Abstract
The quality of super resolution images obtained by stochastic single-molecule microscopy critically depends on image analysis algorithms. We find that the choice of background estimator is often the most important determinant of reconstruction quality. A variety of techniques have found use, but many have a very narrow range of applicability depending upon the characteristics of the raw data. Importantly, we observe that when using otherwise accurate algorithms, unaccounted background components can give rise to biases on scales defeating the purpose of super-resolution microscopy. We find that a temporal median filter in particular provides a simple yet effective solution to the problem of background estimation, which we demonstrate over a range of imaging modalities and different reconstruction methods.
Similar content being viewed by others
Introduction
The diffraction limit, which has traditionally limited the ability of light microscopy to discern biological structures at nanometer resolution, has been circumvented by a number of different super-resolution techniques1. In one class of widely used methods, including (F)PALM, STORM, dSTORM and GSDIM2, individual fluorescent molecules are stochastically switched to a temporary detectable state, during which the location of the individual molecules is determined at higher resolution using image analysis algorithms3,4,5. Several different methodologies for performing stochastic single-molecule super-resolution reconstructions have been described and generally fall into two broad categories: localization based3,4,6,7 and grid based reconstruction methods5,8. Localization based methods typically utilize a Gaussian fit or a center of mass calculation, while grid based reconstruction methods rely on an inverse modeling approach by deconvolution or compressed sensing.
However, a typical super-resolution dataset may contain significant non-sparse, structured background components, complicating the analysis regardless of the method chosen for analysis. This background may accrue for a variety of reasons, such as weakly, continuously emitting fluorescent molecules attached to cellular structures or cellular auto-fluorescence9,10. In order to accurately reconstruct a super-resolution image, all analysis algorithms require that the foreground signal from sparsely distributed emitters (containing the super-resolution information) is sufficiently separated from this background.
For each data frame the observed fluorescence can be modeled as a sparse distribution of emitters that is convolved with a given or estimated point spread function and a spatio-temporal background:

The first term (foreground) contains the super resolution information and is fitted to the PSF model, given a certain estimated or fitted background. We find that the quality of this background estimate is critical to attaining reliable reconstructions; in many practical circumstances this can have a much greater impact on the fidelity of the final image than the specifics of the treatment of the foreground term.
The vast majority of published super-resolution reconstruction algorithms utilize spatial filtering or local background fitting for background estimation. While foreground and background can be distinguished with some limited specificity on the basis of their spatial frequencies and intensity, there typically exists no clear band-gap between these spatial frequencies across the whole data set. This makes spatial filtering a limited tool for robustly separating foreground from complex, structured background.
A key difference between non-specific (background) fluorescence and emitters of interest is that the latter appear and disappear over relatively rapid timescales. In the general signal processing literature, there are many different methods described for background estimation that exploit temporal information, which can greatly differ in computational complexity11. Although a few previous super-resolution studies have mentioned, obliquely, some form of temporal filtering12,13 for estimating the background component, the importance and effect of this type of background estimation has not been rigorously studied or reported. For this reason, we have explored temporal background estimation methods in the context of super-resolution. We found that a running median filter applied to each pixel in the dataset along its temporal axis represents a straightforward and particular effective background estimator that greatly enhances the quality of the reconstruction. The logic behind the median filter as a background estimator is that super-resolution data is always somewhat sparse and insofar as it is sparse, foreground contributions will tend be discarded by the median filter as outliers and therefore readily separated from background components. The running nature of the filter allows for gradual temporal changes in background and an arbitrary spatial shape of the background is permitted.
We have applied temporal median filtering to data obtained from several different stochastic super resolution techniques, reconstruction methods and probes. Furthermore, we have performed a number of simulations that mimic realistic conditions for stochastic super resolution data in order to validate and check the effect of the various techniques.
Results
Estimation of background component using temporal median filter
The ability of a temporal median filter to separate background and foreground is illustrated in Figure 1. The panels in the left column (a,d,g) show raw data frames from LifeAct-mEos3.2, MyosinIIa-Alexa532 and MyosinIIa-Alexa647 data sets. The middle column (b,e,h) shows the background estimated for that frame using the temporal median filter (window size of 101 frames with 10 frame interpolation, see material and methods). The right column (c,f,i) shows the foreground obtained by directly subtracting the estimated background from the raw frames. Static or very slowly varying fluorescence in the image largely ends up in the background. Notable are the fiducial beads visible in panels a and b that are no longer apparent in the foreground image, as well as the ridge at the cell border (arrow). Panel j shows the raw fluorescence trace (MyosinIIa Alexa 647 data set) at two adjacent pixels (arrow in panel h) and the corresponding background estimate. The background level is different for these two adjacent pixels, but after correction for background, the foreground traces, shown in panel k, are now strongly correlated. The two traces exhibit a number of switching events that are now more accurately separated from the background. The estimated background is relatively smooth compared to the noise in the raw data, which depends on the window size used for the running median filter (see material and methods).
