Abstract
Previous studies have reported that the developmental processes of vertebrate eyes are controlled by four Pax-6 splicing variants, each modulating different downstream genes, whereas those of insect eyes are controlled by duplicated Pax-6 genes. Cephalopods belong to the Protostomes but possess a camera-type eye similar to those in vertebrates. We examined Pax-6 variations in the squid and found five types of Pax-6 splicing variants but no duplication of the Pax-6 gene. In the five splicing variants, the splicing patterns were produced by the combination of two additional exons to the ortholog and one jettisoned exon containing most of the Homeobox domain (HD). These five variants show spatio-temporal patterns of gene expression during development in the squid. Our study suggests that cephalopods acquired Pax-6 splicing variants independent of those in vertebrates and that these variants were similarly utilized in the development of the squid eye.
Similar content being viewed by others
Introduction
In bilaterian animals, the presence of splicing variants or duplicated loci of the Pax-6 gene is known to be important for eye formation1,2,3. In vertebrates, Pax-6(5a), an alternative splicing variant, has a distinct role in postnatal iris formation and is indispensable for the structural integrity of the cornea, lens and retina2. Pax-6(5a) has an additional exon to the authentic form and this additional exon encodes an insertion fragment of 14 amino acids in the Paired domain (PD). This insertion acts as “a molecular toggle” and provides a unique target recognition property that differs from that of authentic Pax-61. In the mouse brain, alternative Pax-6(5a) mRNAs are expressed at higher levels than authentic Pax-6 mRNAs, but Pax-6(5a) mRNAs are absent from the mouse lens4. It has also been reported that the ratio of the Pax-6 spliced variants varies in accordance with the developmental stage in the embryonic chick retina5.
In Drosophila, two Pax-6 homologues, namely the eyeless (ey) and twin of eyeless (toy) genes, exist as a result of gene duplication6. From exhaustive studies of the eye determination network, the two Drosophila Pax-6 proteins encoded in the genes demonstrate unique properties and phenotypic dissimilarity has been observed in forced expression assays7. In the Drosophila genome, there are two more Pax-6-like genes, eyegone (eyg) and twin of eyegone (toe), although the evolutionary relationship between these genes and ey and toy is unclear8,9. Although ey and toy act primarily in retinal specification, the primary function of eyg is to promote cell proliferation and, subsequently, to determine the appropriate size of the eye imaginal disc10,11. Each variant exerts its influence on development through different transcriptional mechanisms: ey acts as an activator, whereas eyg possesses a unique property as a dedicated repressor12,13,14. Drosophila eyg has a corrupted PD and a distinct function in eye development similar to vertebrate Pax-6(5a), but eyg is encoded as a separate gene and is not the result of alternative splicing of the ey gene.
Thus, it appears that different types of Pax-6 transcripts are required for the development of eyes in vertebrates and in invertebrates and the strategies for gaining diversity among Pax-6 transcripts appear to differ in different species. Accordingly, we addressed the diversity of strategies for gaining differences in Pax-6 transcripts, with particular emphasis on the analysis of the various Pax-6 genes/transcripts in coleoid cephalopods, which have camera eyes similar to those of vertebrates.
Results
Five Pax-6 variants and their expression patterns in squid embryos and adult eye tissues
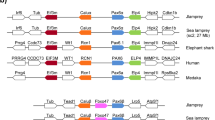
We performed a single 3′-RACE PCR for the pygmy squid Pax-6 gene (designated as IpPax-6) to investigate the splicing variants and multiple loci of Pax-6 in coleoid cephalopods. We found that there were no multiple loci in the pygmy squid, but we identified three Pax-6 variants of discrete lengths. Differences in the amino acid sequences among these Pax-6 variants were confined to limited regions. Hence, they were hypothesized to be the result of alternative splicing events of a single locus. We next validated the presence of splicing variants using RT-PCR and we finally obtained five types of splicing variants, including an apparent ortholog of authentic Pax-6 (Figure 1). The length and structure of authentic IpPax-6 were similar to those of the Pax-6 genes found in other squid species, Euprymna scolopes and Loligo pealei15,16. Authentic IpPax-6 (authentic form, 499 aa) comprises two independent DNA-binding domains, the PD and HD domains and a C-terminal P/S/T-rich domain (PST), which is the dedicated activator with a partner trans-activator protein, as shown in many animals (Figure 1). Both protein sequence similarity and the phylogenetic tree confirmed that IpPax-6 was an ortholog of fly ey and vertebrate Pax-4/6 (Supplementary figure 1). The four identified variants produced proteins with lengths differing from that of authentic IpPax-6 (Figure 1).
