Abstract
Whilst the development of membrane-active antibiotics is now an attractive therapeutic concept, progress in this area is disadvantaged by poor knowledge of the structure-activity relationship (SAR) required for optimizing molecules to selectively target bacteria. This prompted us to explore the SAR of the Lactobacillusreuteri membrane-active antibiotic reutericyclin, modifying three key positions about its tetramic acid core. The SAR revealed that lipophilic analogs were generally more active against Gram-positive pathogens, but introduction of polar and charged substituents diminished their activity. This was confirmed by cytometric assays showing that inactive compounds failed to dissipate the membrane potential. Radiolabeled substrate assays indicated that dissipation of the membrane potential by active reutericyclins correlated with inhibition of macromolecular synthesis in cells. However, compounds with good antibacterial activities also showed cytotoxicity against Vero cells and hemolytic activity. Although this study highlights the challenge of optimizing membrane-active antibiotics, it shows that by increasing antibacterial potency the selectivity index could be widened, allowing use of lower non-cytotoxic doses.
Similar content being viewed by others
Introduction
The widespread dissemination of bacterial pathogens exhibiting resistance to commonly used antibiotics continues to drive the discovery and development of novel treatments for bacterial infections1,2,3. Among the recently validated therapeutic strategies are antimicrobials that act on the bacterial cell membrane, such as the cyclic lipopeptide daptomycin and the lipoglycopeptide telavancin that were approved by the United States FDA in 2003 and 2011, respectively4. Other membrane-active agents are undergoing clinical trials such as XF-73 (a dicationic porphyrin) and the HT-61 (a quinolone-like compound) for nasal decolonization of MRSA4. Antibacterials that target the bacterial cytoplasmic membrane are collectively classified as membrane-active agents and typically demonstrate potent activities against bacteria, namely Gram-positive pathogens. In general, these membrane-active agents display a complex mode of action with multiple cellular effects that may arise from damage of the membrane's physical integrity, dissipation of components of the proton motive force [the transmembrane pH gradient (ΔpH) and the membrane potential (Δψ)] and reduction in ATP synthesis via inhibition of the respiratory chain4. Although membrane-active agents are becoming more attractive as therapeutics, there is still limited information on structure-activity relationships (SAR) to increase their selectivity for prokaryotic cells over mammalian cells. This prompted us to perform a pilot study as described herein, as any information gained could prove valuable in guiding the optimization of other membrane-active molecules to improve selectivity for pathogens. In part, we were inspired by the expanding knowledge that antimicrobial peptides (AMPs) can be systematically optimized by introducing amino acids that alter their overall charge, hydrophobicity and amphipathicity, yielding derivatives that are less toxic to mammalian cells5,6.
We sought to further expand the SAR of the membrane-active agent reutericyclin (1), a naturally occurring, low molecular weight tetramic acid antibiotic produced by sourdough isolates of Lactobacillus reuteri7. The production of reutericyclin in sourdough is thought to prevent stable colonization of competing Gram-positive Lactobacilli. Consequently, reutericyclin is only active against Gram-positive bacteria, including pathogens associated with topical infections such as methicillin-resistant Staphylococcus aureus (MRSA), Streptococcus sp. and Clostridium difficile8,9,10. Initial studies show that reutericyclin specifically dissipates the ΔpH of susceptible Lactobacillus spp.11, but owing to differences in membrane lipid composition and respiratory chain components between Lactobacilli and Staphylococci12, it is unclear if this action also accounts for reutericyclin's activity against S. aureus, which is a key target pathogen for these compounds.
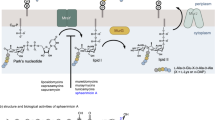
The structure of reutericyclin consists of a tetramic acid core flanked with a trans-2-decenoyl chain at the N-1 position, an acetyl group at the 3-position and an isobutyl group at the 5-position (Fig. 1A). An attractive feature of this structure is that it is readily amenable to advanced chemical optimization, thus providing straight-forward access to SAR information from which to develop novel antibiotics. To this end, we recently reported a series of N-alkyl reutericyclin analogs which retained activity against Gram-positive pathogens and treated an experimental staphylococcal skin infection in mice10,13. In continuation of our efforts to evaluate the therapeutic potential of reutericyclin, herein we report an expanded SAR performed by synthesizing new analogs carrying modifications at the 1, 3 and 5 positions of the tetramic core (Fig. 1A). This also provided new insights to the mode of action of reutericyclins against S. aureus and modifications that augment or hinder their activity and selectivity.
Studies on the membrane active agent reutericyclin.
A] Objectives of the study - 1] Perform structure-activity relationship (SAR) study by modifying the 1, 3 and 5-positions of the tetramic core 2] Determine selectivity index by measuring cytotoxicity against Vero cells and comparing to MIC against S. aureus Newman strain and 3] Determine the mechanism of action against S. aureus. B] Synthetic scheme for the preparation of the reutericyclin analogs.
Results
Synthesis of analogs
Reutericyclin analogs were synthesized using a variation of the procedure reported by Schobert et al.14 for the synthesis of reutericyclin (Fig. 1B). The initial steps of the synthesis allowed us to introduce variations at the 5-position (R1) of the tetramic core. Reaction of the appropriate amino acid benzyl ester with Bestmann ylide in presence of catalytic amounts of benzoic acid followed by debenzylation by catalytic hydrogenation afforded the basic tetramic core containing the R1 substituent. Next, substituents were introduced at the 3-position (R3) of the tetramic core by first esterifying the 4-O position with the appropriate acyl chloride (R3COCl) in the presence of triethylamine followed by the addition of acetone cyanohydrin to the reaction mixture which induced the 4-O to 3-C migration of the acyl group and afforded the 3-acylteteramic acids15. Finally, treatment of the 3-acylteteramic acids with sodium hexamethyldisilazanide and quenching with the desired acyl chloride (R2COCl) allowed for the introduction of the N-1 substituents (R2) and afforded the desired N-acylated 3-acyltetramic acids.
