Abstract
Peptide ligand-induced dimerization of the extracellular region of the epidermal growth factor receptor (sEGFR) is central to the signal transduction of many cellular processes. A small molecule microarray screen has been developed to search for non-peptide compounds able to bind to sEGFR. We describe the discovery of nitro-benzoxadiazole (NBD) compounds that enhance tyrosine phosphorylation of EGFR and thereby trigger downstream signaling pathways and other receptor tyrosine kinases in cancer cells. The protein phosphorylation profile in cells exposed to NBD compounds is to some extent reminiscent of the profile induced by the cognate ligand. Experimental studies indicate that the small compounds bind to the dimerization domain of sEGFR and generate stable dimers providing allosteric activation of the receptor. Moreover, receptor phosphorylation is associated with inhibition of PTP-1B phosphatase. Our data offer a promising paradigm for investigating new aspects of signal transduction mediated by EGFR in cancer cells exposed to electrophilic NBD compounds.
Similar content being viewed by others
Introduction
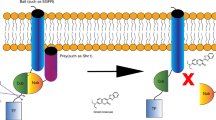
The epidermal growth factor receptor (EGFR) is a membrane-spanning protein that governs major signaling pathways and therefore its overexpression and deregulation have a severe impact on cells, resulting in aggressive tumor growth1. The binding of natural peptide ligands to domains I and III of the extracellular region of EGFR (sEGFR) induces topological rearrangements, exposing the dimerization domain II of two monomers in a conformation favorable for them to associate and form functionally active homodimers or heterodimers with a similar ligand-less ErbB2 or peptide ligand-bound ErbB3 and ErbB42,3,4,5,6. This specific ligand-induced dimerization is responsible for distinct allosteric changes in the cytoplasmic tyrosine kinase domain of EGFR, which lead to direct contacts between the C-lobe and N-lobe required to activate the ATP-binding site and create appropriate docking sites for the recruitment of various effector proteins7,8,9. The phosphorylated EGFR induced by peptide ligands or cytoplasmic proteins undergoes endocytosis and further degradation in cells10.
However, other investigations have shown that dimerization and/or activation of EGFR can also be promoted by non-ligand-bound mechanisms. For example, cytohesins have been demonstrated to behave as cytoplasmic activators of EGFR in human lung adenocarcinoma11. In addition, some point mutations located in the EGFR kinase domain activate auto-phosphorylation of the receptor7,12 and small molecules bound to the ATP-binding site can cause reversible dimerization of the kinase domain and affect TGF-α-induced tyrosine phosphorylation13. Moreover, hydrogen peroxide induces EGFR phosphorylation14,15,16 as proven recently by sulfenylation of the ATP-binding site of the receptor17.
As dimerization plays a key role in the phosphorylation of the receptor, the sEGFR dimerization interface is of huge potential interest for identifying new molecular interactions affecting receptor functions and for a better understanding of the complexity of its behavior in healthy and diseased cells. Small molecule microarrays have opened up a new way for rapid and high throughput screening of compound libraries against desired proteins18. Both chemical and photochemical reactions have been applied to use reactive moieties in different compounds as a means of coupling to functionalized plane surfaces19,20. In this study, we have developed a new microarray screen to detect chemical compounds that bind to the dimerization domain of sEGFR. We have identified compounds enhancing tyrosine phosphorylation of the receptor in cancer cells. Our data indicate that compounds containing the nitro-benzoxadiazole ring can bind to the dimerization domain and allosterically activate the receptor and thereby trigger downstream and lateral signal transduction.
Results
Screening compound library with small molecule microarrays
The strategy of searching for compounds that bind to the sEGFR dimerization domain II and modulate EGFR tyrosine phosphorylation is shown in Fig. 1. First, it entails preparing planar microarrays, representing a structural diversity of 1,364 preselected potential pharmacophores (Diversity Set II library of the National Cancer Institute), by non-covalent immobilization of all compounds on a new formulated hydrogel support. This non-biased immobilization approach enabled us to avoid the chemical reactions usually required to couple the compounds of interest covalently to a functionalized surface, thus making all the moieties of the compounds being tested potentially accessible to a given protein target. Secondly, since protein-protein interaction surfaces, including the protruding dimerization loop, are hidden in the tethered ligand-unbound conformation of the monomeric form of EGFR2,3,4,21, we took advantage of the domain organization of sEGFR to construct a shortened protein, thereby providing small molecule interactions with the whole surface of the dimerization domain II. Thirdly, we used near-infrared fluorescence detection to reduce the interference from auto-fluorescent signals emitted by heterocyclic rings of small molecules at visible wavelengths22,23.
Schema of compound library screening with microarrays and identification of small molecules enhancing protein tyrosine phosphorylation of EGFR.
The structure of the sEGFR is shown in a tethered conformation of four domains: I (yellow), II (green), III (gray) and IV (red). The histogram shows competitive assay data obtained for three selected compounds (for NSC 228155 - column 1). The signal monitored from binding of each molecule to sEGFR (gray column) was used as 100% to assess the binding efficiency to sEGFR in competition with DII/sEGFR (brown column). Protein tyrosine phosphorylation was assessed in MDA MB468 cells exposed to the compounds at 20 μM final concentration for 60 min at 37°C. The proteins were analyzed with anti-pTyr P100 antibody. Lane 1: cells exposed to NSC 228155; lanes 2 and 3: cells exposed to two other selected compounds; lane 4: untreated cells; lane 5: cells treated with 150 ng/ml EGF for 10 min; MM: molecular mass markers, kDa.
The human DNA region corresponding to domain II of sEGFR was sub-cloned from the ultimate ORF clone IOH62670 and overexpressed in E. coli cells (Supplementary Fig. 1). The resulting truncated protein DII/sEGFR formed exclusively inclusion bodies in the heterologous bacterial host. Therefore, the cell extracts were treated under conditions that prevent protein misfolding24. The purified domain II-specific protein was then labeled with IRDye 800CW and probed to the Diversity Set II library on small molecule microarrays.
20 compounds selected from the first screening were spotted in quadruplicate on the hydrogel support and analyzed in a competitive assay between unlabeled DII/sEGFR and IRDye-labeled sEGFR proteins mixed in an 8:1 ratio in the binding buffer. This second screening was intended to help distinguish specific binding signals from false positives. A significant decrease in the fluorescent signals was detected for three compounds (see Fig. 1) suggesting that they might specifically bind to domain II of sEGFR.
