Abstract
We propose a bacteria-based microrobot (bacteriobot) based on a new fusion paradigm for theranostic activities against solid tumors. We develop a bacteriobot using the strong attachment of bacteria to Cy5.5-coated polystyrene microbeads due to the high-affinity interaction between biotin and streptavidin. The chemotactic responses of the bacteria and the bacteriobots to the concentration gradients of lysates or spheroids of solid tumors can be detected as the migration of the bacteria and/or the bacteriobots out of the central region toward the side regions in a chemotactic microfluidic chamber. The bacteriobots showed higher migration velocity toward tumor cell lysates or spheroids than toward normal cells. In addition, when only the bacteriobots were injected to the CT-26 tumor mouse model, Cy5.5 signal was detected from the tumor site of the mouse model. In-vitro and in-vivo tests verified that the bacteriobots had chemotactic motility and tumor targeting ability. The new microrobot paradigm in which bacteria act as microactuators and microsensors to deliver microstructures to tumors can be considered a new theranostic methodology for targeting and treating solid tumors.
Similar content being viewed by others
Introduction
Chemotherapy, the most common type of tumor treatment, is an effective conventional therapy against actively proliferating tumor cells. However, this therapy can damage other types of fast-growing, healthy normal cells, such as blood and hair cells, in the process of treating cancer cells, inducing various adverse reactions, or side effects. Chemotherapy can be the resistance to therapeutic response in the slowly cell proliferating hypoxic region by deficient angiogenesis1,2,3. Therefore, one of the most significant challenges of chemotherapeutic treatments is the continuous, specific delivery of optimal quantities of drugs to target cells. To overcome this challenge, many research groups have investigated the development of a drug delivery system (DDS) by using biocompatible and biodegradable materials, which can control the drug release from microstructures to alleviate side effects4,5,6,7. Although many DDSs have been developed, they still have difficulty in effectively delivering drugs to cancer sites because they lack active motility and because physical barriers, such as malformed blood vessels, elevated interstitial pressure and large transport distances in the tumor interstitium, exist in solid tumors2,3,5,8.
Recently, various types of microrobots have been proposed because of their many advantages in various fields, such as medicine, environment monitoring, space and military. Especially, to overcome the drawbacks of conventional chemotherapy using DDSs, various types of biomedical microrobots have been proposed owing to their potential in biomedical applications9,10,11. In the development of microrobots, the most challenging aspects are the fabrication and integration of microactuators and microsensors with high stability and efficiency. So, many research groups have tried to overcome these problems by combining several technologies, such as the micro/nano electro-mechanical system (MEMS/NEMS), nanotechnology and biotechnology11,12. Although the microrobots have much potential in biomedical applications, difficulties, such as the implementation of sensing and therapeutic functions, the integration of actuation and the control of power supply in small bodies still remain13,14. For effective control and actuation of a microrobot, several researchers suggested the magnetic metal-based microrobot, which is composed of a neodymium-iron-boron, soft-magnetic metal actuated by electromagnetic coil systems15,16. In addition, our group also developed a microrobot by using the magnetic materials and controlled it through external magnetic fields in a two-pair coil system17. However, these microrobot systems require several pairs of complex electromagnetic coil systems. For microrobot actuation, microorganisms with high motility, such as Escherichia coli (E. coli), Salmonella typhimurium (S. typhimurium), Serratia marcescens (S. marcescens) and magnetotactic bacteria (MTB) strains (Magnetospirillum gryphiswaldense strain MSR-1, Magnetospirillum magnetotacticum strain MS-1, Magnetospirillum magneticum strain AMB-1 and Magnetococcus strain MC-1) have been used18,19,20,21,22. However, several bacteria strains, such as E. coli, S. marcescens and MTB strains still have several weak points which limit their use as microactuators for biomedical microrobots: complex incubation procedure, drug resistance against antibiotics and pathogenicity in living animals23. Recently, we applied genetically modified S. typhimurium for treatment and diagnosis of solid tumors, such as colorectal and breast cancers, because of its propensity to naturally accumulate in tumors of small living animals, particularly hypoxic tumors24,25,26,27. The bacteria have been used alone or in combination with conventional therapeutics and have shown antitumor efficacy28,29. Since drug production using engineered bacteria is currently limited, bacteria synthesis must be fine-tuned to achieve therapeutic levels. But, the levels still cause systemic toxicity25.