Effects of changes at the N-1 position (R2) on antibacterial activity
The synthesized compounds were tested against a panel of clinical relevant Gram-positive pathogens including, Enterococcus faecalis, Strep. pyogenes, Strep. pneumoniae, C. difficile and S. aureus (Table 1). To study the SAR at the N1-position (R2) of reutericyclin (1), we synthesized the N-unsubstituted analog 2 and acyl analogs 3-5. The N-unsubstituted analog 2, representing a simple tetramic acid core, was completely inactive against all bacteria in the panel (>200 μg/ml). This indicates that the tetramic acid core alone is insufficient for antibacterial activity and highlights the importance of the trans-2-decenoyl hydrophobic chain of reutericyclin to enable interaction with the bacterial membrane. The need for the hydrophobic substituent was further evident in 3 and 4. Replacement of reutericyclin's trans-2-decenoyl chain at the N1-position by the shorter valeroyl chain (3) led to a considerable loss (≥ 32-fold) in activity against most Gram-positive bacteria in the panel. This observation is in agreement with our previous study where the longer chain N-alkyl analogs were generally more potent than the shorter chain analogs13. As compared to 3, the cinnamoyl analog (4) which also has a short chain but contains a terminal phenyl showed slightly better activity throughout the panel (≤ 8-fold). This is likely due to the higher lipophilicity of 4 (logD = 2.61) as compared to 3 (logD = 2.05). Finally, replacement of the trans-2-decenoyl chain of reutericyclin by a 6-Phenylhexanoyl (5) produced a 4-fold improvement in activity against Strep. pneumoniae and MRSA N315 and a minor 2-fold improvement against Strep. pyogenes. In addition, 5 also displayed excellent activity against E. faecalis and C. difficile (<0.7 μg/ml) even though its activity was 3 to 4-fold less when compared to reutericyclin.
Effects of changes at the 3-acyl position (R3) on antibacterial activity
The 3-acyl group has a significant effect on the acidity of tetramic acids16,17. Accordingly, the importance of the 3-acetyl group of reutericyclin was established in our previous study in which analogs containing a 3-cyano group instead of 3-acetyl were inactive against bacteria. To expand the SAR at this position, compounds 6-13 were synthesized. Replacement of the methyl portion of the 3-acetyl of reutericyclin by the more aliphatic n-butyl (6) had nominal effect (± 2-fold) against most bacteria except for S. aureus and C. difficile where the compound's activity improved by >4-fold and >3-fold, respectively, over reutericyclin. A similar trend against the target pathogen S. aureus was observed when the methyl group was replaced with the larger and more lipophilic but aromatic phenyl group (7). We therefore subjected 7 to further studies. Replacement of its lipophilic phenyl group with a more polar aromatic 4-pyridine (8) caused a significant decrease in activity against the entire panel of Gram-positive pathogens. Similarly, the non-aromatic polar 4-piperidine (9) was also significantly less active. We speculate that 9 exists as a zwitterion in solution, leading to reduced interaction with the charged bacterial membrane and have a lower potential to dissipate the membrane potential. This is further supported by membrane depolarization studies described below. Furthermore, the Boc protected intermediate (i.e. 10) of 9 which prevents the ionization of the amine and also increases the lipophilicity of the molecule showed better activity. A hydroxy substituent at the 4-position of the phenyl group of 7 (in 12) also decreased potency against the entire panel of Gram-positive pathogens. This may be in part due to the electron donating effects of the 4-OH which decreases the acidity of the tetramic portion, potentially affecting its function as an ionophore. As expected, the protected 4-OMEM intermediate (11) which blocks the electron donating effect and also increases lipophilicity had comparatively better activity than 12. Similarly, replacement of the 4-OH of 12 by the isosteric, but electron withdrawing and more lipophilic 4-Fluorine (in 13) led to considerable improvement in activity that was comparable to 7. Thus the R3 substituents have considerable influence on the activity of reutericyclins. Lipophilic substituents tend to improve activity while polar substituents lead to loss of activity most likely due to a decrease in interaction with the membrane. In addition, ionizable amines and electron donating groups which respectively neutralize or decrease the acidity of the tetramic acid core, also leads to loss of activity.
Effects of changes at the 5-position (R1) on antibacterial activity
Reutericyclin contains a hydrophobic isobutyl group at the 5-position which contributes to the overall lipophilicity of the molecule. In order to assess the importance of 5-isobutyl we synthesized the unsubstituted analog (14) and found it to be 8 to 32-fold less active than reutericyclin. We have previously established that the isobutyl can be replaced by other lipophilic groups, namely isopropyl and benzyl, without affecting activity. In order to further explore the SAR at the 5-position, we synthesized analogs containing a polar substituent instead of lipophilic groups. Replacement of the isobutyl with substituents containing a polar group such as hydroxy (15), carboxylic acid (16), amine (17) or 4-phenol (19) substantially decreased activity across the entire test panel. The 5-substituents cannot directly affect the acidity of the tetramic core as the 3-substituents and hence the loss of activity is most likely the result of decreased interaction with the membrane. In case of 17, the loss of activity can also be attributed in part to its existence as a zwitterion, similar to 9. Masking of the polar groups improved the activity of these compounds. For example, the Boc protected amine intermediate (18) and t-Butyl protected 4-phenol intermediate (20) both exhibited better activity than their final products. Amongst all the 5-position analogs synthesized in this study, the protected phenol analog (20) was the most active, displaying >4-fold better anti-staphylococcal activity than reutericyclin; however it was not better than reutericyclin against other bacteria in the test panel.
Relationship between logD and MIC
A plot of Log10 MIC of compounds against S. aureus Newman versus their logD indicates that all active compounds (MIC ≤ 1 μg/ml against S. aureus Newman) in the series had higher logD values (Fig. S1, Supplementary data). However, a higher logD did not necessarily correlate with a lower MIC, as only a moderate correlation (R2 = 0.72) was observed between logD and MIC, suggesting that lipophilicity is not the only parameter contributing to the activity of the compounds. Thus the SAR of these compounds is complex and probably influenced by the interplay between the lipophilicity, solubility, polarity and charge of the molecules.
Activity of synthesized compounds against Gram-negative bacteria
All compounds were inactive against Gram-negative bacteria, indicating that the chemical modulations did not improve the spectrum of activity of reutericyclin (Table S1, Supplementary data). The Gram-negative test panel included Acinetobacter baumannii, Burkholderia cepacia, Escherichia coli (K12), Klebsiella pneumoniae, Proteus mirabilis, Proteus vulgaris, Pseudomonas aeruginosa and Stenotrophomonas maltophilia. This appeared to result from these molecules being substrates for efflux pumps, since an E. coli derivative of strain K12 carrying a deletion of TolC was susceptible to compounds showing activity against Gram-positive bacteria. The TolC protein is part of the AcrAB-TolC efflux system, which has a wide substrate profile18. Intrinsic resistance to reutericyclins in Gram-negatives therefore arises from efflux-mediated mechanisms.