Identification of nitro-benzoxadiazole compounds enhancing tyrosine phosphorylation of EGFR
To explore the ability of the selected compounds to act as potential modulators (both activators and inhibitors) of EGFR activity, we examined the effect of each compound on total protein tyrosine phosphorylation in MDA MB468 breast cancer cells known to overexpress the receptor. Two compounds had no effect on protein tyrosine phosphorylation, whereas compound NSC 228155, 4-nitro-7-[(1-oxidopyridin-2-yl)sulfanyl]-2,1,3-benzoxadiazole (Fig. 2) significantly enhanced the phosphorylation of several proteins and notably that of a large protein with a migration velocity close to that of EGFR as compared to EGF-treated cells (see Fig. 1). Immunoblotting analysis with anti-EGFR and anti-phospho-EGFR pTyr1068 antibodies confirmed that this protein was EGFR.
Structure of NBD compounds NSC 228155 and CN 009543V.
Compound CN 009543V was identified with a Tanimoto similarity coefficient > 0.75 using 2,1,3-benzoxadiazole sub-structure NSC 228155 for searching. Values of cLogP were calculated with free module Marvin Sketch 5.4.1.1 (www.chemaxon.com).
Further immunoblotting studies demonstrated that tyrosine phosphorylation of EGFR was clearly detectable after exposure of MDA MB468 cells to NSC 228155 for 5 minutes and the rate of protein phosphorylation gradually increased with longer exposure times (Supplementary Fig. 2). Meanwhile, the total amount of the receptor (both non-phosphorylated and phosphorylated) decreased at 60 min of exposure to NCS 228155 indicating that the phosphorylated EGFR had undergone degradation and/or down-regulation, as was previously documented for the receptor in EGF-induced cells10.
The effect of the compound was dose-dependent, judging by the enhanced phosphorylation of two EGFR tyrosine residues tested, Tyr1068 (Supplementary Fig. 3) and Tyr1173 (Supplementary Fig. 4). The EC50 value of the compound was evaluated as 52 μM with respect to the phosphorylated Tyr1068.
The protein phosphorylation response to exogenous signals depends on the type of cancer25, so it was interesting to assess the effect of NSC 228155 on EGFR in another cancer. For this purpose, we used the NSCLC-N6-L16 cell line, obtained from a non-small cell lung cancer patient in our laboratory26, in which the expression level of the receptor was found to be more than 100-fold lower than in MDA MB468 cells. Compound NSC 228155 enhanced both total tyrosine phosphorylation and phosphorylation at Tyr1068 and Tyr1173 residues of EGFR. However, the activation effect was lower, apparently because of less abundant receptors in the NSCLC-N6-L16 cells (Supplementary Fig. 5).
To understand better the way in which small compounds enhance tyrosine phosphorylation of EGFR in cancer cells, we were interested in studying other molecules sharing a structural similarity with NSC 228155. Such a compound, CN 009543V, methyl 2-(acetylamino)-3-[(7-nitro-2,1,3-benzoxadiazol-4-yl)sulfanyl]propanoate (see Fig. 2) was identified in the French National Chemical Library and studied in parallel assays with NSC 228155. This new compound also enhanced tyrosine phosphorylation of EGFR in the treated MDA MB468 cells (even at higher extent than NSC 228155 at the same concentration), suggesting that the electrophilic structure of nitro-benzoxadiazole (NBD) could play an important role in the induction of EGFR phosphorylation.
Kinase inhibitors prevent enhanced tyrosine phosphorylation of the receptor
Tyrosine kinase inhibitors AG1478 and PD 153035 are known to inhibit selectively the ATP-binding pocket of EGFR and not ErbB227,28. To find out whether the enhanced tyrosine phosphorylation of EGFR caused by NBD compounds was related to activation of the receptor's kinase domain, the serum-starved cells were first incubated with 10 μM AG1478 or 2 μM PD 153035 for 90 min and then exposed to 100 μM NSC 228155 or CN 009543V for 15 min in the same medium. Western blotting analysis showed that the kinase inhibitors completely prevented both total tyrosine and Tyr1068 phosphorylation of EGFR (Fig. 3a). Similar inhibition was detected in the cells exposed to 150 ng/ml EGF used as a control. Hence, in the MDA MB468 cells exposed to NBD compounds, the ATP-binding site of EGFR adopts an appropriate active conformation required for receptor phosphorylation.
EGFR tyrosine kinase inhibitors prevent tyrosine phosphorylation of the receptor (a) and protein phosphorylation in downstream signaling pathways (b) in cancer cells exposed to compounds NSC 228155 or CN 009543V.
MDA MB468 cells serum-starved overnight were pre-incubated (or not) with 10 μM AG1478 or 2 μM PD 153035 for 90 min and then, where indicated, incubated with 100 μM NSC 228155 or CN 009543V or 150 ng/ml EGF or vehicle (0.2% DMSO) for 15 min.Proteins were blotted to nitrocellulose membrane and analyzed with biotinylated anti-pTyr P100, anti-pEGFR Y1068 and anti-EGFR (epitope in cytoplasmic region) antibodies. The phospho-specific antibodies were anti-phospho-Shc (Tyr239/240), anti-phospho-p44/42 MAPK (ERK1/2) (Thr202/Tyr204), anti-phospho-c-CBL (Tyr774) and anti-Gab1 (Tyr627). The signal intensity of the α-tubulin band was quantitatively evaluated and used as the loading control of protein samples.
NBD compounds trigger protein phosphorylation in EGFR downstream signaling pathways
As mentioned above, the enhanced tyrosine phosphorylation of various molecular weight proteins was also observed in Western blots after longer exposure of cells to NBD compounds (see Fig. 1), suggesting activation of signaling pathways downstream of EGFR. To verify this assumption and to discriminate between signal transduction initiated by EGFR and other receptor tyrosine kinases, we compared the phosphorylation state of some key proteins in the same cell samples, pre-incubated or not with the EGFR-specific kinase inhibitor AG1478.
It is known that activation of the MAPK signaling pathway occurs via auto-phosphorylation at EGFR Tyr1173, which serves as a docking site for the scaffold protein Shc in EGF-induced cells29. Incubation of MDA MB468 cells with NSC 228155 or CN 009543V clearly enhanced tyrosine phosphorylation of Tyr239/Tyr240 residues in 70-kDa and especially in 55-kDa Shc proteins and the rate of protein phosphorylation was comparable to that in cells exposed to EGF (Fig. 3b). However, pre-incubation with AG1478 completely prevented Shc phosphorylation in cells exposed to NBD compounds or to EGF. Similarly, compounds NSC 228155 and CN 009543V induced phosphorylation at Thr202/Tyr204 of 42-kDa ERK1 and 44-kDa ERK2 proteins as did EGF in treated cells (see Fig. 3B) confirming data obtained in another assay (see Supplementary Fig. 4). The inhibitory action of AG1478 toward ERK1/2 phosphorylation was relatively low in cells exposed to small compounds or EGF. These data indicated that NBD compounds induce the MAPK pathway in MDA MB468 cells.
