Abstract
Although apoptosis and necrosis have distinct features, the identification and discrimination of apoptotic and necrotic cell death in vitro is challenging. Immunocytological and biochemical assays represent the current gold standard for monitoring cell death pathways; however, these standard assays are invasive, render large numbers of cells and impede continuous monitoring experiments. In this study, both room temperature (RT)-induced apoptosis and heat-triggered necrosis were analyzed in individual Saos-2 and SW-1353 cells by utilizing Raman microspectroscopy. A targeted analysis of defined cell death modalities, including early and late apoptosis as well as necrosis, was facilitated based on the combination of Raman spectroscopy with fluorescence microscopy. Spectral shifts were identified in the two cell lines that reflect biochemical changes specific for either RT-induced apoptosis or heat-mediated necrosis. A supervised classification model specified apoptotic and necrotic cell death based on single cell Raman spectra. To conclude, Raman spectroscopy allows a non-invasive, continuous monitoring of cell death, which may help shedding new light on complex pathophysiological or drug-induced cell death processes.
Similar content being viewed by others
Introduction
Apoptotic cell death is a highly regulated process that is characterized by stereotypical morphological changes of the cellular architecture1. Cell shrinkage, plasma membrane blebbing, cell detachment, externalization of phosphatidylserine, nuclear condensation and ultimately DNA fragmentation are well-described features of apoptosis1,2. Activated caspases 3 and 6 have been identified as key regulator enzymes that mediate these morphological apoptotic hallmarks1. The frequency of apoptosis-specific molecules is particularly highly dependent on the type of apoptotic stimulus, time-point of analysis as well as the cell type3. Cell populations that potentially contain viable or necrotic cells as well as apoptotic cells cannot be distinguished by standard bulk techniques such as DNA-electrophoresis, Western Blot or colorimetric enzyme assays. Therefore, a detailed analysis of apoptotic cell death requires a series of different assays2,3,4; however, these assays depend upon large numbers of cells and are unable to probe individual apoptotic cells5. Flow cytometry and fluorescence microscopy are alternative techniques for investigating heterogeneous cell populations. Utilization of propidium iodide (PI) and fluorescein isothiocyanate (FITC)-conjugated Annexin V (Annexin V-FITC) is a standard procedure to monitor the progression of apoptosis. Early apoptotic cells are Annexin V-positive and PI-negative (Annexin V-FITC+/PI−), whereas late (end-stage) apoptotic cells are Annexin V/PI-double-positive (Annexin V-FITC+/PI+)3. However, to verify the stages of apoptosis, time-course analyses and additional methods such as caspase assays are necessary2,3,6. Moreover, this method cannot discriminate between late apoptotic and primary necrotic cells, since both of these groups of cells are Annexin V-FITC+/PI+. Other staining approaches use fluorescence-conjugated antibodies, which specifically bind to intracellular apoptotic markers. These tests require cell fixation and permeabilization; therefore a real-time monitoring of apoptotic processes is not possible. Fluorescent dyes that are suitable for live cell imaging are often associated with insufficient photostability and cytotoxic effects, or they interfere with the apoptotic machinery6.
Raman spectroscopy is an optical, marker-free technology that allows the continuous analysis of dynamic death events in single cells by investigating the overall molecular constitutions of individual cells within their physiological environment. Interestingly, this technology is not dependent on defined cellular markers and can be adapted for heterogeneous cell populations7. In Raman spectroscopy, rare events of inelastic light scattering occur on molecular bonds due to the excitation with monochromatic light and generate a fingerprint spectrum of the investigated specimens8,9. Although the effect of Raman scattering is weak, the presence of water does not impact Raman spectra, enabling the examination of native biological samples without the need for fixation or embedding procedures, making the technique superior to infrared spectroscopy. Raman spectroscopic systems are mainly composed of a light source, which is typically a laser that is connected to optical filters, a spectral grating and a detector9,10. The implementation of near-infrared lasers for Raman spectroscopy allowed the characterization of living cells without triggering photo-induced cellular damage11. Coupling of the Raman system to a conventional microscope enabled a combination of morphological and fluorescence screening and allowed spatially-resolved analyses12. Using such systems, Notingher et al. investigated the impact of Triton-X100, ricin and sulphor-mustard on A549 lung epithelial cells13,14. Single cell Raman spectra showed incremental spectral changes dependent on the incubation time of the toxic agents, indicating that death modalities such as apoptosis and necrosis were reflected by specific peak shifts13. Etoposide, which is known to trigger apoptotic cell death, induced a decrease of DNA, RNA and protein bands in Raman spectra of A549 cells15. Kunaparedy et al. exposed a melanoma cell line to oxygen-glucose deprivation and found significant changes in DNA, RNA and protein bands in Raman spectra of these cells. It was concluded that these changes indicate necrotic cell death16. In K562 leukemia cells, apoptosis was induced by adding Triton-X100 and necrosis was mediated by cytosine arabinoside treatment. The resulting Raman spectra of viable, apoptotic and necrotic K562 leukemia cells were distinguished by employing principal component analysis (PCA) and other multivariate methods17. A support vector machine (SVM) model was shown to be a powerful approach to classify spectra to predefined categories of cell death18.
In this study, Raman microspectroscopy was utilized to identify room temperature (RT)-induced early and late apoptotic events in two sarcoma cell lines - Saos-2 and SW-1353 cells. In addition, we investigated the possibility to also diagnose primary heat-mediated necrosis in these cells. A routine fluorescence staining approach was chosen to detect cell viability as well as different apoptotic stages using Annexin V-FITC and PI19. The combination of Raman spectroscopy with conventional fluorescence microscopy allowed the identification of individual viable, early and late apoptotic cells. By correlating the acquired fluorescence images of individual cells to their Raman spectra, we demonstrated that even early apoptotic events are detectable by Raman spectroscopy. Based on fluorescence images and multivariate data analysis, the validation of spectral patterns of viable, apoptotic and primary necrotic cells was accomplished. Consistent spectral shifts that featured either RT-induced apoptosis or heat-mediated primary necrosis were identified in both cell lines. Our findings suggest that Raman microspectroscopy is a powerful single cell analysis platform to detect and discriminate apoptotic and necrotic cell death.