Selectivity studies
One of the issues concerning the development of many membrane-active agents as antibacterial agents is selectivity for bacterial cells over mammalian cells, as these agents have the potential to disrupt human plasma membranes and induce related toxicity19. This is of particular concern for compounds containing the tetramic core as this nucleus is also found in several cytotoxic natural products17. The cytotoxicity of the synthesized compounds was determined against Vero kidney cells and results compared to their activities against S. aureus Newman (Table 2). In general compounds displaying good antimicrobial activity also displayed cytotoxic IC50 values against Vero cells. A strong correlation (R2 = 0.81) was observed between the cytotoxicity and MIC for compounds against S. aureus Newman (Fig. S2, Supplementary data). However, a cytotoxic IC50 did not necessarily translate into a poor selectivity index. For example, the cytotoxic IC50 of reutericyclin was 7.5 μg/ml, representing a low selectivity index of only 10-fold. Conversely, owing to the lower MIC of 7 against S. aureus Newman (Table 1), the selectivity index was more than 125, which is favorable for therapeutic development. Comparison of compounds phenol 19 with its t-Butyl protected intermediate 20, amine 17 with its Boc-protected intermediate 18 and compounds 15 and 16 provide much insight to the close correlation between antimicrobial activity and cytotoxicity (Table 2). For example, compound 19 was less active against S. aureus Newman (MIC = 50 μg/ml) and also had a high cytotoxic IC50 (95.3 μg/ml), whereas 20 displayed good antimicrobial activity (MIC <0.1 μg/ml), but was simultaneously more cytotoxic (11.5 μg/ml), but with a more favorable selectivity index (>115). A plot of Log10 cytotoxicity versus logD (Fig. S3, Supplementary data) displays a modest correlation (R2 = 0.42), at the most, between the two suggesting that the cytotoxicity is influenced by multiple physicochemical properties of the molecule similar to their MIC.
Hemolytic activity
The reutericyclin analogs are structurally similar to the amphipathic anionic surfactants which are known to cause hemolysis of mammalian erythrocytes. Hence, we determined the hemolytic activity of these compounds against defibrinated sheep blood. Compounds with low MICs against S. aureus Newman displayed low to moderate effective concentrations (EC50) against mammalian RBCs i.e. concentrations causing hemolysis. A strong correlation (R2 = 0.81) was found between the hemolytic activity and MIC for compounds against S. aureus Newman (Fig. S4, Supplementary data). However, as compared to the MIC values, the hemolytic EC50 values of the potent compounds were much higher thus providing a large therapeutic window e.g. compound 7 had MIC of <0.1 μg/ml and hemolytic EC50 ≈ 22.
The activity of compounds correlates with effects on membrane potential
In order to examine the effect of reutericyclins on the bacterial membrane, we determined their potential to collapse the proton motive force of S. aureus. Initially we used the fluorescent probe 3,5-dipropylthiacarbocyanine DiSC3(5), which responds to changes in the ΔpH and Δψ and was previously adopted to study the activity of reutericyclin against Lactobacillus spp.11 In our assays, DiSC3(5) was found to be an unsuitable probe as its florescence was quenched by reutericyclin and its analogs in the absence of cells. Therefore, we adopted the flow cytometric approach utilizing the diethyloxacarbocyanine dye DiOC2(3) to determine if reutericyclins altered the membrane potential of S. aureus. DiOC2(3) is known to accurately assess the membrane potential in S. aureus and other bacteria20,21,22, where a larger membrane potential causes a shift in fluorescence of DiOC2(3) from green to red. As expected 2 representing a simple tetramic acid core failed to affect the membrane potential of S. aureus, up to 100 μM, which supports the requirement of a hydrophobic moiety for interaction with the cytoplasmic membrane bilayer. Reutericyclin dissipated the membrane-potential in a concentration dependent manner, validating that the membrane is likely the antibacterial target in S. aureus (Fig. 2A). Indeed, the membrane potential was completely dissipated upon exposure to reutericyclin at 10 μM for 30 min, which was similar to the control proton ionophore CCCP at an equivalent molar concentration (Fig. 2B). Similarly, the highly active 13 also reduced the membrane potential in a manner similar to reutericyclin. Although 5 and 6 displayed equivalent and superior MICs to reutericyclin against S. aureus (Table 1) respectively, their dissipation of the membrane potential was slightly lower than reutericyclin (Fig. 2B). This might suggest that in spite of bearing structural similarity to reutericyclin, the substitutions may have altered the interaction of these compounds with the cytoplasmic membrane or these molecules display a marginally different mechanism of action. Compounds lacking good activity against S. aureus also failed to disrupt the membrane potential (Fig. 2B). For example, compounds 9 and 17 bearing an ionizable amine substitution which diminished the activity of reutericyclin also reduced their ability to dissipate the membrane potential. To examine if reutericyclins disrupted the physical integrity of the S. aureus membrane the uptake of the impermanent fluorescence dye propidium iodide which only stains DNA of cells with damaged membranes was used. Consistent with previous studies10, reutericyclin and other compounds (e.g. 5, 6 and 13) did not disrupt the physical integrity of the S. aureus membrane at concentrations causing dissipation of the membrane potential (Fig. S5, Supplementary data). Therefore, although the compounds lyse the membranes of erythrocytes, they do not similarly damage staphylococcal membranes.
Effects on membrane potential of S. aureus.
A] Reutericyclin (REU) acts in a concentration-dependent with full collapse of the membrane potential at 10 μM, similar to the control CCCP. B] Comparison of the ability of reutericyclin analogs at 10 μM to dissipate the membrane potential; complete dissipation of the membrane potential by CCCP (10 μM) is shown.
Effects on macromolecular synthesis
The compounds reutericyclin, 13 and 5 all reduced the biosynthesis of DNA, RNA and protein, with no process appearing more susceptible to inhibition (Fig. 3). This coincides with the membrane being the cellular target for reutericyclins, including compound 5 which failed to show comparable effects on the membrane potential as reutericyclin (Fig. 2B).
Effect of reutericyclins on the macromolecular synthesis of S. aureus.
The concentrations of compounds corresponding to 1 and 4 × their MICs are: 1 i.e. reutericyclin, (0.78 and 3.12 μg/ml), 5 (0.78 and 3.13 μg/ml), 13 (0.2 and 0.8 μg/ml). Controls were: novobiocin (8 μg/ml), rifampicin (0.08 μg/ml) and tetracycline (1.6 μg/ml).