A prototype of the bacteria-based microrobot was fabricated by combining a polystyrene (PS) microbead and flagellated bacteria14,19,30. However, it was used only simple properties that bacteria can be attached to the surface of a hydrophobic PS microbead. Because one of the most important factors in the fabrication of a bacteria-based microrobot is the method of attaching flagellated bacteria to the microstructure, many researchers have tried to develop attachment and pattering between bacteria and microstructures19,20,22,31. To fabricate selectively bacteria-attached PS microbeads, M. Sitti used the reactive ion etching (RIE) plasma-based patterning technique to induce the hydrophilic condition on the surface of the microbeads19. Moreover, we have focused on the development of various types of bacteria-based microrobots by using several bacteria-patternable materials. First, flagellated S. marcescens could be patterned onto hydrophobic SU-8 microcubes by using non-fouling proteins, such as bovine serum albumin (BSA)22. Second, flagellated S. typhimurium could be regulated to attach to surfaces of hydrophilic poly(ethylene glycol) (PEG) microbeads by using poly-L-lysine (PLL)31. However, these bacterial attachment methodologies were not verified for their applicability in living animals.
In this study, we propose a new methodology for bacteria-based microrobot fabrication and verify the tumor targeting property of a bacteria-based microrobot, referred to as “bacteriobot”, composed of attenuated S. typhimurium attached to a microstructure. These bacteria enable the microrobot to move toward tumors, acting as a combination of microsensor, microactuator and therapeutic agent; the microstructure acts as a therapeutic molecule containing high amounts of drugs. The bacteriobot can be considered as a new type of active DDS with the various useful properties of a microorganism for synergistic therapy against incurable diseases such as cancer. In addition, the bacteriobot based on a new fusion paradigm of RT (robot technology) and BT (biotechnology) presents a novel anti-tumor strategy.
Results
Development of bacteriobots using interaction between biotin and streptavidin
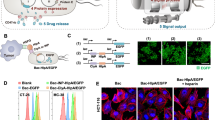
In this study, we used a high-motility strain of attenuated S. typhimurium defective in guanosine 5′-diphosphate-3′-diphosphate (ppGpp) synthesis (ΔppGpp strain) and expressing bacterial luciferase (lux) or green fluorescent protein (gfp)24. The bacteria were strongly attached to the microstructure by exploiting the high-affinity interaction between biotin and streptavidin (Fig. 1a). Biotin is a small molecule that exists in all living organisms and can be easily conjugated to many proteins, such as streptavidin without significant loss of biological activity; conjugation of a protein to biotin permits that protein to interact with streptavidin-conjugated molecules. In this case, bacteria were engineered to display biotin in the outer membrane proteins (omps), which are widely distributed on the bacterial surface31; these bacteria were then attached to microstructures consisting of rhodamine-containing fluorescent PS carboxylated microbeads that were covalently coupled to streptavidin-conjugated tandem fluorochrome composed of peridinin chlorophyll protein (PerCP), which was further labeled by Cy5.5 (PerCP-Cy5.5). The fluorophores were used to obtain a near-infrared fluorescence image for tracking purposes. Figure 1b shows the confocal laser scanning microscopy imaging of the bacteriobots that were fabricated by attaching the attenuated gfp-expressing biotinylated ΔppGpp strain to fluorescent PS microbeads coated with streptavidin-PerCP-Cy5.5. We obtained an actual image of a bacteriobot by collecting the images of stack acquisition using confocal laser scanning microscopy and acquired a 3D bacteriobot image by using the iso-surface algorithm of 3D reconstruction software (Supplementary Movie 1).
Development of bacteriobots using biotin-streptavidin conjugation.
(a) Schematic representation of bacteriobots. Biotin (500 μg) was incubated with omp-expressing S. typhimurium (3 × 108 cells/mL) for 1 hr. Rhodamine-containing fluorescent carboxylated PS microbeads (1 × 108/ml) were covalently coupled to streptavidin-PerCP-Cy5.5 (500 μg). Biotin-displaying S. typhimurium and streptavidin-PerCP-Cy5.5-coated PS microbeads were co-incubated for 30 min at 37°C. (b) S. typhimurium-attached PS microbeads were observed using a confocal laser scanning microscope.