A 90-kDa c-CBL was weakly phosphorylated at Tyr774 in cells exposed to NBD compounds and a similar low level of phosphorylation of this protein was detected in the EGF-exposed cells (see Fig. 3b). In all cases, this phosphorylation was blocked by AG1478. Although weak, small compound-induced post-translational modification of the adaptor protein c-CBL, involved in the ubiquitination/degradation pathway29,30, is in agreement with the degradation of EGFR after longer treatment of the cells with NSC 228155 (see Supplementary Fig. 2).
Tyrosine phosphorylation was also weakly enhanced at Tyr627 of a 60-kDa Gab1 in cells exposed to NSC 228155 or CN 009543V compared to a higher phosphorylation rate of the protein in cells exposed to EGF (see Fig. 3b). Again, in all three cases, AG1478 prevented Tyr627 phosphorylation of Gab1, the protein involved in binding to and activation of the tyrosine phosphatase SHP231, which directly associates with EGFR through its SH2 domain32.
Compound NSC 228155 promotes transactivation of other RTKs
External signals recognized by EGFR can be transduced to other RTKs by transactivation. To assess tyrosine phosphorylation of different RTKs in cancer cells exposed to NBD compounds, we turned to antibody arrays as a more sensitive method than Western blotting.
Significant tyrosine phosphorylation was recorded for EGFR and a relatively lower response was recorded for Insulin R and anti-EphA1 receptors in the MDA MB468 cells exposed to NSC 228155 for 5 min (Fig. 4a and 4c). The phosphorylation rate of these proteins increased after a 10-min exposure. Moreover, five other RTKs, namely ErbB2, ErbB3, IGF-I R, Mer and ROR1, were phosphorylated at this point, suggesting that signal transduction induced by NSC 228155 was distributed among other RTKs. It is worth noting that, except for ROR1, the other RTKs were also illuminated in cells exposed to EGF for 10 min (Fig. 4b and 4c). However, although the tyrosine phosphorylation response of EGFR to its natural ligand was at the same level as to NSC 228155, the response to EGF was clearly weaker from other RTKs as compared to the action of small compounds under the conditions used.
Time-dependent activation of RTKs in MDA MB468 cells exposed to NSC 228155 (a, c) and comparison of protein tyrosine phosphorylation in cells exposed to NSC 228155 versus EGF for 10 min (b, c).
Tyrosine phosphorylation levels of RTKs were assessed after 5-min and 10-min exposure to 100 μM NSC 228155 or 10-min exposure to 150 ng/ml EGF. Histograms show the arbitrary level of protein tyrosine phosphorylation determined in NCS 228155 (white column) and EGF (gray column) treated cells. The rate of tyrosine phosphorylated RTKs was quantified by normalization with control spots according to the manufacturer's protocol (R&D Systems).
Thus, the protein phosphorylation profile of the downstream signaling cascade induced by NS C228155 or CN 009543V as well as the phosphorylation profile of RTKs induced by NSC 228155 in breast cancer cells are to some extent reminiscent of the profiles induced by the cognate ligand EGF suggesting that the NBD compounds behave as allosteric activators at EGFR.
Anti-EGFR neutralizing antibody prevents tyrosine phosphorylation by NBD compounds
Peptide ligand-induced dimerization is considered a canonical process required for allosteric modification and catalytic activation of the receptor33. Therefore, blocking ligand-recognition sites in sEGFR domains I and III with neutralizing antibodies interrupts the dimerization process and inhibits the kinase activity of the receptor.
We compared the action of a neutralizing anti-EGFR antibody LA1 on the receptor tyrosine phosphorylation in MDA MB468 cells exposed to NBD compounds and to EGF. The serum-starved cells were incubated with antibody for 90 min and then exposed to NSC 228155 or CN 009543V or EGF for 15 min in the same medium. Pre-incubation of the cells with the neutralizing antibody significantly, but not completely, decreased total tyrosine phosphorylation and Tyr1068 phosphorylation of EGFR in cells exposed to NBD compounds whereas the antibody completely abolished tyrosine phosphorylation of the receptor in cells exposed to EGF (Fig. 5). The strong inhibition of tyrosine phosphorylation by the neutralizing antibody indicated that the induction mechanism of EGFR by NBD compounds involves the dimerization process of the extracellular region of the receptor.
The neutralizing antibody LA1 in cancer cells prevents NBD compound-promoted EGFR tyrosine phosphorylation.
MDA MB468 cells serum-starved overnight were pre-incubated (or not) with 50 μM anti-EGFR mAb LA1 for 90 min and then, where indicated, treated with 100 μM NSC 228155 or CN 009543V or 150 ng/ml EGF for 15 min.
Induced tyrosine phosphorylation of EGFR is associated with inhibition of protein tyrosine phosphatase
Activation of EGFR in cells exposed to NBD compounds could alternatively originate from inhibition of protein tyrosine phosphatases, which exhibit large substrate specificity. In particular, PTP-1B, known to dephosphorylate EGFR34,35, is inhibited by extracellular hydrogen peroxide generated at EGF interaction with the extracellular region of the receptor36,37.
We measured PTP-1B activity in cell extracts of MDA MB468 cells after buffer exchange of a soluble protein fraction with tyrosine phosphatase assay buffer (see Methods), which provided a better performance than desalting of the samples to be analyzed. It was found that the activity of PTP-1B decreased in the cells exposed to NSC 229155 and CN 009543V by almost 29% and 62%, respectively, as compared to the cells incubated with a vehicle (Fig. 6a, left). Lower activity of PTP-1B was also detected in the cells induced by EGF in agreement with other observations37.
Inhibition of PTP-1B phosphatase is associated with tyrosine auto-phosphorylation of EGFR in cancer cells exposed to NBD-compounds.
Histograms of PTP-1B (a) and total phosphatase (b) activities and images of monomeric and dimeric forms of EGFR detected with Western blot (c). MDA MB468 cells were pre-incubated with a vehicle (0,2% DMSO) or 2 μM PD 153035 for 2 h and then incubated with a vehicle or 500 ng/ml EGF or 100 μM of each of NSC 228155 or CN 009543V for 15 min at 37°C. Buffer-exchanged extracts were used for assays. Protein tyrosine phosphatase activity was measured by dephosphorylation of phosphopeptide substrate. Total phosphatase activity was measured by hydrolysis of para-nitrophenyl phosphate (pNPP) and expressed in % of the enzyme activity in extracts of the cells incubated with a vehicle (taken as 100%). Levels of PTP-1B and total phosphatase activities were normalized to total protein concentration in the extracts to be assayed. Detection of EGFR was carried out with anti-EGFR antibody recognizing the cytoplasmic domain ((c), upper image) and anti-pEGFR Y1068 antibody ((c), lower image). Electrophoretic migration of proteins was carried out in different gels; it was longer for Western blotting with anti-pEGFR Y1068 antibody.
