Abstract
Experimental and bioinformatic studies of transcription initiation by RNA polymerase II (RNAP2) have revealed a mechanism of RNAP2 transcription initiation less uniform across gene promoters than initially thought. However, the general transcription factor TFIIB is presumed to be universally required for RNAP2 transcription initiation. Based on bioinformatic analysis of data and effects of TFIIB knockdown in primary and transformed cell lines on cellular functionality and global gene expression, we report that TFIIB is dispensable for transcription of many human promoters, but is essential for herpes simplex virus-1 (HSV-1) gene transcription and replication. We report a novel cell cycle TFIIB regulation and localization of the acetylated TFIIB variant on the transcriptionally silent mitotic chromatids. Taken together, these results establish a new paradigm for TFIIB functionality in human gene expression, which when downregulated has potent anti-viral effects.
Similar content being viewed by others
Introduction
Basal transcription factors have been considered to be essential for all RNA polymerase II (RNAP2) promoters. The original set of general transcription factors (TFIID subunit TBP, TFIIB, TFIIA, TFIIF, TFIIE, TFIIH), identified in studies of a limited set of promoters appears however to be incomplete1, while at the same time cases of RNAP2 transcription that are independent of TBP, TFIIA, TFIIF, TFIIE and TFIIH have been amply documented2,3,4,5,6,7,8,9,10. Results from the Encyclopedia of DNA Elements (ENCODE) project revealed that factors previously considered specific, such as E2F1, are more prevalent at transcriptional start sites than many of the original general factors11,12, blurring the distinction between ‘general’ and ‘specific’ transcription factors. Supporting the lack of such distinction are cases where a specific factor such as YY1 is needed for basal transcription under certain conditions9. Correspondingly, statistical analysis of core promoters has failed to identify a straightforward sequence pattern for gene core promoters. Instead, numerous complex combinations of sequence elements have been identified1,13, suggesting that the sequence affects the minimal composition of the RNAP2 pre-initiation complex (PIC)14,15. Moreover, the local structure of the promoter has the potential to affect PIC composition as well. For example, negative supercoiling of the DNA template can abolish the requirement for TFIIH2,8 and TBP6,16. Presently, all lines of available evidence point to the existence of various RNAP2 PICs, the composition of which depends on the DNA promoter sequence, its local DNA dynamics17,18 and cellular conditions.
Notably, the general factor TFIIB, unlike all other RNAP2 general transcription factors, has never been reported as dispensable for transcription except in a recent report that demonstrated TFIIB knockdown did not affect expression of histone genes in Drosophila19. The role of TFIIB is still poorly understood20. Besides a critical role in transcription initiation, it has been proposed that TFIIB may influence start site selection and transcription termination by binding to the 3′ ends of genes21,22,23,24,25,26. In PIC chromatographic separations, this factor is known to elute separately from the rest of the known PIC components and its binding to promoter DNA is weak in the absence of TBP or YY19. In the context of the “classical” TATA-box adenovirus major late promoter (AdMLP), TFIIB is known to bind a pair of TFIIB recognition elements up- and downstream of the TATA box (BREu and BREd), stabilizing the TBP-TATA box complex and assisting in recruitment of the polymerase to the precise transcriptional start site (TSS) in vitro10,27,28,29,30,31,32. However, little if anything is known about the role of TFIIB as a general transcription factor in human and other mammalian cells where the large majority of promoters (~90%) are BRE-less/TATA-less13.
Whether results derived from studies of RNAP2 transcription initiation in vitro accurately model the requirements for RNAP2 transcription initiation in vivo, where the DNA is organized as a chromatin template in the highly dense nuclear environment, has not been systematically studied. Here we examine the RNAP2 transcription requirements for TFIIB in vivo, by analyzing data from the Gene Expression Atlas and the Encyclopedia of DNA Elements (ENCODE) project, by applying TFIIB knockdown in various cell types and analyzing cellular gene microarrays, cellular functions, cell cycle transitions, viral replication and viral gene expression in human cells. The results reveal surprising differences in the requirements for transcription between cellular and viral promoters and demonstrate that TFIIB is likely to be dispensable for transcription of many cellular genes. TFIIB however, is essential for HSV transcription and replication. In addition, we report a novel cell cycle TFIIB regulation and localization of acetylated TFIIB on the transcriptionally silent mitotic chromatids.
Results
ENCODE data reveal that overlap of TFIIB binding at TSS with TFIIF and RNAP2 in the human genome is partial and cell specific
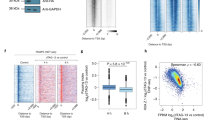
To define the cistrome of TFIIB, first we analyzed chromatin immunoprecipitation-tiling array (ChIP-Chip) data encompassing the human genome from the ENCODE Project11. We compared the distribution of RNAP2 and general transcription factor TFIIB and TFIIF binding sites, as determined by chromatin immunoprecipitation with antibodies specific to the RNAP2 α-subunit (POLR2A), TFIIB (GTF2B) and TFIIF (GTF2F1). The ENCODE data is from the human leukemia cell line K562. Within the 28292 presently known human genes33, 11569 bound to RNAP2. Thus, 41% of ENCODE region genes are actively transcribed by RNAP2 in K562 cells. Figure 1A shows the total number of sites bound to RNAP2, TFIIF and TFIIB and the fraction proximal (+/− 1000 bp) to “known gene” transcriptional start sites (TSS). Of the 11569 sites bound to RNAP2, 5682 mapped to TSS regions. Similarly, 9726 sites bound to TFIIB were detected, of which only 41% (3964) localized to TSS areas. Consistent with this, a study of the rat genome reported TFIIB binding predominantly in regions distinct from the established 5′-termini of protein coding genes34. Additionally, 21882 sites bound to TFIIF, 6647 of which were proximal to TSS sequences. As shown in Figure 1B, 82% (4636) of TSS that bound RNAP2 also bound TFIIF. In contrast, only 60% (3400) of TSS that bound RNAP2 also bound TFIIB. RNAP2 binding to TSS may reflect RNAP2 at TSS as well as paused RNAP2 and RNAP2 actively engaged in transcription in surrounding 1 kb of sequence, which are not expected to contain TFIIB as it is released from the promoter shortly after the initiation of transcription. Whereas 90% (3574) of TSS that bound TFIIB also bound TFIIF, only 53% (3518) of TSS with bound TFIIF also bound TFIIB. Together, these data raise the question of whether RNAP2 may initiate transcription in the absence of TFIIB on at least some genes, such as the recently described Drosophila histones19, under certain cellular conditions. These data support the paradigm that both TFIIB and TFIIF may play other roles besides being accessory proteins for RNAP2 transcription initiation.
