Abstract
Warburg effect is a dominant phenotype of most cancer cells. Here we show that this phenotype depends on its environment. When cancer cells are under regular culture condition, they show Warburg effect; whereas under lactic acidosis, they show a nonglycolytic phenotype, characterized by a high ratio of oxygen consumption rate over glycolytic rate, negligible lactate production and efficient incorporation of glucose carbon(s) into cellular mass. These two metabolic modes are intimately interrelated, for Warburg effect generates lactic acidosis that promotes a transition to a nonglycolytic mode. This dual metabolic nature confers growth advantage to cancer cells adapting to ever changing microenvironment.
Similar content being viewed by others
Introduction
Warburg effect is a metabolic hallmark of most cancer cells, characterized by an excessive conversion of glucose to lactate even with ample oxygen1,2,3. This metabolic mode wastes most, estimated at 80–90%, of glucose carbon. While Warburg effect occupies a central position in cancer metabolism4, we ask if cancer cells might use glucose in a more economic fashion. This idea arises from a fact that glucose levels in solid tumors are typically very low5; e.g., in human stomach and colon cancers, the average glucose levels are only 0.1 and 0.4 mM, respectively. Conceivably, given such low glucose levels, an economic utilization of glucose is strategically important for cancer cells to survive or grow.
Lactic acidosis is common in most solid tumors6,7,8,9. Biochemically, it is an inevitable consequence of the Warburg effect. We postulate that lactic acidosis has a capacity to feedback to glucose utilization/metabolism. In an earlier report, we showed that lactic acidosis could alter glucose metabolism in cancer cells, but did not address the issue in detail10; here we show that it exerts a significant effect on glucose metabolism in cancer cells by converting a usually dominant Warburg effect to a nonglycolytic phenotype. Our findings reveal a dual metabolic nature of cancer cells, which is critical for their survival and growth.
Results and Discussion
Cancer cells exhibit dual metabolic nature – Warburg effect versus a nonglycolytic phenotype
Under regular culture conditions, cancer cells showed a typical Warburg effect, converting most incoming glucose to lactate; however, under lactic acidosis, they became less glycolytic but more ‘oxidative’ (Figure 1a), i.e., while the oxygen consumption rate (OCR) was comparable to that of the control cells, the extracellular acidification rate (ECAR) was reduced significantly, leading to a ~5-fold increase of the OCR/ECAR ratio (Figure 1a). As ECAR and OCR reflects the glycolytic rate and OXPHOS, this 5-fold reduction of the glycolytic rate without obvious change of OCR suggests that cells under lactic acidosis is not glycolytic and relies more on OXPHOS for ATP generation. Supporting this notion is that cellular ATP concentration was not significantly disturbed by the decrease of glycolysis (Figure 1b) - the intracellular ATP level in cells under lactic acidosis (~0.50 mM) was only marginally lower than that in control cells (~0.64 mM). Accordingly, lactic acidosis considerably reduced the glucose consumption rate and virtually blocked net lactate generation, but significantly increased the ratios of cell mass, as well as incorporation of glucose into DNA or RNA, over glucose consumed (Figure 1c). The above results suggested that cancer cells under lactic acidosis are ‘oxidative’ and using glucose more economically, representing a nonglycolytic phenotype.
The effect of lactic acidosis on oxygen consumption, glycolysis and fate of glucose metabolism in 4T1 cells.
(a) The effect of lactic acidosis on oxygen consumption rate (OCR), extracellular acidification rate (ECAR) and OCR/ECAR ratio in 4T1 cells. Vertical lines indicate time points of the administration of corresponding inhibitors. When oligomycin is added, the ATP synthase complex is inhibited such that the respiratory chain associated oxygen consumption is inhibited. When the ATP synthesis uncoupler FCCP is added, oxygen consumption by the respiratory chain is resumed; when rotenone (complex I inhibitor) and antimycin A (complex III inhibitor) are added, the electron transfer as well as oxygen consumption by the respiratory chain is blocked. (b) ATP levels in 4T1 cells with or without lactic acidosis. (c) 4T1 cells are incubated in culture medium with or without lactic acidosis (20 mM lactate, pH 6.7) for 48 hours traced by [14C]-6-D-glucose. The following parameters are determined as described in Methods: glucose consumption, lactate generation, cell counts and incorporation of 14C into DNA and RNA (μmole glucose/μg DNA or RNA/per μmole glucose consumed, see Methods). The data are mean ± SD, n = 3, except n = 6 in Seahorse experiment and are confirmed by 2 independent experiments. * p < 0.05, ** p < 0.01, compared with control.
When cells were exposed to lactic acidosis conditions at varying degrees, they exhibited a metabolic mode between Warburg effect and nonglycolytic phenotype (Figure 2a).
Effect of lactic acidosis on glucose consumption and lactate generation by 4T1 cells.
(a) A titration of exogenous lactic acid on glucose consumption, net lactate production and cell division. 4T1 cells were incubated in RPMI-1640 medium supplemented with lactic acid to desired lactate concentrations as indicated, incubated for 24 hours, followed by cell counting and glucose consumption and net lactate generation determinations. L/G ratio: lactate generated over glucose consumption; C/G ratio, cells produced by consumption of 1 μ mole glucose. (b) Lactic acidosis generated by 4T1 cancer cells transit Warburg effect to a nonglycolytic phenotype. 4T1 cells were incubated in the RPMI-1640 medium containing 18 mM glucose; at indicated hours, medium glucose and lactate levels, medium pH and L/G ratios were monitored. A given L/G ratio denotes μmoles lactate generated from consuming one μmole glucose within an incubation period, e.g., the L/G ratio at 64 hours means the lactate generated between 48 and 64 hours divided by the glucose consumed in the same period. Note that the L/G ratio is correlated with the medium pH and inversely correlated with the lactate concentration. Data are mean ± SD, n = 3 and were confirmed by 2 independent experiments. * p < 0.05, ** p < 0.01, compared with control.
The transition from Warburg effect to a nonglycolytic mode was observed when cancer cells were cultured with high levels of glucose: they were initially glycolytic (converting most glucose to lactate), then, with accumulation of lactate and acidification of medium (lactic acidosis), a corresponding decrease of glucose consumption, lactate generation, as well as the ratio of lactate generated over glucose consumed were observed (Figure 2b).
The transition from glycolytic to nonglycolytic phenotypes was also observed in a number of other cancer cell lines including Bcap37, HeLa and A549: they exhibited a glycolytic phenotype under regular conditions but switched to a nonglycolytic phenotype under lactic acidosis (Figure 3 and Supplementary Fig. S1).