Discussion
There is a need for novel antibiotics in order to combat the growing menace of multidrug resistant bacteria. Over the last decade, membrane active agents have gained considerable attention as antimicrobial agents, owing to clinical introduction of daptomycin, the low resistance potential for these agents and their potent activities against bacterial biofilms that are inherently refractory to most classes of antibiotics4. However, the therapeutic development of most membrane-active agents is complicated by selectivity problems and a lack of SAR information to address this issue. We therefore performed an SAR around the membrane-active agent reutericyclin by synthesizing a series of N-acyl analogs with modifications at three key positions in the tetramic core. The results revealed a complex SAR that is modulated by hydrophobicity, polarity and overall charge. As a general trend, more lipophilic analogs were more active which may be attributed to the ability of these compounds to achieve a higher partitioning coefficient across the bacterial cytoplasmic membrane. Polar substituents generally led to loss of activity most likely due to decreased interaction with the hydrophobic portions of the membrane. Substituents that altered the charge of the tetramic core either by decreasing the acidity or by formation of a zwitterion also led to a loss in activity, possibly due to increased electrostatic repulsion and/or loss of ionophore properties of reutericyclin13. To further rationalize these findings and understand how different substituents affect reutericyclins interaction with biological membranes, biophysical studies that adopt liposomes with varying membrane composition will be required23. Unfortunately, antibacterial activity generally correlated with cytotoxicity in Vero epithelial cells and hemolytic activity, demonstrating the challenge of identifying chemical functionalities to decrease cytotoxicity whilst increasing or retaining antibacterial activity. These challenges are similar to those experienced during the discovery process for non-cytotoxic antimicrobial peptides24, where several rounds of optimization are required to obtain derivatives with suitable physicochemical properties, such as amphipathicity, charge and hydrophobicity6. Therefore, from the small sample set produced in our studies it can be deduced that multiple rounds of synthesis or creation of combinatorial libraries will be required to discover non-cytotoxic membrane-active small molecules and provide information to optimize similar membrane-active molecules.
Noteworthy compounds include 6, 7 and 13 that displayed excellent activities against most of the tested Gram-positive pathogens (<1 μg/ml). Interestingly, despite increased antibacterial potency 6, unlike reutericyclin, did not rapidly dissipate the membrane potential. This broadly suggests that modifications to membrane-active agents could expand their mode of action within an environment that is as biologically complex as the bacterial cell envelope i.e. consisting of membrane lipids, embedded proteins and peptidoglycan structures. The higher selectivity indices for 6, 7 and 13, against S. aureus, might also suggest that membrane-active agents with elevated antibacterial activity could be utilized at a lower therapeutic dose, which is sufficient to eradicate bacteria, but avoids inducing cytotoxicity in mammalian cells. A similar concept referred to as dose sparing has recently been proposed by Farha et al.25, where combinations of two compounds specifically dissipating the Δψ and ΔpH are synergistic at concentrations below their individual MIC and are not cytotoxic at lower levels. Thus, lesser non-cytotoxic doses can be used to achieve antibacterial potency, whilst circumventing cytotoxicity. Compounds 6, 7 and 13 may therefore be better drug leads for treating staphylococcal skin infections, for which the reutericyclin class shows efficacy10. In light of the reutericyclins being highly serum bound10, which is a factor that circumvents hemolysis, topical application currently appears more amenable. Nevertheless, these studies point to the need for much chemical evolution, achievable through further medicinal chemistry efforts, to discover reutericyclins that are less cytotoxic and suitable for systemic use, which could have implications for other small unexplored membrane-active molecules.
Methods
Synthesis of reutericyclins
All reagents and solvents were purchased from commercial sources and were used without further purification. All reactions were performed under an inert atmosphere. The final reaction mixtures were purified by reverse phase flash chromatography using a Biotage Isolera Flash Purification System. The purity of the synthesized compounds were determined on a Waters ACQUITY UPLC-PDA-ELSD-MS system using a C18 reverse phase column and 0.1% formic acid/water - 0.1% formic acid/acetonitrile binary solvent system. All synthesized compounds were at least 95% pure. The structures of the synthesized compounds were confirmed by1H NMR which was recorded on a 400Mhz Varian AVANCE 400-FT NMR. The high resolution mass spectral (HRMS) analysis was performed on a Waters XEVO QTOF LCMS. A representative procedure for the synthesis of these analogs is provided in the supplementary data.
(Z)-1-((E)-dec-2-enoyl)-3-(1-hydroxyethylidene)-5-isobutylpyrrolidine-2,4-dione (1, Reutericyclin)
1H NMR (400 MHz, Acetonitrile-d3) δ 0.69–0.75 (m, 9H), 1.10–1.17 (m, 8H), 1.30 (q, J = 6.8 Hz, 2H), 1.58–1.62 (m, 3H), 2.10 (q, J = 7.1 Hz, 2H), 2.32 (s, 3H), 4.37 (t, J = 5.3 Hz, 1H), 6.89 (dt, J = 14.0, 6.9 Hz, 1H), 7.09 (d, J = 15.4 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C20H31NO4, 350.2325, found 350.2327.
3-acetyl-4-hydroxy-5-isobutyl-1H-pyrrol-2(5H)-one (2)
1H NMR (400 MHz, Methanol-d4) δ 0.97 (d, J = 3.3 Hz, 3H), 0.98 (d, J = 3.2 Hz, 3H), 1.45 (ddd, J = 14.1, 9.3, 5.2 Hz, 1H), 1.63 (ddd, J = 13.6, 9.1, 4.3 Hz, 1H), 1.81–1.88 (m, 1H), 2.44 (s, 3H), 3.93 (d, J = 6.8 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C10H15NO3, 198.113, found 198.1132.
(Z)-3-(1-hydroxyethylidene)-5-isobutyl-1-pentanoylpyrrolidine-2,4-dione (3)
1H NMR (400 MHz, Acetonitrile-d3)1H NMR (400 MHz, Acetonitrile-d3) δ 0.90 (d, J = 6.3 Hz, 3H), 0.93 (d, J = 6.3 Hz, 3H), 0.95(t, J = 7.3 Hz, 3H), 1.34–1.46 (m, 2H), 1.59–1.67 (m, 2H), 1.75–1.88 (m, 3H), 2.51 (s, 3H), 2.85–3.03 (m, 2H), 4.60 (br s, 1H). HRMS (ES+), m/z [M+H] calcd for C15H23NO4, 282.1705, found 282.1704.
(Z)-1-cinnamoyl-3-(1-hydroxyethylidene)-5-isobutylpyrrolidine-2,4-dione (4)
1H NMR (400 MHz, Methanol-d4) δ 0.93 (d, J = 6.2 Hz, 3H), 0.98 (d, J = 6.2 Hz, 3H), 1.85–1.90 (m, 3H), 2.57 (s, 3H), 4.65 (t, J = 5.0 Hz, 1H), 7.44 (dd, J = 5.0, 1.8 Hz, 3H), 7.63–7.71 (m, 2H), 7.80 (d, J = 15.8 Hz, 1H), 8.05 (d, J = 15.8 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C19H21NO4, 328.1549, found 328.1545.