Background and foreground estimation by temporal median filtering.
Panels in the left column (a,d,g) show raw data frames from LifeAct-mEos3.2, MyosinIIa-Alexa532 and MyosinIIa-Alexa647 data sets. Middle column (b,e,h) shows the background estimated for that frame using the temporal median filter (window size of 101 frames with 10 frame interpolation). Right column (c,f,i) shows the foreground calculated by subtracting the estimated background from the raw frames. For display purposes the values were clipped at zero in order to only show the fluorescence that is higher than the estimated background. Panel j shows the raw fluorescence trace (MyosinIIa Alexa 647 data set) at two adjacent pixels (arrow in panel h) and the corresponding background estimate.
Note that the part of the signal that the median filter deems to be background is not sparse, which would confound attempts at deducing its origin of emission with any super resolution algorithm. The implied foreground component, by contrast, is very sparse. The dataset in the top row contains a very dominant and complex background. Many of the strong foreground events are barely recognizable as such in a single raw data frame. The Myosin-Alexa dataset of the third row does not appear to contain much background upon inspection of the raw data, but the temporal median filter nonetheless reveals a substantial non-sparse signal component.
Dual-color GSDIM co-localization experiment
We performed several dual-color GSDIM co-localization experiments using the probes Alexa532 and Alexa647 attached to different secondary antibodies recognizing the same primary antibody, which binds to myosinIIa in one experiment (Fig. 2) and vinculin in another experiment (Supplementary Fig. S1). Hence, a clear colocalization of the Alexa 532 and 647 color channels is expected in the super resolution reconstructions. However, for both experiments we observed large discrepancies between the reconstructed color channels (Fig. 2b,f, Supplementary Fig. S1a,c) using three representative (existing) super-resolution analysis algorithms, including Gaussian fitting3, center of mass localization4 (Supplementary Fig. S2a,d) and a (multi-fitting) grid based method5 (Supplementary Fig. S3a,d). Specifically, in the Alexa532 data-sets, the presence of a moderately high background (Fig. 2a, supplementary video 1) results in a systematic bias such that the reconstruction is skewed towards regions with high fluorescence intensities. This gives the appearance of distinct foci along the fibers and subtle deformations, such as over-sharpening (compare Fig2. b and f, Fig2. b and c, Intensity profile, Fig. 2d). These artifacts were no longer observed when background was accounted for (see material and methods) using the temporal median filter prior to reconstruction (Fig. 2c, Supplementary Figs. S2, 3). Prior application of the temporal background-estimation filter results in a higher co-localization index as determined by cross-correlation analysis of the Alexa532 and Alexa647 data-sets (Fig. 2, insets).
Application of a temporal median filter prior to localization analysis improves fidelity of two-color GSDIM data.
GSDIM imaging of myosinIIa independently labeled with Alexa532 (a–c) and Alexa647 (e–g). Without the utilization of the temporal median filter, the RapidSTORM reconstruction of the Alexa532 data set shows localizations that are skewed towards regions of high fluorescence (b) and exhibit poor co-localization with the Alexa647 (f) based on Pearson's cross-correlation analysis (f, inset). Use of the temporal median filter prior to running the localization eliminates these artifacts in the Alexa532 reconstruction (c) and shows higher correlation with the Alexa647 reconstruction (g, inset). Intensity traces (d, h) are normalized to the area under the trace. Similar analysis for alternative reconstruction methods can be found in supplemental Figs S2,3. Scale bar: 3 μm.
LifeAct-mEos3.2 PALM data with structured background
In PALM data structured-background occurs quite frequently, as a slow buildup of cellular auto-fluorescence with complex spatial characteristics can develop during the acquisition13,14. In a HeLa cell expressing LifeAct-mEos3.215,16 we observed this phenomenon (Fig. 1a,b, arrow and Supplementary video S2), which gives rise to errant super resolution localizations yielding a spurious structure along the edge of the background (Fig 3a,c) and deformations. These types of artifacts are no longer present after improved background correction (Fig 3a–c), regardless of reconstruction method. Improper estimates of structured background of such large relative magnitude may easily lead to artifacts in the reconstruction on a micrometer scale, nullifying the intent of super resolution microscopy.
Reconstructions for LifeAct-mEos3.2 HeLa cell using RapidSTORM, QuickPALM and deconvolution with and without the temporal median filter applied.
An area that shows a high-degree of structured (heterogeneous) background fluorescence indicated with an arrow in Fig S1a-b leads to a spurious structure when using RapidSTORM (median smoothing 5 px setting) or deconvolution if the temporal median was not applied. When the temporal median filter prior to running the reconstruction analysis was applied (b, e, h) the effect of structured background is greatly reduced in the reconstruction such that the intensity profiles are now in much closer agreement for the different methods (c, f, i). This illustrates the relative importance of background estimation in the overall reconstruction process. Scale bar: 3 μm.