Diagrams of splicing variants found in the pygmy squid.
The upper row shows an estimated exon-intron structure of the squid Pax-6 gene. The arrowhead shows an intron confirmed in squid species by genomic PCR analysis and in a previous study. The authentic form (499 aa) is the most abundant and is similar to the Pax-6 gene of other squid species. Variants 1 and 3 lack the exon 4-encoding the N-terminal half of the HD. Variants 2 and 3 have an additional exon, exon 6, in the PST domain. Variant 4 also shows an additional exon 3 encoding 20 amino acids in the linker region between the PD and HD domains.
To explore the stage-specific expression of the squid Pax-6 variants, we performed Q-PCR for various tissues and at various embryonic stages using primers designed to target the additional exons of IpPax-6 (Figure 2 & Figure S2). Squid eggs show epibolic gastrulation and direct development without typical molluscan larval stages17. Embryonic eyes appear from the external epiderm of the blastodisc and are differentiable after stage 18, with retinal pigmentation starting at stage 20. The lens appears as a transparent stick-like structure visible to the unaided eye at stage 25. We first performed Q-PCR utilizing primers targeting exon 2, which covers all five variants. The Q-PCR analysis showed that IpPax-6 was expressed at stage 16 prior to eye vesicle formation (Figure 2A). The expression intensity of IpPax-6 was gradually upregulated with the development of the squid embryo (Figure 2A), with the eyeball showing the highest expression intensities among the tissues tested. As observed in the other bilaterian animals, the authentic and variant forms of IpPax-6 were expressed at markedly depressed levels in the muscle tissue. We then utilized primers targeting variants lacking exon 4 (variants 1 and 3, Figure 2B). The primers detected variants 1 and 3 at low levels in the embryos at stage 16 and in the eyeball tissue. We also used primers targeting variants including exon 6 (variants 2 and 3, Figure 2C). The Q-PCR analysis showed that variants 2 and 3 were expressed in the eyeballs and optic lobes as well as in embryos at stages 16 and 25. As the formation of photoreceptor cells and the lens begins in embryos at stage 25, the variants including exon 6 may contribute to eye development. The results demonstrate that the expression patterns of the IpPax-6 variants differed significantly from that of authentic IpPax-6.
Expression of the pygmy squid Pax-6 variants.
Expression levels of all IpPax-6 variants (A), variants without exon 4 (variants 1 and 3) (B) and variants including exon 6 (variants 2 and 3) (C) were quantified by real-time RT-PCR analysis. The expression level in each body part relative to stage16 (1.0) was calculated and subsequently normalized to the expression level of alpha-tubulin. The quantifications were performed twice on different cDNAs generated independently and geometric means were calculated. The y-axis is arbitrary. Error bars represent standard deviations. (D–G) Whole-mount in situ hybridization analyses with anti-sense RNA probes for IpPax-6 exon 2 (D, F) and IpPax-6 exon 4 (E, G). An RNA probe designed from exon 2 targeting all five variants showed the Pax-6 expression across the brain area of embryos at stage 22 (D) and at stage 25 (F). The RNA probe designed from exon 2 also indicates Pax-6 expression around the eyes (D', side view). An RNA probe designed from exon 4 targeting variants intrinsically as well as variant forms 2 and 4 showed similar expression patterns (E, G) to that of the probe targeting exon 2, except in the tissue around the eyes (E). This result suggests that the variants with an exon 4 deletion (variants 1 and 3) show specific localization in the tissue around the eyes compared with the other variants (arrowhead). Scale bars, 10 μm.