(Z)-3-(1-hydroxyethylidene)-5-isobutyl-1-(6-phenylhexanoyl)pyrrolidine-2,4-dione (5)
1H NMR (400 MHz, Acetonitrile-d3) δ 0.76 (d, J = 6.4 Hz, 3H), 0.80 (d, J = 6.2 Hz, 3H), 1.23–1.32 (p, J = 7.7, 7.2 Hz, 2H), 1.48–1.60 (m, 4H), 1.64 (t, J = 6.4 Hz, 2H), 1.67–1.76 (m, 1H), 2.38 (s, 3H), 2.51 (t, J = 7.7 Hz, 2H), 2.71–2.91 (m, 2H), 4.36 (t, J = 5.8 Hz, 1H), 7.03–7.12 (m, 3H), 7.17 (t, J = 7.6 Hz, 2H). HRMS (ES+), m/z [M+H] calcd for C22H29NO4, 372.2175, found 372.2175.
(Z)-1-((E)-dec-2-enoyl)-3-(1-hydroxypentylidene)-5-isobutylpyrrolidine-2,4-dione (6)
1H NMR (400 MHz, Acetonitrile-d3) δ 0.65–0.75 (m, 12H), 1.05–1.14 (m, 8H), 1.16–1.24 (m, 2H), 1.25–1.36 (m, 2H), 1.44 (p, J = 7.5 Hz, 2H), 1.59 (dd, J = 5.3, 2.6 Hz, 3H), 1.74 (p, J = 2.5 Hz, 1H), 2.02–2.14 (m, 2H), 2.66–2.75 (m, 2H), 4.30–4.41 (m, 1H), 6.82–6.93 (m, 1H), 7.05 (dt, J = 15.4, 1.4 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C23H37NO4, 392.2801, found 392.2798.
(Z)-1-((E)-dec-2-enoyl)-3-(hydroxy(phenyl)methylene)-5-isobutylpyrrolidine-2,4-dione (7)
1H NMR (400 MHz, Acetonitrile-d3) δ 0.74–0.83 (m, 6H), 0.85 (d, J = 6.3 Hz, 3H), 1.14–1.30 (m, 8H), 1.39 (p, J = 7.1 Hz, 2H), 1.71–1.77 (m, 2H), 1.85 (p, J = 2.5 Hz, 1H), 2.18 (q, J = 7.9, 7.4 Hz, 2H), 4.51 (dd, J = 7.0, 3.8 Hz, 1H), 6.94–7.04 (m, 1H), 7.15 (dt, J = 15.4, 1.4 Hz, 1H), 7.41–7.49 (m, 2H), 7.59 (tt, J = 7.0, 1.2 Hz, 1H), 7.92 (dt, J = 8.5, 1.5 Hz, 2H). HRMS (ES+), m/z [M+H] calcd for C25H33NO4, 412.2488, found 412.248.
(Z)-1-((E)-dec-2-enoyl)-3-(hydroxy(pyridin-4-yl)methylene)-5-isobutylpyrrolidine-2,4-dione (8)
1H NMR (400 MHz, Methanol-d4) δ 0.88–0.96 (m, 9H), 1.32–1.40 (m, 8H), 1.50 (q, J = 7.0 Hz, 3H), 1.74–1.94 (m, 3H), 2.25 (p, J = 7.9, 7.5 Hz, 3H), 4.29 (dd, J = 6.7, 3.1 Hz, 1H), 6.93–7.08 (m, 1H), 7.41 (dt, J = 15.4, 1.4 Hz, 1H), 7.99 (d, J = 6.6 Hz, 2H), 8.85 (d, J = 6.6 Hz, 2H). HRMS (ES+), m/z [M+H] calcd for C24H32N2O4, 413.244, found 413.2433.
(Z)-1-((E)-dec-2-enoyl)-3-(hydroxy(piperidin-4-yl)methylene)-5-isobutylpyrrolidine-2,4-dione (9)
1H NMR (400 MHz, Methanol-d4) δ 0.86–0.93 (m, 9H), 1.32–1.37 (m, 10H), 1.44–1.58 (m, 3H), 1.71–1.81 (m, 2H), 1.83–2.06 (m, 4H), 2.25 (dq, J = 14.5, 7.2 Hz, 2H), 3.10 (t, J = 10.5 Hz, 2H), 3.48 (dq, J = 11.9, 3.7 Hz, 2H), 3.74 (tt, J = 9.6, 3.9 Hz, 1H), 4.20 (dd, J = 6.3, 2.9 Hz, 1H), 6.97 (m, 1H), 7.49 (dd, J = 15.6, 1.8 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C24H38N2O4, 419.291, found 419.2901.
tert-butyl4-((Z)-((R)-1-((E)-dec-2-enoyl)-5-isobutyl-2,4-dioxopyrrolidin-3-ylidene)(hydroxy)methyl) piperidine-1-carboxylate (10)
1H NMR (400 MHz, Methanol-d4) δ 0.89–0.98 (m, 9H), 1.32–1.41 (m, 10H), 1.49 (s, 9H), 1.50–1.59 (m, 3H), 1.59–1.69 (m, 2H), 1.81–1.90 (m, 4H), 2.32 (q, J = 6.9 Hz, 2H), 2.91 (br s, 1H), 3.66 (tt, J = 11.6, 3.6 Hz, 1H), 4.17 (d, J = 13.3 Hz, 1H), 4.69 (t, J = 5.2 Hz, 1H), 7.14 (ddt, J = 23.4, 15.3, 6.9 Hz, 1H), 7.34 (dt, J = 15.4, 1.4 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C29H45N2O6Na, 541.3254, found 541.3246.
(Z)-1-((E)-dec-2-enoyl)-3-(hydroxy(4-((2-methoxyethoxy)methoxy)phenyl)methylene)-5-isobutylpyrrolidine-2,4-dione (11)
1H NMR (400 MHz, Chloroform-d) δ 0.80–1.07 (m, 9H), 1.24–1.32 (m, 8H), 1.52 (br s, 2H), 1.89 (br s, 3H), 2.31 (br s, 2H), 3.40 (s, 3H), 3.53–3.61 (m, 2H), 3.81–3.90 (m, 2H), 4.63 (d, J = 109.1 Hz, 1H), 5.39 (s, 2H), 7.17 (d, J = 8.8 Hz, 3H), 7.33 (d, J = 17.5 Hz, 1H), 8.29 (d, J = 15.8 Hz, 2H). HRMS (ES+), m/z [M+H] calcd for C29H41NO7, 516.2961, found 516.2961.
(Z)-1-((E)-dec-2-enoyl)-3-(hydroxy(4-hydroxyphenyl)methylene)-5-isobutylpyrrolidine-2,4-dione (12)
1H NMR (400 MHz, Methanol-d4) δ 0.89–1.01 (m, 9H), 1.30–1.40 (m, 8H), 1.53 (q, J = 7.2 Hz, 2H), 1.88 (td, J = 15.5, 13.6, 5.6 Hz, 3H), 2.31 (d, J = 7.3 Hz, 2H), 4.55–4.66 (m, 1H), 6.90 (d, J = 8.9 Hz, 2H), 7.05–7.18 (m, 1H), 7.34 (d, J = 15.4 Hz, 1H), 8.22 (d, J = 8.9 Hz, 2H). HRMS (ES+), m/z [M+H] calcd for C25H33NO5, 428.2437, found 428.2433.
