8 pathogenic (blue) species and 3 saprophytic (green) species. (b) ML phylogenetic tree of H. oryzae R5-6-1 and related Magnaporthaceae species based on six combined sequences and the results of an ancestral state reconstruction analysis of physiological characters (endophytic, pathogenic and saprophytic) based on ML and Bayesian inference (BI) approaches. The ML bootstrap values, posterior probability values and maximum parsimony (MP) bootstrap values are sequentially indicated above the branches and asterisks indicate that all three values are 100. The results of an ancestral state reconstruction analysis using ML and BI methods were mapped onto the best ML tree (the results of the MP analysis were the same as the ML results, which are not shown). The external small pie charts on the tree represent the relative likelihoods of alternative character states based on the ML analysis and the internal large pie charts are based on Bayesian MCMC methods.
Transposable elements
One of the most striking features observed in the H. oryzae genome, compared with its closest phylogenetic neighbors, was its relatively high number of transposable elements (TEs) (Supplementary Table S3; S4), i.e., 15% in H. oryzae vs. 10.2%, 0.65% and 7.2% in M. oryzae, M. poae and G. graminis, respectively. LTR retrotransposons (i.e., Gypsy, Copia and LINE Tad1) were the most abundant class I TEs in the genomes of H. oryzae, M. oryzae and G. graminis (50.7, 72 and 72.5% of all TEs, respectively). TcMar-Fot1 was the most abundant class II TE in all three genomes. The higher number of TEs in H. oryzae was primarily due to the presence of larger numbers of unclassified TEs (38.3% of TEs, 2912 kb for H. oryzae vs. 8.2% of TEs, 341 kb for M. oryzae). Generally, TEs were more common in regions of the H. oryzae genome with lower gene density (Supplementary Fig. S2). Regions containing either H. oryzae-specific orphan genes (BLASTP: E-value < 10−5) or duplicated genes (paralogs, see below) were located in the vicinity of areas with a high frequency of TEs (Supplementary Fig. S2). Consistent with this finding, most of the 929 TE-disrupted genes and 368 TE-containing genes were H. oryzae specific and had unknown functions (Supplementary Fig. S3). Among these genes, 83 and 15 encode proteins that contain a signal sequence and are therefore likely secreted. These results strongly suggest that the TEs have had an important influence on the dynamics of the evolution of the H. oryzae genome. We also found evidence of RIP activity in all H. oryzae TE families (Supplementary Fig. S4). Because RIP is active only under conditions of sexual recombination, we also searched for the presence of mating type genes. We found orthologs of MAT1-1-1 (HAOR_ 11805), MAT1-1-2 (HAOR_ 11806) and MAT1-1-3 (HAOR_ 11807), but not of a MAT1-2 gene, suggesting that H. oryzae is heterothallic and principally able to perform sexual reproduction. The fact that RIP has not protected H. oryzae against its significant transposon expansions is likely due to the existence of very rare sexual activity, which is similar to M. oryzae18.
Comparative genomic analysis
An MCL analysis identified a total of 19,338 ortholog groups (157,857 genes) that were clustered between H. oryzae and other 16 fungal genomes [comprising phytopathogens (M. oryzae, M. poae, G. graminis, Colletotrichum higginsianum, Fusarium graminearum, Botrytis cinerea, Sclerotinia sclenotiorum and Ustilago maydis), symbionts (Epichloe festucae, Tuber melanosporum, Laccaria bicolor and Piriformospora indica), saprophytes (Aspergillus nidulans, Neurospora crassa and Chaetomium globosum) and the yeast Saccharomyces cerevisiae]. On average, each group contained approximately 8.2 genes. Two ortholog groups were shared only by H. oryzae and other symbiotic fungi and 13 were shared only by H. oryzae and saprophyte genomes; however, 1641 were shared exclusively by H. oryzae and pathogenic fungi, which is consistent with the origin of H. oryzae from pathogenic ancestors (vide supra). No MCL clusters were identified that were shared by all symbiotic fungi but were absent from the genomes of pathogens and saprophytes. Thus, we did not identify any common “symbiosis-determining genes”. This result is in accordance with other observations19 and with the observed “loss and gain” of the nutritional lifestyle (see above), thus suggesting that each lifestyle evolved independently of the others.
A total of 11,895 genes of H. oryzae clustered into 9315 MCL groups (Supplementary Fig. S1; Supplementary Table S2). Of these, 3834 genes (1254 groups) were present in one or more paralogs, considerably more than in M. oryzae, M. poae or G. graminis [1383 genes (570 groups), 1514 genes (663 groups) and 2200 genes (848 groups), respectively]. In addition, most of these genes occurred in duplicated DNA sequences (Supplementary Fig. S2), thus illustrating that the larger genome size of H. oryzae is the result of significant gene duplication. A total of 119 groups (192 genes) in H. oryzae were not found in M. oryzae, M. poae or G. graminis, whereas orthologs (generally fewer than 4) were found in the genomes of some of the 13 fungi (Supplementary Fig. S5). The highest number of orthologs was shared between H. oryzae and C. globosum (Supplementary Fig. S5) and many of them were present only in these two fungi (Supplementary Fig. S5). While it is theoretically possible that these genes were obtained by horizontal gene transfer (HGT), we consider this rather unlikely because of the high number of these genes; the rate of HGT within filamentous fungi is typically very low20. Rather, the close similarity of the nutritional lifestyles of H. oryzae and C. globosum may have led to the maintenance of these genes in their genomes while they were lost in other species. These genes can thus be considered typical saprophytic growth genes.