To distinguish which variants are present in each stage, we performed RT-PCR utilizing primer sets across exon boundaries. Variant 1 was considered to be expressed in all/some embryonic stages but not in the adult eyes (Supplementary figure 2). RT-PCR analysis also showed that variant 4 was strongly expressed in the adult eyes, particularly in the retina, but not in the lenses (Supplementary figure 2A). Variants 2 and 3 were expressed across all embryonic stages and also in adult tissues (Supplementary figure 2B).
To identify the tissue-specific expression of IpPax-6 variants, we performed in situ hybridization using RNA probes designed to bind specifically to each variant (Figure 2D–G). The RNA probe designed from exon 2 targets all five variants identified in this study. The RNA probe designed from exon 4 bound to the authentic form and to variants 2 and 4. IpPax-6 was found to be localized in the brain area, including the dorsal basal lobe, superior frontal lobe, peduncle/olfactory lobes and optic lobes (Figure 2D–G), as described in Hartmann et al.18 The tissue outside the retina (perhaps corresponding to the future iridophore layer) also clearly expressed IpPax-6 at stage 22 (Figures 2D and 2D'). IpPax-6 expression was observed in this layer until stage 25. The in situ hybridization utilizing the probe targeting exon 4 suggested that variants 2 and 4 had similar expression patterns in the brain but not in the eyes (Figure 2E). This finding suggests that variants 1 and 3 (lacking exon 4) are upregulated in the outer layer of the eyes. These consequences imply that each IpPax-6 variant is regulated independently across the processes of eye formation.
Exon-intron structure of Pax-6 in other cephalopods/molluscs
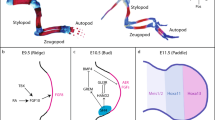
We investigated whether this type of alternative splicing was acquired only in coleoid cephalopods. Applying RT-PCR analysis to Japanese spear squid (Loligo bleekeri) embryonic RNAs, we found three types of mRNAs possibly derived from alternative splicing (exon 4 skipping, exon 3 insertion and exon 6 insertion) in the eyes (Figure 3A, B). The inserted exons 3 and 6 encoded 20 and 40 amino acids, respectively, whereas the skipped exon 4 encoded 51 amino acids. To survey the presence of similar alternative splicing in other molluscan genomes, we examined the exon-intron structures of Pax-6 in the owl limpet and pearl oyster. The complete genome sequence of the owl limpet (Lottia gigantea, obtained from JGI genome portal Lotgi v1.0, e_gw1.86.103.1)19 and of the pearl oyster (Pinctada fucata, obtained from the OIST Marine Genomics Unit genome browser P. fucata_ver1.0, transcript: pfu_aug1.0_8418.1_67856.t1, scaffold8418.1)20 showed that molluscan Pax-6 has five exons. Exon 4 in the squid was conserved across the molluscan species tested. However, exons 3 and 5 were not found in the pearl oyster Pax-6 gene. Thus, we found that variant forms 2 and 4 have been acquired in the coleoid cephalopod lineage (Figure 1).
Indels found in IpPax-6 variants and predicted 3D structures of the HD.
Aligned nucleotide sequences of (A) exon 3 and (B) exon 6 of the pygmy squid and the Japanese spear squid, respectively. Alignment of translated amino acid sequences of the HD used in comparative modeling (C). Variants 1 and 3 of IpPax-6 lack a part of helix 1. The three-dimensional structure of the spliced HD obtained by homology modeling (D, D'). Green sticks indicate proteins of IpPax-6 and gray balls represent target DNA molecules. The dotted circle indicates the part of helix 1 lost by the deletion of exon 4.
To the best of our knowledge, our study is the first to report in-frame splicing variants of squid Pax-6 that were expressed differently according to the embryonic stage. Previous studies isolated discrete types of splicing variants that had lost the N-terminal half of the PD domain in other squid species15,18, but these variants did not show spatio-temporal differences in expression. Our study also suggested that the mechanisms underlying the acquisition of variations in Pax-6 transcripts by alternative splicing have been uniquely acquired in the coleoid cephalopod lineage, as the lower molluscs, such as bivalves, do not possess a corresponding exon-like fragment in their genomes.