The effect of lactic acidosis on oxygen consumption and glycolysis in Bcap 37, Hela and A549 cells.
The effect of lactic acidosis on oxygen consumption rate (OCR), extracellular acidification rate (ECAR) and OCR/ECAR ratio in Bcap37, Hela and A549 cells. The data are mean ± SD, n = 6 and are confirmed by 2 independent experiments. * p < 0.05, ** p < 0.01, compared with control.
The above results taken together represent a transition from Warburg effect to a nonglycolytic phenotype and a revelation of a dual metabolic nature of cancer cells. The lactic acidosis conditions used in this study are within the physiological range found in diverse solid tumors6,7,8,9.
A mechanistic insight into lactic acidosis regulating metabolic phenotype
The effect of lactic acidosis on glucose metabolism can be dissected into 2 effects exerted separately by lactate and proton.
Under varying lactic acidosis conditions, cellular pyruvate levels remained constant (Figure 4a), but lactate levels were proportional to extracellular lactate concentrations (Figure 4b), resulting in an increment of the [lactate]/[pyruvate] ratio (Figure 4c). It was noted that the net lactate production by cancer cells was inversely correlated with the intracellular [lactate]/[pyruvate] ratios (Figure 4e): when the ratio was 171 (under 20 mM lactic acidosis), cells consumed glucose but did not generate net lactate; when the ratio was smaller than 171, cells consumed glucose and generated lactate; when the ratio was larger than 171, cells consumed both glucose and lactate; According to this relationship and the equilibrium constant (9.01 × 103) of the reaction11 (conversion between lactate and pyruvate), we estimated that the cytosolic free NAD/NADH ratio in 4T1 cancer cells was 53. At equilibrium, the mass action ratio (Q, [lactate][NAD]/[pyruvate][NADH]) is ~9.00 × 103 (Figure 4f), which is equal to the equilibrium constant of the reaction, i.e., the forward reaction and reverse reaction rates are equal with no net change of Gibbs free energy (Figure 4g). One important assumption is that, under physiological conditions, the cytosolic free NAD/NADH ratio is considered to be a constant. This issue has been addressed in a very detailed manner in an earlier report12.
The conversion between pyruvate + NADH and lactate + NAD+ in 4T1 cells under varying lactic acidosis conditions.
(a–c) Intracellular concentrations of pyruvate and lactate and lactate/pyruvate ratios under varying lactic acidosis conditions (culture medium supplemented with pure lactic acid as indicated). The data represent mean ± SD, n = 3 and are confirmed by 2 independent experiments. * p < 0.05, ** p < 0.01, compared with control. (d & e) Glucose consumption and lactate generation by cells under varying lactic acidosis conditions. (f) The estimated mass action ratio Q ([lactate][NAD]/[pyruvate][NADH]) values in 4T1 cells cultured under varying lactic acid conditions. (g) Changes of Gibbs free energy (ΔG) for the conversion between pyruvate + NADH and lactate + NAD+ in 4T1 cells, according to the equation in Methods.
Lactic acidosis acidified the cytosol (Figure 5a); low pH significantly decreased the overall glycolytic flux (Figure 5b) and inhibited the activities of all glycolytic enzymes except PGAM and PGK (Figure 5c and Supplementary Fig. S2). Increasing the lactate anion level did not significantly affect the glycolytic rate (Supplementary Fig. S3) nor activities of glycolytic enzymes, except that the GAPDH activity was enhanced (Supplementary Fig. S4).
Effect of pH on glycolytic flux.
(a) Intracellular pH (pHi) and extracellular pH (pHe), when 4T1 cells are cultured with or without lactic acidosis (culture medium with 20 mM lactate, pH 6.7). (b) A time course glycolysis assay containing 250 μg/ml 4T1 cell lysate along with all essential cofactors and 10 mM glucose, a titration glycolysis assay with indicated protein concentrations for 60 minutes and pH-dependent inhibition of the glycolytic flux in a reaction containing 250 μg/ml protein of 4T1 cell lysate (60 minutes). Reaction mixtures containing boiled 4T1 cell lysates serve as controls. (c) pH-dependent inhibition of individual glycolytic enzymes and LDH in 4T1 cell lysate. The data are mean ± SD, n = 3 and are confirmed by 3 independent experiments. * p < 0.05, ** p < 0.01, compared with control.
Taken together, we came to a notion that, while proton inhibits glycolytic enzymes leading to a reduced glycolytic flux, lactate level is crucial in dictating the status (equilibrium or non-equilibrium) of the LDH-catalyzed reaction. A plausible explanation for why there is no net lactate generation under lactic acidosis (20 mM lactate with pH ≤ 6.7) is: first, the equal forward and backward rates of the LDH-catalyzed reaction allow pyruvate generated from glycolysis flow to metabolic branches other than to lactate; second, the pyruvate generation rate from glycolysis is equal to its removal rate (e.g., pyruvate carboxylation, entering Krebs cycles, etc.). When condition changes either way, there would be positive or negative net lactate production (Figure 4e).
The effect of lactic acidosis on glucose transport and glycolytic enzyme expression
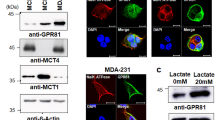
We measured glucose transport into cells under regular and lactic acidosis conditions. Lactic acidosis significantly inhibited [3H]-2-DG transport into cells but did not alter Glut1 expression (Figure 6a & b). However, the intracellular glucose concentration in cells under lactic acidosis was significantly higher than that in control cells (Figure 6c), suggesting that glycolysis was impaired more severely than glucose transport.
Effect of lactic acidosis (20 mM/pH 6.7) on glucose uptake, intra/extracellular glucose levels, glucose consumption and lactate generation.
(a) Glut1 expression. 4T1 cells incubated in RPMI-1640 medium with lactic acidosis for indicated hours were lysed, total proteins resolved by SDS-polyacrylamide gel electrophoresis and Glut1 expression analyzed by immuno-blot. (b) Lactic acidosis significantly inhibits [3H]-2DG uptake by 4T1 cells. (c & d) Intracellular and extracellular glucose levels of 4T1 cells incubated with or without lactic acidosis. (e) Lactate generation by 4T1 cells with or without lactic acidosis. Data represent mean ± SD, n = 3 and were confirmed by at least 2 independent experiments. * p < 0.05, ** p < 0.01, compared with control.