The search for genes involved in the clusters of orthologous group (COG) classifications revealed many more genes in the ‘Lipid transport and metabolism’ cluster for the M. oryzae (1490 genes) genome than in the H. oryzae (395 genes), M. poae (274 genes) and G. graminis (342 genes) genomes (Fig. 2). Lipid transport and metabolism are necessary for M. oryzae to complete appressorium-mediated infection, especially the prepenetration of the leaf cuticle21. The loss of most genes in the ‘Lipid transport and metabolism’ cluster is consistent with the loss of the capability for appressorium-mediated leaf infection in root-infection-only species (see below).
COG function classification of genes in the genomes of H. oryzae and related species.
8763, 7319, 6502 and 7593 genes of H. oryzae, M. oryzae, M. poae and G. graminis genomes, respectively, have COG classifications among the 24 categories. More genes of M. oryzae are in the cluster for ‘Lipid transport and metabolism’.
A total of 573 gene families were shown by a CAFE22 analysis to have undergone expansion in H. oryzae (p < 0.01, at least 4 genes in total), whereas only 10 gene families have undergone contraction (Supplementary Table S5). The most expanded gene families were transposons, as noted above and genes that encode proteins involved in DNA binding, such as centromere protein B (CENP-B). CENP-B proteins are transposase-derived centromeric proteins; in S. pombe, they localize to and recruit histone deacetylases to silence Tf2 retrotransposons23,24. CENP-Bs can also repress solo long terminal repeats (LTRs) and LTR-associated genes, which prevents ‘extinct’ Tf1 retrotransposons from re-entering the host genome23,24. The strong expansion of this gene family in H. oryzae reveals the activity of a retrotransposon surveillance pathway in H. oryzae, which may reflect an attempt to counteract the strong transposon activity.
Other genes for which expansion was demonstrated included those encoding chitinases, the transport and metabolism of carbohydrates (and other solutes), kinesin light chains, serine/threonine protein kinases, secondary metabolite biosynthesis and inorganic ion transport and metabolism. This expansion of genes involved in signaling and transport indicates that H. oryzae has developed a complex regulatory machinery to sense signals received from the external environment and couple them with intracellular signaling and transport pathways.
The root colonization strategies and transcriptome analyses during fungal infection
To identify which of these gene expansions might be related to the shift of H. oryzae from a plant pathogen to a plant endosymbiont, we performed genome-wide expression profiling using RNA-seq (Supplementary Note 3) and we verified selected genes using qPCR (Supplementary Note 4). To obtain detailed information about the root colonization strategy employed by H. oryzae and its differences from that of M. oryzae and thus to choose the most appropriate time points for RNA isolation, we first inoculated roots of in vitro-cultivated rice plants with conidia of DsRed2-tagged H. oryzae strain Ho19red and eGFP-tagged M. oryzae strain Ho31gfp9 and documented infection using fluorescence and confocal microscopy (Supplementary Fig. S6).
By 1–2 days after inoculation (DAI), numerous runner hyphae of H. oryzae were present along the longitudinal axis of the root surface and the fungal penetration of the rhizodermis was observed. Melanized appressoria, which are typically associated with leaf infection, were not observed on the roots. Instead, hyphopodia were present on the surface and penetrated the epidermal cells. Upon penetration (4 DAI), fungal growth was visible in the epidermal cells of the root, but not in the cortical cell layers. Some invasive hyphae (IH) were strictly confined to certain epidermal cells, which contained large numbers of IH. At 6 DAI, mild colonization of cortical cell layers was observed. No IH were detected in vascular tissue, even at a late stage (20 DAI). At 20 DAI, the biomass of the aerial part of the rice colonized by H. oryzae was significantly increased compared with the control plants (Supplementary Fig. S7).
Similar to H. oryzae, the conidia of M. oryzae germinated within 2 DAI and produced fungal hyphae (Supplementary Fig. S6). Numerous runner hyphae were present and formed infectious structures (hyphopodia) and penetrated epidermal cells via penetration pegs12. However, no strictly confined IH were observed in M. oryzae at 4 DAI. The IH had rapidly colonized the vascular tissues by 6 DAI, when the systemic infection of leaves and stems commenced (Supplementary Fig. S6). Based on these results, we chose roots infected by H. oryzae at 2, 6 and 20 DAI and roots infected by M. oryzae at 2 and 6 DAI for the RNA-seq analysis.