TFIIB binding sites in relation to RNAP2 and transcriptional start sites.
Analysis of ENCODE chromatin immunoprecipitation-tiling microarray (ChIP-chip) data. (A) TFIIB and RNAP2 binding sites within “Known Gene” gene start sites in the hg19 human genome release from human K562 cells and fraction within +/−1 kb of TSS; (B) Overlap between TFIIB-, TFIIF- and RNAP2-bound sites within +/−1 kb of TSS in K562 cells. (C) Overlap between TFIIB- and RNAP2-bound sites within +/−1 kb of TSS in HL60 cells from the hg18 release (D) TFIIB protein immunofluoresence in mouse blastocyst; TFIIB (green) and DAPI (blue). The blastocyte domains are shown on the diagram at the right: red arrow - inner cell mass; blue arrow- outer thin layer.
We also determined the cistrome of TFIIB in the human promyelocytic leukemia cell line HL60 after stimulation with retinoic acid (32 h) to promote differentiation to mature granulocytes. The available pilot ENCODE ChIP-Chip data encompassing 1% of the human genome11 (Figure 1C) shows that, of 762 known TSSs, 21% (160) bound RNAP2 within +/−1000 bp of flanking sequence. Only 6% of TSS contained bound TFIIB. Of the 160 TSSs bound to RNAP2, only 17% also contained TFIIB within +/−1000 bp. Taken together, these data raise the possibility of potentially important gene and cell type differences in TFIIB utilization.
TFIIB expression varies widely in different mammalian cell types
To further understand transcription factor TFIIB function in cells, we examined TFIIB availability in different mammalian cell types. Analyzing the mRNA levels published in the Gene Expression Atlas and Allen Brain Atlas demonstrated that the ratio of transcripts for the large subunit of RNAP2 to TFIIB varied widely from 2 (in human fetal lung) to 32 (in alveolar cell lineRH41) (Supplemental Figure 1). Although transcript amounts may not reflect protein expression, in our own immunostaining experiments, TFIIB protein level was also found to vary significantly between cell types during murine embryogenesis. As shown for a mouse embryo at the blastocyst stage (Figure 1D), a high level of TFIIB protein was observed in the inner cell mass forming the embryoblast, whereas TFIIB protein was undetectable in cells from the outer thin layer trophoblast, which becomes the placenta. Though it is possible that TFIIB epitope accessibility may vary by cell type or during mouse development, a similar variation in TFIIB expression is observed during Xenopus embryogenesis (Supplemental Figure 2). Furthermore, TFIIB protein expression is induced in the adult rat central and peripheral nervous system upon injury35,36,37, decreased in primate brain after estadiol treatment38 and markedly up- or down-regulated by differentiation in some cell types36,39. These data prompted us to examine the functional consequences of TFIIB deficiency on RNAP2 transcription initiation in intact cells.
Loss of TFIIB is not lethal for human cells
We compared the effects of depleting TFIIB to depletion of transcription factor YY1, a ubiquitously distributed transcriptional repressor, in several human cultured cell lines, including HeLa, primary human umbilical vein endothelial cells (HUVEC) and primary human coronary smooth muscle cells by siRNA gene knockdown. Achieving transfection efficiency greater than 95% with two different siRNAs, the level of TFIIB protein was brought to less than 200 molecules per cell by 90 h post transfection (estimate based on Western blot, Figure 2A, Supplemental Figure 3). Such a level of a universally required factor would be vastly insufficient to sustain cellular transcription from all active genes. At 72 h (Figure 2) and 96 h (not shown) post siRNA transfection, all cell types showed viability (Figure 2B), active replication (Figure 2C) and no detectable TFIIB protein (Figure 2A). As a control, we knocked down the specific transcription factor YY1 and tested the effects of transcription factor deficiency on the cells. The loss of YY1 induced apoptosis in all cell types, while TFIIB knockdown cells were indistinguishable from cells transfected with a negative control siRNA (Figure 2B). Thymidine incorporation assay (Figure 2C) showed that DNA synthesis was significantly more affected by YY1 (60–80% decrease) than TFIIB knockdown (>10% decrease) in all cells.
Cellular response to TFIIB knockdown in human cells.
(A) TFIIB-siRNA knockdown in HeLa cells 90 h post transfection. TFIIB protein in cell lysate by Western blot is shown (lane 2); control siRNA (CsiRNA, lane 1), BsiRNA (lane 2) and transcription factor YY1siRNA (lane 3) are shown for comparison. Antibodies are indicated on the left and position of protein markers are on the right. TFIIB knockdown efficiency was also assessed by immunofluorescence: TFIIB (red) and DAPI (blue). The unprocessed image is shown in Supplemental Figure 6. (B) Apoptosis in cells with B-, YY1- and CsiRNA knockdown. The diagram presents the % of apoptotic cells from three independent experiments for the indicated cell types: C - control, B - BsiRNA, YY1 – YY1siRNA. The total amount of cells is taken as 100%; (C) Thymidine incorporation assay in siRNA-treated cells. Seventy hours after siRNA transfection, cells were given [3H]-thymidine for 12 hours: C-control, B- The data were calculated as the mean plus standard deviation from three experiments in triplicates BsiRNA, YY1 siRNA.