Development of flow-free chemotactic microfluidic chamber for motility assessment of bacteriobot
We expect that the proposed bacteriobot can show tumor targeting performance similar to that of the bacteria, which can move toward a solid tumor. The motility of the bacteria can strongly give influence the motility of the bacteriobot as a microactuator and microsensor. To evaluate the tumor-targeting performance of the bacteriobot, a flow-free chemotactic microfluidic chamber containing cell lysates or spheroids was developed (Fig. 2a). The device contains three aligned chambers connected by micro-tubing. The central chamber contains a solution of bacteria and/or bacteriobots, while the chambers on both sides contain solutions of cell lysates or spheroids. First, the spatial and temporal blue (Trypan blue (GIBCO BRL, Gaithersburg, MD)) and red (Safranin (Sigma-Aldrich Chemical Co. St. Louis, MO)) dyes concentration profiles within the gradient-generating microfluidic chamber were computed using COMSOL. For the COMSOL simulation, we used a diffusion coefficient D = 1 × 10−9 m2/sec for the trypan blue and the safranin red dye solutions and assumed the diffusion of the dyes in deionized water at 25°C. Second, we carried out a preliminary test using the gradient-generating microfluidic chamber with 0.1% trypan blue dye and 0.28% safranin red dye of molecular weights (MW) of 960.8 and 350.9, respectively. The chemical concentration gradients in the microfluidic chamber were established with the colored dyes (blue and red) and maintained for 30 min (Fig. 2b and c, middle figures). Bottom figures of Fig. 2b and c show the detailed concentration line profiles and the actual gradient of the colored dyes across the center channel after the colored dyes were filled at t = 30 min. Moreover, the concentration gradient of the colored dyes was established 10 min after injecting the chemicals and maintained for over the 30 min in the chemotactic microfluidic chamber, which was enough time for the measurement of the bacteria and/or bacteriobot chemotaxis. Consequently, the microfluidic device allowed the diffusion of cell lysate molecules or cell spheroids from each side chamber through the micro-tubing toward the central chamber in a manner that generated a chemical concentration gradient in the preliminary test.
Schematic representation of gradient-generating microfluidic device for motility and velocity evaluations of bacteria and/or bacteriobots.
(a) The microfluidic device includes two side chambers, loaded with cell lysates or cell spheroids (left or right) and a central chamber, loaded with bacteria or bacteriobots. (b) COMSOL model showing colored dyes gradients in the gradient-generating microfluidic channel for 30 min. (c) Continuous concentration gradient formation by colored dyes (blue and red) in the gradient-generating microfluidic channel for 30 min.
Tumor targeting and localization of bacteriobot
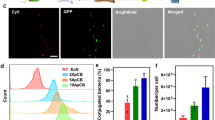
In the chemotactic microfluidic chamber containing cell lysates or spheroids, the chemotactic response of the bacteria and/or bacteriobots to the concentration gradients of various cells, such as NIH/3T3, CT-26 (colorectal cancers) and 4T1 (breast cancers) lysate molecules or spheroids was detected by measuring the migration of the bacteria and/or bacteriobots out of the central chamber toward the side chambers. Bacteria showed significantly greater motility toward tumor cell lysates or spheroids than toward normal cell lysates (Fig. 3a) or spheroids (Fig. 3b). In addition, bacteriobots showed higher average velocity when migrating toward tumor cell lysates or spheroids than when migrating toward normal cell lysates or spheroids. Cy5.5-containing PS microbeads without bacterial attachment (microbeads only) showed no accumulation in tumor lysates or spheroids (Fig. 3c, d and Supplementary Movie 2). Next, to validate the tumor targeting and localization of the bacteriobots in tumor-bearing mice, bacteriobots (bacteria:microbead ratio = 3:1) were injected systemically into CT-26 tumor-bearing mice via tail veins. As controls, S. typhimurium (3 × 107 colony-forming units (CFU)) or Cy5.5 fluorescence-coating PS microbeads alone (1 × 107) were injected into CT-26 tumor-bearing mice in a similar fashion. Bacterial bioluminescence and near-infrared fluorescence (Cy5.5) were measured from the extracted tumors using a cooled charge-coupled device (CCD) camera. Bacterial bioluminescent signal was detected in the tumors from the bacteria- and bacteriobot-injected animal groups in both tumor models, but not in the tumors from the microbead-injected control animals (Fig. 4a, b and c, left figures). Subsequently, Cy5.5 fluorescence was observed in the tumors from the bacteriobot-injected animals, but not in the tumors from the bacteria- or microbead-injected control animals (Fig. 4a, b and c, right figures), indicating successful tumor targeting by the bacteriobots. For a more in-depth investigation of the targeting and localization of the bacteriobots in tumors, a histological immunofluorescence study was performed after intravenous injection of S. typhimurium, Cy5.5 fluorescence-coating PS microbeads, or bacteriobots in CT-26 tumor-bearing mice; tumors were collected 3 days post-inoculation (dpi). Results showed that gfp-expressing S. typhimurium were observed in the tumors from the bacteriobot-injected and bacteria-injected animals (Fig. 5a and c). The near-infrared fluorescence emitted by Cy5.5 was measured only in the tumors from the bacteriobot-injected animals (Fig. 5c) and not in the tumors from control animals injected with bacteria or microbeads (Fig. 5a and b). These results show that the S. typhimurium bacteriobots were successfully delivered to the tumor region in the CT-26 tumor mouse model.