The deficiency of the appressorium-mediated leaf infecting ability of H. oryzae
The genes controlling appressorium-mediated penetration in M. oryzae have been extensively studied25. These genes involve four key signaling pathways: the cyclic AMP-protein kinase A (cAMP-PKA) pathway, the MAP kinase (MAPK) pathway, the cell wall integrity MAPK pathway and the osmoregulation pathway. H. oryzae contains orthologs of all these genes, in addition to genes involved in ROS25 and autophagy26, which are also important for the formation and function of appressoria (Supplementary Table S10). Furthermore, most of the experimentally verified virulence-associated genes of M. oryzae are also present in H. oryzae (Supplementary Table S11). However, all three of the fungi that only infect roots (H. oryzae, M. poae and G. graminis) lack 19 virulence genes, including mpg1, moact and morgs7, which are important for interactions with the leaf surface or the penetration ability of appressoria27,28,29. Tucker30 proposed that the ability to develop appressorium-mediated infection was acquired after the ability to infect roots and the former ability required the acquisition of new genes, such as mpg1, or gene functions. Because the ancestor of H. oryzae was a plant pathogen, it is likely that H. oryzae lost the ability to infect leaves owing to the loss of genes indispensable for appressorium-mediated infection. It has been shown that H. oryzae could restrict root infection by M. oryzae9, suggesting that competition between these relatives may have partly resulted in obligate root infection by H. oryzae.
The differences in transduction of the extracellular signal
A total of 73 G-protein-coupled receptors (GPCRs) were found in the M. oryzae genome, including 60 pth11-like GPCRs, whereas only 25, 26 and 27 GPCRs were found in the H. oryzae, M. poae and G. graminis genomes, with 16, 19 and 17 pth11-like GPCRs, respectively (Supplementary Table S12). pth11-like GPCRs were present in all the main clades in the phylogenetic tree of H. oryzae (Supplementary Fig. S11), suggesting that H. oryzae possesses orthologs of all of them. However, there were obvious significant differences in the expression of the abundantly expressed pth11-like GPCRs (Supplementary Fig. S11). Similar findings were obtained for cAMP receptor-like GPCRs (Supplementary Fig. S11). In addition, the orthologous gene (determined by bidirectional best hits) of the M. oryzae Gα subunit magC was expressed much more strongly in H. oryzae. Taken together, these results suggest that M. oryzae and H. oryzae differ in their responses to extracellular signals from the host, which may be part of the difference between endophytic and pathogenetic interactions. These results also make it likely that signals produced by the same host may be received differently by M. oryzae and H. oryzae.
Different nutritional preferences of H. and M. oryzae
H. oryzae possesses the highest number of carbohydrate-active enzymes (CAZymes; 606 genes) compared with its relatives, which is due to the high numbers of glycoside hydrolases, carbohydrate-binding domains and polysaccharide lyases (Supplementary Table S15). A total of 109 plant cell wall-degrading enzymes (CWDEs) are shared by H. oryzae and M. oryzae (Supplementary Fig. S10; S12). Genes that were particularly enhanced in H. oryzae include the GH55 exo-β-1,3-glucanases, GH78 α-rhamnosidases, GH95 α-fucosidases, PL1 polygalacturonate lyases and the CBM67 rhamnose-binding module. Except for GH55, which may be involved in H. oryzae cell wall turnover or antagonism against competing fungi, the other genes all function in hydrolyzing the hemicellulose side chains and pectin of the plant and, thus, in penetrating the roots. These data are also in accordance with the findings that, compared with necrotrophic and hemibiotrophic fungi, plant pathogenic fungi tend to have fewer CWDEs, such as enzymes of GH61, GH78, PL1 and PL3 as well as enzymes containing CBM1, CBM18 and CBM50 domains31. Genome-wide expression profiling using RNA-seq revealed that most CWDEs were expressed in both H. oryzae and M. oryzae at all stages of their interactions with rice. However, there were remarkable differences in the expression patterns of these CWDEs: 44 genes clustered into six groups (Fig. 3a) and were regulated in essentially opposite directions in the two fungi. Most of them were downregulated in H. oryzae but upregulated in M. oryzae except genes in the group B and D (Fig. 3a). While the expressions of genes in the group B were not changed in M. oryzae, they were suppressed in H. oryzae. Similarly, the gene expressions in the group D kept stable in H. oryzae but upregulated in M. oryzae. These results show that the expression levels of CWDEs were tightly restricted at the onset of the biotrophic phase in H. oryzae. A similar pattern was recently reported in a comparison of the infection of living and dead roots by Piriformospora indica32. Those of the present study are consistent with the scenario that following successful colonization, M. oryzae is inclined to kill the root cells and commence necrotrophic nutrition, whereas H. oryzae prefers to keep the root cells alive and exhibits biotrophy. Quantification using GC/MS revealed that the glucose content of roots infected with H. oryzae was significantly higher than that of roots infected with M. oryzae and with sterile water (control samples) at all infection stages (Fig. 3b). Glucose could inhibit the expression of CWDEs by plant-colonizing fungi, which is known as glucose repression33,34,35. Differences in the allocation of carbon by the plant during endophytic and pathogenetic interactions suggest that the host response may play an important role in the evolutionary process of the pathogen, which is consistent with the finding that differentiation among H. oryzae and related species was accompanied by the divergence of their hosts (vide supra).
Different expression patterns of fungal genes encoding plant cell wall-degrading enzymes and different plant carbon allocation in endophytic and pathogenetic interactions.