Intricate structures in LifeAct-Venus GSDIM data sets
Figure 4 shows two additional GSDIM data sets obtained with LifeAct-Venus, a different fluorescent probe. Panel 4a shows the sum of all frames in the data sets; panel 4b, shows the sum of all background subtracted frames, both represent a diffraction limited image. Panels 4c and 4d show the RapidSTORM3 reconstruction obtained without and with application of the temporal median filter. The analysis of these data sets revealed that temporal median filtering reduces the presence of strong foci at filament crossings, which appear to induce deformations that are not apparent in the diffraction limited images (arrow 4a,d). The fact that panel 4c has features that deviate from panels 4a, 4b and 4d can be attributed to the inappropriate application of a super resolution algorithm to a dataset that is not sufficiently sparse.
GSDIM data of two HeLA cells with LifeAct-Venus Panel a shows the sum of all frames in the data sets; panel b, shows the sum of all background subtracted frames, both represent a diffraction limited image. Panels c and d show the RapidSTORM reconstruction obtained without and with application of the temporal median filter. The analysis of these data sets revealed that temporal median filtering reduces the presence of strong foci at filament crossings, which appear to induce deformations that are not apparent in the diffraction limited images. Panels e and f show the RapidSTORM reconstruction obtained without and with application of the temporal median filter of another HeLa cell with an intricate F-actin network. The used threshold for all reconstructions was the same and the Gaussian smoothing filter was selected in this case (1 sigma). Color scale for images in panels c,d and e,f were chosen to be equal (scale bar 3 μm).
Panels 4e and 4f show the RapidSTORM reconstruction obtained without and with application of the temporal median filter of another HeLa cell with LifeAct-Venus. The background-corrected reconstruction reveals more intricate details in the F-actin structures, which are otherwise lost to background-related localization errors. Also the deformation induced by a hot-spot (arrow 4e,f) is greatly reduced in the background corrected reconstruction.
Analysis of synthetic datasets
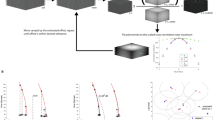
A number of simulations that mimic realistic conditions for stochastic super resolution data were performed in order to validate and check the effect of the various techniques. Key parameters such as event amplitude, event density and background conditions were varied. For the latter, uniform background conditions and structured background conditions were used. These parameter variations were applied to ring structures and filament structures, for which the ring size and the distance between filaments was varied respectively.
Previous work in synthetic analysis has aimed to characterize the performance of a method in terms of accuracy and a false positive and false negative rate. A different approach is taken here, as these measures are not necessarily adequate to characterize the effect of structured background on reconstruction quality. Specifically, the method described here is aimed to better quantify the systematic bias induced by correlated errors in the reconstruction, which is not conveyed by localization standard error of the mean alone. An image may appear very sharp, which in the absence of bias may be regarded as an indication of high accuracy, but in the presence of systematic biases, the result may in fact be substantially distorted.
A quantification of localization events in isolation makes it fundamentally difficult to consider correlations between localization errors. But by quantifying the appearance of structures as a whole, a measure of correlated errors can be derived. Line and ring patterns can be used to characterize the behavior of an algorithm. Line patterns (Fig. 5) test the ability to resolve nearby features, whereas ring patterns (Fig. 6) additionally test the influence of curved geometries as well. In order to quantitatively characterize the accuracy of a method, the symmetry axis of these 2D patterns is utilized to obtain a 1D distribution of localizations (profiles). The repeatability of the localizations can be characterized in term of the width of these distributions and the systematic bias is given by the offset between the distribution of localizations and the underlying structure. It is in this measure of bias that the different methods investigated here show the biggest performance differences and where the necessity of accurate background estimation becomes most apparent.
Thin filaments at different inter-filament distances (a) simulated with structured background (b). The structured background leads both artifacts and distorted structures (c), which are mitigated by the utilization of a temporal median filter prior to performing the localization analysis (d), resulting in a more accurate rendering of the structures. (e) Quantification of synthetic filaments shown in c-d. Pairs of filaments, each with thickness 10 nm (black bar). The profiles shown in red and green represent the quantified mean profile of the reconstructed filaments measured from the midline outward. Reconstructions were obtained from RapidSTORM, with settings median smoothing (5 px) and threshold 100 and rendered using the obtained amplitude blurred with a Gaussian with a SEM of 2 nm. Prior to reconstruction by RapidSTORM the background was accounted for using the temporal median filter, with a filter size of 101 frames and 10 frame interpolation.