Results
Induction of apoptosis and primary necrosis in Saos-2 and SW-1353 cells
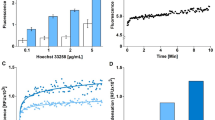
To avoid dose- and time-dependent adverse effects of chemical agents and their possible ambiguous induction of death pathways5, exposure to RT has been described as valid method to induce caspase-dependent apoptotic cell death in several cell lines20,21. In this study, Saos-2 and SW-1353 cells were exposed to RT for four to seven days to induce apoptosis. We determined significantly increased activities of caspases 3 and 6 after four days at RT in Saos-2 cells and after five days in SW-1353 cells. Caspase activities further increased in both cell lines when exposure to RT was continued (Figure 1a and b).
Verification of apoptotic and necrotic cell death using standard methods.
(a), (b) After exposure to room temperature (RT), activities of caspase 3 and caspase 6 are determined in (a) Saos-2 and (b) SW-1353-cells (n = 3, * P < 0.05 compared to non-treated control). (c–f) Flow cytometric analysis of Annexin V/PI-stained, apoptotic (c) Saos-2 and (d) SW-1353 cells as well as heat shock-treated necrotic (e) Saos-2 and (f) SW-1353 cells (n = 1) after four days at RT.
To determine the percentage of early and late apoptotic cells in RT-exposed cultures, conventional flow cytometry was performed using Annexin V-FITC and PI. Viable cells remained unstained (Annexin V-FITC−/PI−). Early apoptotic cells showed Annexin V-FITC+/PI− staining patterns; whereas late apoptotic cells exhibited Annexin V-FITC+/PI+ staining patterns due to a loss of plasma membrane integrity3,22,23. In non-treated control cultures, 6% of the Saos-2 cells and 14% of the SW-1353 cells were Annexin V-FITC+/PI−; whereas 2% of the Saos-2 cells and 11% of the SW-1353 cells were Annexin V-FITC+/PI+ (Supplemental Figure S1). After four days of exposure to RT, a significant increase of early and late apoptotic cells was identified within both Saos-2 and SW-1353 cell populations (Figure 1c, d). 38% of the Saos-2 cells and 62% of the SW-1353 cells were Annexin V-FITC+/PI− and 59% of the Saos-2 cells and 29% of the SW-1353 cells were Annexin V-FITC+/PI+. In contrast, after induction of heat-triggered necrosis, 97% of the Saos-2 cells and 60% of the SW-1353 cells exhibited strong PI staining (Figure 1e and f), which identifies primary necrotic cells24. Due to extensive membrane rupture, 85% of the Saos-2 and 52% of the SW-1353 cells expressed also Annexin V, a phenomenon which was seen by others before3.
Cellular death events define the Raman spectral fingerprint
As shown by Annexin V and PI staining, heterogeneous cell populations that included viable and dead cells were present after inducing apoptosis by exposure to RT as well as after inducing necrosis by exposure to high temperatures. To selectively collect and analyze Raman spectra of viable cells (Annexin V-FITC−/PI− population), early apoptotic cells (Annexin V-FITC+/PI−), late apoptotic cells (Annexin V-FITC+/PI+) and heat-treated primary necrotic cells (Annexin V-FITC+/PI+), spectral acquisition and fluorescence imaging was combined. Raman spectroscopy and imaging of Annexin V-FITC and PI fluorescence were conducted simultaneously on stained cells using our custom-built Raman microspectrometer system8,25,26,27,28. Cellular uptake of the fluorescent dyes did not impact the Raman spectral profiles (Supplemental Figure S2). Viable, non-stained Saos-2 cells exhibited a Raman spectral signature with a defined peak pattern displaying a biochemical cellular fingerprint that is composed of complex overlapping signals from proteins, nucleic acids, lipids and carbohydrates (Figure 2a)8,12,14. Raman spectra of early apoptotic Annexin V-FITC+/PI− Saos-2 cells featured a new Raman band at 1375 cm−1 (Figure 2b, Supplemental Figure S3). This signal became more prominent in spectra of late apoptotic Saos-2 cells. Moreover, late apoptotic Saos-2 cells exhibited considerable decreasing relative spectral intensities when compared to viable cells, particularly observable at the Raman shifts 1003 cm−1 and 1450 cm−1 (Figure 2c, Supplemental Figure S3). Raman spectra of heat-treated necrotic Annexin V-FITC+/PI+ Saos-2 cells exhibited an increasing relative intensity at 1003 cm−1. Furthermore, the amide I peak, which was at the position 1658 cm−1 in viable and apoptotic cells, was shifted in necrotic cells. This shift is related to a higher intensity at the 1658 cm−1 signal in viable cells when compared to necrotic cells (Figure 2d, Supplemental Figure S3). Higher standard deviations were detected for the Raman band at 780–800 cm−1 in both apoptotic and necrotic Saos-2 cells. In addition, all treated Saos-2 cells showed a diminishing peak at 1052 cm−1 (Supplemental Figure S3).
Raman spectra and fluorescence microscopy of Saos-2 cells.
Spectra of (a) viable (Annexin V-FITC−/PI−) (n = 90), (b) early apoptotic (Annexin V-FITC+/PI−) (n = 87) and (c) late apoptotic (Annexin V-FITC+/PI+) (n = 64) Saos-2 cells. (d) Spectra of heat-induced necrotic Saos-2 cells (n = 88). Spectral standard deviations are depicted as grey lines. Bright-field (BF) and fluorescence images (left panel) display morphology and Annexin V/PI staining of single cells. White arrows identify representative cells for each group.