Evaluation of the tumor-targeting properties of bacteria and/or bacteriobots using cell lysates and 3D cell spheroids in a gradient-generating microfluidic device.
(a–d) Evaluation of the tumor-targeting properties of bacteria (a, b) and bacteriobots (c, d) using cell lysates (a, c) and 3D cell spheroids (b, d) in a gradient-generating microfluidic device. Cell lysates (1 × 107 cells/mL) were loaded in the left or right chamber of the microfluidic device for 30 min after bacteria or bacteriobots were loaded in the central chamber; the density of the bacteria (a) and the velocities of the bacteriobots (b) were recorded. *, P < 0.001 compared to NIH/3T3 cell lysate. Values are expressed as means ± S.D. Cell spheroids were loaded in the left or right chamber of the microfluidic device for 30 min after the bacteria or bacteriobots were loaded in the central chamber; density of the bacteria (b) and the velocities of the bacteriobots (d) were recorded. *, P < 0.001 was compared to spheroids prepared from normal NIH/3T3 fibroblasts. Values are expressed as means ± S.D.
Tumor targeting and localization of bacteriobots in a syngeneic mouse tumor model.
Mice (n = 6) were injected subcutaneously with CT-26 cells (1 × 106). When the tumors reached a volume of approximately 130 mm3, the tumor-bearing mice were injected with bacteria (3 × 107 CFU/100 μL), microbeads (1 × 107/100 μL) or bacteriobots (bacteria:microbeads ratio = 3:1 per 100 μL). Representative in-vivo and ex-vivo bioluminescence and NIR images (Cy5.5 image) were captured 3 days post-injection. (a) In-vivo bioluminescence and NIR imaging of mouse tumor models. (b) Ex-vivo bioluminescence and NIR imaging of the dissected tumors. (c) Ex-vivo bioluminescence and NIR imaging: quantification of photons emitted. The data represent the mean ± S.D. (n = 6) (*, P < 0.001).
Localization of bacteriobots in the dissected tumor masses at 3 days post infection.
Tumor-bearing mice were injected with (a) bacteria, (b) microbeads and (c) bacteriobots, the tumor masses were fixed and investigated histologically and bacteria, microbeads and bacteriobots were localized by indirect fluorescence. (a–c) DAPI staining of the same tissue sections and mergence of the DAPI-stained slides (blue); (a and c) bacteria were detected by indirect immunofluorescence (green); (b and c) microbeads and bacteriobot were detected by indirect fluorescence (red) in the dissected tumor masses. Scale bars, 10 μm.
Discussion
The fabrication and integration of microactuators and microsensors with high stability and efficiency are at the forefront of microrobot developlemt13,14. Among the various microorganisms that could be used for the microactuator and microsensor, bacteria with high motility have been considered as promising candidates. Magnetotactic bacteria (MTB) strains, such as Magnetococcus strain MC-1 and watermelon-shaped magnetotactic bacterium (MWB-1), have been used by some researchers because of their high motility of about 200–300 μm/sec in a magnetic field21,32. In addition, MTB strains can be easily controlled by a combination of magnetotaxis and other taxis methods, such as chemotaxis and aerotaxis. However, MTB strains have several disadvantages in their application to biomedical microrobots because of the difficult incubation procedure and complex electromagnetic coil systems that are required23. As an alternative, we focused on high motility flagellar bacteria (S. marcescens and S. typhimurium) for use as microactuators22,31. These bacterial strains can be controlled by various external chemical stimuli in specific environmental conditions because of their sensitive receptors and therefore can be used as microsensors21,33. However, because S. marcescens has acute pathogenicity36, it cannot be applied to a biomedical microrobot. In this study, we used the attenuated S. typhimurium strain defective in the synthesis of guanosine 5′-diphosphate-3′-diphosphate (ppGpp) (ΔppGpp strain)17,24,32 because it has shown unique diagnostic and therapeutic characteristics against various solid tumors, such as those of colorectal and breast cancers24,25,26,27,34,35.