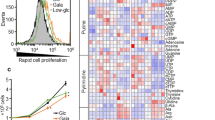
Apart from direct inhibition of glycolytic enzymes by proton, impaired glycolysis might be owing to reduced expression of enzymes. We incubated 4T1 or Bcap37 cells with lactic acidosis for 0, 1, 2 or 3 days and used derived cell lysates to measure the expression levels by immuno-blots and activities of individual enzymes assayed under standard condition at pH 7.3. Although the qualities of commercial antibodies impeded us from detecting murine HK, PGI, PGAM, ENO2 in 4T1 cells, a general lack of decreased activities or expression of most glycolytic enzymes over time in both murine and human cancer cells was observed (Figure 7). In contrast to the activities of most enzymes including 3 rate-limiting enzymes HK, PFK and PK increased over the incubation time of cells under lactic acidosis, this trend was not observed in Bcap37 cells (Figure 7a).
The effect of lactic acidosis on the expression and activities of glycolytic enzymes.
Cells were incubated in RPMI-1640 medium supplemented with lactic acidosis. At 0, 1, 2 or 3 day, individual enzyme activity and expression level in derived cell lysates were measured. (a) Enzyme activity for each glycolytic enzyme in the lysates of Bcap37 cells (upper panel) and 4T1 cells (lower panel) incubated with lactic acidosis over the indicated time course. Data represent mean ± SD, n = 3. * p < 0.05, ** p < 0.01, compared with control. Data were confirmed by 2 independent experiments. (b) Immuno-blots analyses for indicated glycolytic enzyme in the lysates of Bcap37 cells (left panel) or 4T1 cells (right panel) incubated at the same condition as in (a).
The results demonstrated so far taken together suggest that regulation of glucose utilization/metabolism by lactic acidosis is mainly via direct effects of proton and lactate on the glycolytic status.
The effect of acidosis and lactosis on glycolysis
Under acidosis alone, the cytosol was acidified hence glucose consumption and lactate generation was significantly impaired (Supplementary Fig. S5a–c). However, without providing extracellular lactate, the intracellular [lactate]/[pyruvate] ratio was 28 (Supplementary Fig. S5d–f) and cytosolic free [NAD]/[NADH] ratio was 53. Thus, the Q value [lactate][NAD]/[pyruvate][NADH] is 1.48 × 103, which was significantly smaller than the equilibrium constant (9.00 × 103). Hence, acidosis alone allowed net lactate production in cancer cells (Supplementary Fig. S5c).
Lactosis (20 mM, pH 7.4) only slightly changed the glucose consumption and lactate generation rates (Supplementary Fig. S5b & c) and it alone did not decrease cytosolic pH (Supplementary Fig. S5a) thus would not significantly inhibit the glycolytic rate. Lactosis only moderately increased the cellular [lactate]/[pyruvate] ratio (Supplementary Fig. S5d–f) hence would not significantly affect the forward and reverse reaction rates between lactate and pyruvate according to the Q value and the change in Gibbs free energy (Supplementary Fig. S5g & h).
Lactate transport into cells under lactic acidosis differed significantly from that under lactosis (Supplementary Fig. S6 and S5e). This is due to a proton gradient across the cellular membrane. With lactic acidosis, the higher proton level at the medium side than at the cytosol side (Supplementary Fig. 5a) facilitates the lactate transport into cells by monocarboxylic acid transporter (MCT1)13, because MCT1 is a symporter for lactate and proton14. Under lactosis, a reversed gradient (Supplementary Fig. S5a) hampers lactate transport into cells.
The advantage of dual metabolic nature of cancer cells
Warburg effect confers growth advantage to cancer cells in diverse ways1,2,3,4. When glucose supply is sufficient, glucose overconsumption as a result of Warburg effect generates lactic acidosis, which may confer another advantage to cancer cells: excessive glucose consumption may lead to glucose starvation and a Warburg effect to nonglycolytic phenotype transition by the lactic acidosis may then allow cells to more economically use glucose that is limiting. We tested this idea both in a cell culture system and in an animal model, as shown below.
When cultured with a low glucose level (0.5 mM), cancer cells remained proliferative with lactic acidosis but rapidly die without it (Figure 8a). Cancer cells with limited supply of glucose (6 mM) rapidly exhaust glucose followed by a quick death; however, with lactic acidosis, they consume glucose considerably slower and maintain a progressive proliferation for a significantly longer period (Figure 8b). We then used 4T1 tumor as a model to further test the above idea in mice. We proposed that local delivery of basic bicarbonate to tumor may convert intratumoral lactic aicdosis to lactosis, which in principle would switch cancer cells back from a nonglycolytic phenotype to Warburg effect. When glucose is limiting, Warburg effect potentially would deprive glucose and lead to even severer metabolic stress. In 4T1 tumors with or without bicarbonate treatment (Figure 8c–e), glucose and lactate levels were comparable, but intratumoral pH was significantly higher in bicarbonate group than in control group, suggesting a conversion from lactic acidosis to lactosis. Additionally, bicarbonate, while imposing virtually no effect on the overall body weight (Figure 8f), significantly inhibited 4T1 tumor growth (Figure 8g & h) and caused a severer necrosis (Figure 8i), as compared to the control. These results were consistent with our previous data: under severe glucose deprivation, cancer cells survive under lactic acidosis but die otherwise10. Taken together, we suggest that Warburg effect confers growth advantage to cancer cells when glucose supply is sufficient, whereas nonglycolytic mode, promoted by lactic acidosis, renders cancer cells adaptive to metabolic microenvironments with limited glucose supply.
Impacts of lactic acidosis on cancer cell proliferation or survival in vitro and in vivo.
(a) Lactic acidosis allows 4T1 cells to grow under moderate glucose deprivation. Cells incubated in RPMI-1640 medium containing 0.5 mM glucose without lactic acidosis or with lactic acidosis (20 mM lactate and pH 6.7) are counted at indicated time points. Data represent mean ± SD, n = 3 and are confirmed by 2 independent experiments. (b) Lactic acidosis allows a significantly longer sustainable proliferation with limited supply of glucose. Cells incubated in RPMI-1640 medium containing 6 mM glucose without lactic acidosis or with lactic acidosis (20 mM lactate and pH 6.7) are counted at indicated time points. Data represent mean ± SD, n = 3 and are confirmed by 2 independent experiments. (c–i) Elevating intratumoral pH inhibits tumor growth and promotes tumor necrosis. Sodium bicarbonate administration surrounding tumor areas inhibits 4T1 tumor growth and increases their necrosis. (c–e) Intratumoral glucose and lactate concentration and pH; (f) Body weight curves; (g) Tumor growth curves; (h) Tumor weights on the day of sacrifice; (i) Necrosis in tumors. The n is 5 for each group at the starting point; however, one mouse in the control group dies on day 19. Hence, the images of 4 tumor samples from each group are shown. For detailed experimental procedures refer to Methods. The statistical data represent mean ± SEM. * p < 0.05, ** p < 0.01, *** p < 0.001, compared with control.