RapidSTORM reconstructions of a ring with radius 75 nm with a threshold of 10 photons and using the standard spatial median filter without (a) and with (b) application of the temporal median filter prior to reconstruction.
Background corrected reconstruction no longer show artifacts in the reconstructions and in all cases reliably reconstruct the ring. For these simulations the event cycle amplitude A = [200, 800, 3200] was varied and the structured background, where the peak of the structured background was varied using the values bs = [0, 10, 20, 30, 40, 50] photons using the same event list for each case with simulations settings as described in the supplementary information. The radial profiles of the rings were calculated for each panel and are shown together with the original ring in the right hand side column. The intensity scale in each panel was adjusted to show the full intensity range independent of the other panels. The localizations were obtained from RapidSTORM with settings median smoothing (5 px) and threshold 100 and rendered using the obtained amplitude blurred with a Gaussian with a SEM of 2 nm. Prior to reconstruction by RapidSTORM the background was accounted for using the temporal median filter, with a filter size of 101 frames and 10 frame interpolation.
Figure 5 shows the results of simulations of 10 nm wide filament pairs separated at decreasing distances (Fig. 5a) that were positioned at different distances (d) and placed on structured background (see material and methods). A simulated data set containing structured background (Fig. 5b) can result in reconstructions that show both artifacts and distorted structures (Fig. 5c). These effects are mitigated by the utilization of a temporal median filter prior to performing the localization analysis (Fig. 5d), resulting in a more accurate rendering of the structures. Fig. 5e shows the quantification of synthetic filaments shown in Fig. 5c–d. The profiles shown in red and green represent the quantified mean profile of the reconstructed filaments measured from the midline outward. The red profiles were obtained using RapidSTORM without applying the temporal median filter prior to analysis and the green profiles were obtained using RapidSTORM after application of the temporal median filter. Application of the temporal median filter removes the artifactual structure in between the filaments (zero position), which was caused by the structured background. Furthermore, the application of the temporal median filter removed the inward bias visible in the top row panels (red), which is no longer present in the bottom row panels (green).
Figure 6 shows the effect of application of the temporal median filter prior to reconstruction with RapidSTORM on rings with a radius of 75 nm (see material and methods). The left column in both figures shows the reconstruction of the ring when there is no structured background. For all amplitudes, we find that without the presence of background the ring can be reconstructed reliably (left column), independent of the temporal median filter. However, if the structured background increases, localizations are increasingly biased by the structured background, not only skewing the localizations towards the center of the ring, yielding a smaller ring, but also generating false positives caused by the structured background. When the background, estimated using the temporal median filter, is accounted for prior to analysis with RapidSTORM, in all cases it was observed that the ring can be constructed without any bias towards the center of the ring giving a similar result independent of the background level. As expected from the Thompson equation17, at higher background levels and lower amplitudes the accuracy is reduced, yielding a wider ring wall but the result remains unbiased and its position is conserved.
We have also explored the effect of increasing event densities on the reconstruction of the rings with a radius of 150 nm using the same simulation parameters as used for the 75 nm ring. It was observed that an increasing overlap of events can introduce a reconstruction bias towards the center of the ring (Figs. S7). Application of the temporal median filter will compensate for this bias when using RapidSTORM (Fig. S8) because the temporal median filter removes part of the fluorescence that stems from overlapping events. However, this result is particular to RapidSTORM. The median filter in combination with a deconvolution reconstruction in fact shows a slight outward bias of the ring at extreme event density (Fig S9). This highlights the importance of choosing an appropriate combination of background estimator and localization algorithm (see Discussion).
Discussion
We have shown that appropriate background estimation is of pivotal importance for obtaining reliable super resolution reconstructions, which we illustrate with data obtained from (d)STORM, GSDIM and PALM using several different probes representing a range of data types and qualities. Especially in case of spatially complex background, the advantages of temporal median background filtering can be profound. Application of the proposed background estimator eliminates the large discrepancies that are otherwise observed when analyzing the same data with different super-resolution algorithms (Fig. 7), underlining the relative importance of background estimation in the overall reconstruction process. By using an appropriate background estimator, a wider range of imaging conditions, type of probes and samples are tolerated for stochastic super-resolution microscopy.
Line scan profiles for the LifeAct-mEos3.2, MyosinIIa-Alexa532 and MyosinIIa-Alexa647 data sets from (Fig. 2, 3,S2, S3) put side by side, using three reconstruction methods without and with application of the temporal median filter.
The panels a, c and e reveal that the different methods give different results for the same data set when no temporal median filter is applied. The panels b, d and f reveals that application of the temporal median filter yields results that are in close agreement for the three reconstruction methods used.