A complex composition of intracellular molecules formed the cell-specific Raman fingerprint of viable SW-1353 cells (Figure 3a)8,12,14. When SW-1353 cells entered the apoptotic pathway as identified by Annexin V-FITC expression, spectral signals within this fingerprint were altered. In detail, the Raman signal at 1375 cm−1 increased significantly due to early apoptosis in SW-1353 cells (Figure 3b, Supplemental Figure S4). Late apoptotic SW-1353 cells showed decreasing peaks at 1003 cm−1 and 1658 cm−1 when compared to viable SW-1353 cells (Figure 4c, Supplemental Figure S4). Raman spectra of heat-treated necrotic Annexin V-FITC+/PI+ SW-1353 cells featured an increasing Raman signal at 1375 cm−1 (Supplemental Figure S4). A shift of the Raman band at 1658 cm−1 was detected in necrotic SW-1353 cells (Figure 3d, Supplemental Figure S4). Consistent with the Raman spectra measured from collapsing Saos-2 cells, the peak at 1052 cm−1 was neither detected in apoptotic nor necrotic SW-1353 cells (Supplemental Figure S4).
Raman spectra and fluorescence microscopy of SW-1353 cells.
Spectra of (a) viable (Annexin V-FITC−/PI−) (n = 90), (b) early apoptotic (Annexin V-FITC+/PI−) (n = 82) and (c) late apoptotic (Annexin V-FITC+/PI+) (n = 74) SW-1353 cells. (d) Heat-treated necrotic SW-1353 cells (n = 76). Spectral standard deviations are depicted as grey lines. Bright-field (BF) and fluorescence images (left panel) display the cell morphology and Annexin V/PI staining of single cells. White arrows depict representative cells for each group.
Score plots of principal component analysis (PCA).
(a–c) Score plots comparing the Raman spectra of viable Saos-2 cells versus (a) early apoptotic, (b) late apoptotic and (c) necrotic Saos-2 cells. (d–f) Raman spectra of viable SW-1353 cells versus spectra of: (d) early apoptotic (e) late apoptotic and (f) necrotic Saos-2 cells. Mean score values, standard deviations and p-values of each sample group compared to viable cells are indicated on the left side of each plot.
Multivariate scores of Raman spectra reflect the modality and the stages of cell death
Analysis of single peak intensities is not sufficient to resolve complex spectral peak shifts. For this purpose, PCA was employed. PCA is a powerful multivariate analytical approach to reveal spectroscopic variations and peak correlations by calculating principal components (PCs), which dissolve the variances within the spectral data set29,30. Related to these PCs, individual Raman spectra are described by few score values29. Raman spectra of viable Saos-2 or SW-1353 cells and their respective early apoptotic, late apoptotic or necrotic derivatives were subsequently analyzed by PCA. Score values were used to validate significant differences in the Raman spectra due to apoptotic or necrotic cell death (Figure 4). The corresponding PC loading spectra reflected the actual spectral changes due to different cell death modalities in Saos-2 and SW-1353 cells (Figure 5). As a result of PCA, Raman spectra of viable and early apoptotic Saos-2 cells exhibited significantly different PC 2 score values. The PC 2 score values represented 11% of the total spectral variances (Figure 4a). When plotting the score values of Raman spectra of viable Saos-2 cells and late apoptotic Saos-2 cells, PC 1 score values, which explained 40% of the total spectral variance, differed significantly (Figure 4b). Using PC 2 scores, viable Saos-2 cells were clearly separable from necrotic Saos-2 cells based on their Raman spectral pattern. Here, PC 2 accounted for 22% of the total spectral variance (Figure 4c).
Loading spectra of principal component analysis (PCA).
Loadings exhibit the spectral differences of viable Saos-2 cells compared to (a) early apoptotic, (b) late apoptotic and (c) necrotic Saos-2 cells. Loadings of PCA depict specific peak shifts due to (d) early apoptosis, (e) late apoptosis and (f) necrosis in SW-1353 cells.
Comparable results were observed when performing PCA on the analogous data of SW-1353 cells. PC 2 score values demonstrated that the Raman spectra acquired from viable SW-1353 cells differed significantly from Raman spectra of early apoptotic SW-1353 cells. These differences in PC 2 score values reflected 13% of the total spectral variances (Figure 4d). PC 2 score values revealed vast differences between the spectra of viable and late apoptotic SW-1353 cells. Here, the explained spectral variance in PC 2 was 32% and indicated that the spectral differences correlate with the progressing apoptotic cell death (Figure 4e). Raman spectra of viable and necrotic SW-1353 cells formed two distinct clusters in the scores plot. PC 1 score values separated these clusters based on 42% of the total spectral variance (Figure 4f).
Loadings of the PC score values and exhibited the spectral changes that occurred due to early apoptotic, late apoptotic cell death or necrosis. Loadings have a spectral dimension, where positive and negative peaks can be observed. If the original spectrum is assigned to positive score values, positive peaks in the loadings indicate rising Raman signals in the original spectra. In contrast, negative loading bands denote a reduced contribution of the respective signals in the measured Raman spectra29. In our study, loadings of PC 2 indicated increasing Raman bands (795 cm−1, 1375 cm−1) and decreasing signals (1003 cm−1, 1658 cm−1) for early apoptotic Saos-2 cells when compared to viable Saos-2 cells (Figure 5a). When the later stage of apoptosis was reached, some of these spectral shifts were preserved. In addition, a reduction of further signals (1047 cm−1) was indicative for late apoptotic Saos-2 cells (Figure 5b). In contrast, loadings of PC 2, which depicted the shifts for Raman spectra of necrotic Saos-2 cells, showed increasing 1003 cm−1 signals, decreasing bands (784 cm−1, 1315 cm−1) and a reduced contribution of the broad band at 1658 cm−1 (Figure 5c). In the PCA of viable and early apoptotic SW-1353 cells, PC 2 loadings indicated decreasing signals at 1047 cm−1, but an increasing band at 1375 cm−1 for early apoptotic SW-1353 cells (Figure 5d). Compared to viable SW-1353 cells, a decreasing band at 786 cm−1 and an increasing signal at 1437 cm−1 were identified in the spectra of late apoptotic SW-1353 cells based on the PC 2 loadings (Figure 5e). PC 1 loading signals denoted that several peaks (1003 cm−1, 1244 cm−1) increased and signals at 1437 cm−1, 1658 cm−1 decreased due to necrosis (Figure 5f).