Concluding remarks
We show that a given cell type can have dual metabolic (Warburg effect or nonglycolytic phenotype) natures under different conditions, which confers survival and/or growth advantage to cancer cells adapting to changing environments. This finding is different from a current concept15, which classifies cancer cells into 2 distinct subgroups, either Warburg effect or oxidative.
Our study may deliver a novel, feasible and effective way to manipulate cancer metabolism. Lactic acidosis and low glucose levels are common in many solid tumors. In principle, cancer cells in these tumors may exhibit nonglycolytic phenotype, which is an economical mode to use glucose hence is important for cancer cells to survive or to proliferate when tumor glucose level is low. Increasing the tumor pH, e.g., by bicarbonate infusion, can convert lactic acidosis to lactosis. Importantly, under lactosis, cancer cells switch back to Warburg effect, which rapidly use glucose and waste most glucose carbon. Consequently, this may lead to glucose deprivation and kill cancer cells. Therefore, manipulation of tumor pH may deliver an effective way to intervene cancer metabolism for therapeutic purpose. Besides regulation of glycolysis, many previous reports have confirmed that lactic acidosis played multifaceted roles in promoting cancer progression including cancer cell metabolism16,17 and survival10,18, cancer cell chromosomal instability19, tumor angiogenesis7,20. Getenby and Gillies groups confirmed that bicarbonate infusion into tumors can significantly attenuate metastasis. and cancer metastasis19,21. Clinical studies demonstrated that high level of lactate was a strong prognostic indicator of increased metastasis and poor overall survival7,15,22,23,24,25,26. Therefore, neutralizing intratumoral lactic acidosis is potentially important in cancer therapy.
While complex signaling networks, which regulate glucose transporter and glycolytic enzymes, plays important roles in glucose metabolism27,28,29, the direct effect of biochemical factors in tumor environment on enzyme activity and reaction status (equilibrium or non-equilibrium), which has been somehow overlooked, represents a rapid and effective mode of regulation.
Methods
Cancer cell lines
Murine breast cancer cell line 4T1, human breast cancer cell line Bcap37, human cervical cancer cell line HeLa, human lung cancer cell line A549 are used in this study. In pilot experiments, we measured the glucose consumption and lactate generation rates of these cells and noted that over 80% of incoming glucose was converted to lactate by these cells even with the presence of ample oxygen. As these cell lines exhibit typical Warburg effect, they were chosen for the study. In addition, all the cell lines are characterized to be tumorigenic and 4T1 cells are highly metastatic.
Lactic acidosis, lactosis and acidosis
A lactic acidosis condition is generated by adding pure lactic acid into the cell culture medium (RPMI-1640, supplemented with 10% FCS) to a final lactate level at 20 mM and final pH at 6.7. A lactosis condition is generated by adding pure lactic acid into the medium to a final lactate level at 20 mM and the pH is adjusted to 7.4 by 0.1 N NaOH. An acidosis condition is generated by adding 0.1 N HCl into the culture medium until pH reaches 6.7.
Incorporation of glucose into nucleic acids
Half million (5 × 105) cells per well in complete RPMI are seeded into culture flasks and allowed to attach overnight. The culture media are replaced with 10 ml fresh complete RPMI plus 6 mM glucose containing 0.1 μCi [14C]-6-D-glucose (specific radioactivity 55 mCi/mmol, ARC0121A, American Radiolabeled Chemicals, Inc, St. Louis, MO, USA) and supplemented with or without 20 mM pure lactic acid. After incubation at 37°C in a humidified CO2 incubator for 48 hours, the medium is collected for glucose and lactate determination and the cells are washed with ice-cold PBS and trypsinized, DNA and RNA extracted with AllPrep® DNA/RNA/Protein Mini Kit (QIAGEN) for radioactivity count by a liquid scintillation counter (Wallac 1414, Finland). The overall calculation is in following steps. Step 1, calculation of μmole glucose incorporation into DNA or RNA: DPM (decay per minute) of total extracted DNA or RNA divided by specific radioactivity of glucose; Step 2, calculation of μmole glucose incorporation into per μg DNA or RNA: the number calculated from step 1 is divided by total DNA or RNA; Step 3, calculation of μmole glucose incorporation into per μg DNA or RNA by using per μmole glucose: the number calculated from step 2 is divided by consumed glucose. Therefore, glucose incorporation into DNA or RNA is expressed by μmole glucose/μg DNA or RNA/per μmole glucose consumed.
Cell proliferation, glucose consumption and lactate generation
4T1, Bcap37, A549 and HeLa cells are maintained in RPMI-1640 (Invitrogen) supplemented with 10% fetal bovine serum (FBS), 100 U/ml penicillin/streptomycin and 2 mM L-glutamine (complete RPMI). Unless otherwise indicated, 1 × 106 cells are seeded into 25 cm2 culture flask (Corning) to allow attachment overnight in a humidified CO2 incubator. The culture media are then replaced with fresh complete RPMI plus 6 mM glucose. To examine the effects of lactate (0–25 mM) and pH (7.4–6.5), lactic acid is added to desired lactate concentrations and pH. The cell count is carried out with a hematocytometer under optical microscope, glucose level measured automatically by the HK colorimetric method using an Olympus AU2700 system and the lactate production determined by the VITROS Chemistry Product LAC Slides using the VITROS 5.1 FS system.
Glucose uptake
A million (1 × 106) cells per well in complete RPMI plus 6 mM glucose are seeded into 6-well plates and allowed to attach overnight. The culture media are then replaced with 2 ml fresh complete RPMI containing 0.2 μCi [3H]-2-deoxy-D-glucose (specific radioactivity 60 Ci/mmole, ART0200, American Radiolabeled Chemicals, Inc, St. Louis, MO, USA) and supplemented with or without 20 mM pure lactic acid. After incubation at 37°C in a humidified CO2 incubator for 1 hour, cells are washed 6 times with ice-cold PBS, trypsinized and collected for radioactivity count by a liquid scintillation counter (Wallac 1414, Finland).