Attenuated strains of S. typhimurium have many unique features in cancer therapy compared to other conventional methods. They have been shown to specifically target and proliferate in tumor24,25,27,28,29,35. Because of chemotactic receptors and flagella, S. typhimurium can direct chemotaxis toward molecular signals in the tumor microenvironment and enable penetration of cancer tissue25. With self-propulsion, they can actively swim away from the vasculature and penetrate deep into tumor tissue, which can have a higher density far from the vascular source. Most importantly, genetic manipulation of bacteria is easy, particularly for the S. typhimurium, which will have the greatest effect on a therapeutic strategy because they enable precise tuning of drug production in deep tumor tissue that is resistant to other therapeutic modalities, such as chemotherapy and radiotherapy29,34. However, limited drug production often remains a challenge when bacteria fail to produce a sufficient concentration of drugs to induce a therapeutic effect25. Because of these characteristics of S. typhimurium, we used this strain as the microactuator and microsensor and partly as the therapeutic agent to propel microrobot structures toward tumor tissue.
In the development of a bacteriobot, the strong attachment of the bacteria to the microstructure is very important for its motility and stability in living animals. Researchers have recently adopted a bacterial patterning technique using RIE O2 plasma19. However, this technique can only be applied to microstructures with a hydrophobic surface and cannot actively regulate the bacterial attachments on the microstructures because of the unidirectional stream of RIE plasma. We also proposed applying a bacterial adhesion to the surface of the SU-8 microstructure by using a selective coating with BSA, a non-fouling protein22. In addition, we proposed a new bacterial patterning methodology on a selective region of microbeads using the submerged properties of microbeads on agarose gel31. Consequently, we can actively regulate bacterial attachment to microstructures using various patterning methods. Despite being able to regulate bacterial adhesion to microstructures, however, non-specific binding of bacteria should be considered in these methods because all bacteria can easily attach to the coated surface of a microstructure. In this study, we proposed a new attachment methodology based on the high-affinity interaction between biotin and streptavidin. For the complete attachment of bacteria and microstructure, biotin molecules were attached to the outer membrane proteins of S. typhimurium and streptavidin protein was coated on the surface of the microstructure. As a result, a bacteriobot was fabricated by the powerful conjugation of streptavidin and biotin (Fig. 1 and Supplementary Movie 1).
For evaluating the motility of bacteria and/or a bacteriobot, a stable concentration gradient of chemo-effectors is a key condition in the development of chemotactic microfluidic. However, it is difficult to maintain a stable and uniform gradient of chemo-effectors without flow in a microfluidic chamber. We proposed the new flow-free chemotactic microfluidic chamber and carried out chemotaxis motility tests using cell lysates or spheroids (Figs. 2, 3 and Supplementary Movie 2). Compared with conventional methods, such as an agarose-in-plug method, which is a capillary assay, the proposed flow-free chemotactic microfluidic chamber can quickly measure bacterial motility in the liquid medium, characterize the chemotaxis phenomena of the low-motility bacteria and quantify the evaluation of bacteria and/or bacteriobot chemotaxis37,38,39,40,41. Evaluating bacteria chemotaxis using flow-based microfluidic channels has been proposed previously38; however, it is difficult to generate a stable concentration gradient because of the flow in the microfluidic channels.
The S. typhimurium strain has intrinsic advantages: it can sense, move, accumulate and replicate in solid tumors, such as colorectal and breast cancers24,25,26,27. Because of the remarkable abilities of S. typhimurium against solid tumors, this strain has been previously used for tumor targeting and therapy24. We fabricated new types of bacteriobot using the targeting and motile specificities of S. typhimurium and evaluated the targeting and localization properties of these bacteriobots against solid tumors by using the proposed flow-free microfluidic channels (Fig. 3) and tumor-bearing mice (Figs. 4, 5).