TFIIB deficiency produces specific alterations in the human transcriptome
Changes in cellular transcript levels in TFIIB knockdown HeLa were analyzed by gene-chip microarray at 90 h post transfection. Consistent with the observed resilience of the cells to TFIIB knockdown, only ~1% of the ~33,000 gene probes analyzed were differentially regulated >2.02-fold in response to TFIIB knockdown (Figure 3A). The data show 91 up-regulated genes and 150 down-regulated genes. Most of the up- and down-regulated genes with known promoter sequences have several TFIIB consensus binding sites (BRE) upstream and downstream of the TSS (Supplemental Figure 4). The BRE free HSP70 promoter confirmed to be unaffected. Pathway analysis using the Ingenuity™ Knowledge Base clustered the downregulated genes to specific cellular functions related to viral infectious diseases and cell growth (Supplemental Figure 5). In addition, members of the integrin family, actin cytoskeleton and calcium signaling pathways, including several proteins involved in cell adhesion, were dysregulated more than two-fold (Supplemental Tables S1, S2).
Gene expression profiling of HeLa cells in response to TFIIB knockdown.
(A) Of 33,000 genes in the Agilent Human Genome Microarray, 201 were down-regulated (green) and 91 were up-regulated (red) more than 2-fold (Supplemental Tables 1 and 2) as compared to control transfection. (B) Protein-protein binding network of TFIIB, according to the Ingenuity™ Knowledge Base (Ingenuity.com): Genes of proteins interacting with TFIIB binding partners that were up- (red) or down- (green) regulated by more than 2-fold are indicated. For clarity, only nodes connecting the affected genes to TFIIB are shown. (C) The mRNA content (in relative units on the vertical) of HDAC4, DR1, CLOCK, H2B, H2A, H1X, PC4, MTPN, TFIIB in TFIIB deficient cells (red) and wild type cells (blue) was analysed by qRT-PCR. The identity of the gene specific mRNA transcripts is indicated below the bars. The data were calculated as the mean plus standard deviation from three experiments in triplicates with normalization by 18S rRNA and presented as a comparison with the cells that received control siRNA. The blue bar represents the cells transfected with control siRNA and the values have been normalized to 1. * refers to statistical significance (t test, P < 0.05). (D) ChIP of the DR1 promoter region and TFIIB-specific antibody (αΒ) in HeLa cells. Lanes 1 and 3 show amplification of the DR1 promoter fragment before ChIP (total); lane 2- amplification of DR1 promoter fragments captured by the TFIIB antibody from CsiRNA treated cells; lane 4- amplification from BsiRNA treated cells. Lane 5 shows the captured and PCR amplified CsiRNA fragments with control pre immune antibody (αΧ). The unprocessed CHIP image is shown in Supplemental Figure 6. (E) RNA stability in response to CsiRNA (blue bars) and BsiRNA (red bars) was compared in actinomycin D treated cells (10 μg/ml). RNA was isolated at different time points (min) after adding actinomycin D as shown below the bars. The isolated total RNA (0.1 μg) was probed for H2B, H2A and PC4 mRNA by RT-PCR. The relative units values are normalized to 1 (control siRNA values). * refers to statistical significance (t test, P < 0.05).
Transcription factors that could substitute for human TFIIB on certain promoters may exist and remain to be established, but a universal replacement for TFIIB is unlikely as revealed by some in vitro transcriptional experiments (data not shown). In yeast, overexpression of the transcriptional co-activator PC4 homolog protein has been shown to rescue a cold-sensitive TFIIB mutant40. According to the gene microarray data, TFIIB knockdown caused human PC4 up-regulation by 2-fold, the threshold for statistically significant change in gene expression for this data set. To search for candidate TFIIB-redundant factors, we mapped the gene expression data onto the known TFIIB protein binding network, according to the Ingenuity™ Database (Figure 3B). Statistically significant changes in expression were identified by the microarray and RT-PCR (Figure 3C) for five genes of the TFIIB binding network. A 3-fold decrease was observed in the transcripts of histone deacetylase HDAC4, histone H1 variant X, down-regulator of transcription DR1 (NC2), CLOCK homolog and cAMP-responsive element binding protein 5. The only up-regulated gene was MTPN, a gene that encodes myotrophin and leucine zipper protein 6. No difference was observed in the expression of PC4 or histones H2A or H2B. As expected, the ChIP with TFIIB antibody revealed absence of TFIIB in the proximity of the DR1 promoter upon TFIIB knockdown (Figure 3D). Microarray analyses did not reveal significant changes in the expression of genes involved in global mRNA stability. Moreover, mRNA levels of H2B, H2A and PC4 in control siRNA and TFIIB siRNA knockdown cells after treatment with actinomycin D showed that TFIIB deficiency does not alter the mRNA stability of the poly(A)-less H2B and H2A mRNAs or the poly(A) containing PC4 mRNA (Figure 3E). The data suggest possible effects on chromatin remodeling or relief of DR1-mediated repression of transcription41 as two potential mechanisms of coping with TFIIB deficiency.
TFIIB silencing selectively inhibits HSV-1 transcription and viral plaque formation
Although in cell transcription from many cellular promoters appears to be resilient to TFIIB depletion (Figures 3), in vitro observations by us (not shown) and others42 show that TFIIB is an essential factor for several HSV-1 promoters. This difference in the requirement for TFIIB may enable selective inhibition of viral transcription and therefore viral replication, by TFIIB silencing. To test this directly, we infected TFIIB knockdown HeLa cells with HSV-142. Plaque assays of HSV-1 infected cells revealed that viral replication was reduced >7-fold in the TFIIB knockdown cells (Figure 4A). To examine the global effect of TFIIB knockdown on HSV-1 gene transcription we used a HSV-1 viral gene microarray (Figure 4B). Comparing the gene expression profiles of TFIIB knockdown and control HeLa cells in three independent array experiments, we observed that all herpes genes that produced a consistently readable signal in the microarray were downregulated 1.5 to 10-fold in response to TFIIB knockdown. Results from HSV-1 gene microarray were verified by RT-PCR on select genes (Figure 4C) as well as at the protein level with protein-specific antibody and immunofluorescence (Figure 4D). Consistent with our unpublished in vitro experiments, the HSV-1 gene array and RT-PCR experiments (Figure 4B) revealed that the viral TK (UL23) promoter was repressed more than 5 times in the absence of TFIIB. In contrast, the immediate early gene promoter ICP0 remained unperturbed at 3 hours post infection (Figure 4C), the time of peak activity during normal viral infection42. The ICP4 gene, which requires TFIIB in vitro (not shown) is essential for viral replication42. In addition to its role in basal transcription, TFIIB mediates HSV-1 transcriptional activation via interactions with the HSV proteins VP16 and ICP442, a mechanism of regulation presumably shared among the majority of herpes genes42. TFIIB knockdown may thus have a doubly inhibitory effect on herpes viral transcription in cells.