Function of squid Pax-6 variants and their putative role in eye development
The addition and deletion of an amino acid fragment encoded in the alternatively used exons is expected to cause structural changes in the IpPax-6 protein variants, which may alter their function in the developmental process. Two of its variants (variants 1 and 3) lack a 153mer in the middle of authentic Pax-6 and half of the HD (Figure 1). To explore whether the deletion influences their functional properties, we performed three-dimensional (3D) structural predictions of the proteins based on comparative modeling. The putative 3D structures of the HDs of authentic IpPax-6 and the variant lacking the segment coded by exon 3 were constructed. The template structure was identified by DNA-bound form so that we could predict the structure of IpPax-6 and the variant in DNA-bound form. The putative 3D structure of the authentic form was reasonably well modeled; the core residues, namely, Phe on the loop before the first helix of the HD, Leu on the first helix, Leu on the second helix and Trp and Phe on the third helix of the model structure, were conserved and the three helices of the HD were apparently tightly packed into one another (Figure 3C). The residues important for DNA binding, namely, two Arg residues at the N-terminal arm and polar residues on the surface of the third helix, were located reasonably close to the DNA interface (Figure 3C, D). However, the putative 3D structure of the variant presented a number of problematic issues. In the modeled structure, the loss of the region encoded by exon 3, which encodes the N-terminal part of the first helix, was compensated by 15 residues encoded in exon 2. Thus, the amino acid sequences of the authentic and variant forms differed only in the region containing the 15 residues of the N-terminal side. This difference, however, significantly increased the structural energy of the variant and apparently destabilized the overall structure. This instability may result from a lack of the Phe on the loop before the first helix and of the Leu on the first helix. These components are evidently important for packing the three helices. In addition, two Arg residues at the N-terminal loop that bind to the DNA bases in the minor groove in the authentic form were missing in the variant. These problems in stability and DNA binding in the variant strongly suggest that the HD of the variant is unstable and that the domain has little binding affinity for DNA (Figures 3D and 3D'). The lack of a stable HD further suggests that variants 1 and 3 have different DNA target sites from that of authentic IpPax-6 in squid species.
Two variants (variants 2 and 3) also exhibited a 120 mer insertion within the PST domain (Figure 1). The inserted sequence was found to be specific to squid (Figure 2). This insertion might change the trans-activation activities of the PST domain. Variant 4 showed a unique insertion (57 mer) between the PD and HD. The Motif program (http://www.genome.jp/tools/motif/) found no known domains or signatures in the inserted sequence. This insertion elongates a linker between the PD and HD domains.
Discussion
Role of the IpPax-6 variants in squid eye development
In this study, we found four alternative splicing variants of IpPax-6 in the pygmy squid. To examine the role of the IpPax-6 in squid eye development, we first confirmed the expression of splicing variants in four developmental stages of the pygmy squid embryo and four distinct tissues. We then predicted the 3D structure of the splicing variants to test the functionality of the DNA-binding domain. The 3D structure prediction suggests that two of the squid Pax-6 variants with a deletion in the HD (variants 1 and 3) have distinct DNA-binding activities from that of authentic IpPax-6 due to the probable loss of HD stability. The variants with a deletion in the HD domain were expressed at the early stage of squid embryonic development and in the eyeball of adult squids. The variants were also specifically localized in the membrane surrounding the eyes, which corresponds to the future eye reflector at the middle stage of eye development. This localization of the variants strongly implies that the alternatively spliced variants play a role in the trans-activation of eye structural proteins. The target of the activation remains unknown, but the eye reflector protein, reflectin21, is one of a number of candidates. It has been reported that the ectopic expression of ey ΔHD was able to induce the formation of a compound eye in Drosophila12. The DNA-binding activity of the PD domain is sufficient to induce ectopic eyes in Drosophila through activation of the retinal determination gene network. The tissue specificity of ΔHD variants in the squid and the role of the PD domain shown in Drosophila strongly suggest that the Pax-6 variants play a part in the placement of the eyes. In addition to its influence on DNA-binding activity, protein-protein interaction between Pax-6 and other proteins is clearly important for changing gene regulatory networks in eye development. In fact, after the formation of the optic disc controlled by Pax-6, cell lineages in the eye will be determined by co-localization of Pax-6 and its partner transcription factors; Sox/Six/Rx. However, there is no existing three-dimensional structure of proteins that interact with Pax-6 and it is difficult to estimate a protein-protein interaction between Pax-6 splicing variants and other proteins.