Use of a temporal median filter enables background estimation for each pixel in a given data frame from the temporal distribution; an estimate of the background is obtained in this way without interfering with the spatial resolution of the original image. Ideally, the use of temporal median filtering and estimation of the background is directly incorporated in the reconstruction software, taking into account the appropriate statistical model. However, we observe that background subtraction with uniform offset can be applied as a preprocessing step and that reconstruction on the background corrected data set using existing software in general works robustly and the added uncertainty generally is negligible compared to the detrimental effects structured background can have. In the case of localization based algorithms, special care could be taken when calculating the localization error as the background estimate will introduce some extra noise. The standard error of the median is by good approximation proportional to the standard error of the mean, which scales with α × σ/√N, where N denotes the median filter window size, σ2 the variance and α denotes a proportionality factor that depends on the distribution. For Poissonian data, the variance scales with the mean intensity μ, σ2 = k2 × μ, where k denotes the detector gain. Then the added variance due to the background estimate becomes σb2 = α2 × σ2/N. The relative increase in the variance is approximately σb2/σ2 = α2/N. The approximate proportionality factor for a normal like distribution is about 1.253, hence for a window size of ~100 frames the variance increases by 1.6%.
We have compared a variety of existing widely used background estimation methods. The local fitting of background as employed in3 generally turns out to be the least robust method (Fig 3). Understandably, this method cannot discern the peak of a background feature from a foreground event, nor can one effectively discriminate a peak on a slope, from a displaced peak. Therefore, using such a method leads to both false positives and biases, respectively.
A spatial (Gaussian) filter as a background estimate4 works somewhat better, but is sensitive to sharp background features, which get mistaken for foreground and high density areas, which readily get mistaken for background in a distortive manner (Fig S2).
The published temporal filters have been implemented and are found to be generally superior to the spatial filters. The method in12 proposes a ten frame running mean filter. This works reasonably well for guarding against errors in localization accuracy, but we find that for high density, high S/N data, the median filter captures up to four times as much of the foreground signal, compared to the short mean filter, which may result in substantial distortions of appearance (supplemental Fig. S10). The differences of frames method employed in18 can be viewed as a 1-frame mean filter and consequently shares similar shortcomings, but to a stronger degree.
The background estimation method employed in13 may also be considered a temporal filter. In essence, the first principal component of the dataset is taken as a background estimate. This may be regarded as a truncated eigenbackground11. For datasets with simple background behavior, this method works well. But according to expectation, we find a single principal component as a background estimate breaks down for datasets where the background dynamics are too complex to be well represented by a single component (supplemental Fig. S11).
We find that the temporal median filter generally works best over the range of parameter conditions tested. That said, at extreme event densities background estimation becomes fundamentally more difficult and potentially significant errors may remain in the final reconstruction (supplemental Fig. S9–11).
Temporal median filtering can be easily integrated into existing workflows and consistently shows profound improvements in combination with several different classes of reconstruction algorithms and over a wide-range of data types and qualities. As our results demonstrate, without a conscious choice of background estimator appropriate for the given data quality, one is not guaranteed to achieve any super resolution at all. Because of the significant impact background can have on the fidelity of the final reconstructed image, it is highly recommended that an appropriate consideration of background estimation be a fundamental part of any stochastic super-resolution analysis workflow.
Methods
Estimation of the background component
For most datasets with slow varying mean frame intensities the temporal median filter can be applied directly. However, because the average frame intensity can sometimes vary significantly due, for example, to an overall loss of fluorescence caused by depletion of fluorescent molecules or variations in the intensity of the excitation laser, a direct application of a temporal median filter on the raw data may be filtering other temporal signal components than event switching. To correct for entire frame intensity fluctuations, first the mean fluorescence intensity for each frame is determined and subsequently the data is scaled according to this mean fluorescence profile13. A temporal moving median filter is then applied to the scaled data. The obtained median values are then rescaled by multiplying the median values with the mean frame intensity.



Where Dt denotes the mean frame intensity for frame t, Nx,y,t denotes the normalized data frame and w denotes the window size for applying the temporal median filter. The size of the time window should be chosen such that it is significantly longer than the typical slow-switching events (≫10). For our datasets, we typically use a window size of about 100 frames to calculate the moving median.
We have implemented two ways of computing the running temporal median. One way of efficiently computing the filter is by incrementally updating a list of sorted values for each pixel and taking the center of the list. Another way of increasing efficiency is by means of computing the median only at certain keyframes and linearly interpolating those. Such keyframe interpolation is well justified by the slowly varying nature of the background itself. Both methods produce virtual indistinguishable end-results. We find that the keyframe method is roughly equally efficient as the incremental method, given keyframes spaced 12 frames apart. With a 50 frame median filter radius (101 frames), of 512x512 pixels, this processing takes ~0.1 seconds per frame on an Intel i7-2700k at 3.8 GHz. This makes the keyframe method somewhat preferable; it could be made faster still by increasing the distance between keyframes without significantly affecting quality and it does not rely on low level language extensions to attain this speed, making it easier to integrate as a technique (a python script is provided). It is preferable that each stochastic super resolution software package integrates an optimized version of calculating the median filter, depending on the used programming language or hardware architecture (e.g. parallel computing or GPU) different approaches can be followed for calculating the running median19.