Effect of TFIIB knockdown on viral growth and viral gene transcription in HeLa cells.
(A) HSV-1 plaque assay in TFIIB (B) and control (C) siRNA knockdown HeLa cells (60 h post transfection). Mean viral growth (PFU/cell) is indicated (n = 6). (B) Genomic HSV-1 microarray analysis of viral gene expression in TFIIB knockdown HeLa cells transfected with siRNA and 60 h later infected with HSV-1 strain KOS at 2.5 PFU/cell. RNA was isolated 12 h post infection; control (dark blue bars) and TFIIB (sky blue). Genes are shown at right. (C) RT-PCR of the HSV-1 genes ICP0, ICP22, ICP4 in infected HeLa cells 12 h post infection relative to ARPP0 mRNA from negative control transfection. C-control siRNA knockdown cells; B- TFIIB knockdown cells. Results were consistent in four independent experiments. (D). The TFIIB and ICP4 immunofluorescence in virus-infected cells. Cells received either TFIIB or control siRNA and were stained with (a, d) anti-TFIIB antibody (green), (b, e) anti-ICP4 antibody (red) or (c, f) DNA with DAPI (blue).
In addition to the direct effects of TFIIB on HSV-1 gene transcription, transactivation42 and replication, TFIIB may indirectly affect viral gene expression and growth via alterations in the cellular transcriptome. Supporting this idea, an analysis of the changes in the human transcriptome in the absence of TFIIB using the Ingenuity™ Knowledge Base predicted inhibition of replication across a broad range of viral taxa, with the utmost statistical significance (Supplemental Figure 5a).
TFIIB expression and acetylation is governed by the cell cycle
Viruses commonly arrest cell cycle progression to facilitate replication, in part via cyclins and their interaction partners. TFIIB has high structure and sequence similarity to cyclin A31. Like the cyclins, TFIIB knockdown is not lethal to the cells43,44. Whether TFIIB also has cell cycle phase-specific regulation and functionality like cyclin A has not previously been explored. The experimental evidence we present here suggests that like the cyclins, the expression of TFIIB oscillates during the cell cycle in proliferating cells. In cultured HeLa cells released from nocodozole arrest, total TFIIB expression is elevated when a majority of cells are in the S phase and lowered as the cells progress to the G2 and M phase (Figure 5A).
Functions of TFIIB and the acetylated TFIIB variant in cell cycle transitions.
(A) TFIIB immunofluorescence (red) after release from nocodazole block (hours). DNA is stained with DAPI (blue). (B) DAPI flow cytometry of control siRNA (wt) or TFIIB siRNA (−B) treated cells after release from nocodazole block (hours). DNA content is shown below (2N, 4N). (C) Confocal microscopy of nonacetylated and acetylated TFIIB in HeLa cells visualized with nonacetylated TFIIB specific (αB) or acetyl-TFIIB (αAcB) specific antibodies by immunofluorescence (red) with DAPI staining (blue) during the indicated above the plots cell cycle phase. (D) A western blot (left) shows that the TFIIB antibody (αB) in a mixture with the 12 amino acids acetyl Lysine TFIIB peptide (acp) recognises only the nonacetylated recombinant TFIIB (rhnB, lane 1) and not acetylated recombinant TFIIB (rhacB, lane 2). The anti acetyl-TFIIB andibody (acB) (right blot) recognises the acetylated recombinant protein (lane 4) and not the nonacetylated TFIIB protein (lane 3). Ponceau S (S) staining of each blot before immunostainning is shown beneath the respective westerns. The unprocessed images are shown in Supplemental Figure 6.
FACS analyses of cells synchronized with nocodozole revealed that TFIIB knockdown altered the cell cycle by delaying the exit from G2/M and progression to S phase (Figure 5B) suggesting yet another similarity to cyclin A, which is required for entry into M and the completion of S43,44.
We have previously reported that TFIIB is an autoacetylase45. Protein acetylation plays a key role in mitotic progression46. To determine whether TFIIB is acetylated in a cell cycle phase-specific manner, we examined the subcellular distribution of non-acetylated and acetylated TFIIB during the cell cycle and mitosis by confocal microscopy (Figure 5C), using antibodies specific for each variant (Figure 5D). Non-acetylated TFIIB colocalized with DNA in the G0/1, S and G2 phases of cell cycle, but not during mitosis, consistent with studies showing GFP-tagged TFIIB is not bound to DNA during mitosis47. In contrast, the acetylated TFIIB variant colocalized to DNA during mitosis at prophase, metaphase, anaphase and telophase in addition to the other phases of the cell cycle, consistent with detection of TFIIB bound to the transcriptionally silent mitotic DNA by more sensitive ChIP analysis48.
The presence of the acetylated TFIIB variant during the different states of mitosis when RNAP2 ceases active transcription raises the question of its functional meaning. Acetylated TFIIB could serve as a “memory (epigenetic) imprint” for RNAP2 recruitment and bulk RNA synthesis of specific genes at the mitotic exit, for transcriptional reactivation of the genome. The exact function of the acetylated TFIIB variant during mitosis and the mitotic exit remains to be established. Nevertheless, the TFIIB function with its autocatalytic acetylation activity adds a new metabolic aspect to the elaborate cell cycle control system.