(a) Significantly regulated CAZymes in either H. oryzae or M. oryzae with enzymatic domains functioning in degrading cellulose, hemicellulose and pectin. Orthologous genes are aligned in the same lines and the short line refers to orthologous genes that have been lost. DAI2, DAI6 and DAI20 refer to transcripts expressed by H. oryzae (RDAI) or M. oryzae (GDAI) infecting rice roots at 2, 6 and 20 days after inoculation, respectively. Red, increase in transcript abundance; green, decrease in transcript abundance. (b) GC-MS quantification of glucose and fructose in rice roots infected with H. oryzae (HOR), M. oryzae (MOR) or sterile water (Control, CON). Blue and red asterisks indicate significant differences of HOR when compared with CON and MOR, respectively (p < 0.01). The y-axis indicates the contents (mg) of D-glucose/D-fructose in 1 mg dry weight of rice roots.
Differences in triggering of host defense responses of H. and M. oryzae
Among the 173 orthologs of virulence-associated genes shared by M. oryzae and H. oryzae, 21 (12.1%) genes were significantly regulated in either M. oryzae or H. oryzae (Supplementary Fig. S15). However, most of these genes were downregulated in H. oryzae but upregulated in M. oryzae, suggesting that the expression of virulence-associated genes in H. oryzae was suppressed. The reduced expression of these genes was also important to decrease plant immune responses.
167 small secreted cysteine-rich proteins (SSCRPs) were identified in the H. oryzae genome, more than the numbers found in related species (Supplementary Table S13). These proteins are considered to facilitate infection by reprogramming host cells and modulating plant immune responses36. Interestingly, none of the significantly regulated SSCRP orthologs in H. oryzae were significantly regulated in M. oryzae and vice versa (Supplementary Fig. S15; Supplementary Table S14). Thus, the differential expression of these genes in H. oryzae and M. oryzae may be a mechanism involved in their different infection strategies.
The H. oryzae CAZyme also encodes an increased arsenal of enzymes and proteins that interact with chitin (82 genes vs. 61 in M. oryzae, Supplementary Fig. S12; Supplementary Table S14). Interestingly, only 40 of these proteins are orthologs (Supplementary Fig. S10). A phylogenetic analysis showed that the CBM50 (LysM) chitin-binding domains had undergone species-specific expansion, with tandem repeats arising from gene duplication (Supplementary Fig. S13). Most of the H. oryzae-specific genes (42 genes) were secreted extracellularly (Supplementary Table S16; Supplementary Table S17). Similar to the GH55 endo-β-1,3-glucanases, these genes could function in H. oryzae cell wall turnover or antagonism against competing fungi. However, LysM-bearing proteins function to scavenge the chitin of the plant host and therefore prevent the recognition of the fungus by the plant or other hosts37,38. In M. oryzae, LysM effectors also suppress chitin-triggered immunity39,40, although the mechanism by which fungal LysM effectors compete with plant receptors for chitin binding remains unknown.
Genes involved in the secondary metabolite
As is typical of fungi, H. oryzae and M. oryzae also encode genes (37 and 35, respectively) for secondary metabolite (SM) synthesis, numbers that are much higher than those of M. poae and G. graminis (Supplementary Table S18). A total of 60% of SMs were shared between H. oryzae and M. oryzae (Supplementary Fig. S10). A phylogenetic analysis revealed the specific expansion of polyketide synthase (PKS) genes in H. oryzae (Supplementary Fig. S13). In addition, cytochrome P450 (CYP) monooxygenases were expanded in H. oryzae (211 genes) and M. oryzae (228 genes) compared with G. graminis and M. poae (Supplementary Table S19). Similar to SM genes, only few of the orthologous CYPs were shared between the M. oryzae and H. oryzae genomes when orthologs were considered (Supplementary Fig. S10). Interestingly, genes involved in abscisic acid signaling, which might play roles in the manipulation of plant hormone metabolism, exhibited lower (or downregulated) expression in H. oryzae, whereas genes involved in the synthesis of auxin and gibberellin exhibited higher (or upregulated) expression in H. oryzae (Supplementary Table S20). We note, however, that H. oryzae possesses no orthologs of the cps/ks genes, which are essential for gibberellin biosynthesis in Fusarium fujikuroi (Supplementary Table S20). Thus, our results suggest that H. oryzae may influence the biosynthesis of this plant hormone instead of producing it directly. The rice growth-promoting function of H. oryzae may therefore be related to auxins and/or gibberellins, which exhibit remarkable promotional effects on plants41.
Horizontal gene transfer (HGT) in the H. oryzae genome
HAOR_13605, which encodes a periplasmic phosphate ABC transporter, was highly expressed (Supplementary Table S24). Interestingly, this gene appears to have been obtained by horizontal gene transfer from a Granulicella sp. (a bacterium belonging to the phylum Acidobacteria42) (Supplementary Fig. S14). However, since this protein lost its function as a phosphate-specific transporter (see Supplementary Note), HAOR_13605 is more likely to function as a sensor of nutrients that regulate carbon metabolism, which requires further investigation.