Constructs and sample preparation
The LifeAct-mEos3.2 was a kind gift from Tao Xu15,16. HeLa cells were obtained from the American Tissue Culture Collection (ATCC). Cells were maintained in Dulbecco's Modified Eagle Medium supplemented with GlutaMAX and 10% Fetal bovine serum (Invitrogen, Bleiswijk NL) and grown for at least 2 days in phenol-free media before imaging. Cells were plated on 24 mm #1 round cover-glasses (Menzel-Gläser) in six well plates. Transfections were done with Lipofectamine 2000 reagent according to the manufacturer's protocol (Invitrogen, Bleiswijk NL). Twenty-four hours after transfection, cells were transferred to an Attofluor sample chamber (Invitrogen) and imaged live in microscopy medium (137 mM NaCl, 5.4 mM KCl, 1.8 mM CaCl2, 0.8 mM MgSO4, 20 mM D-glucose, 20 mM HEPES).
Optical setup and imaging
PALM imaging was performed on an inverted Nikon Eclipse Ti microscope equipped with a TIRF system using a 60x ApoTIRF 1.49 oil objective. A Coherent OBIS 488 50 mW laser was used to locate cells producing pre-converted mEos3.2. Photo-conversion of mEos3.2 was elicited by continuous illumination with a Coherent 50 mW 405 Cube laser with power settings of <1 mW, while the converted FP was excited and bleached with a 1 W 561 Coherent OPSL with power settings typically in the range of 200–300 W/cm2. Excitation light was passed through a quad-band dichroic 405/488/561/640 (Chroma). Emission light was passed through a 561 nm RazorEdge ultrasteep long-pass edge filter (Semrock). Images were recorded with an Andor iXon 897 EMCCD with 50 ms exposure times at a frame rate at 12.2 Hz. Pixel size in the image was 67 nm.
Super-resolution imaging of Non-muscle MyosinIIA filaments and vinculin
A7r5 cells were cultured on #1.5 coverslips. After 48 hours cells were washed briefly with PBS, fixed with 4% PFA for 10 min at room temperature and extracted with 0.1% Triton X-100. Samples were extensively washed with PBS and blocked with 5% BSA for 30 min at room temperature. MyosinIIA was labeled with a monoclonal primary antibody raised in rabbit (Sigma-Aldrich) diluted to a final concentration of 1 mg/ml, for one hour at room temperature, washed and incubated with anti-rabbit IgG polyclonal antibody conjugated to Alexa Fluor 647 dye molecules (Invitrogen) and anti-rabbit IgG polyclonal antibody conjugated to Alexa Fluor 532 dye molecules (Invitrogen) both at a final concentration of 0.01 mg/ml, for 30 minutes at room temperature. Cells were imaged in the presence of an oxygen scavenging system (10% glucose, 0.5 mg/ml glucose oxidase, 40 μg/ml catalase, 50 mM MEA).
HeLa cells were cultured on #1.5 coverslips. After 24 hours cells were washed briefly with PBS, fixed with 4% PFA for 10 min at room temperature and extracted with 0.1% Triton X-100. Samples were extensively washed with PBS and blocked with 5% BSA for 30 min at room temperature. Vinculin was labeled with a monoclonal primary antibody raised in mouse (abcam) diluted 1:400, for one hour at room temperature, washed and incubated then with anti-mouse IgG polyclonal antibody conjugated to Alexa Fluor 647 dye molecules (Invitrogen) and-anti mouse IgG polyclonal antibody conjugated to Alexa Fluor 532 dye molecules (Invitrogen) both at a final concentration of 0.01 mg/ml, for 30 minutes at room temperature. Cells were imaged in the presence of an oxygen scavenging system (10% glucose, 0.5 mg/ml glucose oxidase, 40 μg/ml catalase, 50 mM MEA).
Imaging of the samples was carried out on a Leica SR-GSD microscope. Images were taken in TIRF mode at 100 frames per second. Colors were sequentially imaged in order of decreasing wavelength. The setup consisted of the following components: an inverted microscope (DMI6000 B, Leica Microsystems GmbH), a 1.47-NA TIRF objective (HCX PL APO 100× NA 1.47), a tube lens providing an extra factor of 1.6× in magnification, a 488-nm fiber laser (2RU-VFL-P-300-488), a 532-nm fiber laser (2RU-VFL-P-1000-532-B1R, MPB Communications), a 642-nm fiber laser (2RU-VFL-P-1000-642-B1R, MPB Communications) and an EMCCD camera (iXon DU-897, Andor) with an effective EM gain of 148. Images were taken in TIRF mode at 100 frames per second for ~5100 time frames, giving a total measurement time of about 1 min for each color. Colors were imaged in order of decreasing wavelength. The filter cube (642HP-T) for imaging with the 642-nm laser consisted of an excitation filter (zet405/642x), a dichroic mirror (t405/642rpc) and emission filters (et710 100lp and ET650LP). The epifluorescence filter cube (532HP-T) for imaging with the 532-nm laser consisted of an excitation filter (zet405/532x), a dichroic mirror (t405/532rpc) and emission filters (et600/100 m and ET550LP). Pixel size in the image was 93.11 nm.