(Z)-1-((E)-dec-2-enoyl)-3-((4-fluorophenyl)(hydroxy)methylene)-5-isobutylpyrrolidine-2,4-dione (13)
1H NMR (400 MHz, Acetonitrile-d3) δ 0.70–0.77 (m, 6H), 0.79 (d, J = 6.1 Hz, 3H), 1.10–1.21 (m, 8H), 1.34 (p, J = 7.3 Hz, 2H), 1.66–1.77 (m, 3H), 2.13 (q, J = 6.8 Hz, 2H), 4.40–4.54 (m, 1H), 6.89–6.99 (m, 1H), 7.05–7.17 (m, 3H), 7.96–8.07 (m, 2H). HRMS (ES+), m/z [M+H] calcd for C25H32FNO4, 430.2393, found 430.2401.
(Z)-1-((E)-dec-2-enoyl)-3-(1-hydroxyethylidene)pyrrolidine-2,4-dione (14)
1H NMR (400 MHz, Methanol-d4) δ 0.93 (t, J = 6.2 Hz, 3H), 1.25–1.42 (m, 8H), 1.54 (p, J = 6.3, 5.8 Hz, 2H), 2.32 (q, J = 7.1 Hz, 2H), 2.54 (s, 3H), 4.24 (s, 2H), 7.13 (dt, J = 14.0, 6.9 Hz, 1H), 7.41 (dd, J = 15.4, 1.5 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C16H23NO4, 294.1705, found 294.1698.
(Z)-1-((E)-dec-2-enoyl)-3-(1-hydroxyethylidene)-5-(hydroxymethyl)pyrrolidine-2,4-dione (15)
1H NMR (400 MHz, Methanol-d4) δ 0.92 (t, J = 6.9 Hz, 3H), 1.30–1.41 (m, 8H), 1.54 (p, J = 7.0 Hz, 2H), 2.32 (q, J = 6.6, 6.2 Hz, 2H), 2.55 (s, 3H), 3.99 (dd, J = 11.6, 1.7 Hz, 1H), 4.21 (dd, J = 11.6, 2.8 Hz, 1H), 4.51 (s, 1H), 7.12 (dt, J = 15.3, 6.9 Hz, 1H), 7.43 (dt, J = 15.3, 1.3 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C17H25NO5, 324.1811, found 324.1803.
3-((Z)-1-((E)-dec-2-enoyl)-4-(1-hydroxyethylidene)-3,5-dioxopyrrolidin-2-yl)propanoic acid (16)
1H NMR (400 MHz, Methanol-d4) δ 0.92 (t, J = 6.9 Hz, 3H), 1.30–1.41 (m, 8H), 1.54 (p, J = 7.1 Hz, 2H), 2.23–2.39 (m, 6H), 2.56 (s, 3H), 4.61–4.68 (m, 1H), 7.07–7.17 (m, 1H), 7.35 (d, J = 15.4 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C19H27NO6, 366.1917, found 366.1916.
(Z)-5-(4-aminobutyl)-1-((E)-dec-2-enoyl)-3-(1-hydroxyethylidene)pyrrolidine-2,4-dione (17)
1H NMR (400 MHz, Methanol-d4) δ 0.92 (t, J = 6.7 Hz, 3H), 1.20–1.30 (m, 2H), 1.30–1.42 (m, 8H), 1.51 (p, J = 7.2 Hz, 2H), 1.62 (p, J = 7.7 Hz, 2H), 1.87–1.98 (m, 1H), 2.11 (tt, J = 11.0, 5.7 Hz, 1H), 2.27 (q, J = 7.0 Hz, 2H), 2.35 (s, 3H), 2.89 (t, J = 7.5 Hz, 2H), 4.18 (dd, J = 5.0, 2.7 Hz, 1H), 6.98 (dt, J = 15.0, 6.9 Hz, 1H), 7.50 (d, J = 15.4 Hz, 1H). HRMS (ES+), m/z [M+H] calcd for C20H32N2O4, 365.244, found 365.244.
tert-butyl(4-((Z)-1-((E)-dec-2-enoyl)-4-(1-hydroxyethylidene)-3,5-dioxopyrrolidin-2-yl)butyl) carbamate (18)
1H NMR (400 MHz, Chloroform-d) δ 0.88 (t, J = 6.7 Hz, 3H), 1.12–1.23 (m, 1H), 1.23–1.40 (m, 10H), 1.42 (s, 9H), 1.45–1.60 (m, 4H), 1.94–2.13 (m, 2H), 2.24–2.36 (m, 2H), 2.56 (s, 2H), 3.07 (q, J = 6.7 Hz, 2H), 4.52 (s, 1H), 7.11–7.21 (m, 1H), 7.30 (s, 1H). HRMS (ES+), m/z [M+H] calcd for C25H39N2O6Na, 487.2784, found 487.2779.
(Z)-1-((E)-dec-2-enoyl)-5-(4-hydroxybenzyl)-3-(1-hydroxyethylidene)pyrrolidine-2,4-dione (19)
1H NMR (400 MHz, Chloroform-d) δ 0.90 (t, J = 6.6 Hz, 3H), 1.23–1.39 (m, 8H), 1.51 (p, J = 7.3 Hz, 2H), 2.30 (td, J = 7.3, 3.7 Hz, 2H), 2.41 (s, 3H), 3.23 (dd, J = 14.1, 2.7 Hz, 1H), 3.43 (dd, J = 14.3, 5.6 Hz, 1H), 4.78 (br s, 1H), 6.67 (d, J = 8.3 Hz, 2H), 6.84 (d, J = 8.3 Hz, 2H), 7.16–7.27 (m, 2H). HRMS (ES+), m/z [M+H] calcd for C23H29NO5, 400.2124, found 400.2122.
(Z)-5-(4-(tert-butoxy)benzyl)-1-((E)-dec-2-enoyl)-3-(1-hydroxyethylidene)pyrrolidine-2,4-dione (20)
1H NMR (400 MHz, Chloroform-d) δ 0.69 (t, J = 6.8 Hz, 3H), 1.08 (s, 9H), 1.12 (ddd, J = 14.0, 7.0, 2.9 Hz, 8H), 1.32 (p, J = 7.3 Hz, 2H), 2.04–2.13 (m, 2H), 2.17 (br s, 3H), 3.03 (dd, J = 14.0, 2.7 Hz, 1H), 3.28 (d, J = 8.6 Hz, 1H), 4.57 (d, J = 83.7 Hz, 1H), 6.59–6.68 (m, 4H), 6.94–7.06 (m, 2H). HRMS (ES+), m/z [M+H] calcd for C27H37NO5, 456.275, found 456.2759.