Glycolysis assay in a cell free system
4T1 cells cultured in complete RPMI with 6 mM glucose and at 70% confluence are washed with ice-cold PBS twice and lysed with MPER supplemented with Halt™ protease inhibitor cocktail (both from Thermo Scientific). The resultant crude 4T1 cell lysates when reconstituted with a buffer (200 mM Hepes [pH 7.3], 5 mM MgCl2, 0.5 mM EDTA, 5 mM Na2HPO4, 50 mM KCl, 10 mM Glucose, 1.5 mM ATP, 1.5 mM ADP, 2 mM NAD and 0.2 mM NADH), empirically determined to be suitable for all 11 relevant enzymes, are able to fulfill an entire conversion of glucose to lactate in the cell free system. Unless otherwise specified, each reaction is initiated by adding 4T1 cell lysates to the above buffer in a total volume of 1 ml; because the volumes of added cell lysates were small as compared to total reaction volumes, the levels of endogenous glycolytic metabolites and cofactors are negligible.
For a time course experiment, reactions are initiated by adding 4T1 cell lysates to a final protein concentration at 250 μg/ml and terminated by 1 ml acetonitrile at 15 min intervals for 1 hr. For the titration experiments, the reactions are initiated by 4T1 cell lysates to final protein concentrations at 250, 500 or 1000 μg/ml and terminated by acetonitrile at 60 min. For testing sensitivity to pH, the buffer pH is adjusted to desired values; reactions are initiated by adding 4T1 cell lysates to a final protein concentration at 250 μg/ml and terminated at 1 hr by acetonitrile. To determine the effect of lactate on glycolysis, we add lactate to the buffer to desired levels and re-adjust the pH to 7.3 and reactions are initiated by adding 4T1 cell lysates to a final protein concentration at 1000 μg/ml and terminated at 1 hr by acetonitrile. In all cases, the control reactions contain boiled cell lysates with equal amounts of total proteins. Upon termination, samples are cleared by centrifugation and levels of in vitro generated lactate are determined using the HPLC method as described below.
Intracellular pH determination
Intracellular pH is determined according to previous reported methods30,31,32. To create a standard curve, we incubate half million (5 × 105) cells/well (in complete RPMI plus 6 mM glucose) in 6-well plates and let the cells to attach overnight. Cells are then pre-incubated with 2.5 μM BCECF-AM (Invitrogen) in PBS for 30 min at 37°C, washed twice with PBS, added with solution A (140 mM KCl, 1.2 mM MgSO4, 1.2 mM KH2 PO4, 11.7 mM D-glucose, 1.3 mM CaCl2, 10 mM HEPES, pH 6.0 to pH 8.0, 20 μM nigericin [Sigma]) and incubated at 37°C for 10 min. Afterwards, cells are observed under a Zeiss LSM710 laser confocal microscope (535 ± 15 nm emission signal excited by 488 or 405 nm excitation). A standard curve is created by plotting the ratio of emission signal excited by 488 nm over that by 405 nm against the above-defined range of buffer pHs. To measure intracellular pHs of cells exposed to different lactate concentrations and pH, cells are pre-incubated and washed as above, added with solution B (0.42 mM Ca(NO3)2, 0.41 mM MgSO4, 5.33 mM KCl, 17.86 mM NaHCO3, 103.45 mM NaCl, 5.63 mM Na2HPO4, 11.7 mM D-Glucose,) supplemented with lactic acid or HCl to desired lactate concentrations and pH and incubated for 1 hr at 37°C. Cells are then observed by a confocal microscope as above specified and the ratios of emission signals excited by 488 nm over that by 405 nm are compared with corresponding readings on the standard curve to obtain pHi values.
Determination of intracellular glucose, pyruvate, lactate and ATP levels
4T1 cells (6 × 105) are seeded and allowed to attach overnight. The media are replaced with fresh complete RPMI with 6 mM glucose, supplemented with lactic acid to desired lactate concentrations (0–25 mM) and pH (7.4–6.5). For time courses, cells are collected at 0, 5, 10, 20, 30, 60, 120, 240, 360 min; for titrations, cells are collected at 60 min.
We use a published method to extract cellular glucose33. In brief, 0.5 ml 1 M perchloric acid is employed to sequentially extract 5 flasks of 4T1 cells (total 107 cells). Then, the extracts are neutralized by 3 M K2CO3 and centrifuged (4°C, 20000 g); the glucose levels in the supernatants are measured by a standard enzymatic method34.
For intracellular pyruvate and lactate concentration determinations, cells are washed with ice-cold PBS 3 times swiftly, lysed with 500 μl acetonitrile and transferred to Eppendorf tubes for centrifugation (20,000 g/15 min/4°C). Then 40 μl of supernatants are subjected to quantitative analyses by HPLC: an ICSEP ICE-ION-300 Column (Transgenomic) is attached to a HEWLETT PACKARD series 1100 HPLC system maintained at 57°C. The mobile phase is 0.005 M H2SO4 and the flow rate at 0.4 ml/min. The peak UV traces (210 nm) corresponding to pyruvate (15.43 min) and lactate (22.72 min) are quantified by plotting against standard curves.
Intracellular concentrations of lactate, pyruvate are calculated based on measured mass values in 1 × 106 cells and converted to micromoles given that the average volume of a cell is 2.346 × 10−6 μl calculated by microscopically measuring diameters of >500 cells.
For cellular ATP quantitation, cells were washed as above, lysed with 500 μl of 0.5 M perchloric acid, neutralized with 2 M K2CO3 and the lysates centrifuged (20,000 g/15 min/4°C). The lysates were then neutralized with 1 M HCl and centrifuged as above. 50 μl of supernatants were subjected to HPLC analyses; a Sunchrom C18 column (250 × 4.6 mm, 5 μm particle size, 300 Å; The Great Eur-Asia Sci-Tech Development Co., Ltd, Beijing, China) was attached to a HEWLETT PACKARD series 1100 HPLC system that was pre-equilibrated in 20 mM triethylamine, pH 6.5 (adjusted with phosphoric acid) and 0.1% acetonitrile. After loading, the column was washed with starting solvent for 5 min and developed by an acetonitrile gradient at a flow rate of 0.5 ml/min. The peak UV traces (260 nm) were quantified by plotting against standard curves.