Super-resolution imaging of Actin
HeLa cells were cultured on #1.5 coverslips. After 24 hours cells were transiently transfected using PolyEthylene Imine (PEI) using 1 μg of DNA and 3 μg of PEI per well on a 6-well plate, with a plasmid bearing LifeAct tagged with the yellow fluorescent protein variant Venus. After 24 hours cells were washed briefly with PBS and fixed with PFA for 10 min at room temperature. Samples were extensively washed with PBS and imaged in the presence of PBS.
Imaging of the samples was carried out on a Leica SR-GSD microscope. Images were taken in TIRF mode at 100 frames per second. The setup consisted of the following components: an inverted microscope (DMI6000 B, Leica Microsystems GmbH), a 1.47-NA TIRF objective (HCX PL APO 100× NA 1.47), a tube lens providing an extra factor of 1.6× in magnification, a488-nm fiber laser (2RU-VFL-P-300-488), a 532-nm fiber laser (2RU-VFL-P-1000-532-B1R, MPB Communications), a 642-nm fiber laser (2RU-VFL-P-1000-642-B1R, MPB Communications) and an EMCCD camera (iXon DU-897, Andor) with an effective EM gain of 148. Images were taken in TIRF mode at 100 frames per second for ~5100 time frames, giving a total measurement time of about 1 min for each color. Colors were imaged in decreasing wavelength order. The filter cube (642HP-T) for imaging with the 642-nm laser consisted of an excitation filter (zet405/642x), a dichroic mirror (t405/642rpc) and emission filters (et710 100lp and ET650LP). The epifluorescence filter cube (532HP-T) for imaging with the 532-nm laser consisted of an excitation filter (zet405/532x), a dichroic mirror (t405/532rpc) and emission filters (et600/100 m and ET550LP). Pixel size in the image was 93.11 nm.
Synthetic data
We have performed a number of simulations that mimic realistic conditions for stochastic super resolution data in order to validate and check the effect of the various techniques. Key parameters, such as event amplitude, event density and background (both uniform and structured) were varied. These parameter variations have been applied to ring structures and filament structures, with variable ring size and the distance between filaments respectively (see Supplementary Material for more details).
Reconstruction algorithms and rendering
For analysis of stochastic real and simulated data two localization based methods and a direct reconstruction, deconvolution based method were used. For the localization techniques, we utilized RapidSTORM 33,7, which performs a Gaussian based fit and QuickPALM 1.14, which performs a center of mass based calculation. Background was corrected for as a preprocessing step for these two algorithms, the corrected data was offset with a constant value to prevent negative values. For the grid based method we implemented the deconvolution variant without regularization as described in5,20. For this algorithm the background estimate was directly incorporated into the statistical model. Prior to application of the reconstruction algorithms any drift was corrected if present. The user defined parameters for each of these methods, such as width of the PSF and intensity thresholds, were estimated from the experimental data or obtained directly from the synthetic input. The threshold was chosen in such a way that the number of detected events was minimal outside the cell or in regions without structures. Other user defined settings are mentioned in the appropriate figure legends. In order to ensure consistency and to allow direct comparison between the final images, we utilized our own rendering program written in Matlab R2010a (code made available in supplementary material). From the localization lists obtained from RapidSTORM or QuickPALM the location and amplitude of each event was extracted and rendered; the zoom factor of the final image was set to 8. Each localization was rendered using a Gaussian (integral normalized to unity) with a FWHM of 23.55 nm (σ = 10 nm) and localizations from simulated data were rendered with a FWHM of 4.71 nm (σ = 2 nm).
References
Gould, T. J., Hess, S. T. & Bewersdorf, J. Optical Nanoscopy: From Acquisition to Analysis. Annual Review of Biomedical Engineering 14, 231–254 (2012).
Ewers, H. 4. Nano Resolution Optical Imaging Through Localization Microscopy. 1–20 (2012). 10.1016/b978-0-12-385872-6.00004-0.
Wolter, S., Löschberger, A., Holm, T. & Aufmkolk, S. rapidSTORM: accurate, fast open-source software for localization microscopy. Nature (2012).