Compound preparation
Stocks of reutericyclin and test compounds were dissolved in DMSO at a concentration of 10 mg/ml. From this stock dilutions were made in the biological media of respective experiments described below.
Determination of minimum inhibitory concentrations (MICs)
The MICs of compounds were determined in Mueller-Hinton broth according to the Clinical Laboratory Standards Institute (CLSI), using the microbroth dilution method in 96-well round bottom plates. The final test concentration of compounds ranged from 200-0.1 μg/ml and after incubation for 18–20 h the MICs were recorded as the lowest concentration of test compound inhibiting visual growth13. Microbroth dilution MICs against C. difficile were described elsewhere26, save for the use of Brain-heart infusion broth containing 0.1% w/v of L-cysteine, a compound concentration of 64-0.03 μg/ml and incubation of plates for 24 h under anaerobic conditions in a DG250 Anaerobic Chamber (Don Whitley, UK).
Cytotoxicity
The in vitro cytotoxicity of compounds was evaluated against Vero cells (kidney epithelial cells; ATCC CCL-81) as described previously27. The final test concentration of compounds ranged from 200-0.1 μg/ml and was prepared in Dulbecco's Modified Eagle's Medium (Hyclone). After exposing cells to compounds for 72 h, cell viability was determined using CellTiter-Glo® Luminescent Cell Viability assay (Promega). The concentration of compounds causing 50% decrease in viability (IC50 values) was determined using GraphPad prism version 6 [GraphPad Software]. Selectivity indices were calculated as the Cytotoxic IC50 divided by the MIC of compound against S. aureus Newman.
Hemolysis assay
Defibrinated sheep blood was purchased from Colorado Serum Company. The blood was centrifuged at 350 rcf for 20 min and the buffy coat and plasma layers discarded. Erythrocytes were washed three times in sterile phosphate-buffered solution (PBS), pH 7.4 and then diluted to a final v/v concentration of 5% in PBS. 100 μl of washed erythrocytes were added to two fold serial dilutions of test compound or positive control Triton X 100 (100 μl, prepared in PBS) in 96-well, round bottom plates. After incubating at 37°C for 90 min, the plates were centrifuged and the supernatant transferred to a white wall 96-well plate taking care not to disturb the unlysed cells. Absorbance of supernatants was then read at an optical density (OD) of 540 nm. Percent hemolysis was calculated using the formula; Percent lysis = (OD540 of sample – OD540 of blank)/(OD540 of positive control). The EC50 values were determined using GraphPad prism version 6 [GraphPad Software].
Analysis of the membrane potential and permeability
The effects of compounds on the membrane potential of S. aureus was assayed by flow cytometry to access the ratio of red to green fluorescence of cells stained with the fluorescent probe diethyloxacarbocyanine dye DiOC2(3). The amount of red fluorescence emitted is dependent on the membrane potential, whereas the emitted green fluorescence is independent on the membrane potential. These experiments were performed as previously described28. Cultures of S. aureus Newman were grown to the exponential phase (OD600nm ≈ 0.3) in Mueller-Hinton II broth and compounds were added at 100, 10 or 1 μM. Cultures were exposed to compounds for 30 min at 37°C and then stained for 5 min with 30 μM of DiOC2(3). Samples were then analyzed using BD LSR II flow cytometer, with collection of 10,000 events for each sample ran at a low flow rate and acquisition of signals in the logarithmic scale. DiOC2(3) was excited using the 488-nm excitation laser and its green fluorescence emission was detected using FITC filters and its red fluorescence was detected using PI-A filters. The effects of compounds on the membrane potential were analyzed in the software FlowJo X 10.0.7 as follows: a parameter for the ratio of red-to-green fluorescence for each sample was applied and the histograms comparatively analyzed on the same logarithmic scale. The control carbonyl cyanide m-chlorophenyl hydrazone (CCCP; Sigma-Aldrich), which completely dissipates the membrane potential was used as a positive control. A minimum of two independent samples were evaluated. Membrane permeability was also evaluated by flow cytometry using the membrane impermanent dye propidium iodide (PI). Cells of S. aureus Newman were grown to OD600nm ≈ 0.3 as above and compound added at 10 μM for 30 min. PI was then added at 10 μM and incubated in the dark for 10 min before analyzing samples with BD LSR II flow cytometer, with collection of 10,000 events at a low flow rate and acquisition of signals in the logarithmic scale. Red fluorescence was detected using PI-A filters, with excitation at 488-nm. Histogram plots of number of events against PI fluorescence of the population were comparatively analyzed using FlowJo X 10.0.7. Thioridazine at 100 μg/mL which affects cell envelope integrity was used as a positive control.
Macromolecular synthesis
The effects of reutericyclin and two other active compounds, bearing significant modifications, (5 and 13) on macromolecular synthesis in S. aureus Newman were evaluated as previously described29,30. The following radiolabelled precursors from American Radiolabeled Chemicals, Inc were adopted: methyl-3H]thymidine, [5,6-3H]uridine and [4,5-3H]leucine for DNA, RNA and protein respectively. Three independent cultures were grown to mid-logarithmic phase (OD600nm ≈ 0.3), before the addition of radiolabelled precursors at 1 μCi/mL. After 5 min compounds were added at 1 and 4 × their MIC against S. aureus Newman and cells incubated for 20 min at 37°C. Cells were then lysed with cold 10% TCA and the precipitated macromolecules collected on GF/C filters, washed with 95% ethanol and analyzed by liquid scintillation counting. The relative percent activity in cells exposed to compounds was determined. The antibiotics novobiocin (DNA; from Sigma-Aldrich), rifampicin (RNA; from TCI America) and tetracycline (protein; from Calbiochem) were used as positive controls.
References
Theuretzbacher, U. Global antibacterial resistance: The never-ending story. J. Global Antibiot. Resist. 1, 63–69 (2013).
Bush, K. et al. Tackling antibiotic resistance. Nat. Rev. Microbiol. 9, 894–896 (2011).
Boucher, H. W. et al. 10 x '20 Progress-development of new drugs active against gram-negative bacilli: an update from the Infectious Diseases Society of America. Clin. Infect. Dis. 56, 1685–1694 (2013).
Hurdle, J. G., O'Neill, A. J., Chopra, I. & Lee, R. E. Targeting bacterial membrane function: an underexploited mechanism for treating persistent infections. Nat. Rev. Microbiol. 9, 62–75 (2011).
Zhu, X. et al. Design of imperfectly amphipathic alpha-helical antimicrobial peptides with enhanced cell selectivity. Acta Biomater. 10, 244–257 (2014).