To find out whether this inhibition was associated with the activation of EGFR by NBD compounds, the enzyme assays were carried out with cells pre-incubated with PD 153035. At 2 μM concentration of the kinase inhibitor, the activity of PTP-1B in cells treated with NSC 228155 or CN 009543V remarkably increased, close to levels observed for cells incubated with a vehicle (Fig. 6a, right). Similarly, an increase in the protein tyrosine phosphatase activity was detected in EGF-induced cells. Meanwhile, no phosphorylation of EGFR at Tyr1068 was observed in cells pre-incubated with PD 153035 and then treated with NBD compounds or EGF (Fig. 6c). Hence, abolition of the kinase activity of EGFR by a specific tyrosine kinase inhibitor prevented the inhibitory effect of PTP-1B to dephosphorylate tyrosine residues of the receptor in the cells exposed to NBD compounds or to the cognate ligand.
We also measured total phosphatase activity in the buffer-exchanged extracts of MDA MB468 cells. No diminution of the activity was detected in the cells treated with EGF or NSC 228155, whereas the level of p-nitrophenol formed in the cells exposed to CN 009543V was almost 30% lower compared to the cells treated with a vehicle (Fig. 6b, left). Pre-incubation with PD 153035 increased weakly total phosphatase activity in the cells exposed to CN 009543V suggesting that the latter also inhibits or inactivates another major phosphatase(s) that might participate in protein tyrosine dephosphorylation (Fig. 6b, right).
Thus, this data showed that tyrosine auto-phosphorylation of EGFR at Tyr1068 is mainly associated with inhibition of PTP-1B tyrosine phosphatase in cancer cells exposed to NBD compounds.
Compound NSC 228155 stimulates dimerization of sEGFR domain II
To gain further insight into the mechanism(s) of the enhanced phosphorylation of EGFR, we sought to determine whether dimeric forms of the receptor could be detected in cells lysed under soft conditions to prevent disruption of fragile protein complexes. Aliquots of extract samples used in tyrosine phosphatase assays were separated by SDS-PAGE and analyzed by Western blot. Thin and accurate bands corresponding to dimeric forms of a full-length protein were detected in the cells pre-incubated with a vehicle or PD 153035 and then exposed to NSC 228155 or CN 009543V (Fig. 6c). Weak fluorescent bands of dimers could also be detected for the phosphorylated receptor in the cells pre-incubated with PD 153035 and then incubated with NBD compounds. On the contrary, dimers of EGFR were practically undetectable in EGF-induced cells. It is worth noting that dimeric forms of the receptor were detected only in protein extracts obtained with a moderate-strength IP lysis buffer and not with a high-strength RIPA lysis buffer. Hence, NBD compounds bind to EGFR by forming more stable dimers than those induced by EGF.
Next, to extend our findings with cells, we incubated the purified domain II-specific protein DII/sEGFR with NSC 228155 at different molar ratios in non-reducing conditions and analyzed the reaction products by gel electrophoresis. As the truncated protein did not enter the non-denaturing gel, the migration was performed under denaturation conditions in the presence of 0.1% SDS and β-mercaptoethanol.
A small pool of spontaneously-formed stable dimers of DII/sEGFR, estimated at around 0.5% of the total monomer/dimer protein forms, was detected by Western blotting (Fig. 7a). It is worth mentioning that full-length EGFR dimers can be formed spontaneously, in the absence of peptide ligands, due to dynamic association and dissociation of monomers in the cells38,39,40,41. However, addition of NSC 228155 clearly increased the yield of stable dimers, by almost three times at a 16-fold molar excess of the small compound over the protein in the reaction mixture (Fig. 7b). Surprisingly, no appreciable effect was detected when NSC 228155 was incubated with the dimerization domain II of ErbB2, suggesting that NBD compound binding is provided by distinct amino acids (Fig. 7c).
Compound NSC 228155 stimulates in vitro dimerization of sEGFR domain II and not that of sErbB2.
The binding reaction was carried out in the absence of β-mercaptoethanol. Detection of dimeric proteins by Western blotting was carried out using anti-His primary antibody and DyLight 800-labeled secondary antibody (a). The curve shows the percentage of dimers in a total pool of monomeric and dimeric forms of DII/sEGFR detected at different protein-to-compound ratios (b). Sequence alignment between domain II of sEGFR and sErbB2 (c).
These results indicated that NBD compounds bind to EGFR and most probably to domain II of the extracellular region, leading to an increase in a dimeric form of the protein stable to denaturation during electrophoretic migration.
Discussion
The cognate ligand EGF binds to sEGFR at the site shared between domain I and domain III in the close (monomer) conformation of the receptor. This binding leads to fundamental structural changes by positioning two protruding dimerization loops asymmetrically face to face in two opposing monomers of the open (dimer) conformation33. In this respect, by analogy with activating modulators of seven transmembrane receptors (for a review, see42), these peptide ligands behave as natural orthosteric agonists of EGFR to trigger signaling pathways. Our data shed light on non-peptide nitro-benzoxadiazole-ring carrying compounds, some derivatives of which have been used in bio-conjugational applications43, as possible candidates for small allosteric agonists of EGFR, which activate tyrosine phosphorylation of the receptor and trigger downstream and lateral signaling in cancer cells.
Consistent with this proposal, compounds NSC 228155 and CN 009543V enhance tyrosine auto-phosphorylation at Tyr1068 and Tyr1173 leading to the activation of the MAPK/ERK cascade by involving ERK1/2, Shc and to a lesser extent Gab1. Such an action of small activators is reminiscent of that of cognate ligands44. In addition, phosphorylation of c-CBL in cells treated by NBD compounds results in the activation of the ubiquitination/degradation cascade in a way similar to EGF29,30. Lastly, NSC 228155 promotes transactivation of several RTKs, including ErbB2 and ErbB3, Insulin R and IGF-1 R receptors in the cells, which can be explained by their heterodimerization with EGFR as shown in EGF-induced cells45,46.
To put these observations into the context of the sEGFR dimerization mechanism, we postulated that the NBD compounds bind to the extracellular region and affect the dimerization process. First, compound NSC 228155 was selected by probing to a truncated protein having only domain II of sEGFR. Secondly, blockage of the orthosteric site by neutralizing antibody remarkably prevented tyrosine phosphorylation of the receptor in cancer cells exposed to NBD compounds. Thirdly, the kinase inhibitor PD 153035 prevented tyrosine auto-phosphorylation of EGFR, but not the formation of full-length dimers of the receptor in the cells exposed to NBD compounds. Fourthly, incubation of the purified domain II of sEGFR with NSC 228155 generated dimeric forms of the protein apparently stable to denaturation by SDS. These results show that NBD compounds could bind to an allosteric site(s) located on the extracellular region of EGFR and stimulate the formation of dimers as a prerequisite for further allosteric activation of the catalytic site in the cytoplasmic domain.