Seahorse metabolic flux analysis
Oxygen consumption rate (OCR) and extracellular acidification rate (ECAR) are measured at 37°C using XF96 extracellular analyzer (Seahorse biosciences). 4T1 cells are seeded in XF96 cell culture microplate at a density of 10,000 cells/well with normal culture medium and allowed to attach overnight. Prior to the assay, cells are prewashed 3 times with XF assay medium supplemented with 1 mM pyruvate and 12 mM glucose and then equilibrated in XF assay medium with or without lactic acidosis at 37°C in a non-CO2 incubator for 1 hour. All the medium is adjusted to pH 7.4 (control) or pH 6.7 (lactic acidosis) on the day of the assay. Three baseline measurements of OCR and ECAR were taken before sequential injection of mitochondrial inhibitors. The respiratory chain inhibitors are ATP synthase complex inhibitor oligomycin (1 μM), ATP synthesis uncoupler Carbonyl cyanide-p- trifluoromethoxyphenylhydrazone (FCCP, 1 μM), complex I inhibitor rotenone (5 μM) and complex III inhibitor antimycin A (1 μM).
Calculation of the free NAD/NADH ratio and mass action ratio Q
Cellular [pyruvate]/[lactate] ratios are inferred from the HPLC analyses (above) and the free NAD/NADH ratios are based on a reported method11 with modifications12. Briefly, cells are incubated in the presence of serial concentration of exogenous lactate. At equilibrium, [Lactate]eq[NAD]eq/[pyruvate]eq[NADH]eq[H+] = Keq, where Keq = 9.00 × 1010, or Apparent Keq = Keq [H+] = 9.00 × 103 at neutral pH35,36; the free NAD/NADH ratio is equal to apparent Keq[pyruvate]eq/[lactate]eq. The 0.13 mM pyruvate and 21.8 mM lactate levels in cells exposed to 20 mM lactic acid are at equilibrium (Results), thus the free NAD/NADH ratio is 53, which as argued in the same section does not significantly differ in diverse cell samples. The Q value is the free NAD/NADH ratio multiplied by the measured [lactate]/[pyruvate] ratio of each sample; thus, e.g., in cells exposed to 25 mM lactic acid, the Q is 53 × 25.0/0.125 = 10.6 × 103 (Fig. 2f), greater than the Apparent Keq (9.00 × 103) hence favoring a nonglycolytic phenotype (Results).
Calculation of the Gibbs free energy change (ΔG) of the conversion between pyruvate + NADH and lactate + NAD+
ΔG can be calculated according to the equation:  , where
, where  denotes the standard transformed Gibbs free energy at 37°C and Q = ([Lactate][NAD+])/([Pyruvate][NADH]). The question is that
denotes the standard transformed Gibbs free energy at 37°C and Q = ([Lactate][NAD+])/([Pyruvate][NADH]). The question is that  is not available.
is not available.
We calculate the  from
from  according to the equation:
according to the equation:

Since the change of ΔH and ΔS is negligible37,38 between 298 K (25°C) and 310 K (37°C), the equation can be expressed as:

Where  = −25.1 kJ/mol39,
= −25.1 kJ/mol39,  = −61.9 kJ/mol40,41. Thus
= −61.9 kJ/mol40,41. Thus  = −23.6 kJ/mol.
= −23.6 kJ/mol.
Activity assays on individual glycolytic enzymes
4T1 or Bcap37 cells maintained in complete RPMI with 6 mM glucose and at 70% confluence are washed and lysed as above-described. The cell lysates are then cleared by centrifugation (14,000 rpm/15 min/4°C) to avoid interference with the absorbance measurements (below) and supernatants collected. The activities of each of 11 enzymes are individually measured according to previously reported methods42,43,44,45,46,47,48,49,50,51,52. Briefly, a reaction is initiated by reconstituting the cleared cell lysate (2–20 μg protein) with the reaction buffer, reactant and, when appropriate, cofactor, in 1 ml. The amounts of the lysates used and the reaction times are carefully tested for each enzyme to maintain the linearity for each reaction. The absorbance at wavelength of 260 nm (PGAM, Enolase) or 340 nm (HK, PGI, PFK1, Aldolase, TPI, GAPDH, PGK, PK, LDH) are monitored at 25°C with a spectrophotometer (TU - 1901 Beijing Purkinje General Instrument Co., Ltd).
To determine the pH effect on enzyme activities, we use 50 mM PIPES (pKa 6.76) for pH 7.3–6.5 and 50 mM MES (pKa 6.09) for pH 6.5 to 6.0; this maximizes the buffering capacities at each pH range. The pH is adjusted to 7.3, 7.1, 6.9, 6.7, 6.5, 6.3 and 6.0 with 1 M KOH. To determine the effect of lactate anion on enzyme activities, we prepare 50 mM PIPES with serial levels of lactate and adjust pH to 7.3 with 1 M KOH.
Determination of the effect of lactic acidosis on glycolytic enzyme expression and activities
4T1 and Bcap37 cells (1 × 106) are seeded and allowed to attach overnight. The media are then replaced with fresh complete RPMI with 6 mM glucose, supplemented by lactic acid to a lactate level at 20 mM and pH at 6.7. Cells are collected at 0, 24, 48 and 72 hours.
For measuring the activities of individual glycolytic enzymes, whole cell lysates are prepared and the activity of each glycolytic enzyme assayed as above-described.
For immuno-blot assays, we employ a reported method53. Cells are lysed in 625 mM Tris-HCl (pH 6.8), 10% SDS, 25% glycerol, 5% β-mercaptoethanol and 0.015% bromphenol blue supplemented with a protease inhibitor cocktail (Pierce, Rockford, IL), followed by sonication. Samples are stored at −80°C after heating and protein concentrations scored by the BCA protein assay (Pierce, Rockford, IL) to guide equal loading of each sample to 6% to 15% SDS-polyacrylamide gels. Resolved proteins are transferred to PVDF membranes, probed with proper primary and secondary antibodies and visualized by the Western Lighting Plus ECL kit (Perkin Elmer, Waltham, MA). The film images are scanned and densitometry determined with the Quantity One Software (Bio-Rad). The antibodies against murine and human glycolytic enzymes are purchased from Cell Signaling Technology.
Inhibition of 4T1 tumor growth by sodium bicarbonate (NaHCO3)
The animal studies were performed in accordance with the regulations and guidelines of institutional animal ethical committee of Zhejiang University. The animal study was approved by the institutional animal ethical committee of Zhejiang University. Female BALB/c mice (18–22 g) are purchased from Shanghai SLAC Laboratory Animal Co., Ltd (Shanghai, China) and housed in a standard polypropylene cage containing sterile bedding under a controlled condition (temperature: 23 ± 2°C; humidity: 50 ± 5%) with an alternating 10/14 hr light/dark cycle in Zhejiang University Laboratory Animal Center (No. SYXK [Zhe] 2007-0029).