Fjell, C. D., Hiss, J. A., Hancock, R. E. & Schneider, G. Designing antimicrobial peptides: form follows function. Nat. Rev. Drug Discov. 11, 37–51 (2012).
Holtzel, A., Ganzle, M. G., Nicholson, G. J., Hammes, W. P. & Jung, G. The first low molecular weight antibiotic from lactic acid bacteria: reutericyclin, a new tetramic acid. Angew. Chem. Int. Edit. 39, 2766–2768 (2000).
Ganzle, M. G., Holtzel, A., Walter, J., Jung, G. & Hammes, W. P. Characterization of reutericyclin produced by Lactobacillusreuteri LTH2584. Appl. Environ. Microbiol. 66, 4325–4333 (2000).
Hurdle, J. G., Heathcott, A., Yang, L., Yan, B. & Lee, R. E. Reutericyclin and related analogues kill stationary phase Clostridiumdifficile at achievable colonic concentrations. J. Antimicrob. Chemother. 66, 1773–1776 (2011).
Hurdle, J. G., Yendapally, R., Sun, D. & Lee, R. E. Evaluation of analogs of reutericyclin as prospective candidates for treatment of staphylococcal skin infections. Antimicrob. Agents Chemother. 53, 4028–4031 (2009).
Ganzle, M. G. & Vogel, R. F. Studies on the mode of action of reutericyclin. Appl. Environ. Microbiol. 69, 1305–1307 (2003).
Kaneda, T. Iso- and anteiso-fatty acids in bacteria: biosynthesis, function and taxonomic significance. Microbiol. Rev. 55, 288–302 (1991).
Yendapally, R., Hurdle, J. G., Carson, E. I., Lee, R. B. & Lee, R. E. N-substituted 3-acetyltetramic acid derivatives as antibacterial agents. J. Med. Chem. 51, 1487–1491 (2008).
Schobert, R., Dietrich, M., Mullen, G. & Urbina-Gonzalez, J.-M. Phosphorus ylide based functionalizations of tetronic and tetramic acids. Synthesis 3902–3914 (2006).
Abe, M., Imai, T., Ishii, N. & Usui, M. Synthesis of quinolactacide via an acyl migration reaction and dehydrogenation with manganese dioxide and its insecticidal activities. Biosci. Biotechnol. Biochem. 70, 303–306 (2006).
Stickings, C. E. Studies in the biochemistry of micro-organisms. 106. Metabolites of Alternaria tenuis auct.: the structure of tenuazonic acid. Biochem. J. 72, 332–340 (1959).
Royles, B. J. L. Naturally-occurring tetramic acids - structure, isolation and synthesis. Chem. Rev. 95, 1981–2001 (1995).
Piddock, L. J. Multidrug-resistance efflux pumps - not just for resistance. Nat. Rev. Microbiol. 4, 629–636 (2006).
Wu, X. & Hurdle, J. G. Screening for a diamond in the rough. Chem. Biol. 20, 1091–1092 (2013).
Novo, D., Perlmutter, N. G., Hunt, R. H. & Shapiro, H. M. Accurate flow cytometric membrane potential measurement in bacteria using diethyloxacarbocyanine and a ratiometric technique. Cytometry 35, 55–63 (1999).
Novo, D. J., Perlmutter, N. G., Hunt, R. H. & Shapiro, H. M. Multiparameter flow cytometric analysis of antibiotic effects on membrane potential, membrane permeability and bacterial counts of Staphylococcus aureus and Micrococcus luteus. Antimicrob. Agents Chemother. 44, 827–834 (2000).
Shapiro, H. M. Membrane potential estimation by flow cytometry. Methods 21, 271–279 (2000).
Peetla, C., Stine, A. & Labhasetwar, V. Biophysical interactions with model lipid membranes: applications in drug discovery and drug delivery. Mol. Pharm. 6, 1264–1276 (2009).
Takahashi, D., Shukla, S. K., Prakash, O. & Zhang, G. Structural determinants of host defense peptides for antimicrobial activity and target cell selectivity. Biochimie 92, 1236–1241 (2010).
Farha, M. A., Verschoor, C. P., Bowdish, D. & Brown, E. D. Collapsing the proton motive force to identify synergistic combinations against Staphylococcus aureus. Chem. Biol. 20, 1168–1178 (2013).
Wu, X. et al. Prospects for flavonoid and related phytochemicals as nature-inspired treatments for Clostridium difficile infection. J. Appl. Microbiol. 116, 23–31 (2014).
Rakesh et al. Antitubercular nitrofuran isoxazolines with improved pharmacokinetic properties. Bioorg. Med. Chem. 20 (2012).
Silverman, J. A., Perlmutter, N. G. & Shapiro, H. M. Correlation of daptomycin bactericidal activity and membrane depolarization in Staphylococcusaureus. Antimicrob. Agents Chemother. 47, 2538–2544 (2003).
Oliva, B. et al. Biological properties of novel antistaphylococcal quinoline-indole agents. Antimicrob. Agents Chemother. 47, 458–466 (2003).
Sun, D. et al. Evaluation of flavonoid and resveratrol chemical libraries reveals abyssinone II as a promising antibacterial lead. ChemMedChem 7, 1541–1545 (2012).
Acknowledgements
This work was funded by Grant 5R01AT006732 from the National Center for Complementary and Alternative Medicine at the National Institutes of Health, with additional funds from the American Lebanese Syrian Associated Charities (ALSAC), St Jude Children's Research Hospital. We wish to thank Dr. Jerrod Scarborough and the St. Jude High-Throughput Analytical Chemistry (HTAC) core for performing the HRMS analysis of the synthesized compounds.
Author information
Authors and Affiliations
Contributions
R.E.L. and J.G.H. conceived, designed and supervised the study. P.T.C. designed and synthesized the compounds. X.W. conducted MIC tests in C. difficile, macromolecular tests and FACS. M.M.M. performed MIC testing against all other organisms and A.P.S. performed cytotoxicity and hemolysis assays. All authors discussed the results. The manuscript was written by P.T.C., R.E.L. and J.G.H.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Cherian, P., Wu, X., Maddox, M. et al. Chemical Modulation of the Biological Activity of Reutericyclin: a Membrane-Active Antibiotic from Lactobacillusreuteri. Sci Rep 4, 4721 (2014). https://doi.org/10.1038/srep04721
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04721
This article is cited by
-
Design, synthesis and microbiological evaluation of ampicillin–tetramic acid hybrid antibiotics
The Journal of Antibiotics (2017)
-
In vitro toxicity determination of antifungal constituents from Combretum zeyheri
BMC Complementary and Alternative Medicine (2016)
-
The biofilm inhibitor Carolacton inhibits planktonic growth of virulent pneumococci via a conserved target
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.