Classification of single cell Raman spectra to predefined cell death modalities
A SVM model was developed, which enabled the assignment of Raman spectra according to the categories of cell death. For the prediction of newly acquired Raman data, it was shown that SVMs are superior to PCA, since SVMs can handle different cell types as well as spectra of multiple different categories27,31. Within this SVM model, classes of Raman spectra from viable, early and late apoptotic cells as well as necrotic cells were specified. The SVM algorithm was sufficient to classify spectra from different cell types. Accordingly, spectra of Saos-2 and SW-1353 cells were pooled. In total, 335 random spectra were used as training data to generate the SVM. The resulting model achieved a total training accuracy of 92.8% and a cross validation accuracy of 91.0%. Independent validation of the SVM model was achieved by predicting the cell death modalities of 332 Raman spectra that served as testing data. In this validation, all Raman spectra of viable cells were correctly classified. Spectra of early apoptotic cells were assigned with an accuracy of 94.3%. The model assigned several late apoptotic cells to the group of early apoptosis, which reduced the sensitivity for detecting late apoptosis to 75.0%. For classifying necrotic cells, a remarkable sensitivity of 94.1% and a specificity of 97.2% were reached. To summarize, the SVM model exhibited a total sensitivity of 91.4 ± 11.2%, a specificity of 96.2 ± 3.0% and a total accuracy of 96.2 ± 3.3% for identifying viable cells and the different modalities of cell demise based on Raman spectra (Table 1).
Discussion
In this study, we demonstrated that Raman microspectroscopy is a powerful non-contact, non-invasive and marker-free single cell analysis platform for the identification of apoptotic and necrotic cell death. The combined analysis of individual Saos-2 and SW-1353 cells with fluorescence microscopy and Raman spectroscopy enabled a targeted validation of spectral patterns for early and late apoptotic cells. Although we have not used different methods for inducing apoptosis in this study, we proved that exposure to RT was sufficient to induce apoptosis by measuring caspase activities and apoptosis-specific fluorescence staining. Progressing apoptosis was reflected by minor shifts of the Raman spectra at several positions, which was observed in both cell lines. Heat-treated necrotic cells were characterized by increased protein signals in their Raman profiles, which is in contrast to apoptotic cells where decreasing protein bands were observed. Multivariate analysis identified viable, apoptotic and necrotic cells, based on their specific spectral features.
Raman spectroscopy was employed in previous studies to identify cell death using various cell types including K562 leukemia cells, T-lymphocytes, gastric carcinoma cells, MEL-28 cells and A549 lung epithelial cells13,15,16,17,32,33. In these studies, physical effects, chemotherapeutics and other pharmaceutical substances were used to induce variants of cell death events revealing the differing impact on the cellular molecular constitution13,15,16,17,32,33. All of these studies identified significant differences in Raman spectra of individual cells when compared to untreated cells. In our study, two cell lines were analyzed using the same Raman settings. Instead of using chemical agents to induce apoptotic or necrotic cell death, we accomplished RT-induced apoptosis and heat-induced necrosis and a targeted analysis of early and late apoptotic cells was performed. PCA score values of single cell Raman spectra demonstrated that the molecular profiles of cells in an early apoptotic stage were significantly different to those of viable cells. The explained variance of the PC 2, which reflects these significant differences, was similar in both cell lines (11% for Saos-2 cells and 13% for SW-1353 cells). An increasing Raman signal at 1375 cm−1 was identified in early apoptotic Annexin V-expressing Saos-2 and SW-1353 cells. This peak has not been described in dying cells before, although a Raman signal at 1375 cm−1 was previously found in spectra of peripheral blood cells and isolated chromosomes, in which highly condensed chromatin structures are known to be present11,34. Indeed, incipient nuclear condensation was detected in detaching Annexin V-positive MDKC cells35. In our study, spectral differences of late apoptotic versus viable cells progressed according to the stage of apoptosis. The Raman signal at 1375 cm−1 increased further in late apoptotic Saos-2 cells, indicating progression of nuclear condensation. In SW-1353 cells, the contribution of this band to the late apoptotic cell stage was less clear. For both cell lines, nucleic bands at 780–800 cm−1 decreased during apoptosis. It has been suggested before that the decrease of the 780–800 cm−1 bands may correlate with the progress of internucleosomal DNA cleavage36. Raman spectra of late apoptotic Saos-2 and SW-1353 cells featured diminishing protein signals, which were especially evident at wavenumber 1003 cm−1. In late-apoptotic SW-1353 cells, an increased contribution of lipids at 1437 cm−1 is exhibited, which is likely to correlate to reduced protein signals14. Active intracellular apoptotic proteases could mediate a decreased contribution of protein signals to these Raman spectra1. In total, the spectral changes that were detected due to RT-induced apoptosis in Saos-2 cells and SW-1353 cells were consistent with other reports where apoptosis was triggered by other factors13,14,15,36. However, we did not investigate if the treatment had a different or similar effect on the cellular Raman spectra compared to treatment with apoptosis-inducing chemical agents. Future studies should focus on this issue.