Conclusion
In this study, we generated a new model of a mutualistic endophytic fungus that originated from pathogenic ancestors and this model provided us with the first chance to study this evolution by comparative genomic and transcriptomic analyses. Our research highlights the importance of both internal dynamics (such as transposable elements) and external pressures (such as competing relationships with relatives and different responses from hosts) in the evolutional transformation of H. oryzae from a pathogen to a mutualistic endophyte. Furthermore, horizontal gene transfer may have also partly affected the H. oryzae genome. H. oryzae has also been shown to be useful for agricultural management9 and the availability of its genome will greatly promote the progress of research into the fundamental mechanisms of mutualistic symbiosis and will pave the way for future roles of H. oryzae in agricultural applications.
Methods
Fungal strains and plant materials
The endophytic Harpophora oryzae strain R5-6-1 and the rice blast fungus Magnaporthe oryzae strain Guy11 have been studied in our laboratory for several years8,9,43. The DsRed2-tagged H. oryzae and eGFP-tagged M. oryzae9 were used to monitor the infection process. The rice cultivar CO-39 (Oryzae sativa), for which the genome sequence is available44, was used as a compatible host plant for inoculation experiments. All the fungal strains were cultured on complete medium (CM)45 at 25°C. The H. oryzae strains were cultured in the dark, whereas the M. oryzae strains were cultured under a 16-h light/8-h dark photoperiod. Conidia were harvested from 10-d-old cultures of M. oryzae. For H. oryzae, germinating phialidic conidia were harvested from 4-d-old potato dextrose broth (PDB, with 5 g glucose/L)46. Rice seeds were surface sterilized as previously described9 and planted in half-strength Murashige & Skoog (1/2 MS) solid medium at 30°C in the dark for 4 days to promote root growth. The plants were then grown vertically under a 16-h light/8-h dark photoperiod at 28/24°C for an additional 6 days. Inoculations were performed as previously described12. Infection assays were conducted using roots laid on sterile filter paper placed on 1/2 MS solid medium. Each plate of four plants received a total of 105 conidia of M. oryzae or 106 conidia of H. oryzae, which were distributed randomly on top of the root system. The inoculated plants were grown horizontally at 26°C under a16-h light/8-h dark photoperiod, with the roots in darkness at all times. The root infection process was monitored using an LSM780 laser-scanning confocal microscope (Carl Zeiss Inc., Jena, Germany). Genomic DNA was extracted from 100 mg of fungal material grown in PDB (with 5 g glucose/L) using a DNeasy Plant Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer's instructions.
Phylogenetic analysis and ancestral state reconstruction (ASR)
Six loci were used in this phylogenetic analysis (Supplementary Table S25). All sequences were aligned, concatenated and manually adjusted using Geneious Pro v4.8.3 (http://www.geneious.com/). GTR + G + I model was selected as the best-fit model for the datasets using jModelTest v247. The phylogenetic analyses were performed using the maximum-likelihood (ML) criterion implemented in RAxML48 through the RAxML-HPC BlackBox web server at CIPRES (http://www.phylo.org/) with the best-fit model. Maximum parsimony (MP) analyses were performed using PAUP v4.0b10. Bayesian inference analyses (BI) were performed with the best-fit model using the Markov chain Monte Carlo method in MrBayes v3.2.149. Information about the studied characters in ASR were retrieved from Luo and Zhang11 and from Yuan et al8. MP- and ML-based ASRs were conducted in Mesquite v2.75 (http://mesquiteproject.org.). To account for phylogenetic mapping uncertainty, we also employed a BI approach to analyze ancestral states using the ‘Multistate’ option in BayesTraits v.2.050. More details are provided in the Supplementary Notes.
Genome sequencing, assembly and analysis
The genome of H. oryzae R5-6-1 was sequenced with the Roche/454 Pyrosequencing Platform and the Illumina Hiseq2000 sequencing platform. Low-quality data that had a QUAL value of less than 30 and consisted of short reads (length < 50 bp) were filtered from the raw data. The high-quality reads underwent primary assembly using the GS De Novo Assembler (Newbler v2.9; Roche)51 and ALLPATHS-LG (version:allpathslg-43984)52 and were then scaffolded using SSPACE v2.053. Finally, gaps were filled using SOAP GapCloser v1.054. The completeness of the H. oryzae genome was assessed using CEGMA v 2.013. The H. oryzae R5-6-1 genome sequence has been deposited to GenBank under accession number JNVV00000000. De novo analysis was employed to examine the repetitive sequences. The repetitive elements were identified and classified using a de novo repetitive sequence search with RepeatModeler v1.07 (http://www.repeatmasker.org/RepeatModeler.html). For repeat annotation, the repeat library produced by RepeatModeler was used directly with RepeatMasker v4-0-3 (http://repeatmasker.org). The RIP indices were determined with the software RIPCAL by comparisons with the non-repetitive genome55,56.
Gene prediction and annotation
The annotation of the genomic sequences of H. oryzae was performed with Augustus v2.757 and trained with the assembled RNA-seq transcripts and the annotated information from M. oryzae was incorporated as a reference. Genes were annotated using BLASTP v2.2.27 against the NR database and InterproScan (http://www.ebi.ac.uk/interpro/). Gene ontologies were classified based on InterproScan annotation IDs. Exon junctions were obtained from RNA-seq using TopHat v2.0.958.