We also found that the squid Pax-6 gene uniquely acquired an additional exon in the PST domain. Previous reports have suggested that vertebrate Pax-6 is a dedicated transcriptional activator2,21,22,23,24,25 and several reports have proposed the PST domain as a trans-activation domain. Weasner et al.7 demonstrated that the expression of mutant Ey and Toy proteins lacking the non-conserved PST segments failed to induce ectopic eye formation in flies. One of the differences between ey and toy has been attributed to the PST domain, which interacts with disparate binding partners13. Thus, the insertion of the polypeptide encoded by exon 6 into the PST domain in the IpPax-6 variants is likely to change the transactivation activity of that domain.
The linker between the PD and HD domains in the variant protein encoded by the transcript with exon 3 was longer than that in authentic Pax-6. The length of this linker is well known to vary among vertebrates and invertebrates. This linker region and almost one-half of the following PST domain are contained in an intrinsically disordered region, which is a frequent feature of most transcription factors26. This variant was specifically found in the adult eyeballs. These two results suggest the conjecture that the insertion into the linker region changes the transcriptional activity through disruption of the interactions between Pax-6 and other molecules and that the target DNA sequence may also be changed by elongation of the linker between the PD and HD domains. This speculation is corroborated by our findings showing that the IpPax-6 variants had significant tissue- and embryonic stage-specificities.
Acquisition of the Pax-6 variants in eye formation
As found in several bilaterian species, the basic function of animal Pax-6 appears to involve the regulation of the anterior part of the head and the initiation of photoreceptor formation, regardless of the photoreceptor cell type. In insect Pax-6, ey has at least three additional variants generated through gene duplication. The duplicates are thought to have originated in a pancrustacean ancestor (insect + crustacea), although the exact origin remains uncertain9. The roles of two of these duplicates, toy and eyg, have diverged in the compound eye formation processes of the fly and beetle27,28 and functional changes in toy and eyg have significantly contributed to the evolution of the size and shape of the eyes in the beetle27. These results indicate that the diversification of the Pax6 isoforms by duplication is required for eye development in insects. Diversification of Pax-6 variants is also found in vertebrates. However, the acquisition of variations in Pax-6 is completely distinct between these two taxa; vertebrate Pax-6(5a) originated from the alternative splicing of the locus, whereas insect eyg originated from gene duplication29. Pax-6(5a) shows functional similarity to the Drosophila eyg, which also lacks the PD required for HD function30. In mice, another type of protein variation, protein sumoylation onto the Pax-6 splicing variants, is expressed in the embryonic optic and lens vesicles and shows different DNA-binding and transcriptional activities from the authentic protein3. The bilaterian common ancestor (UBA) is considered to have had a Pax-6 gene for the development of photoreceptors (Figure 4). Thus, eye development has been achieved by alternative splicing or gene duplication, either of which leads to variations in the protein isoforms.
Schematic views of Pax-6 variants from metazoan animals.
Squid was shown to have Pax-6 splicing variants involved in the eye formation processes. Three Pax6 genes are known to be involved in insect compound eye development. Although it is difficult to determine ey, toy and eyg duplication events precisely, toy and eyg are known to play roles independent to ey in compound eye formation in flies. In mammals, it has been demonstrated that at least four forms of Pax-6 are generated by alternative splicing and sumoylation. UBA, urbilaterian ancestor.