Finally, in this study we proposed a new microrobot paradigm to target and treat solid tumors in which attenuated S. typhimurium acts as a microactuator and microsensor to deliver microstructures to solid tumors. A strong streptavidin/biotin-based conjugation method between bacteria and microstructures was proposed and used for the fabrication of the bacteriobot. In addition, the bacteriobot's motility and targeting were assessed in vitro using novel chemotactic microfluidic channels and in vivo using syngeneic mouse tumor models. The feasibility of bacteriobot technology as a drug delivery agent for tumor therapy needs to be explored further. The findings of our study suggest that the bacteriobot concept will be of great influence in the development of biomedical theranostic microrobots that can carry out versatile functions, such as the detection and eradication of incurable malignancies.
Methods
Bacterial strains
The ΔppGpp S. typhimurium strain SHJ2037 (relA::cat, spoT::kan) was described previously17. Salmonellae were grown in Luria-Bertani broth medium (Becton Dickinson, Franklin Lakes, NJ) with vigorous aeration at 37°C. For the imaging of bioluminescence, the bacterial luciferase gene (lux) from S. typhimurium-Xen26 (Caliper Life Sciences, Hopkinson, MA) was transduced into strain SHJ2037 by P22HT into transduction. For fluorescence microscopic imaging, pEGFP (Clontech, Palo Alto, CA) was transformed into strain SHJ203742.
Cell culture
The NIH/3T3 mouse embryo fibroblast cell line and the murine CT-26 colorectal and 4T1 mammary carcinoma cell lines were obtained from the American Type Culture Collection (CRL-1658, 2638 and 2539, respectively, Manassas, VA) and cultured in DMEM (Gibco-BRL/Invitrogen, Carlsbad, CA) supplemented with 10% v/v FBS and 1% v/v antibiotic-antimicotic solution (Gibco-BRL/Invitrogen). Cells were cultured in a humidified CO2 incubator (37°C in atmosphere of 5% CO2). Cell suspensions were prepared by dissociation in 0.25% trypsin-EDTA (Gibco-BRL/Invitrogen), centrifugation at 1000 rpm for 3 min at room temperature and re-suspension in growth medium.
Preparation of cell lysates
NIH/3T3, CT-26 and 4T1 were suspended in PBS. The cells suspension was frozen in liquid nitrogen for 1 min and then thawed in a 37°C water bath for 4 min. The freeze-thaw cycle was repeated five times in rapid succession. Protein concentration was determined using the Bio-Rad Protein Assay (Bio-Rad Laboratories, München, Germany).
3D cell spheroid generation
Confluent cultures of NIH/3T3, CT-26 and 4T1 cells were trypsinized, washed in PBS and resuspended in DMEM. Drops of each cell suspension (between 5,000 and 10,000 cells/20 μL) were placed onto the inside surface of lids of 10 cm cell culture dishes and the lids were then inverted over dishes containing 10 mL DMEM. The hanging drop cultures were incubated and after a sufficient sedimentation time, the resulting cellular aggregates were harvested using a Pasteur pipette under a dissecting microscope.
Fabrication of gradient-generating microfluidic device
The gradient-generating microfluidic device was designed to generate a continuous concentration gradient using simple diffusion phenomena without flow; the device consists of two chambers (left and right) for loading cell lysates or spheroids and a central chamber for loading bacteria and/or bacteriobots. The microfluidic device was fabricated by conventional photo- and soft-lithography procedures43. In photolithography, a photo-resistor (SU-8 2050, MicroChem Corp., Microchem, Newton, MA) was spin-coated onto the wafer. The thickness of the photoresistor was approximately 100 μm and the wafer was soft-baked on a hotplate at 110°C for 25 min. Next, the designed pattern was transferred by photo mask and ultraviolet exposure. After hard baking and a developing step, the embossed pattern SU-8 mold was fabricated. In soft-lithography, PDMS (polydimethylsiloxane) solution (Sylgard 184 Silicone Elastomer Kit, Dow Corning, Midland, MI) was slowly poured on the top of the SU-8 mold; the PDMS solution is a mixture of PDMS prepolymer (Sylgard 184 A) and a curing agent (Sylgard 184 B) in a 10:1 volume ratio. After curing in a dry oven, the PDMS was detached from the SU-8 mold, revealing the intaglio pattern PDMS. After punching the inlet and outlet, the PDMS was bonded with glass using an O2 plasma asher and a dry oven.