Orthologous genes and evolution analysis
Orthologs were identified using OrthoMCL v2.0 (http://orthomcl.org/orthomcl/). Ortholog, co-ortholog and inparalog pairs were identified with OrthoMCL. The pairs and their weights were used to construct an OrthoMCL graph for clustering with the MCL algorithm59. A total of 844 clusters of 1:1 orthologs of the 17 fungi were obtained and approximately one-third (270) of the clusters were randomly chosen to infer the phylogenomic relationships among these taxa. The proteins of the 270 ortholog groups were aligned using CLUSTALW v260 and were concatenated. The alignments were then analyzed with Gblocks v0.91b61 using the default parameters to select conserved regions. The best amino acid substitution model (LG + I + G model) was chosen using ProtTest v3.262. The divergence times between species were estimated using the Langley-Fitch method with r8s63 by calibrating them against the reassessed origins of the Ascomycota 500–650 million years ago64. The evolution of protein family size variation was analyzed using CAFE v2.222. Protein families from the OrthoMCL analysis were used that contained at least 4 proteins from H. oryzae and related species (M. oryzae, M. poae and G. graminis). The bidirectional best hits (BBHs) of H. oryzae and M. oryzae proteins were considered orthologous genes for the RNA-seq analysis.
Protein family classification
To identify potential pathogenicity and virulence, whole-genome BLAST searches were performed against protein sequences of M. oryzae in the pathogen-host interaction (PHI) database (version 3.4, http://www.phibase.org/) (E-value < 10−5, coverage >55%). G-protein coupled receptors (GPCRs) were predicted based on a previous report65. GPCR-like proteins were evaluated to verify the presence of seven transmembrane helices using TMHMM v2.0 and Phobius (http://www.cbs.dtu.dk/services/TMHMM/ and http://phobius.sbc.su.se/). Carbohydrate-active enzymes were classified using the dbCAN HMMer-based classification system (http://csbl.bmb.uga.edu/dbCAN/). Secreted proteins were identified using WoLF-PSORT (http://wolfpsort.seq.cbrc.jp/). SSCRPs shorter than 200 aa in length and containing at least 4% cysteine residues were identified among the predicted secreted proteins. Cytochrome P450s were classified based on de novo BLASTP searches (E-value < 10−15, coverage >55%) against protein sequences in the Cytochrome P450 Database (FCPD) (http://p450.riceblast.snu.ac.kr/). To identify fungal secondary metabolite pathways, the genome annotation data were analyzed using the web-based prediction tool SMURF (http://jcvi.org/smurf/index.php). Phytohormone-related genes were identified by performing BLASTP against the Arabidopsis Hormone Database (AHD) v2.0 (http://ahd.cbi.pku.edu.cn/).
Transcriptome analysis
Roots infected with H. oryzae were harvested in liquid nitrogen 2, 6 and 20 days after infection (DAI); the same procedure was applied to M. oryzae, except that the harvesting occurred 20 DAI. Roots from 12 independent rice plants were considered an experimental replicate. Total RNA was extracted using TRIzol reagent (Invitrogen) according to the manufacturer's protocol. The RNA integrity of all the samples was verified on an Agilent 2100 Bioanalyzer. Nine H. oryzae libraries (three developmental stages and three biological replicates) and six M. oryzae libraries (two developmental stages and three biological replicates) were prepared with the Illumina TruSeq RNA Sample Preparation Kit and were sequenced using the Illumina HiSeq 2000 based on 100 bp paired-end read sequencing. The insert sizes of all the libraries were 180 bp for both H. oryzae and M. oryzae. All the clean reads were mapped to the genome sequence using TopHat v 2.0.958 and an expression profile was created using Cufflinks v2.0.266. The abundances were reported as normalized fragments per kb of transcript per million mapped reads. Transcripts with a significant P value (<0.05) and a greater than twofold change (log2) in transcript level were considered differentially expressed. All the P values were corrected for false discoveries resulting from multiple hypothesis testing using the Benjamini-Hochberg procedure. Heatmaps of gene expression profiles were generated using R (www.R-project.org) based on significant expression changes (log10 FPKM plus 1).
Quantification of glucose and fructose in rice roots
The contents of glucose and fructose in rice roots were quantified using gas chromatography-mass spectrometry (GC-MS). All the conditions were as described for the RNA-seq preparation. GC-MS was performed in parallel with cultures harvested for RNA-seq, with three additional control samples. The control samples for 2 DAI, 6 DAI and 20 DAI were prepared using sterile water instead of a conidia suspension for inoculation. Each mixture of roots from 12 independent rice plants was considered an experimental repetition and six biological replicates were performed for each sample. The sample preparation and parameters and the procedures for the GC-MS analysis were as previously described67 with slight modifications (more details are provided in the Supplementary Notes).
References
Valent, B. & Chumley, F. G. Molecular genetic analysis of the rice blast fungus, Magnaporthe grisea. Annu. Rev. Phytopathol. 29, 443–467 (1991).
Talbot, N. J. On the trail of a cereal killer: exploring the biology of Magnaporthe grisea. Annu. Rev. Microbiol. 57, 177–202 (2003).
Ebbole, D. J. Magnaporthe as a model for understanding host-pathogen interactions. Annu. Rev. Phytopathol. 45, 437–456 (2007).
Zamioudis, C. & Pieterse, C. M. Modulation of host immunity by beneficial microbes. Mol. Plant-Microbe Interact. 25, 139–150 (2012).