Authentic IpPax-6 is the major form in terms of expression intensity and ubiquity in tissues, whereas the variants 1 and 3, which lack exon 4, were exclusively expressed in the eye reflector. Based on gene expression studies, an authentic transcript tends to be maintained the major function, whereas a minor transcript tends to be free from selective pressure and ready to acquire a novel function31. Therefore, these minor variants might be responsible for the acquisition of novel tissue specificity and target protein. A similar case can be observed in the lineage-specific alternative splicing events of vertebrate Pax-6(5a), which plays a conspicuous role in postnatal iris formation and the structural integrity of the cornea, lens and retina.
In summary, we identified the acquisition of splicing variations of Pax-6 in cephalopod eyes (Figure 4) and found that the acquisition occurred independently in vertebrates and cephalopods. These Pax-6 splicing variations in cephalopods were controlled spatio-temporally during eye formation. Although the acquisition of camera eyes in the cephalopods is yet a problem, Pax-6 variants in cephalopods have been acquired in a lineage-specific manner.
Methods
Sample Collection and RNA Extraction
Specimens of the pygmy squid, Idiosepius paradoxus, were collected from Chita Peninsula, Nagoya, Japan and kept in a laboratory, as previously described32, where the embryos were staged17. Eggs of Loligo bleekeri were obtained from Shimoda Bay, Kanagawa, Japan and kept in a laboratory until hatching. Total RNA was extracted from RNA later-fixed tissue using a Molluscan RNA kit (Omega bio-tech) with on-column DNase digestion. Tissue collection was performed using seawater containing 2% ethanol.
Gene Cloning
Pygmy squid Pax-6 cDNAs were obtained by 5′- and 3′-RACE PCR using a SMARTer RACE kit (Takara) and then sub-cloned into a T-vector (Promega). RT-PCRs were performed using squid embryonic and adult RNAs to confirm the alternative splicing forms. The RNAs were reverse transcribed into cDNA using PrimeScript® RT Reagent (Takara) and amplified using Advantage2 PCR (Takara) kits. The primers used in the cloning steps are listed in Table S1. The nucleotide sequences obtained in this study are available under the following accession numbers: [DDBJ: AB716343-AB716348].
Comparative Modeling for Authentic Pax-6 and its Variant Lacking Exon 3
Three-dimensional structures of the HD of the authentic Pax-6 gene of I. paradoxus and its variant lacking the segment coded by exon 3 were modeled based on the structure of the Drosophila melanogaster Aristaless HD (PDB ID: 3lnq)33. The template structure had 58 residues and the alignments were built without gaps. The sequence identity between the authentic form and the template was 74% and that between the variant and the template was 59%. Three-dimensional structures of each of the sequences were built using MODELLER34 with five different random number seeds and the structure with the lowest DOPE energy was chosen as the best model.
Real-Time PCR
Real-time RT-PCR expression analysis was carried out using iQ5 (Bio-Rad) and THUNDERBIRD™ SYBR® qPCR Mix (Toyobo). Template cDNAs were obtained using a PrimeScript® RT reagent kit (Takara). Primers are listed in Table S2. A control gene (alpha-tubulin) was selected according to the method of Sirakov et al.35.
In Situ Hybridization
DIG-labeled RNA probes targeting exons 2 and 4 were made using an in vitro transcription kit. The primer pairs used to generate exon 2- and 4-targeted probes are listed in Table S3. RT-PCR fragments obtained from previous primers were sub-cloned into a T-vector (Promega) and used as templates for in vitro transcription. Whole-mount in situ hybridization was performed according to the protocol published previously32.
Molecular Phylogenetic Tree Analysis
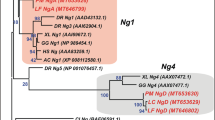
The multiple alignment of the Pax gene family of metazoan animals was built based on the full-length amino acid sequences of the proteins using the clustalW2 program36. We inferred the molecular phylogenetic trees of the Pax gene family by the maximum likelihood (ML) method using the RAxML program37 with the WAG model of amino acid substitutions and the discrete Γ distribution of heterogeneity of evolutionary rates among sites. Topology searches were performed using the fast bootstrap version of the RAxML program, which analyzes 200 bootstrap replicates.