Computational modeling of dye concentration gradients in chemotactic microfluidic chamber
The chamber model in 2D was imported into COMSOL (COMSOL Inc., Burlington, MA) and extruded to obtain a full 3D model of the chamber and the gates. The 3D model was meshed to obtain 119399 elements. Molecular transport calculations involved the species conservation law and Fick's diffusional transport with a diffusion coefficient of 1 × 10−9 m2 sec−1. In addition, the convection of the species was ignored.
Fabrication of bacteriobots
For the surface modification of S. typhimurium, we used EZ-Link NHS-LC-Biotin (Thermo Scientific, Rockford, IL). The outer membrane protein (omp) of S. typhimurium (3 × 108 CFU/mL) was exposed to 500 μg biotin for 1 hr. For the surface modification of polystyrene (PS) microbeads, we used rhodamine-containing fluorescent PS microbeads of 3 μm diameter (Polyscience, Warrington, PA). The PS microbeads (1 × 108/mL) were covalently coupled to 500 μg streptavidin-conjugated tandem fluorochrome composed of peridinin chlorophyll protein (PerCP), which was further labeled by Cy5.5 (PerCP-Cy5.5) (BD Biosciences, San Diego, CA) to allow near-infrared (NIR) imaging of the bacteriobots in a syngeneic mouse tumor model. Bacteriobots were then fabricated by co-incubation of 500 μg biotin-labeled S. typhimurium and 500 μg streptavidin-PerCP-Cy5.5-coated PS microbeads for 30 min at 37°C. The bacteriobots were observed using a laser confocal scanning microscope (TCS SP5/AOBS/Tandem, Leica, Germany).
3D reconstruction and animation of bacteriobot
For cross-section and whole-construct imaging of the 3D bacteriobot, the fabricated bacteriobots were scanned using a laser confocal scanning microscope (TCS SP5/AOBS/Tandem, Leica, Germany) with a 40× oil immersion lens (NA1.25). After confocal stack acquisition, Imaris (3D Imaging Analysis software, Bitplane) was available for 3D reconstruction and animation.
Motility of bacteriobots
Motility of bacteriobots was analyzed using a Nikon Ti-U microscope (Nikon USA, Melville, NY). A tracking algorithm using MATLAB evaluated Bacteriobot velocity. The algorithm recognizes a PS microbead from a movie frame and calculates its speed and velocity. First, the algorithm searches for the edges of the object to assess whether it is a PS microbead. It changes the original images to binary images and filters out noise objects using a threshold size. As the centroid coordinates of a PS microbead can be recorded in pixel values and the real length per pixel can be calculated, the difference in coordinates between frames was used to calculate the total distance and speed of a bacteriobot. In addition, the final difference in the coordinates between the first frame and the last frame was used to calculate the total displacement and final velocity of the bacteriobot.
Animal models
Five- to six-week-old male BALB/c mice (20–30 g body weight) were purchased from the Orient Company (Seongnam, Korea). All animal care, experiments and euthanasia were performed in accordance with protocols approved by the Chonnam National University Animal Research Committee (Gwangju, Korea). Anesthesia was performed using isoflurane (2%) for in vivo imaging. CT-26 cells (1 × 106 cells) were harvested, suspended in 100 μL PBS and injected subcutaneously into the right thigh of each mouse. Tumor volume (mm3) was calculated using the following formula: (L × H × W)/2, where L is the length, W is the width and H is the height of the tumor in millimeters. When the tumors reached a volume of approximately 130 mm3, mice were injected intravenously with luciferase-emitting bacteria (3 × 107 CFU/100 μL), microbeads (1 × 107/100 μL) or bacteriobots (bacteria:microbeads ratio = 3:1 in 100 μL) through the tail vein.
Immunofluorescence staining
The tumor was extracted, fixed in 4% paraformaldehyde in PBS for 2 hours at room temperature and cryopreserved using OCT compound (Sakura Finetek USA Inc., Torrance, California, USA) at −80°C. Sections of the tissues (5 μm thick) were mounted on glass slides, fixed and stained as described as follows. For immunofluorescence staining, the slides were incubated with primary anti-salmonella antibody overnight at 4°C (1:100, Abcam, Cambridge, UK). After washing three times with PBST (with 0.01% Tween-20), goat anti-rabbit 488 secondary antibody was used (1:100, Invitrogen, Eugene, OR) for 2 hours at room temperature. The sample was stained with DAPI/Antifade (1:10000, Invitrogen). The stained tissue was subsequently washed with PBS and mounted on glass slides for analysis using a FV1000D laser confocal scanning microscope (Olympus, Tokyo, Japan).