Henriques, R. et al. QuickPALM: 3D real-time photoactivation nanoscopy image processing in ImageJ. Nature Methods 7, 339–340 (2010).
Mukamel, E., Babcock, H. & Zhuang, X. Statistical Deconvolution for Superresolution Fluorescence Microscopy. Biophysical Journal 102, 2391–2400 (2012).
Cheezum, M., Walker, W. & Guilford, W. Quantitative comparison of algorithms for tracking single fluorescent particles. Biophysical Journal (2001).
Wolter, S., Schüttpelz, M. & Tscherepanow, M. Real-time computation of subdiffraction-resolution fluorescence images. Journal of 1–11 (2010).
Zhu, L., Zhang, W., Elnatan, D. & Huang, B. Faster STORM using compressed sensing. Nature Methods 9, 721–723 (2012).
Betzig, E. et al. Imaging Intracellular Fluorescent Proteins at Nanometer Resolution. science 313, 1642–1645 (2006).
Hess, S., Gould, T. & Gunewardene, M. Ultrahigh resolution imaging of biomolecules by fluorescence photoactivation localization microscopy. Micro and Nano (2009).
Piccardi, M. Background subtraction techniques: a review. 2004 IEEE International Conference on Systems, Man and Cybernetics (IEEE Cat. No.04CH37583) 3099–3104 (2004). 10.1109/ICSMC.2004.1400815.
Baddeley, D., Jayasinghe, I. D., Cremer, C., Cannell, M. B. & Soeller, C. Light-Induced Dark States of Organic Fluochromes Enable 30 nm Resolution Imaging in Standard Media. Biophysical Journal 96, L22–L24 (2009).
Hess, S. T. et al. Dynamic clustered distribution of hemagglutinin resolved at 40 nm in living cell membranes discriminates between raft theories. Proceedings of the National Academy of Sciences 104, 17370–17375 (2007).
Betzig, E. et al. Imaging Intracellular Fluorescent Proteins at Nanometer Resolution. science 313, 1642–1645 (2006).
Riedl, J. et al. Lifeact: a versatile marker to visualize F-actin. Nature Methods 5, 605–607 (2008).
Zhang, X., Chu, X., Wang, L., Wang, H. & Liang, G. Rational Design of a Tetrameric Protein to Enhance Interactions between Self-Assembled Fibers Gives Molecular Hydrogels. Angewandte 1–5 (2012).
Thompson, R. E., Larson, D. R. & Webb, W. W. Precise Nanometer Localization Analysis for Individual Fluorescent Probes. Biophysical Journal 82, 2775–2783 (2002).
Cremer, C. et al. Superresolution imaging of biological nanostructures by spectral precision distance microscopy. Biotechnology Journal 6, 1037–51 (2011).
Juhola, M., Katajainen, J. & Raita, T. Comparison of algorithms for standard median filtering. Signal Processing (1991).
Wang, G. & Smith, S. J. Sub-diffraction Limit Localization of Proteins in Volumetric Space Using Bayesian Restoration of Fluorescence Images from Ultrathin Specimens. PLoS Computational Biology 8, e1002671 (2012).
Acknowledgements
We thank M.A. Hink for useful discussions. This research was supported by the Netherlands Research Organization (NWO-ALW VIDI 864.09.015 to M.P. and E.H.), a Middelgroot investment grant (834.07.003), the European Science Foundation (EuroMembrane, TraPPs to K.C.C. and T.W.J.G.) and the Dutch Technology Foundation STW (to T.W.J.G., D.L.P. and K.J.)
Author information
Authors and Affiliations
Contributions
E.H. and M.P. conceived and implemented the proposed method in close collaboration with K.C.C. E.H., M.P. and K.C.C. participated in the data analysis. K.C.C., R.M.P.B., T.W.J.G., D.L.P. and K.J. designed and executed the experiments. All authors contributed to the preparation of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplemental information
Supplementary Information
Video 1
Supplementary Information
Video 2
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Hoogendoorn, E., Crosby, K., Leyton-Puig, D. et al. The fidelity of stochastic single-molecule super-resolution reconstructions critically depends upon robust background estimation. Sci Rep 4, 3854 (2014). https://doi.org/10.1038/srep03854
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03854
This article is cited by
-
Near-infrared PAINT localization microscopy via chromophore replenishment of phytochrome-derived fluorescent tag
Communications Biology (2024)
-
Nanoscale imaging of CD47 informs how plasma membrane modifications shape apoptotic cell recognition
Communications Biology (2023)
-
Enhanced super-resolution microscopy by extreme value based emitter recovery
Scientific Reports (2021)
-
Highly photostable fluorescent labeling of proteins in live cells using exchangeable coiled coils heterodimerization
Cellular and Molecular Life Sciences (2020)
-
Visualisation of dCas9 target search in vivo using an open-microscopy framework
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.