Protein peaks (e.g. phenylalanine at 1003 cm−1) increased significantly in both, necrotic Saos-2 and necrotic SW-1353 cells - a phenomenon that has been previously described in necrotic cells16. This increase of protein signals in Raman spectra could be triggered by the necrotic release of nuclear proteins to the cytosol. Conformational changes of proteins as well as forming protein aggregates, which occur intracellular due to excessive high temperature exposure37, might further contribute to altered protein signals in Raman spectra of necrotic Saos-2 and SW-1353 cells. The detected shift in the amide I signal at 1658 cm−1 emphasized the presence of structurally modified proteins. This signal was identified before in Raman spectra of dying cells, where necrosis was induced by chemical treatment and photo degradation11,14. Since both, the increase of the phenylalanine signal as well as the shift of the amide I signal have been identified as necrotic hallmarks in cellular Raman spectra before14,16, we concluded that these spectral shifts were not specific for heat-induced necrosis, but rather for necrosis in general, although we did not investigate to which extent such necrotic hallmarks depend on the method that is used to induce necrosis. Interestingly, in necrotic Saos-2 and SW-1353 cells an increasing 1375 cm−1 signal may indicate changing chromatin architectures, triggered by the release of the HGMB1, a chromatin-binding protein that is relocated in necrotic cells38.
Employing a SVM model showed that based on the Raman spectra of individual cells, cell viability and apoptotic as well as necrotic cell death can be assessed with high accuracy, sensitivity and specificity. This model served as a powerful database for the recognition and discrimination of the Raman spectra. A similar algorithm showed best results for classifying Raman spectra of cancerous versus non-cancerous breast cell lines39. Pyrgiotakis et al. investigated the effects of different chemical treatments on A549 lung epithelial cells using Raman spectroscopy and SVM classification18. When employing such a model in the future, steps of fluorescence staining and imaging could be omitted and analysis of newly acquired data could be attained based on the previously established multivariate model7,27. Then, Raman spectroscopy could facilitate the non-invasive continuous monitoring of drug-induced death signals in single cells, which is of particular interest for the field of drug development and targeting many disease scenarios such as tumor progression, Alzheimer's disease and autoimmunity. However, in order to transform Raman microspectroscopy into an online high throughput assay, spectra acquisition times must be reduced7. Shorter spectra acquisition times should not impact the specificity of multivariate classifications and could therefore be employed once classification models are established. Cellular Raman imaging has then the potential to provide novel insights into dynamic intracellular biochemistry. Using Raman microspectroscopy for cellular imaging, Zoladek et al. resolved subcellular modifications due to apoptotic cell death40. Moreover, adaptions of Raman spectroscopic systems have been accomplished in order to analyze living cells at a single cell level within minimal sample volumes41,42. This could further facilitate the monitoring of cell death processes in various experimental settings. Importantly, Raman-based assays could potentially allow the detection of cell viability within rare cell populations, which is important for tissue engineering and regenerative medicine applications. Automated cell culture monitoring systems could benefit from the implementation of Raman spectroscopy as a method to screen cell viability in real-time43,44. Raman spectroscopy may offer the unique possibility to monitor cell death and cytotoxic effects marker-free within tissues and three-dimensional (3D) environments in their native, unprocessed state26,27. Moreover, in future studies, Raman spectroscopy could be combined with more advanced fluorescence-based techniques, such as fluorescence lifetime imaging microscopy (FLIM) or fluorescence resonance energy transfer (FRET) - a technology that has been shown to enable real-time monitoring of caspase-activities in single cells45,46,47.
To conclude, we validated single cell Raman spectra of individual early and late apoptotic as well as necrotic cells in this study. By combining Raman spectroscopy and fluorescence microscopy, highly reproducible data sets were generated, which showed specificity for RT-triggered apoptotic and heat-induced primary necrotic cells. A multivariate classification model successfully identified viable, RT-induced early and late apoptotic cells as well as heat-treated necrotic cells based on Raman spectra. This approach shows the potential of the Raman microspectroscopy, which could facilitate non-invasive monitoring of drug-induced death scenarios in single cells.
Methods
Cell culture
Saos-2 cells and SW-1353 cells (both ATCC-LGC Standards GmbH, Wesel, Germany) were cultured in DMEM/Hams F12 medium supplemented with 10% FBS (Invitrogen, Carlsbad, CA, US) under standard cell culture conditions (37°C, 5% CO2). Cell numbers were assessed using a CASY cell counter (Schärfe Systems, Reutlingen, Germany). Trypsin/EDTA (0.05%, Invitrogen) was employed to detach the cells from the flask, either for passaging, heat shock induction, flow cytometry or Raman microspectroscopy.
Induction of apoptotic cell death
To induce apoptosis, cells were exposed to room temperature (20°C) for a period of four to seven days. RT-treated cells were harvested from the supernatant at different time points for caspase assays, flow cytometry and Raman spectroscopic analyses.
Induction of primary necrosis
Exposing the cells to non-physiological heat, which spawns a harsh death stimulus and causes immediate cell death due to protein denaturation, induced primary necrosis24. Cells were treated with Trypsin/EDTA and placed into a sterile tube. After centrifugation for 5 minutes at 300 g, the supernatant was removed and the cells were re-suspended in DMEM/Hams F12 medium. The cell suspension was then introduced to a 55°C-heated water bath for 90 minutes. The heat-treated cells were immediately analyzed using Raman microspectroscopy or stained and analyzed using flow cytometry.
Caspase assay
A Caspase Colometric Protease Assay Sampler Kit (Invitrogen, Carlsbad, US) was used for colorimetric detection of active caspases 3 and 6. The assay was performed as recommended by the manufacturer. Total protein concentration was determined using a Micro BCA Protein Assay Kit (Pierce, Rockford, US). Caspase assays were performed on cell lysates, which contained 1.5 μg/μl protein of Saos-2 cells and 2 μg/μl protein for SW-1353 cells. Absorption values were measured at 405 nm using a Tecan spectra reader (Tecan, Crailsheim, Germany). Background absorbance was subtracted from the absorbance values of all samples. Absorbance values of different experiments were normalized to the absorbance values measured in either non-treated Saos-2 cells or non-treated SW-1353 cells, which were cultured under standard conditions (37°C, 5% CO2). The results are provided as relative enzyme activity. Accordingly, the absorbance values of non-treated control cells (0 days of RT-exposure) refer to relative caspase activities of 1.