Porras-Alfaro, A. & Bayman, P. Hidden fungi, emergent properties: endophytes and microbiomes. Phytopathology 49, 291–315 (2011).
Collins, S. L. et al. Pulse dynamics and microbial processes in aridland ecosystems. J. Ecol. 96, 413–420 (2008).
Green, L. E., Porras-Alfaro, A. & Sinsabaugh, R. L. Translocation of nitrogen and carbon integrates biotic crust and grass production in desert grassland. J. Ecol. 96, 1076–1085 (2008).
Yuan, Z. L., Lin, F. C., Zhang, C. L. & Kubicek, C. P. A new species of Harpophora (Magnaporthaceae) recovered from healthy wild rice (Oryza granulata) roots, representing a novel member of a beneficial dark septate endophyte. FEMS Microbiol. Lett. 307, 94–101 (2010).
Su, Z. Z. et al. Evidence for biotrophic lifestyle and biocontrol potential of dark deptate endophyte Harpophora oryzae to Rice Blast Disease. PloS One 8, e61332 (2013).
Zhang, N., Zhao, S. & Shen, Q. A six-gene phylogeny reveals the evolution of mode of infection in the rice blast fungus and allied species. Mycologia 103, 1267–1276 (2011).
Luo, J. & Zhang, N. Magnaporthiopsis, a new genus in Magnaporthaceae (Ascomycota). Mycologia 105, 1019–1029 (2013).
Marcel, S., Sawers, R., Oakeley, E., Angliker, H. & Paszkowski, U. Tissue-adapted invasion strategies of the rice blast fungus Magnaporthe oryzae. Plant Cell 22, 3177–3187 (2010).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Hane, J. K. et al. A novel mode of chromosomal evolution peculiar to filamentous Ascomycete fungi. Genome Biol. 12, R45 (2011).
Jacobs, B. F., Kingston, J. D. & Jacobs, L. L. The origin of grass-dominated ecosystems. Ann. Missouri Bot. Gard. 86, 590–643 (1999).
Kellogg, E. A. Evolutionary history of the grasses. Plant Physiol. 125, 1198–1205 (2001).
Gaut, B. S. Evolutionary dynamics of grass genomes. New Phytol. 154, 15–28 (2002).
Dean, R. A. et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434, 980–986 (2005).
Soanes, D. M. et al. Comparative genome analysis of filamentous fungi reveals gene family expansions associated with fungal pathogenesis. PLoS One 3, e2300 (2008).
Fitzpatrick, D. A. Horizontal gene transfer in fungi. FEMS Microbiol. Lett. 329, 1–8 (2012).
Wang, Z. Y., Soanes, D. M., Kershaw, M. J. & Talbot, N. J. Functional analysis of lipid metabolism in Magnaporthe grisea reveals a requirement for peroxisomal fatty acid β-oxidation during appressorium-mediated plant infection. Mol. Plant Microbe Interact. 20, 475–491 (2007).
De Bie, T., Cristianini, N., Demuth, J. P. & Hahn, M. W. CAFE: a computational tool for the study of gene family evolution. Bioinformatics 22, 1269–1271 (2006).
Cam, H. P., Noma, K. I., Ebina, H., Levin, H. L. & Grewal, S. I. Host genome surveillance for retrotransposons by transposon-derived proteins. Nature 451, 431–436 (2008).
Mikheyeva, I. V. et al. CENP-B cooperates with Set1 in bidirectional transcriptional silencing and genome organization of retrotransposons. Mol. Cell Biol. 32, 4215–4225 (2012).
Li, G., Zhou, X. & Xu, J. R. Genetic control of infection-related development in Magnaporthe oryzae. Curr. Opin. Microbiol. 15, 678–684 (2012).
Liu, X. H. et al. Autophagy vitalizes the pathogenicity of pathogenic fungi. Autophagy 8, 1415–1425 (2012).
Talbot, N. J. et al. MPG1 encodes a fungal hydrophobin involved in surface interactions during infection-related development of Magnaporthe grisea. Plant Cell 8, 985–999 (1996).
Guo, M. et al. The bZIP transcription factor MoAP1 mediates the oxidative stress response and is critical for pathogenicity of the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 7, e1001302 (2011).
Zhang, H. et al. Eight RGS and RGS-like proteins orchestrate growth, differentiation and pathogenicity of Magnaporthe oryzae. PLoS Pathog. 7, e1002450 (2011).
Tucker, S. L. et al. Common genetic pathways regulate organ-specific infection-related development in the rice blast fungus. Plant Cell 22, 953–972 (2010).
Zhao, Z., Liu, H., Wang, C. & Xu, J. R. Comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genomics 14, 274 (2013).
Zuccaro, A. et al. Endophytic life strategies decoded by genome and transcriptome analyses of the mutualistic root symbiont Piriformospora indica. PLoS Pathog. 7, e1002290 (2011).
Nadal, M., Garcia-Pedrajas, M. D. & Gold, S. E. The snf1 gene of Ustilago maydis acts as a dual regulator of cell wall degrading enzymes. Phytopathology 100, 1364–1372 (2010).