However, the overall amount of EGFR auto-phosphorylation in the cell is governed by both protein kinase and phosphatase activities and this can be modulated by intracellular hydrogen peroxide via different mechanisms47. Generation of exogenous H2O2 has also been demonstrated by the binding of EGF to the extracellular region of the receptor by an as yet unknown mechanism36. Exogenous H2O2 appears to enter the cells through membrane aquaporin channels48, where it inhibits the receptor-associated protein tyrosine phosphatases, including PTP-1B37 and thereby shifts the balance between kinase and phosphatase activities toward phosphorylation of EGFR. Recently, it has been shown that both intracellular and extracellular H2O2 sulfenylate Cys797 at the active site of EGFR leading to an increase in the receptor auto-phosphorylation17. Sulfenylation has also been detected in signaling phosphatases, including PTP-1B17, all containing reactive cysteine in a catalytic site49.
Our findings that both NSC 228155 and CN 009543V inhibit the activity of PTP-1B in cells, which is associated with enhancing tyrosine phosphorylation of EGFR, prompt us to suggest, by analogy with the paradigm of cognate ligand-EGFR interactions, that another mechanism is also involved in the activation of the receptor by NBD compounds. Indeed, the fact that tyrosine kinase inhibitor PD 153035 prevents auto-phosphorylation of EGFR, while removing the inhibitory effect of NBD compounds on the activity of PTP-1B, reflects the inaccessibility of Cys797 in the bound catalytic site to H2O2 generated by these compounds, rather than inactivation of the phosphatase by H2O2, or both. Recent toxicological studies of a patented anticancer compound, 7-nitro-4-(phenylthio)benzofurazan50, support this possibility. The reduction of the 7-nitro to a 7-amine in its NBD-ring rapidly generates hydrogen peroxide and superoxide in the presence of molecular oxygen and the formed reactive intermediates with electrophilic scavenging activity react with cellular structures.
Elucidation of electrophilic NBD compound interactions with proteins, resulting in various cellular responses, is at an emerging stage. NBD-ring carrying derivatives have been shown to target the active site of glutathione S-transferase (GST) and affect apoptosis through dissociation of the JNK-GSTP1-1 complex51. In the resolved GST/glutathione/NBD-hexanol structure, the NBD compound covalently attaches to the GST sulfur, forming a σ-complex52. The extracellular region of EGFR contains a large number of cysteine residues involved in intra- and inter-molecular interactions, which are potentially good targets for reactive molecules in a reducing environment. In particular, nitric oxide reversibly inhibits the tyrosine activity of EGFR by S-nitrosylation53 and, as shown recently, by targeting Cys267, Cys287 and Cys446 all located in sEGFR54. However, in vitro stimulation of dimeric forms of the truncated protein DII/sEGFR and not DII/sErbB2 (both harbor 20 cysteines at the same positions, see Fig. 7c), in a non-reducing environment, suggests that other nucleophilic amino acids might be involved in the binding to the reactive intermediates of NBD compounds. To define the chemical basis of the NBD-ring binding to sEGFR and to understand better the activation of the receptor by dimerization-mediated allosteric and secondary messenger H2O2-mediated mechanisms requires further investigation.
Despite the similar behavior of EGFR after induction by NBD compounds and EGF, our results also show differences between their actions. Peptide ligands at nanomolar concentrations induce EGFR rapidly and a delicate balance of ligand-protein interactions accurately governs signal transduction and gene expression in cells55. On the contrary, NBD compounds at higher concentrations induce EGFR relatively slowly and provoke overphosphorylation of the receptor and especially of Eph1 and Mer in MDA MB468 cells. Moreover, the enhanced tyrosine phosphorylation of ROR1 (receptor tyrosine kinase orphan receptor 1) was detected only after treatment with compound NSC 228155 and not with EGF. Expression of this protein, considered a pseudokinase56, is associated with the aggressive growth of tumor cells in breast cancer57. Therefore, further understanding the small molecule action on ROR1 phosphorylation becomes an intriguing task. We also noted that neutralizing antibody does not completely prevent tyrosine phosphorylation of EGFR in cells exposed to NBD compounds. Given the nuclear location of EGFR58, this escape from the antibody means that the lipophilic small molecules (see Fig. 2) rapidly cross the hydrophobic barrier of the cytoplasm membrane and so bind the repressor of the nuclear compartment as well. These differences show that NBD compounds cause aberrant protein phosphorylation in exposed cells.
This study highlights the vulnerability of the extracellular region of EGFR in the perception of external signals by binding of small electrophilic molecules that leads to enhanced modulation of the receptor activity in cancer cells. Our data indicate that NBD compounds, currently used as bio-conjugation agents, are of great interest for human health both to understand as yet unknown aspects of tumor development and progression and to devise alternative therapeutic strategies targeting EGFR and related signaling proteins in diseased cells.
Methods
Recombinant DNA constructions
The ultimate ORF clone IOH62670 coding EGFR (Invitrogen) and TrueORF coding ErbB2 (OriGene Technologies, Inc.) were used as templates to amplify DNA corresponding to domain II of the extracellular region of EGFR and ErbB2, respectively, using the Gateway recombination system purchased from Invitrogen (see Supplementary Fig. 1). Other DNA manipulations were performed as described previously22.
Protein purification
The plasmids carrying domain II of sEGFR or sErbB2 fused with the N-terminal His-tag were transformed into the E. coli BL21 AI strain to provide protein overexpression. Truncated proteins DII/sEGFR and DII/sErbB2 formed exclusively inclusion bodies in this bacterial host. Among different approaches tested to solubilize these proteins, the most appropriate conditions were found to incubate inclusion bodies with N-lauroylsarkosine as described by other authors24. The His-tagged proteins were purified by affinity chromatography on a Ni++-NTA resin as recommended (QIAexpressionist, Qiagen), but using imidazole and N-lauroylsarcosine for protein elution. The concentration and the purity of domain-specific proteins were determined by a capillary electrophoresis system (Agilent Technologies).
Cell lysate preparation
MDA MB468 and NSCLC-N6-L16 cells were grown in DMEM and RPMI 1640 media, respectively, with 10% fetal bovine serum (FBS) and then starved of FBS for 24 h at 37°C in a humidified atmosphere of 5% CO2 in air. For sub-culturing, the cells were rinsed twice in PBS and detached using a trypsin-EDTA solution (0.25%/0.05% v/v in HBSS) and then approximately 1 × 106 cells were incubated in 6-well culture plates. Cancer cells were treated with anti-EGFR neutralizing antibody LA1 (Millipore-Upstate), tyrosine kinase inhibitors AG1478 or PD 153035 (Merck-Calbiochem), EGF (R&D Systems) or NBD compounds at 37°C for various times as indicated in corresponding experiments. The cells were lysed with RIPA buffer in the presence of protease and phosphatase inhibitors (Pierce) and the cell-free supernatant was collected by centrifugation for further studies. Proteins were also extracted from cells with a moderate-strength lysis buffer for enzyme assays (see below). Total protein concentration was determined by the bicinchoninic acid assay (Thermo Scientific) under conditions ensuring sample component compatibility.