References
Epstein, J. A. et al. Two independent and interactive DNA-binding subdomains of the Pax-6 paired domain are regulated by alternative splicing. Genes Dev. 8, 2022–2034 (1994).
Singh, S. et al. Iris hypoplasia in mice that lack the alternatively spliced Pax-6(5a) isoform. Proc Natl Acad Sci USA. 99, 6812–6815 (2002).
Yan, Q. et al. Sumoylation activates the transcriptional activity of Pax-6, an important transcription factor for eye and brain development. Proc Natl Acad Sci USA. 107, 21034–21039 (2010).
Richardson, J., Cvekl, A. & Wistow, G. Pax-6 is essential for lens-specific expression of zeta-crystallin. Proc Natl Acad Sci USA. 92, 4676–4680 (1995).
Carriere, C. et al. Characterization of quail Pax-6 (Pax-QNR) proteins expressed in the neuroretina. Mol Cell Biol. 13, 7257–7266 (1993).
Czerny, T. et al. twin of eyeless, a second Pax-6 gene of Drosophila, acts upstream of eyeless in the control of eye development. Mol Cell. 3, 297–307 (1999).
Weasner, B. M. et al. Transcriptional activities of the Pax-6 gene eyeless regulate tissue specificity of ectopic eye formation in Drosophila. Dev Biol. 334, 492–502 (2009).
Jun, S. et al. Lune/eye gone, a Pax-like protein, uses a partial paired domain and a homeodomain for DNA recognition. Proc Natl Acad Sci USA. 95, 13720–13725 (1998).
Rivera, A. S. et al. Gene duplication and the origins of morphological complexity in pancrustacean eyes, a genomic approach. BMC Evolutionary Biology. 10, 123 (2010).
Chao, J. L., Tsai, Y. C., Chiu, S. J. & Sun, Y. H. Localized Notch signal acts through eyg and upd to promote global growth in Drosophila eye. Development. 131, 3839–3847 (2004).
Dominguez, M. et al. Growth and specification of the eye are controlled independently by Eyegone and Eyeless in Drosophila melanogaster. Nat Genet. 36, 31–39 (2004).
Punzo, C., Kurata, S. & Gehring, W. J. The eyeless homeodomain is dispensable for eye development in Drosophila. Genes Dev. 15, 1716–1723 (2001).
Punzo, C. et al. Functional divergence between eyeless and twin of eyeless in Drosophila melanogaster. Development. 131, 3943–3953 (2004).
Yao, J. G. & Sun, Y. H. Eyg and Ey Pax proteins act by distinct transcriptional mechanisms in Drosophila development. EMBO J. 24, 2602–2612 (2005).
Gehring, W. J. & Ikeo, K. Pax 6: mastering eye morphogenesis and eye evolution. Trends Genet. 15, 371–377 (1999).
Tomarev, S. I. et al. Squid Pax-6 and eye development. Proc Natl Acad Sci USA. 94, 2421–2426 (1997).
Yamamoto, M. Normal embryonic stages of the pygmy cuttlefish, Idiosepius pygmaeus paradoxus Ortmann. Zool Sci. 5, 989–999 (1988).
Hartmann, B. et al. Pax-6 in the sepiolid squid Euprymna scolopes: evidence for a role in eye, sensory organ and brain development. Mech Dev. 120, 177–183 (2003).
Simakov, O. et al. Insights into bilaterian evolution from three spiralian genomes. Nature. 493, 526–31 (2013).
Takeuchi, T. et al. Draft genome of the pearl oyster Pinctada fucata: a platform for understanding bivalve biology. DNA Res. 19, 117–30 (2012).
Crookes, W. J. et al. Reflectins: the unusual proteins of squid reflective tissues. Science. 303, 235–238 (2004).