Yi, M., Park, J. H., Ahn, J. H. & Lee, Y. H. MoSNF1 regulates sporulation and pathogenicity in the rice blast fungus Magnaporthe oryzae. Fungal Genet. Biol. 45, 1172–1181 (2008).
Tonukari, N. J., Scott-Craig, J. S. & Walton, J. D. The Cochliobolus carbonum SNF1 gene is required for cell wall-degrading enzyme expression and virulence on maize. Plant Cell 12, 237–248 (2000).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Bolton, M. D. et al. The novel Cladosporium fulvum lysin motif effector Ecp6 is a virulence factor with orthologues in other fungal species. Mol. Microbiol. 9, 119–136 (2008).
de Jonge, R. & Thomma, B. P. Fungal LysM effectors: extinguishers of host immunity? Trends Microbiol. 17, 151–157 (2009).
Marshall, R. et al. Analysis of two in planta expressed LysM effector homologs from the fungus Mycosphaerella graminicola reveals novel functional properties and varying contributions to virulence on wheat. Plant Physiol. 156, 756–769 (2011).
Mentlak, T. A. et al. Effector-mediated suppression of chitin-triggered immunity by Magnaporthe oryzae is necessary for rice blast disease. Plant Cell 24, 322–35 (2012).
Davies, P. J. Plant Hormones (Springer, Netherlands, 2010).
Männistö, M. K., Rawat, S., Starovoytov, V. & Häggblom, M. M. Granulicella arctica sp. nov., Granulicella mallensis sp. nov., Granulicella tundricola sp. nov. and Granulicella sapmiensis sp. nov., novel acidobacteria from tundra soil. Int. J. Syst. Evol. Microbiol. 62, 2097–2106 (2012).
Dong, B. et al. MgAtg9 trafficking in Magnaporthe oryzae. Autophagy 5, 946–953 (2009).
Goff, S. A. et al. A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296, 92–100 (2002).
Talbot, N. J., Ebbole, D. J. & Hamer, J. E. Identification and characterization of MPG1, a gene involved in pathogenicity from the rice blast fungus Magnaporthe grisea. Plant Cell 5, 1575–1590 (1993).
Sivasithamparam, K. Phialophora and Phialophora-like fungi occurring in the root region of wheat. Aust. J. Bot. 23, 193–212 (1975).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9, 772–772 (2012).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Pagel, M. & Meade, A. Bayesian analysis of correlated evolution of discrete characters by reversible-jump Markov Chain Monte Carlo. Am. Nat. 167, 808–825 (2006).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Gnerre, S. et al. Highquality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl Acad. Sci. USA 108, 1513–1518 (2011).
Boetzer, M., Henkel, C. V., Jansen, H. J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Watters, M. K., Randall, T. A., Margolin, B. S., Selker, E. U. & Stadler, D. R. Action of repeat-induced point mutation on both strands of a duplex and on tandem duplications of various sizes in Neurospora. Genetics 153, 705–714 (1999).
Hane, J. K. & Oliver, R. P. RIPCAL: a tool for alignment-based analysis of repeat-induced point mutations in fungal genomic sequences. BMC Bioinformatics 9, 478 (2008).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Castresama, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011).
Taylor, J. W. & Berbee, M. L. Dating divergences in the Fungal Tree of Life: review and new analyses. Mycologia 98, 838–849 (2006).
Lücking, R., Huhndorf, S., Pfister, D. H., Plata, E. R. & Lumbsch, H. T. Fungi evolved right on track. Mycologia 101, 810–822 (2009).
Kulkarni, R. D., Thon, M. R., Pan, H. & Dean, R. A. Novel G-protein-coupled receptor-like proteins in the plant pathogenic fungus Magnaporthe grisea. Genome Biol. 6, R24 (2005).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Zhang, L. et al. Metabolic profiling of Chinese tobacco leaf of different geographical origins by GC-MS. J. Agric. Food Chem. 61, 2597–2605 (2013).
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (30970097 and 30925025) and the Program for Changjiang Scholars and Innovative Research Team in University (PCSIRT0943) of Ministry of Education of the People's Republic of China. We acknowledge the Hangzhou Woosen Biotechnology Co. Ltd (Zhejiang, China) for the genome and transcriptome sequencing of H. oryzae R5-6-1 and M. oryzae Guy11.
Author information
Authors and Affiliations
Contributions
C.L.Z. and F.C.L. initiated and coordinated the project. X.H.X. performed comparative genomics and transcriptomics analyses, Z.Z.S., C.W., C.C. and J.Y.W. performed transcriptomics analyses, X.H.X. and C.P.K. wrote and edited the paper, X.X.F. coordinated genome and transcriptome sequencing, C.W. and L.J.M performed GC-MS analysis.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Supplementary Table S5
Supplementary Information
Supplementary Table S10
Supplementary Information
Supplementary Table S11
Supplementary Information
Supplementary Table S12
Supplementary Information
Supplementary Table S22
Supplementary Information
Supplementary Table S24
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Xu, XH., Su, ZZ., Wang, C. et al. The rice endophyte Harpophora oryzae genome reveals evolution from a pathogen to a mutualistic endophyte. Sci Rep 4, 5783 (2014). https://doi.org/10.1038/srep05783
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05783
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.