Discussion
The general transcription factor TFIIB is widely reported as an essential component for transcription initiation by RNA polymerase II, although RNAP2 transcription initiation independent of general transcription factors TBP, TFIIA, TFIIF, TFIIE and TFIIH is well reported2,3,4,5,6,7,8,9,10. Our data show that knockdown of TFIIB in human cells does not affect cell viability, DNA synthesis, or yield ≥2-fold changes in expression of most human genes in either primary or transformed cell lines. Though we cannot exclude that TFIIB knockdown globally affects mRNA stability, TFIIB knockdown in the presence of actinomycin D did not alter levels of either total RNA (not shown) or poly(A) and poly(A)-less mRNAs of select genes. Together, these data suggest that TFIIB is dispensable for in vivo RNAP2 transcription initiation on most human genes. In agreement with this, previous reports of TFIIB knockdown in trypanosome, Drosophila and mammalian cell lines did not report an impact of cell viability or large changes in expression of the few genes examined19,36,49. This is consistent with the viability of transgenic mice expressing mutant polyglutamine tract-expanded TBP, which causes low TFIIB occupancy of the HSPB1 promoter50 and presumably those of other genes, but in contrast to early RNAP2 in vitro studies which established the minimal requirements for RNAP2 transcription51.
In contrast to human gene transcription, we found TFIIB is required for transcription of HSV-1 genes as well as HSV-1 viral replication. The disparity between TFIIB requirements in human and herpesviral transcription may reflect differences in promoter organization, TFIIB autoacetylation45, and/or secondary interactions between TFIIB with human or viral transcription machinery20 and/or chromatin components52. Moreover, TFIIB may have indirect effects on viral replication via alterations in the human transcriptome. Indeed, in cells subjected to TFIIB knockdown, Ingenuity™ Pathway Analysis predicts impaired replication of a wide range of viruses. Taken together with previous studies showing transcription factors from multiple viruses complex with TFIIB to transactivate viral transcription, our data suggest that blockade of TFIIB may have broad anti-viral effects.
Our data support the idea that cellular TFIIB expression is dynamically regulated. TFIIB expression changes in a cell cycle dependent manner and, relative to that of RNAP2, varies widely between cell types. Of note, TFIIB promotes certain transcriptome changes that drive cell growth and cell cycle progression. Consistent with this, cell differentiation markedly downregulates TFIIB expression39 and TFIIB-homolog depletion decreases growth rate in a trypanosome-derived cell line49. Similarly, this effect may also underlie the recently described association between TFIIB overexpression and tissue regeneration36, human hepatocellular carcinogenesis53 and the essential role of TFIIB in specific cases of viral transcription. In contrast, TFIIB expression also increases during Muller glia cell differentiation in vitro, whereas TFIIB knockdown decreased expression of proliferation and differentiation markers, suggesting that the impacts of TFIIB on cell growth may be cell specific, context dependent and complex36. Indeed, while TFIIB may not be necessary for cell viability and proliferation in higher eukaryotes, it may be required for cell differentiation, dedifferentiation, and/or development.
Both archaeabacteria and plants contain multiple copies of TFIIB homologs54,55. In archaeabacteria, individual isoforms of the TFIIB homolog TFB coordinate discrete environment-specific gene regulatory programs54. These data, together with our own suggest that TFIIB is likely to be a tissue and cell type specific determinant of cell growth via its effects on the cellular transcriptome. In addition, viruses may have evolved a dependence on TFIIB as a “sensor” of cellular resources to support viral growth and replication, a notion that remains to be established. In yeast, TFIIB is thought to be essential for cell viability and a recent genome-wide analysis revealed TFIIB crosslinking to TSSs of a vast majority of transcriptionally active genes56. As previous work on yeast and archaeal homologs of TFIIB assayed the impact of TFIIB mutagenesis on colony formation, future studies are needed to address whether the yeast or archaeabacterial TFIIB homologs affects cell viability or only cell growth and whether TFIIB utilization in these organisms is environment-specific.
Our data also support a role of TFIIB in regulating gene expression beyond its classically recognized function within the PIC. Analysis of ENCODE Project ChIP-Chip data encompassing the human genome11 revealed 59% of TFIIB bound outside TSS areas, consistent with previous findings in the rat genome that showed TFIIB binding was not restricted to promoter regions34 and in yeast and human cells that TFIIB localized to the 3′ end of genes21,22,23. Taken together, these data suggest that TFIIB has functions beyond transcription initiation and apart from the TTS. In support of this, recent work shows TFIIB localization to the 3′ ends of genes links transcription initiation and termination via gene looping or promotes transcription of anti-sense RNA initiating from the 3′ end of genes21,22,23,24,25,26. Indeed, targeting of yeast chromatin remodeling enzyme Isw2 to chromatin was recently reported to be mediated in part by TFIIB dependent DNA looping57. Moreover, our data reveal that acetylated TFIIB is bound to mitotic chromosomes, when transcription ceases and that TFIIB loss alters the exit from G2/M and progression to S phase, suggesting TFIIB may have functions beyond its classical role in gene expression.
The presented data, taken together with recent work from others, constitutes a fundamental paradigm shift in the understanding of TFIIB functionality in gene expression. While the precise functions of TFIIB in cellular gene expression remain to be established, our findings support the concept that the composition of the PIC is variable and likely reflects promoter sequence specific DNA dynamics and availability of factors that can bind to and regulate the promoter1,14,15,58. Factor availability may be regulated by cell-environment interactions, including metabolism. The acetylation of TFIIB may be a new regulatory pathway in chromatin regulation and DNA replication. Uncovering cell cycle-specific functions of the TFIIB autoacetylatase reveals new aspects of mechanistic integration of RNAP2 transcription machinery with DNA replication, the cell cycle and metabolism. The implications of such connections would be far-reaching in cellular biology. The territories or chromosomal compartments that are maintained by the acetylated TFIIB variant may orchestrate coordinated entry into the next phase of the cell cycle and remain to be discovered.
Methods
Cellular TFIIB and YY1 knockdown
TFIIB knockdown in HeLa cells was accomplished by electroporation with the siRNA-specific reagent (MediMAbs) of endotoxin treated (DETOX, MediMabs, Canada) 1 nmol TFIIB-specific siRNA mix (Invitrogen) per 106 cells or YY1-specific siRNA (Invitrogen) and Silencer® negative control siRNA (Ambion) as previously reported45. All transfections are performed with the siRNA specific reagent TRANS siRNA(MMM-103, MediMabs, Canada). siRNA is purified from endotoxin contaminations with the DETOS reagent (MMM-104, MediMAbs, Canada).