The animal model is based on a previous report54. Each mouse is inoculated with 5 × 104 viable 4T1 cells subcutaneously in the second right mammary fat pad area (day 0); mice are randomly assigned (day 1) to the NaHCO3 or the control group. Two independent experiments have been performed: in the first experiment, there are 5 mice for each group; in the second experiment, 10 mice for each group. Starting from day 7, tumor masses manifest and NaHCO3 (5%, 100 μl) or saline (0.9% sodium chloride for the control group, 100 μl) is injected subcutaneously surrounding the tumors every other day. Starting from day 15, the volume of injected NaHCO3 or saline is increased to 300 μl. The tumor volume is measured by vernier calipers twice or thrice every other day and calculated by an ellipse volume formula V = (L × W2) × 0.5, where L is length and W is width. On day 22 mice are executed by cervical dislocation and tumors weighed. The tumors in the first experiment are severed into 2 halves, which are photographed and the necrotic areas estimated with pixels of necrotic region by Photoshop. The intratumoral pH is measured by an InLab® Micro pH electrode (Mettler-Toledo, Swiss)at non-necrotic region. Tumor tissue is grounded under liquid nitrogen, extracted with 1 M HClO4, the extract centrifuged and the supernatant neutralized with 2 M K2CO3. The supernatant after another centrifugation is used for glucose and lactate level determinations by above-described methods.
Ethics statement
The animal study is approved by the institutional animal ethical committee of Zhejiang University with approval No. zju-2009-1-01-028. The methods were carried out in accordance to the approved guidelines.
Statistical analyses
Unless otherwise stated, each experiment is performed with at least three parallel samples and the data expressed as mean ± SD; regardless whether it requires statistics, each experiment is repeated twice or more to ensure reproducibility.
References
DeBerardinis, R. J., Lum, J. J., Hatzivassiliou, G. & Thompson, C. B. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab 7, 11–20 (2008).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674.
Hirayama, A. et al. Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res 69, 4918–4925 (2009).
Gallagher, F. A. et al. Magnetic resonance imaging of pH in vivo using hyperpolarized 13C-labelled bicarbonate. Nature 453, 940–943 (2008).
Gatenby, R. A. & Gillies, R. J. Why do cancers have high aerobic glycolysis? Nat Rev Cancer 4, 891–899 (2004).
Thistlethwaite, A. J., Leeper, D. B., Moylan, D. J., 3rd & Nerlinger, R. E. pH distribution in human tumors. Int J Radiat Oncol Biol Phys 11, 1647–1652 (1985).
van Sluis, R. et al. In vivo imaging of extracellular pH using 1H MRSI. Magn Reson Med 41, 743–750 (1999).
Wu, H. et al. Central role of lactic acidosis in cancer cell resistance to glucose deprivation-induced cell death. J Pathol 227, 189–199 (2012).
Williamson, D. H., Lund, P. & Krebs, H. A. The redox state of free nicotinamide-adenine dinucleotide in the cytoplasm and mitochondria of rat liver. Biochem J 103, 514–527 (1967).
Sun, F., Dai, C., Xie, J. & Hu, X. Biochemical issues in estimation of cytosolic free NAD/NADH ratio. PLoS One 7, e34525 (2012).
Poole, R. C. & Halestrap, A. P. Transport of lactate and other monocarboxylates across mammalian plasma membranes. Am J Physiol 264, C761–782 (1993).
Halestrap, A. P. & Price, N. T. The proton-linked monocarboxylate transporter (MCT) family: structure, function and regulation. Biochem J 343 Pt 2, 281–299 (1999).
Sonveaux, P. et al. Targeting lactate-fueled respiration selectively kills hypoxic tumor cells in mice. J Clin Invest 118, 3930–3942 (2008).
Broquist, H. P. Buthionine sulfoximine, an experimental tool to induce glutathione deficiency: elucidation of glutathione and ascorbate in their role as antioxidants. Nutr Rev 50, 110–111 (1992).
Fath, M. A., Ahmad, I. M., Smith, C. J., Spence, J. & Spitz, D. R. Enhancement of carboplatin-mediated lung cancer cell killing by simultaneous disruption of glutathione and thioredoxin metabolism. Clin Cancer Res 17, 6206–6217 (2011).
Miller, A. C., Gafner, J., Clark, E. P. & Samid, D. Posttranscriptional down-regulation of ras oncogene expression by inhibitors of cellular glutathione. Mol Cell Biol 13, 4416–4422 (1993).
Wardman, P. Fluorescent and luminescent probes for measurement of oxidative and nitrosative species in cells and tissues: progress, pitfalls and prospects. Free Radic Biol Med 43, 995–1022 (2007).
Vegran, F., Boidot, R., Michiels, C., Sonveaux, P. & Feron, O. Lactate influx through the endothelial cell monocarboxylate transporter MCT1 supports an NF-kappaB/IL-8 pathway that drives tumor angiogenesis. Cancer Res 71, 2550–2560 (2011).
Mi, L., Di Pasqua, A. J. & Chung, F. L. Proteins as binding targets of isothiocyanates in cancer prevention. Carcinogenesis 32, 1405–1413 (2011).
Chen, J. L. et al. The genomic analysis of lactic acidosis and acidosis response in human cancers. PLoS Genet 4, e1000293 (2008).
Chen, J. L. et al. Lactic acidosis triggers starvation response with paradoxical induction of TXNIP through MondoA. PLoS Genet 6, e1001093 (2010).
Dai, C., Sun, F., Zhu, C. & Hu, X. Tumor environmental factors glucose deprivation and lactic acidosis induce mitotic chromosomal instability--an implication in aneuploid human tumors. PLoS One 8, e63054 (2013).
Silva, A. S., Yunes, J. A., Gillies, R. J. & Gatenby, R. A. The potential role of systemic buffers in reducing intratumoral extracellular pH and acid-mediated invasion. Cancer Res 69, 2677–2684 (2009).
Trachootham, D., Alexandre, J. & Huang, P. Targeting cancer cells by ROS-mediated mechanisms: a radical therapeutic approach? Nat Rev Drug Discov 8, 579–591 (2009).
Semenza, G. L. Targeting HIF-1 for cancer therapy. Nat Rev Cancer 3, 721–732 (2003).