Small molecule microarrays
1,364 compounds of the Diversity Set II library provided by the Drug Synthesis and Chemistry Branch of NCI (http://dtp.nci.nih.gov) were spotted from 10-μM dilutions in DMSO onto a hydrogel support developed by ProtNeteomix (http://www.protneteomix.com) with a GMS 417 contacting arrayer (Affymetrix, USA) or a 32-needle manual replicator (VP Scientific, USA). The proteins were conjugated to IRDye 800CW (Li-COR Biosciences, USA) and purified from free dye as described previously59. Small molecule microarrays were incubated with protein probes at 4°C for 90 min and then rinsed with PBS. Scanning of microarrays was carried out at 21-μ pixel density and 800-nm absorbance with the Odyssey infrared imagery system (Li-COR Biosciences, USA) and the signal intensity was monitored with GenePix Pro 4.0 software (http://www.moleculardevices.com). In competition assays, the IRDye-labeled sEGFR (the recombinant protein rhEGFR purchased from R&D Systems) and unlabeled protein DII/sEGFR corresponding to domain II of sEGFR were mixed in a 1:8 ratio and incubated at 4°C for 90 min. The ratio of the fluorescent signal from spots to background signal intensity was used to assess the binding of small compounds to the protein probes. A fluorescent signal greater than 2 was accepted as a significant level of binding efficiency between spotted compound and a labeled protein. Other details of the data analysis on microarrays have been described previously59.
Western blotting
Proteins were separated on an SDS-polyacrylamide gel and transferred to a nitrocellulose membrane with a Turbo system (Bio-Rad). Antibodies used for immunoblotting were anti-pEGFR Y1068, anti-pEGFR Y1173 and anti-EGFR (cytoplasmic region) (purchased from Thermo Scientific), anti-pTyr 100 (total phosphorylated tyrosine), anti-phospho-Shc, anti-phospho-p44/42 MAPK, anti-phospho-c-CBL and anti-Gab1 (purchased from Cell Signaling). Non-phosphorylated EGFR and proteins specifically phosphorylated at given amino acids were first captured with primary antibody and then detected with fluorescent goat anti-mouse anti-IgG secondary antibody conjugated to AlexaFluor-680 (Invitrogen). Detection of total tyrosine phosphorylated proteins was carried out by capturing with the biotinylated anti-pTyr 100 antibody and then with Neutravidin-DyLight 800 (Thermo Scientific). Human anti-α-tubulin (Sigma Aldrich) was used as a loading control for fluorescence immunoblotting analysis. Fluorescent signals of protein bands were recorded at 700 and 800 nm with an Odyssey infrared imagery system. Phosphorylation of ERK1/2 (Thr202/Tyr204) and EGFR (Tyr1173) was also evaluated by chemiluminescence, and, in this case, anti-ERK2 antibody was used to monitor gel loading as described previously60. To assess the signal intensity of protein bands, values of p < 0.05 were considered statistically significant.
Antibody arrays
Profiling of tyrosine phosphorylation of 42 RTKs was performed with a human phospho-RTK array kit (R&D Systems). MDA MB468 cells were treated with NSC 228155 or EGF and cell lysates (400 μg of total protein) were loaded on antibody arrays and incubated overnight at 4°C. Phosphorylated proteins captured by the corresponding antibodies were detected with a pan anti-phospho-tyrosine antibody conjugated to horseradish peroxidase by chemiluminescence as described by the manufacturer. Phosphorylation levels of proteins were estimated by quantifying the mean pixel densities from spots using the image analysis software GenePix Pro 4.0. For each spot, local background was subtracted from pixel density and normalized by the ratio of treated/non treated samples obtained for controls. The pixel density of negative controls was subtracted from the average normalized pixel density of duplicate spots and used to generate histogram profiles of phosphorylated RTKs.
Protein tyrosine phosphatase assay
The activity of PTP-1B was evaluated with a PTP assay kit 1 (Millipore-Upstate). MDA MB468 cells were serum-starved overnight and pre-incubated with a vehicle (DMSO 0.2%) or with EGFR kinase inhibitor PD103035 (2 μM) for 2 h. Cells (2 × 106) were washed twice with PBS and treated with a vehicle or with NBD compounds NSC 228155 or CN 009543V (each 100 μM) or EGF (250 ng/ml) for 15 min at 37°C. The cultures were washed with cold TBS and rapidly lysed with Pierce® IP lysis buffer (25 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% Nonidet P-40, 1 mM EDTA, 5% glycerol) in the presence of a protease inhibitor cocktail (Sigma Aldrich) and then centrifuged at 13,000 g for 15 minutes at 4°C. A portion of the recovered supernatant was submitted to the exchange with tyrosine assay buffer (25 mM HEPES, pH 7.2, 50 mM NaCl, 5 mM dithiothreitol, 2.5 mM EDTA) on Zeba columns, 7K (Thermo Scientific) to assess protein dephosphorylation. Another portion of the supernatant, after addition of phosphatase inhibitor cocktails I and II (Sigma Aldrich), was used to assess tyrosine phosphorylation of EGFR with anti-pEGFR Y1068 antibody by Western blot. Tyrosine phosphatase activity of PTP-1B was quantified using the phosphopeptide substrate RRLIEDAEpYAARG by adding 10 mM dithiothreitol (15 mM in total) to the assay buffer and performing the reaction at 37°C for 45 min. The reaction was stopped by addition of malachite green solution and the absorbance was measured at 630 nm. Total phosphatase activity was also evaluated by pNPP hydrolysis in tyrosine assay buffer and measuring the absorbance at 405 nm. The performance of hydrolysis was checked with a purified λ phosphatase (Santa Cruz Biotechnology). Arbitrary activities of PTP-1B and total phosphatases hydrolyzing pNPP were normalized to total protein concentrations in the extracts to be assayed.
Protein dimerization assays
Dimerization of EGFR in MDA MB468 cells was tested using a moderate-strength IP lysis buffer for protein extraction to avoid disruption of protein complexes. Western blot detection of full-length monomeric and dimeric forms of the receptor separated by SDS-PAGE was performed with anti-EGFR polyclonal antibody recognizing the cytoplasmic domain (Thermo Scientific) and then revealed with anti-rabbit IgG conjugated to DyLight 680 (Thermo Scientific). Dimerization of a truncated extracellular region of EGFR and ErbB2 was assessed in vitro by incubating 1 μmole of purified DII/sEGFR or DII/sErbB2 with increasing concentrations of NSC 2155 at 18°C for 30 min. The reaction was quenched with Tris-HCl pH 8.0 for 15 min, then proteins were migrated by electrophoresis and analyzed by Western blotting using anti-His antibody (Sigma Aldrich).