Apoptosis, RNA stability assays and ChIP
Apoptotic cells were identified by incubation with 5 μM propidium iodide, followed by fluorescence microscopy with a Texas Red emission filter (632 nm) to detect nuclear staining. The mRNA stability for some genes that did not change level of expression in response to TFIIB deficiency was identified by treating the cells with actinomycin D (10 μg/ml). Total RNA was isolated from cells at different time points. Content of specific mRNAs was measured by RT-PCR. ChIP with anti TFIIB antibody was conducted as we previously reported49. The ChIP primers for the human DP1 encompass 704 bp including 100 bp downstream of the TSS: plus primer 5′-TCCTACTTAGCCATTAGGCGAC-3′; minus primer 5′-CCAGATGCCAGGGAAGGTTT-3′.
Cellular and viral microarrays analysis
Total RNA isolated from HeLa cells 80 h post transfection with TFIIB-specific or Silencer® negative control siRNA were analyzed. Two identical microarrays (Agilent Whole Human Genome Oligo Microarray Kit) were used to compare the two pools. RNA was quantified and the quality of the samples was examined using RNA 6000 Nano LabChip® Kit on Agilent 2100 Bioanalyzer. Cyanine 3- or 5-labeled CTP (10.0 mM) were purchased from Perkin–Elmer/NEN Life Science. Fluorescently labeled cRNA targets were generated from 500 ng of total RNA for each reaction using the Agilent Fluorescent Linear Amplification Kit. Each RNA mix was labeled with both cyanine 3 and cyanine 5 to allow dye reversal experiments and minimize dye bias. Hybridization was performed using the Agilent's in situ Hybridization Plus kit with 1 μg labeled cRNA. The arrays were scanned using the Agilent dual-laser DNA microarray scanner and data were extracted with Feature Extraction 9.1. The GeneSpring software (Agilent) was used to generate lists of selected genes and for different statistical and visualization methods. Lowess normalization was applied to correct for artifacts caused by nonlinear rates of dye incorporation as well as inconsistencies of the relative fluorescence intensity between some red and green dyes. Lists of differentially expressed genes considering a 2-fold expression cutoff were generated by filtering on expression level using the data from all independent experiments for each condition. The genes in the gene lists were classified according to their function using the Gene Ontology (GO SLIMS) classification system. The accession code of the array data is GSE48847.
A microarray (SABiosciences, Frederick, MD) printed with all HSV ORFs and its application to compare viral genes levels in siRNA treated cells was previously described in detail59,60,61. The data were analyzed with GEArray Expression Analysis Suite 2.0. Arrays were standardized by using the interquartile method and the data are presented as the fold change in expression of heat-shocked versus mock heat-shocked control. The data represent the averages of three independent experiments.
Infection with HSV-1
HeLa cells were infected with the wild-type virus HSV-1 strain KOS at passage 13 that was propagated as described previously59.
RT-PCR
The ICP0, ICP4, ICP22 primers for the RT-PCR have been published elsewhere59,60,61. The following primers have been used to assess the expression of genes in TFIIB deficient and control HeLa cells: H2A (NM_003528) forward (f) atgtctggacgtggccaagca, reverse (r) agcttgttgagctcctcgtg; histone H2B/j (AF531291) f - ccgaagaagggctccaagaa, r - ttatttggagctggtgtacttg; clock f-gcaaaatgtcatgagcacttaatg, r - ctgcagcccctgaccatggacc; CREB5 (NM 001011666) f- aggtctgggtgatgtcattg, r - atggctgttattgggcagtc; HDAC4 f- aatctgaaccactgcatttcca, r- ggtggttataggaggtcgacact; protocadherin 9 (PCDH9) f –acagccaccacggtcctcta, r – cccttgttgttcccgctcac; GTF2B f-ctggaggagcccccatc, r- cagcaatatctccaatctctttttg; histone H1X (BC000426)f-gatctacaccgaggccaaga, r-cttcttgcggttgagcttg; PC4 (NM_ 006713.3) f- tcaagctcttctggcagtga, r - atctctgctgctgctgctct. Three sets of control siRNA and BsiRNA isolations were assayed in triplicates. Relative gene expression values were calculated after normalization to 18S rRNA that did not change significantly in response to siRNA treatment and adjust the value of the control siRNA sample as 1. Results obtained in individual experiments were expressed as mean ± SEM.
Immunofluorescence and Western blotting
The nocodazole-synchronized and serum stimulated cells were stained with rabbit anti-acetylTFIIB antibody (Abcam) or with anti-nonacetylated TFIIB and Cy3-labeled secondary antibody. The polyclonal TFIIB antibody was pre incubated with the acetylated TFIIB peptide (Abcam). Such treatment resulted in specific recognition of the nonacetylated TFIIB eliminating crossreactivity with the acetylated TFIIB (Figure 5C). ICP4 was visualized with HSV-1 ICP4 (H943) antibody (Santa Cruz) and YY was visualized with YY1-MM-0205-P chicken antibody (MediMabs, Canada).
References
Juven-Gershon, T., Hsu, J. Y., Theisen, J. W. & Kadonaga, J. T. The RNA polymerase II core promoter - the gateway to transcription. Curr Opin Cell Biol 20, 253–9 (2008).
Goodrich, J. A. & Tjian, R. Transcription factors IIE and IIH and ATP hydrolysis direct promoter clearance by RNA polymerase II. Cell 77, 145–56 (1994).
Holmes, M. C. & Tjian, R. Promoter-selective properties of the TBP-related factor TRF1. Science 288, 867–70 (2000).
Jacobi, U. G. et al. TBP paralogs accommodate metazoan- and vertebrate-specific developmental gene regulation. Embo J 26, 3900–9 (2007).
Jallow, Z., Jacobi, U. G., Weeks, D. L., Dawid, I. B. & Veenstra, G. J. Specialized and redundant roles of TBP and a vertebrate-specific TBP paralog in embryonic gene regulation in Xenopus. Proc Natl Acad Sci U S A 101, 13525–30 (2004).