Annexin V-FITC/PI-staining
Cells (3 × 106) were stained using the FITC Annexin V/Dead Cell Apoptosis Kit (V13242, Invitrogen) according to the manufacturer's instructions. Stained cells were diluted in Annexin V-binding buffer (Invitrogen). Suspended cells were used to perform flow cytometry or Raman microspectroscopy.
Flow cytometry
Annexin V-FITC/PI-stained cells were analyzed using a BD FACSCalibur flow cytometer (BD Biosciences, Heidelberg, Germany). In total 10,000 cells were analyzed per measurement. Data was analyzed using FlowJo 10.0.7 software (Treestar Inc, Ashland, US).
Raman microspectroscopy
A custom-built Raman microspectroscopy system was employed for all measurements as previously described8,28,48. The spectroscopic unit features a 784 nm laser diode (Toptica Photonics AG, Munich, Germany) and a conventional fluorescence microscope (IX71, Olympus, Hamburg, Germany), which was equipped with a 60× water immersion objective (NA 1.2, Olympus) and a conventional mercury vapor lamp to excite fluorescent dyes (U-RFL-T, Olympus). The laser was focused through the objective, resulting in a spot diameter of approximately 1 μm. As previously described, when employing this system on detached cells, the collected Raman signals predominantly reflect molecular bonds of the cell nuclei27. Both, Raman spectra and fluorescence images of Annexin V-FITC/PI-staining were recorded of viable, early and late apoptotic as well as necrotic cells. All cells were suspended in solution as described above. Glass bottom dishes (Greiner Bio-one, Frickenhausen, Germany) were used for all measurements. Fluorescence of the Annexin V-FITC and PI staining was visualized using a F-View camera and Cell∧B 2.4 software (both Olympus). The FITC and PI fluorescence channels were recorded separately. Images were processed using Adobe Photoshop CS3. After acquiring the fluorescence images, the optical filters were switched to the Raman measurement mode and the laser beam was focused into the center of an individual cell. A Raman spectrum of an individual cell was accumulated and assigned according to the previously acquired Annexin V-FITC/PI image of the same cell. An acquisition time of 100 seconds with 85 mW laser power were used for each spectrum and cell. Per treatment and per cell line 20–30 spectra of individual cells were conducted per experiment. Three independent experiments were performed.
Processing of Raman spectra and multivariate analysis
The raw spectra were loaded into OPUS software (Opus® 4.2, Bruker Optics, Billerica, MA, USA) to remove cosmic rays manually and subtract the glass background from every sample spectrum. The spectra were cut into the range of 600–1800 cm−1. A baseline correction (64-data point rubber band method, OPUS software) was sufficient to correct background shifts, as the spectra of Saos-2 and SW-1353 cells were not impaired by intrinsic fluorescence signals. Details on these processing steps were previously published8,28,48. The processed spectra were imported to the Unscrambler software (Unscrambler®X10.2, CAMO, Oslo, Norway). Vector normalization was performed on all spectra to correct intensity variations caused by technical effects. Here, for each single spectrum the absolute intensity values of each wavenumber were normalized to the total intensity of the same spectrum. Spectral intensities were divided by a vector that constitutes the length of the spectrum49. Vector-normalized spectra were comparable in their total intensity. Signal intensities are provided as relative values in arbitrary units [a.u.]49. After vector-normalization, the relative intensities between two sample groups were analyzed50. Background-corrected and vector-normalized Raman spectra were used for all further analyses.
Each spectrum was assigned to one of the following categories: viable, early apoptotic, late apoptotic and necrotic. Successive PCAs were calculated as described before8,28,48 in order to visualize the spectral differences between viable cells and either early apoptotic, late apoptotic or necrotic cells.
The Unscrambler®X10.2 software was used to calculate a SVM model, as described previously27. The software-implemented SVM algorithm allowed a multi-class classification51. The spectral wavenumbers 600–1800 cm−1 were used as an input for the SVM. Based on the training data set, a ν-SVM model with a linear kernel type and a ν-value of 0.5 was computed. The training data set consisted of 90 spectra of viable cells, 90 spectra of early apoptotic cells, 69 spectra of late apoptotic cells and 86 spectra of necrotic cells (335 spectra total). Cross validation using 10 cross validation segments was employed. A total cross validation accuracy of 91.04% was reached. To estimate the performance of the model, a new data set ( = testing data) was predicted with the SVM model. This testing data included a total number of 332 spectra. Based on the results for the testing data, sensitivity, specificity and accuracy of the SVM model were analyzed for each category.
Statistical analysis
Vector-normalized Raman spectra are presented as mean ± standard deviation. Significance analysis was employed to compare relative spectral intensities at single wavenumbers as well as PC score values of spectra using OriginPro 9.0.0G (OriginLab Corporation, Northampton, MA, USA). Prior to significance tests, normality distribution of data sets was tested using Kolmogorov-Smirnov test. All investigated data sets were normality distributed. Statistical significance was determined by Fisher's analysis of variance (ANOVA). P-values less than 0.05 (p < 0.05) were defined as statistically significant.
References
Taylor, R. C., Cullen, S. P. & Martin, S. J. Apoptosis: controlled demolition at the cellular level. Nat Rev Mol Cell Biol 9, 231–41 (2008).
Henry, C. M., Hollville, E. & Martin, S. J. Measuring apoptosis by microscopy and flow cytometry. Methods 61, 90–97 (2013).
Wlodkowic, D., Telford, W., Skommer, J. & Darzynkiewicz, Z. Apoptosis and beyond: cytometry in studies of programmed cell death. Methods Cell Biol 103, 55–98 (2011).