Manning, B. D. & Cantley, L. C. AKT/PKB signaling: navigating downstream. Cell 129, 1261–1274 (2007).
Vousden, K. H. & Ryan, K. M. p53 and metabolism. Nat Rev Cancer 9, 691–700 (2009).
Bergman, J. A., McAteer, J. A., Evan, A. P. & Soleimani, M. Use of the pH-sensitive dye BCECF to study pH regulation in cultured human kidney proximal tubule cells. Journal of tissue culture methods 13, 205–209 (1991).
Salvi, A., Quillan, J. M. & Sadee, W. Monitoring intracellular pH changes in response to osmotic stress and membrane transport activity using 5-chloromethylfluorescein. AAPS PharmSci 4, E21 (2002).
Thomas, J. A., Buchsbaum, R. N., Zimniak, A. & Racker, E. Intracellular pH measurements in Ehrlich ascites tumor cells utilizing spectroscopic probes generated in situ. Biochemistry (Mosc) 18, 2210–2218 (1979).
Singh, V. N., Singh, M., August, J. T. & Horecker, B. L. Alterations in glucose metabolism in chick-embryo cells transformed by Rous sarcoma virus: intracellular levels of glycolytic intermediates. Proc Natl Acad Sci U S A 71, 4129–4132 (1974).
Bergmeyer, H. U., Bergmeyer, J. & Grassl, M. Metabolites 1, Carbohydrates. 3rd edn, Vol. 6 (Verlag Chemie, 1984).
Bisswanger, H. Enzyme Kinetics: Principles and Methods. (Wiley-VCH, 2002).
Ward, L. D. & Winzor, D. J. Thermodynamic studies of the activation of rabbit muscle lactate dehydrogenase by phosphate. Biochem J 215, 685–691 (1983).
Alberty, R. A. Biochemical thermodynamics: applications of Mathematica. Methods Biochem Anal 48, 1–458 (2006).
Mendez, E. Biochemical thermodynamics under near physiological conditions. Biochem Mol Biol Educ 36, 116–119 (2008).
Nelson, D. L. & Cox, M. M. Lehninger principles of biochemistry. 4th edn, (Freeman, W.H. 2005).
Donnovan, L. K., Barclay, K. O. & Jespersen, N. Thermochemistry of the reaction catalyzed by lactate dehydrogenase. Thermochim Acta 11, 151–156 (1975).
Rekharsky, M. V., Egorov, A. M., Gal'chenko, G. L. & Berezin, I. V. Thermodynamics of redox reactions involving nicotinamide adenine dinucleotide. Thermochim Acta 46, 89–101 (1981).
Bueding, E. & Mackinnon, J. A. Studies of the phosphoglucose isomerase of Schistosoma mansoni. J Biol Chem 215, 507–513 (1955).
Cowgill, R. W. & Pizer, L. I. Purification and some properties of phosphorylglyceric acid mutase from rabbit skeletal muscle. J Biol Chem 223, 885–895 (1956).
Deng, H. et al. Phosphorylation of Bad at Thr-201 by JNK1 promotes glycolysis through activation of phosphofructokinase-1. J Biol Chem 283, 20754–20760 (2008).
Ferdinand, W. The isolation and specific activity of rabbit-muscle glyceraldehyde phosphate dehydrogenase. Biochem J 92, 578–585 (1964).
Gracy, R. W., Lacko, A. G. & Horecker, B. L. Subunit structure and chemical properties of rabbit liver aldolase. J Biol Chem 244, 3913–3919 (1969).
Ikeda, Y. & Noguchi, T. Allosteric regulation of pyruvate kinase M2 isozyme involves a cysteine residue in the intersubunit contact. J Biol Chem 273, 12227–12233 (1998).
Pagliaro, L., Kerr, K. & Taylor, D. L. Enolase exists in the fluid phase of cytoplasm in 3T3 cells. J Cell Sci 94 (Pt 2), 333–342 (1989).
Plaut, B. & Knowles, J. R. pH-dependence of the triose phosphate isomerase reaction. Biochem J 129, 311–320 (1972).
Scopes, R. K. Crystalline 3-phosphoglycerate kinase from skeletal muscle. Biochem J 113, 551–554 (1969).
Sekine, N. et al. Low lactate dehydrogenase and high mitochondrial glycerol phosphate dehydrogenase in pancreatic beta-cells. Potential role in nutrient sensing. J Biol Chem 269, 4895–4902 (1994).
Wilson, J. E. Brain hexokinase. A proposed relation between soluble-particulate distribution and activity in vivo. J Biol Chem 243, 3640–3647 (1968).
Han, W. et al. Shikonin circumvents cancer drug resistance by induction of a necroptotic death. Mol Cancer Ther 6, 1641–1649 (2007).
Heppner, G. H., Miller, F. R. & Shekhar, P. M. Nontransgenic models of breast cancer. Breast Cancer Res 2, 331–334 (2000).
Acknowledgements
This work has been supported in part by the China National 973 project (2013CB911303), China National 863 project (2007AA02Z143), China Natural Sciences Foundation projects (81272456) and the Fundamental Research Funds for the Central Universities, National Ministry of Education, China, to X Hu. We thank Prof. M.X. Guan, Zhejiang University College of Life Sciences, for allowing us to use the Seahorse XF96 extracellular analyzer for metabolic flux measurements. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
X.H. conceived the concept and designed the study; J.X., H.W., C.D., Q.P., Z.D. and D.H. carried out the experiments; X.H., J.X., H.W. analyzed the data and prepared the figures; B.J. assisted in experiment. X.H. and Y.L. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Xie, J., Wu, H., Dai, C. et al. Beyond Warburg effect – dual metabolic nature of cancer cells. Sci Rep 4, 4927 (2014). https://doi.org/10.1038/srep04927
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04927
This article is cited by
-
Novel cancer treatment paradigm targeting hypoxia-induced factor in conjunction with current therapies to overcome resistance
Journal of Experimental & Clinical Cancer Research (2023)
-
Tumor metabolism rewiring in epithelial ovarian cancer
Journal of Ovarian Research (2023)
-
Alkalization of cellular pH leads to cancer cell death by disrupting autophagy and mitochondrial function
Oncogene (2022)
-
Acetoacetate protects macrophages from lactic acidosis-induced mitochondrial dysfunction by metabolic reprograming
Nature Communications (2021)
-
Lactate dehydrogenases amplify reactive oxygen species in cancer cells in response to oxidative stimuli
Signal Transduction and Targeted Therapy (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.