Leblanc, B. P., Benham, C. J. & Clark, D. J. An initiation element in the yeast CUP1 promoter is recognized by RNA polymerase II in the absence of TATA box-binding protein if the DNA is negatively supercoiled. Proc Natl Acad Sci U S A 97, 10745–50 (2000).
Parvin, J. D., Shykind, B. M., Meyers, R. E., Kim, J. & Sharp, P. A. Multiple sets of basal factors initiate transcription by RNA polymerase II. J Biol Chem 269, 18414–21 (1994).
Parvin, J. D. & Sharp, P. A. DNA topology and a minimal set of basal factors for transcription by RNA polymerase II. Cell 73, 533–40 (1993).
Usheva, A. & Shenk, T. YY1 transcriptional initiator: protein interactions and association with a DNA site containing unpaired strands. Proc Natl Acad Sci U S A 93, 13571–6 (1996).
Van Dyke, M. W., Roeder, R. G. & Sawadogo, M. Physical analysis of transcription preinitiation complex assembly on a class II gene promoter. Science 241, 1335–8 (1988).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Bieda, M., Xu, X., Singer, M. A., Green, R. & Farnham, P. J. Unbiased location analysis of E2F1-binding sites suggests a widespread role for E2F1 in the human genome. Genome Res 16, 595–605 (2006).
Gershenzon, N. I. & Ioshikhes, I. P. Synergy of human Pol II core promoter elements revealed by statistical sequence analysis. Bioinformatics 21, 1295–300 (2005).
Muller, F., Demeny, M. A. & Tora, L. New problems in RNA polymerase II transcription initiation: matching the diversity of core promoters with a variety of promoter recognition factors. J Biol Chem 282, 14685–9 (2007).
Juven-Gershon, T., Hsu, J. Y. & Kadonaga, J. T. Perspectives on the RNA polymerase II core promoter. Biochem Soc Trans 34, 1047–50 (2006).
Usheva, A. & Shenk, T. TATA-binding protein-independent initiation: YY1, TFIIB and RNA polymerase II direct basal transcription on supercoiled template DNA. Cell 76, 1115–21 (1994).
Alexandrov, B. S. et al. DNA dynamics play a role as a basal transcription factor in the positioning and regulation of gene transcription initiation. Nucleic Acids Res 38, 1790–5 (2010).
Alexandrov, B. S. et al. DNA breathing dynamics distinguish binding from nonbinding consensus sites for transcription factor YY1 in cells. Nucleic Acids Res 40, 10116–23 (2012).
Guglielmi, B., La Rochelle, N. & Tjian, R. Gene-specific transcriptional mechanisms at the histone gene cluster revealed by single-cell imaging. Mol Cell 51, 480–92 (2013).
Deng, W. & Roberts, S. G. TFIIB and the regulation of transcription by RNA polymerase II. Chromosoma 116, 417–29 (2007).
Singh, B. N. & Hampsey, M. A transcription-independent role for TFIIB in gene looping. Mol Cell 27, 806–16 (2007).
El Kaderi, B., Medler, S., Raghunayakula, S. & Ansari, A. Gene looping is conferred by activator-dependent interaction of transcription initiation and termination machineries. J Biol Chem 284, 25015–25 (2009).
Wang, Y., Fairley, J. A. & Roberts, S. G. Phosphorylation of TFIIB links transcription initiation and termination. Curr Biol 20, 548–53 (2010).
Mapendano, C. K., Lykke-Andersen, S., Kjems, J., Bertrand, E. & Jensen, T. H. Crosstalk between mRNA 3′ end processing and transcription initiation. Mol Cell 40, 410–22 (2010).
Medler, S. et al. Evidence for a complex of transcription factor IIB with poly(A) polymerase and cleavage factor 1 subunits required for gene looping. J Biol Chem 286, 33709–18 (2011).
Goel, S., Krishnamurthy, S. & Hampsey, M. Mechanism of start site selection by RNA polymerase II: interplay between TFIIB and Ssl2/XPB helicase subunit of TFIIH. J Biol Chem 287, 557–67 (2012).
Bushnell, D. A., Westover, K. D., Davis, R. E. & Kornberg, R. D. Structural basis of transcription: an RNA polymerase II-TFIIB cocrystal at 4.5 Angstroms. Science 303, 983–8 (2004).
Evans, R., Fairley, J. A. & Roberts, S. G. Activator-mediated disruption of sequence-specific DNA contacts by the general transcription factor TFIIB. Genes Dev 15, 2945–9 (2001).
Deng, W. & Roberts, S. G. Core promoter elements recognized by transcription factor IIB. Biochem Soc Trans 34, 1051–3 (2006).
Elsby, L. M., O'Donnell, A. J., Green, L. M., Sharrocks, A. D. & Roberts, S. G. Assembly of transcription factor IIB at a promoter in vivo requires contact with RNA polymerase II. EMBO Rep 7, 898–903 (2006).
Nikolov, D. B. et al. Crystal structure of a TFIIB-TBP-TATA-element ternary complex. Nature 377, 119–28 (1995).
Lagrange, T., Kapanidis, A. N., Tang, H., Reinberg, D. & Ebright, R. H. New core promoter element in RNA polymerase II-dependent transcription: sequence-specific DNA binding by transcription factor IIB. Genes Dev 12, 34–44 (1998).
Hsu, F. et al. The UCSC Known Genes. Bioinformatics 22, 1036–46 (2006).
Yochum, G. S., Rajaraman, V., Cleland, R. & McWeeney, S. Localization of TFIIB binding regions using serial analysis of chromatin occupancy. BMC Mol Biol 8, 102 (2007).
Liu, Z. et al. Increased expression of transcription initiation factor IIB after rat traumatic brain injury. J Mol Histol 42, 265–71 (2011).
Xu, Y. et al. Muller Glia Cells Activation in Rat Retina After Optic Nerve Injury: Spatiotemporal Correlation with Transcription Initiation Factor IIB. J Mol Neurosci 51, 37–46 (2013).
Yang, J. et al. Transcription initiation factor IIB involves in Schwann cell differentiation after rat sciatic nerve crush. J Mol Neurosci 49, 491–8 (2013).