Krysko, D. V., Vanden Berghe, T., Parthoens, E., D'Herde, K. & Vandenabeele, P. [Chapter 16 Methods for Distinguishing Apoptotic from Necrotic Cells and Measuring Their Clearance] Methods in Enzymology: Programmed Cell Death, General Principles for Studying Cell Death, Part A [Khosravi-Far, R., Lockshin, Z. Z. R. A. & Piacentini, M. (eds.)] [307–341] (Academic Press, 2008).
Ledda-Columbano, G. M. et al. Induction of two different modes of cell death, apoptosis and necrosis, in rat liver after a single dose of thioacetamide. Am J Pathol 139, 1099–109 (1991).
Wlodkowic, D., Skommer, J. & Darzynkiewicz, Z. Cytometry of apoptosis. Historical perspective and new advances. Exp Oncol 34, 255–62 (2012).
Li, M., Xu, J., Romero-Gonzalez, M., Banwart, S. A. & Huang, W. E. Single cell Raman spectroscopy for cell sorting and imaging. Curr Opin Biotechnol 23, 56–63 (2012).
Votteler, M., Carvajal Berrio, D. A., Pudlas, M., Walles, H. & Schenke-Layland, K. Non-contact, label-free monitoring of cells and extracellular matrix using Raman spectroscopy. J Vis Exp 29 (2012).
Das, R. S. & Agrawal, Y. K. Raman spectroscopy: Recent advancements, techniques and applications. Vib Spectrosc 57, 163–176 (2011).
Mariani, M. M. & Deckert, V. Raman Spectroscopy: Principles, Benefits & Applications. Bunsen-Magazin 14, 136–147 (2012).
Puppels, G. J. et al. Laser irradiation and Raman spectroscopy of single living cells and chromosomes: Sample degradation occurs with 514.5 nm but not with 660 nm laser light. Exp Cell Res 195, 361–367 (1991).
Puppels, G. J. et al. Studying single living cells and chromosomes by confocal Raman microspectroscopy. Nature 347, 301–3 (1990).
Notingher, I. et al. Discrimination between ricin and sulphur mustard toxicity in vitro using Raman spectroscopy. J Ro Soc Interface 1, 79–90 (2004).
Notingher, I., Selvakumaran, J. & Hench, L. L. New detection system for toxic agents based on continuous spectroscopic monitoring of living cells. Biosens Bioelectron 20, 780–9 (2004).
Owen, C. A. et al. In vitro toxicology evaluation of pharmaceuticals using Raman micro-spectroscopy. J Cell Biochem 99, 178–86 (2006).
Kunapareddy, N., Freyer, J. P. & Mourant, J. R. Raman spectroscopic characterization of necrotic cell death. J Biomed Opt 13, 054002 (2008).
Ong, Y. H., Lim, M. & Liu, Q. Comparison of principal component analysis and biochemical component analysis in Raman spectroscopy for the discrimination of apoptosis and necrosis in K562 leukemia cells. Opt Express 20, 22158–71 (2012).
Pyrgiotakis, G. et al. Cell death discrimination with Raman spectroscopy and support vector machines. Ann Biomed Eng 37, 1464–73 (2009).
Vermes, I., Haanen, C., Steffens-Nakken, H. & Reutelingsperger, C. A novel assay for apoptosis. Flow cytometric detection of phosphatidylserine expression on early apoptotic cells using fluorescein labelled Annexin V. J Immunol Methods 184, 39–51 (1995).
Shimura, M. et al. Characterization of room temperature induced apoptosis in HL-60. FEBS Lett 417, 379–384 (1997).
Shimura, M. et al. Room temperature-induced apoptosis of Jurkat cells sensitive to both caspase-1 and caspase-3 inhibitors. Cancer Lett 132, 7–16 (1998).
Martin, S. J. et al. Early redistribution of plasma membrane phosphatidylserine is a general feature of apoptosis regardless of the initiating stimulus: inhibition by overexpression of Bcl-2 and Abl. J Exp Med 182, 1545–56 (1995).
Walsh, G. M., Dewson, G., Wardlaw, A. J., Levi-Schaffer, F. & Moqbel, R. A comparative study of different methods for the assessment of apoptosis and necrosis in human eosinophils. J Immunol Methods 217, 153–63 (1998).
Janko, C. et al. Navigation to the graveyard-induction of various pathways of necrosis and their classification by flow cytometry. Methods Mol Biol 1004, 3–15 (2013).
Pudlas, M., Brauchle, E., Klein, T. J., Hutmacher, D. W. & Schenke-Layland, K. Non-invasive identification of proteoglycans and chondrocyte differentiation state by Raman microspectroscopy. J Biophotonics 6, 205–11 (2013).
Brauchle, E., Johannsen, H., Nolan, S., Thude, S. & Schenke-Layland, K. Design and analysis of a squamous cell carcinoma in vitro model system. Biomaterials 34, 7401–7 (2013).
Pudlas, M. et al. Raman spectroscopy: a noninvasive analysis tool for the discrimination of human skin cells. Tissue Eng Part C Methods 17, 1027–40 (2011).
Pudlas, M. et al. Non-contact discrimination of human bone marrow-derived mesenchymal stem cells and fibroblasts using Raman spectroscopy. Med Laser Appl 26, 119–125 (2011).
Næs, T., Isaksson, T., Fearn, T. & Davies, T. A user-friendly guide to Multivariate Calibration and Classification (NIR Publications, Chichester, UK, 2002).
Bonnier, F. & Byrne, H. J. Understanding the molecular information contained in principal component analysis of vibrational spectra of biological systems. Analyst 137, 322–32 (2012).
Cortes, C. & Vapnik, V. Support-vector networks. Machine Learning 20, 273–297 (1995).
Yao, H. et al. Raman spectroscopic analysis of apoptosis of single human gastric cancer cells. Vib Spectrosc 50, 193–197 (2009).