Plaza, S., Dozier, C. & Saule, S. Quail Pax-6 (Pax-QNR) encodes a transcription factor able to bind and trans-activate its own promoter. Cell Growth Differ. 4, 1041–1050 (1993).
Glaser, T. et al. Pax-6 gene dosage effect in a family with congenital cataracts, aniridia, anophthalmia and central nervous system defects. Nature Genet. 7, 463–471 (1994).
Czerny, T. & Busslinger, M. DNA-binding and transactivation properties of Pax-6: Three amino acids in the paired domain are responsible for the different sequence recognition of Pax-6 and BSAP (Pax-5). Mol Cell Biol. 15, 2858–2871 (1995).
Altmann, C. R., Chow, R. L., Lang, R. A. & Hemmati-Brivanlou, A. Lens induction by Pax-6 in Xenopus laevis. Dev Biol. 185, 119–123 (1997).
Minezaki, Y., Homma, K., Kinjo, A. R. & Nishikawa, K. Human transcription factors contain a high fraction of intrinsically disordered regions essential for transcriptional regulation. J Mol Biol. 359, 1137–1149 (2006).
Yang, X. et al. Probing the Drosophila retinal determination gene network in Tribolium (II): The Pax-6 genes eyeless and twin of eyeless. Dev Biol. 333, 215–227 (2009).
Zarinkamar, N. et al. The Pax gene eyegone facilitates repression of eye development in Tribolium. Evodevo. 2, 8 (2011).
Yao, J. G. et al. Differential requirements for the Pax-6(5a) genes eyegone and twin of eyegone during eye development in Drosophila. Dev Biol. 315, 535–551 (2008).
Mishra, R. et al. Pax-6, paired domain influences sequence recognition by the homeodomain. J Biol Chem. 277, 49488–49494 (2002).
Kim, E., Magen, A. & Ast, G. Different levels of alternative splicing among eukaryotes. Nucleic Acids Res. 35, 125–131 (2007).
Yoshida, M., Shigeno, S., Tsuneki, K. & Furuya, H. Squid vascular endothelial growth factor receptor: a shared molecular signature in the convergent evolution of closed circulatory systems. Evol Dev. 12, 25–33 (2010).
Miyazono, K. et al. Cooperative DNA-binding and sequence-recognition mechanism of aristaless and clawless. EMBO J. 29, 1613–1623 (2010).
Sanchez, R. & Sali, A. Evaluation of comparative protein structure modeling by MODELLER-3. Proteins. 29, 50–58 (1997).
Sirakov, M. et al. Selection and validation of a set of reliable reference genes for quantitative RT-PCR studies in the brain of the Cephalopod Mollusc Octopus vulgaris. BMC Mol Biol, 10, 70 (2009).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics. 23, 2947–2948 (2007).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 22, 2688–2690 (2006).
Acknowledgements
We are truly grateful to Ms. S. Takagi at Ochanimizu University for her assistance with the analyses. We also thank Dr. T. Kasugai for his kind help with the pygmy squid collection and to Dr. N. Hirohashi for his kind provision of Loligo eggs. This study was supported by the Program to Disseminate Tenure Tracking System of the Ministry of Education, Culture, Sports, Science and Technology (MEXT) and a Grant-in-Aid for Young Scientists (B). K.Y. was supported by the Platform for Drug Discovery, Informatics and Structural Life Science program from MEXT.
Author information
Authors and Affiliations
Contributions
A.O. proposed the concept. M.Y. and A.O. designed the research. K.Y. computed the three-dimensional structure of the gene. M.Y. conducted the experiments and analysis. All authors wrote the main manuscript text and prepared figures. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Yoshida, Ma., Yura, K. & Ogura, A. Cephalopod eye evolution was modulated by the acquisition of Pax-6 splicing variants. Sci Rep 4, 4256 (2014). https://doi.org/10.1038/srep04256
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04256
This article is cited by
-
Validation of Reference Genes for Quantitative Gene Expression Studies in Octopus minor
Ocean Science Journal (2020)
-
Ancestral and novel roles of Pax family genes in mollusks
BMC Evolutionary Biology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.