Wang, J. et al. Estradiol alters transcription factor gene expression in primate prefrontal cortex. J Neurosci Res 76, 306–14 (2004).
Shiraishi, S., Tamamura, N., Jogo, M., Tanaka, Y. & Tamura, T. A. Rapid proteasomal degradation of transcription factor IIB in accordance with F9 cell differentiation. Gene 436, 115–20 (2009).
Knaus, R., Pollock, R. & Guarente, L. Yeast SUB1 is a suppressor of TFIIB mutations and has homology to the human co-activator PC4. Embo J 15, 1933–40 (1996).
Masson, P., Leimgruber, E., Creton, S. & Collart, M. A. The dual control of TFIIB recruitment by NC2 is gene specific. Nucleic Acids Res 36, 539–49 (2008).
Smith, C. A., Bates, P., Rivera-Gonzalez, R., Gu, B. & DeLuca, N. A. ICP4, the major transcriptional regulatory protein of herpes simplex virus type 1, forms a tripartite complex with TATA-binding protein and TFIIB. J Virol 67, 4676–87 (1993).
Gong, D. & Ferrell, J. E., Jr The roles of cyclin A2, B1 and B2 in early and late mitotic events. Mol Biol Cell 21, 3149–61 (2010).
Kalaszczynska, I. et al. Cyclin A is redundant in fibroblasts but essential in hematopoietic and embryonic stem cells. Cell 138, 352–65 (2009).
Choi, C. H., Hiromura, M. & Usheva, A. Transcription factor IIB acetylates itself to regulate transcription. Nature 424, 965–9 (2003).
Gabrielli, B. & Brown, M. Histone deacetylase inhibitors disrupt the mitotic spindle assembly checkpoint by targeting histone and nonhistone proteins. Adv Cancer Res 116, 1–37 (2012).
Chen, D., Hinkley, C. S., Henry, R. W. & Huang, S. TBP dynamics in living human cells: constitutive association of TBP with mitotic chromosomes. Mol Biol Cell 13, 276–84 (2002).
Christova, R. & Oelgeschlager, T. Association of human TFIID-promoter complexes with silenced mitotic chromatin in vivo. Nat Cell Biol 4, 79–82 (2002).
Palenchar, J. B., Liu, W., Palenchar, P. M. & Bellofatto, V. A divergent transcription factor TFIIB in trypanosomes is required for RNA polymerase II-dependent spliced leader RNA transcription and cell viability. Eukaryot Cell 5, 293–300 (2006).
Roizman, B. in Fields Virology, 4th Edition (eds. Knipe, D. M. & Howley, P. M.) 345–377 (Lippincott-Raven, Philadelphia, 2001).
Thomas, M. C. & Chiang, C. M. The general transcription machinery and general cofactors. Crit Rev Biochem Mol Biol 41, 105–78 (2006).
Sharp, P. A. Gene transcription. TFIIB or not TFIIB? Nature 351, 16–8 (1991).
Li, L. et al. General Transcription Factor IIB Overexpression and a Potential Link to Proliferation in Human Hepatocellular Carcinoma. Pathol Oncol Res 19, 195–203 (2013).
Turkarslan, S. et al. Niche adaptation by expansion and reprogramming of general transcription factors. Mol Syst Biol 7, 554 (2011).
Knutson, B. A. Emergence and expansion of TFIIB-like factors in the plant kingdom. Gene 526, 30–8 (2013).
Rhee, H. S. & Pugh, B. F. Genome-wide structure and organization of eukaryotic pre-initiation complexes. Nature 483, 295–301 (2012).
Yadon, A. N., Singh, B. N., Hampsey, M. & Tsukiyama, T. DNA Looping Facilitates Targeting of a Chromatin Remodeling Enzyme. Mol Cell 50, 93–103 (2013).
Parvin, J. D., Timmers, H. T. & Sharp, P. A. Promoter specificity of basal transcription factors. Cell 68, 1135–44 (1992).
Kushnir, A. S., Davido, D. J. & Schaffer, P. A. Role of nuclear factor Y in stress-induced activation of the herpes simplex virus type 1 ICP0 promoter. J Virol 84, 188–200 (2010).
Chen, S. H. et al. Neither LAT nor open reading frame P mutations increase expression of spliced or intron-containing ICP0 transcripts in mouse ganglia latently infected with herpes simplex virus. J Virol 76, 4764–72 (2002).
Danaher, R. J., Jacob, R. J. & Miller, C. S. Establishment of a quiescent herpes simplex virus type 1 infection in neurally-differentiated PC12 cells. J Neurovirol 5, 258–67 (1999).
Acknowledgements
This paper is dedicated to the memory of Dr. Priscilla Schaffer, an extraordinary virologist and a wonderful colleague and mentor. We thank Jose Antao and Joseph Garlick from the Kingston laboratory for providing the HSP70 pIR17-84 plasmid. We acknowledge support from the Milton Fund (AU).
Author information
Authors and Affiliations
Contributions
A.U. conceived the studies and supervised the research. V.G., J.M.Z. and P.A.S. provided conceptual advice. V.G., J.M.Z., P.A.S. and A.U. designed the experiments and V.G., M.L., M.H., J.S.O., A.K., N.K., D.B. and A.U. performed the experiments. All authors conducted data analyses. V.G., J.M.Z. and A.U. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Accession codes: GSE48847.
Electronic supplementary material
Supplementary Information
Supplementary Info File #1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Gelev, V., Zabolotny, J., Lange, M. et al. A new paradigm for transcription factor TFIIB functionality. Sci Rep 4, 3664 (2014). https://doi.org/10.1038/srep03664
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03664
This article is cited by
-
Beyond the canonical role of TFIIB in eukaryotic transcription
Current Genetics (2022)
-
Characterization and mitigation of gene expression burden in mammalian cells
Nature Communications (2020)
-
Consensus transcriptional regulatory networks of coronavirus-infected human cells
Scientific Data (2020)
-
An osmolality/salinity-responsive enhancer 1 (OSRE1) in intron 1 promotes salinity induction of tilapia glutamine synthetase
Scientific Reports (2020)
-
Gene looping facilitates TFIIH kinase-mediated termination of transcription
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.