Moritz, T. J., Taylor, D. S., Krol, D. M., Fritch, J. & Chan, J. W. Detection of doxorubicin-induced apoptosis of leukemic T-lymphocytes by laser tweezers Raman spectroscopy. Biomed Opt Express 1, 1138–1147 (2010).
Pully, V. V., Lenferink, A. T. M. & Otto, C. Time-lapse Raman imaging of single live lymphocytes. J Raman Spectrosc 42, 167–173 (2011).
Corfe, B. M., Dive, C. & Garrod, D. R. Changes in intercellular junctions during apoptosis precede nuclear condensation or phosphatidylserine exposure on the cell surface. Cell Death Differ 7, 234–235 (2000).
Verrier, S., Notingher, I., Polak, J. M. & Hench, L. L. In situ monitoring of cell death using Raman microspectroscopy. Biopolymers 74, 157–162 (2004).
Tyedmers, J., Mogk, A. & Bukau, B. Cellular strategies for controlling protein aggregation. Nat Rev Mol Cell Biol 11, 777–88 (2010).
Bianchi, M. E. & Manfredi, A. Chromatin and cell death. Biochim Biophys Acta 1677, 181–186 (2004).
Fenn, M. B., Pappu, V., Georgeiv, P. G. & Pardalos, P. M. Raman spectroscopy utilizing Fisher-based feature selection combined with Support Vector Machines for the characterization of breast cell lines. J Raman Spectrosc 44, 939–948 (2013).
Zoladek, A., Pascut, F. C., Patel, P. & Notingher, I. Non-invasive time-course imaging of apoptotic cells by confocal Raman micro-spectroscopy. J Raman Spectrosc 42, 251–258 (2011).
Pudlas, M., Koch, S., Bolwien, C. & Walles, H. Raman spectroscopy as a tool for quality and sterility analysis for tissue engineering applications like cartilage transplants. Int J Artif Organs 33, 228–37 (2010).
Brauchle, E. & Schenke-Layland, K. Raman spectroscopy in biomedicine - non-invasive in vitro analysis of cells and extracellular matrix components in tissues. Biotechnol J 8, 288–97 (2013).
Whelan, J., Craven, S. & Glennon, B. In situ Raman spectroscopy for simultaneous monitoring of multiple process parameters in mammalian cell culture bioreactors. Biotechnol Prog 28, 1355–62 (2012).
Pully, V. V. et al. Microbioreactors for Raman microscopy of stromal cell differentiation. Anal Chem 82, 1844–50 (2010).
Rehm, M. et al. Single-cell Fluorescence Resonance Energy Transfer Analysis Demonstrates That Caspase Activation during Apoptosis Is a Rapid Process: Role of Caspase-3. J Biol Chem 277, 24506–24514 (2002).
Morgan, M. J. & Thorburn, A. Measurement of caspase activity in individual cells reveals differences in the kinetics of caspase activation between cells. Cell Death Differ 8, 38–43 (2001).
Johnson, C. E., Freel, C. D. & Kornbluth, S. Features of programmed cell death in intact Xenopus oocytes and early embryos revealed by near-infrared fluorescence and real-time monitoring. Cell Death Differ 17, 170–9 (2010).
Votteler, M. et al. Raman spectroscopy for the non-contact and non-destructive monitoring of collagen damage within tissues. J Biophotonics 5, 47–56 (2012).
Shaver, J. M. [Chemometrics for Raman Spectroscopy.]. Handbook of Raman Spectroscopy [Lewis, I. R. & Edwards, H. G. M. (eds.)] (CRC Press, New York, 2001).
Afseth, N. K., Segtnan, V. H. & Wold, J. P. Raman spectra of biological samples: A study of preprocessing methods. Appl Spectrosc 60, 1358–67 (2006).
Chang, C.-C. & Lin, C.-J. LIBSVM: A library for support vector machines. ACM Trans. Intell. Syst. Technol. 2, 1–27 (2011).
Acknowledgements
We thank Shannon Lee Layland for his helpful comments on the manuscript. Financial support was provided by the Fraunhofer-Gesellschaft Internal programs (Attract 692263 to K.S.L.), the Ministry of Science, Research and the Arts of Baden-Württemberg (33-729.55-3/214 and SI-BW 01222-91, both to K.S.L.) and IZST-Industry on Campus (IOC) funds (IOC-105 to S.Y.B. and IOC-102 to K.S.L.).
Author information
Authors and Affiliations
Contributions
E.B., S.Y.B. and K.S.-L. wrote the manuscript. S.T. conducted the flow cytometer experiments. E.B. performed all other experiments. S.Y.B. and K.S.-L. provided conceptual input. K.S.-L. designed the experiments and supervised the research.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figures
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Brauchle, E., Thude, S., Brucker, S. et al. Cell death stages in single apoptotic and necrotic cells monitored by Raman microspectroscopy. Sci Rep 4, 4698 (2014). https://doi.org/10.1038/srep04698
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04698
This article is cited by
-
Organotin complexes with Schiff’s base ligands: insights into their cytotoxic effects on lung cancer cells
Journal of Umm Al-Qura University for Applied Sciences (2024)
-
Anti-cancer potentiality of linoelaidic acid isolated from marine Tapra fish oil (Ophisthopterus tardoore) via ROS generation and caspase activation on MCF-7 cell line
Scientific Reports (2023)
-
Sonodynamic therapy of pancreatic cancer cells based on synergistic chemotherapeutic effects of selenium-PEG-curcumin nanoparticles and gemcitabine
Applied Physics A (2023)
-
Synthesis, characterization, and anticancer evaluation of 1,3-bistetrahydrofuran-2yl-5-FU as a potential agent for pancreatic cancer
BMC Cancer (2022)
-
Non-invasive cell classification using the Paint Raman Express Spectroscopy System (PRESS)
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.