References
Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 103, 211–225 (2000).
Ogiso, H. et al. Crystal structure of the complex of human epidermal growth factor and receptor extracellular domains. Cell 110, 775–787 (2002).
Garrett, T. P. et al. Crystal structure of a truncated epidermal growth factor receptor extracellular domain bound to transforming growth factor alpha. Cell 110, 763–773 (2002).
Ferguson, K. M. et al. EGF activates its receptor by removing interactions that autoinhibit ectodomain dimerization. Mol. Cell 11, 507–517 (2003).
Burgess, A. W. et al. An open-and-shut case? Recent insights into the activation of EGF/ErbB receptors. Mol. Cell 12, 541–552 (2003).
Dawson, J. P. et al. Epidermal growth factor receptor dimerization and activation require ligand-induced conformational changes in the dimer interface. Mol. Cell. Biol. 25, 7734–7742 (2005).
Zhang, X., Gureasko, J., Shen, K., Cole, P. A. & Kuriyan, J. An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 125, 1137–1149 (2006).
Zhang, X. et al. Inhibition of the EGF receptor by binding of MIG6 to an activating kinase domain interface. Nature 450, 741–744 (2007).
Jura, N. et al. Catalytic control in the EGF receptor and its connection to general kinase regulatory mechanisms. Mol. Cell 42, 9–22, 10.1016/j.molcel.2011.03.004 (2011).
Sorkin, A. & Goh, L. K. Endocytosis and intracellular trafficking of ErbBs. Exp. Cell. Res. 314, 3093–3106 (2008).
Bill, A. et al. Cytohesins are cytoplasmic ErbB receptor activators. Cell 143, 201–211 (2010).
Yun, C. H. et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc. Natl. Acad. Sci. USA 105, 2070–2075 (2008).
Arteaga, C. L., Ramsey, T. T., Shawver, L. K. & Guyer, C. A. Unliganded epidermal growth factor receptor dimerization induced by direct interaction of quinazolines with the ATP binding site. J. Biol. Chem. 272, 23247–23254 (1997).
Gamou, S. & Shimizu, N. Hydrogen peroxide preferentially enhances the tyrosine phosphorylation of epidermal growth factor receptor. FEBS Lett. 357, 161–164 (1995).
Bae, Y. S. et al. Epidermal growth factor (EGF)-induced generation of hydrogen peroxide. Role in EGF receptor-mediated tyrosine phosphorylation. J. Biol. Chem. 272, 217–221 (1997).
Goldkorn, T. et al. EGF-Receptor phosphorylation and signaling are targeted by H2O2 redox stress. Am. J. Respir. Cell. Mol. Biol. 19, 786–798 (1998).
Paulsen, C. E. et al. Peroxide-dependent sulfenylation of the EGFR catalytic site enhances kinase activity. Nat. Chem. Biol. 8, 57–64 (2012).
MacBeath, G., Koehler, A. N. & Schreiber, S. L. Printing small molecules as microarrays and detecting protein-ligand interactions en masse. J. Am. Chem. Soc. 121, 7967–7968 (1999).
Kemp, M. M., Weïwer, M. & Koehler, A. N. Unbiased binding assays for discovering small-molecule probes and drugs. Bioor. Med. Chem. 15, 1979–1989, 10.1016/j.bmc.2011.11.071 (2012).
Foong, Y. M., Fu, J., Yao, S. Q. & Uttamchandani, M. Current advances in peptide and small molecule microarray technologies. Cur. Opin. Chem. Biol. 16, 234–42, 10.1016/j.cbpa.2011.12.007 (2012).
Dawson, J. P., Bu, Z. & Lemmon, M. A. Ligand-induced structural transitions in ErbB receptor extracellular domains. Structure 15, 942–954 (2007).
Snapyan, M. et al. Dissecting DNA-protein and protein-protein interactions involved in bacterial transcriptional regulation by a sensitive protein array method combining a near-infrared fluorescence detection. Proteomics 3, 647–657 (2003).
Gyulkhandanyan, A. et al. Assessment of new cationic porphyrin binding to plasma proteins by planar microarray and spectroscopic methods. J. Biomol. Str. Dynamics 31, 363–375, 10.1080/07391102.2012.703063 (2013).
Peternel, S., Grdadolnik, J., Gaberc-Porekar, V. & Komel, R. Engineering inclusion bodies for non denaturing extraction of functional proteins. Microb. Cell. Fact. 7, 34, 10.1186/1475-2859-7-34 (2008).
Carlson, S. M. & White, F. M. Using small molecules and chemical genetics to interrogate signaling networks. ACS Chem. Biol. 6, 75–85 (2011).
Moreau, D. et al. Original triazine inductor of new specific molecular targets, with antitumor activity against nonsmall cell lung cancer. Int. J. Cancer 123, 2676–2683 (2008).
Fry, D. W. et al. A specific inhibitor of the epidermal growth factor receptor tyrosine kinase. Science 265, 1093–1095 (1994).
Bridges, A. J. et al. Tyrosine kinase inhibitors. 8. An unusually steep structure-activity relationship for analogues of 4-(3-bromoanilino)-6,7-dimethoxyquinazoline (PD 153035), a potent inhibitor of the epidermal growth factor receptor. J. Med. Chem. 39, 267–276 (1996).
Levkowitz, G. et al. Ubiquitin ligase activity and tyrosine phosphorylation underlie suppression of growth factor signaling by c-Cbl/Sli-1. Mol. Cell 4,1029–1040 (1999).
Ettenberg, S. A. et al. Cbl-b-dependent coordinated degradation of the epidermal growth factor receptor signaling complex. J. Biol. Chem. 276, 27677–27684 (2001).
Weidner, K. M. et al. Interaction between Gab1 and the c-Met receptor tyrosine kinase is responsible for epithelial morphogenesis. Nature 384, 173–176 (1996).
Agazie, Y. M. & Hayman, M. J. Molecular mechanism for a role of SH in epidermal growth factor signaling. Mol. Cell. Biol. 23, 7875–7886 (2003).
Lemmon, M. A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Liu, F. & Chernoff, J. Protein tyrosine phosphatase 1B interacts with and is tyrosine phosphorylated by the epidermal growth factor receptor. Biochem. J. 327, 139–145 (1997).
Ferrari, E. et al. Identification of new substrates of the protein-tyrosine phosphatase PTP1B by Bayesian integration of proteome evidence. J. Biol. Chem. 286, 4173–4185 (2011).
DeYulia, G. J., Jr, Cárcamo, J. M., Bórquez-Ojeda, O., Shelton, C. C. & Golde, D. W. Hydrogen peroxide generated extracellularly by receptor-ligand interaction facilitates cell signaling. Proc. Natl. Acad. Sci. USA 102, 5044–5049 (2005).
DeYulia, G. J., Jr & Cárcamo, J. M. EGF receptor-ligand interaction generates extracellular hydrogen peroxide that inhibits EGFR-associated protein tyrosine phosphatases. Biochem. Biophys. Res. Commun. 334, 38–42 (2005).
Cochet, C. et al. Demonstration of epidermal growth factor-induced receptor dimerization in living cells using a chemical covalent cross-linking agent. J. Biol. Chem. 263, 3290–3295 (1988).
Gadella, T. W., Jr & Jovin, T. M. Oligomerization of epidermal growth factor receptors on A431 cells studied by time-resolved fluorescence imaging microscopy. A stereochemical model for tyrosine kinase receptor activation. J. Cell. Biol. 129, 1543–1558 (1995).
Yu, X., Sharma, K. D., Takahashi, T., Iwamoto, R. & Mekada, E. Ligand-independent dimer formation of epidermal growth factor receptor (EGFR) is a step separable from ligand-induced EGFR signaling. Mol. Biol. Cell 13, 2547–2557 (2002).
Chung, I. et al. Spatial control of EGF receptor activation by reversible dimerization on living cells. Nature 464, 783–787 (2011).
Kenakin, T. & Miller, L. J. Seven transmembrane receptors as shapeshifting proteins: the impact of allosteric modulation and functional selectivity on new drug discovery. Pharmacol. Rev. 62, 265–304 (2010).
Lavis, L. D. & Raines, R. T. Bright ideas for chemical biology. ACS Chem. Biol. 3, 142–155 (2008).
Normanno, N. et al. Epidermal growth factor receptor (EGFR) signaling in cancer. Gene 366, 2–16 (2006).
Earp, H. S., Dawson, T. L., Li, X. & Yu, H. Heterodimerization and functional interaction between EGF receptor family members: a new signaling paradigm with implications for breast cancer research. Breast Cancer Res. Treat. 35, 115–132 (1995).
He, L. & Hristova, K. Physical-chemical principles underlying RTK activation and their implications for human disease. Biochim. Biophys. Acta 1818, 995–1005, 10.1016/j.bbamem.2011.07.044 (2012).
Kamata, H., Shibukawa, Y., Oka, S.-I. & Hirata, H. Epidermal growth factor receptor is modulated by redox through multiple mechanisms. Eur. J. Biochem. 267, 1933–1944 (2000).
Truong, T. H. & Carroll, K. S. Redox regulation of epidermal growth factor receptor signaling through cysteine oxidation. Biochemistry 51, 9954–9965, 10.1021/bi301441e (2012).
Lohse, D. L., Denu, J. M., Santoro, N. & Dixon, J. E. Roles of aspartic acid-181 and serine-222 in intermediate formation and hydrolysis of the mammalian protein-tyrosine-phosphatase PTP1. Biochemistry 36, 4568–4575 (1997).
Patridge, E. V. et al. 7-Nitro-4-(phenylthio)benzofurazan is a potent generator of superoxide and hydrogen peroxide. Arch. Toxicol. 86, 1613–1625. 10.1007/s00204-012-0872-9 (2012).
Turella, P. et al. Proapoptotic activity of new glutathione S-transferase inhibitors. Cancer Res. 65, 3751–3761 (2005).
Federici, L. et al. Structural basis for the binding of the anticancer compound 6-(7-nitro-2,1,3-benzoxadiazol-4-ylthio)hexanol to human glutathione s-transferases. Cancer Res. 69, 8025–8034 (2009).
Estrada, C. et al. Nitric oxide reversibly inhibits the epidermal growth factor receptor tyrosine kinase. Biochem. J. 326, 369–376 (1997).
Lam, Y. W. et al. Comprehensive identification and modified-site mapping of S-nitrosylated targets in prostate epithelial cells. PLoS One 5, e9075. 10.1371/journal.pone.0009075 (2010).
Wilson, K. J., Gilmore, J. L., Foley, J., Lemmon, M. A. & Riese, D. J., 2nd Functional selectivity of EGF family peptide growth factors: implications for cancer. Pharmacol. Ther. 122, 1–8 (2009).
Gentile, A., Lazzari, L., Benvenuti, S., Trusolino, L. & Comoglio, P. M. Ror1 is a pseudokinase that is crucial for Met-driven tumorigenesis. Cancer Res. 71, 3132–3141 (2011).
Zhang, S. et al. ROR1 is expressed in human breast cancer and associated with enhanced tumor-cell growth. PLoS One 7, e31127, 10.1371/journal.pone.0031127 (2012).
Wang, Y.-N. & Hung, M.-C. Nuclear functions and subcellular trafficking mechanisms of the epidermal growth factor receptor family. Cell & Bioscience 2, 13, http://www.cellandbioscience.com/content/2/1/13 (2012).
Yeretssian, G., Lecocq, M., Lebon, G., Hurst, H. C. & Sakanyan, V. Competition on nitrocellulose-immobilized antibody arrays: from bacterial protein binding assay to protein profiling in breast cancer cells. Mol. Cell. Proteomics 4, 605–617 (2005).
Dupuy, L. et al. A highly sensitive near-infrared fluorescent detection method to analyze signalling pathways by reverse-phase protein array. Proteomics 9, 5446–5454 (2009).
Acknowledgements
This work was funded by the Agence National de la Recherche grants ANR-07-RIB-012 and ANR-07-PNANO-051-02. The authors acknowledge the Drug Synthesis and Chemistry Branch of NCI for providing the Diversity Set II compound library (http://dtp.nci.nih.gov/dtpstandard/ChemData) and the French National Chemical Library for providing compound CV009543V (http://chimiotheque-nationale.enscm.fr/index.php). We are grateful to C. Gautier, A. Brossard and C. Tomasoni for technical assistance.
Author information
Authors and Affiliations
Contributions
V.S. designed and managed the project and wrote the manuscript. All authors V.S., M.A., M.L.B., M.F.L., F.B., B.R., A.G., L.G., C.L., E.R., C.R. and F.F. participated in the experiments performed and reviewed the manuscript.
Ethics declarations
Competing interests
V.S. and M.A. are the authors of a patent describing small molecule microarrays.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Sakanyan, V., Angelini, M., Le Béchec, M. et al. Screening and discovery of nitro-benzoxadiazole compounds activating epidermal growth factor receptor (EGFR) in cancer cells. Sci Rep 4, 3977 (2014). https://doi.org/10.1038/srep03977
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03977
This article is cited by
-
Discovery of a novel EGFR ligand DPBA that degrades EGFR and suppresses EGFR-positive NSCLC growth
Signal Transduction and Targeted Therapy (2020)
-
Compounds with capacity to quench the tyrosyl radical in Pseudomonas aeruginosa ribonucleotide reductase
JBIC Journal of Biological Inorganic Chemistry (2019)
-
Activation of EGFR by small compounds through coupling the generation of hydrogen peroxide to stable dimerization of Cu/Zn SOD1
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.