Optical bioluminescence imaging
To obtain images of bacterial bioluminescence, animals were anesthetized and placed in a light-tight chamber in the in-vivo imaging system (IVIS, Caliper Life Sciences, Hopkinton, MA) equipped with a cooled CCD camera. Photons emitted from luciferase-expressing bacteria were collected and integrated over 1 min periods. Pseudocolor images indicating photon counts were overlaid on photographs of the mice using the Living Images software v. 2.25 (Caliper Life Sciences). A region of interest was selected manually based on signal intensity. The area of the region of interest was kept constant and the signal intensity was recorded as maximum radiance within each region of interest.
NIR imaging
In-vivo NIR imaging was performed using IVIS. After anesthetizing each mouse with 1.5–2% isoflurane, each experimental sample (bacteria, microbead, or bacteriobot) was injected into the mouse through the tail vein. NIR fluorescence imaging was performed 3 days post-injection using Cy5.5 filters with the following settings: exposure time 1 sec., f/stop 8, binning 8 and field of view 12.8. After in-vivo imaging, the mice were sacrificed and tumors were imaged immediately upon dissection.
Statistics
The data are presented as means with standard deviations and statistical significance was assessed using ANOVA (StatView; Abacus Concepts Inc., Berkeley, CA). All experiments were performed at least three times on separate days and the data presented herein are representative of all the repetitions. P < 0.001 was considered significant.
References
Liu, S. C., Minton, N. P., Giaccia, A. J. & Brown, J. M. Anticancer efficacy of systemically delivered anaerobic bacteria as gene therapy vectors targeting tumor hypoxia/necrosis. Gene Ther. 9, 291–296 (2002).
Jain, R. K. & Stylianopoulos, T. Delivering nanomedicine to solid tumors. Nat. Rev. Clin. Oncol. 7, 653–664 (2010).
Trédan, O., Galmarini, C. M., Patel, K. & Tannock, I. F. Drug resistance and the solid tumor microenvironment. J. Natl. Cancer Inst. 99, 1441–1454 (2007).
Minko, T. Drug targeting to the colon with lectins and neoglycoconjugates. Adv. Drug Del. Rev. 56, 491–509 (2004).
Gong, J. et al. Microparticles and their emerging role in cancer multidrug resistance. Cancer Treat. Rev. 38, 226–234 (2012).
Panyam, J. & Labhasetwar, V. Biodegradable nanoparticles for drug and gene delivery to cells and tissue. Adv. Drug Deliv. Rev. 55, 329–347 (2003).
Brannon-Peppas, L. & Blanchette, J. O. Nanoparticle and targeted systems for cancer therapy. Adv. Drug Deliv. Rev. 56, 1649–1659 (2004).
Akin, D. et al. Bacteria-mediated delivery of nanoparticles and cargo into cells. Nat. Nanotechnol. 2, 441–449 (2007).
Von Maltzahn, W. W. (ed.) Medical Instruments and Devices, The Biomedical Engineering Handbook, 3rd Edition., 52–73, (CRC Taylor & Francis Group, USA, 2006).
Requicha, A. A. Nanorobots, NEMS and nanoassembly. Proc. IEEE. 91, 1922–1933 (2003).
Abbott, J. J., Nagy, Z., Beyeler, F. & Nelson, B. J. Robotics in the Small, Part I: Microbotics. Robotics & Automation Magazine, IEEE. 14, 92–103 (2007).
Sharma, N. & Mittal, R. Nanorobot movement: Challenges and biologically inspired solutions. Int. J. Smart sensor Intel. Systems 1, 87–109 (2008).
Nelson, B. J., Kaliakatsos, I. K. & Abbott, J. J. Microrobots for minimally invasive medicine. Annu. Rev. Biomed. Eng. 12, 55–85 (2010).
Sitti, M. Miniature devices: Voyage of the microrobots. Nature 458, 1121–1122 (2009).
Pawashe, C., Floyd, S. & Sitti, M. Modeling and experimental characterization of an untethered magnetic micro-robot. Int. J. Robotics Res. 28, 1077–1094 (2009).
Zhang, L. et al. Artificial bacterial flagella: Fabrication and magnetic control. Appl. Phys. Lett. 94, 064107 (2009).
Choi, H. et al. Two-dimensional locomotion of a microrobot with a novel stationary electromagnetic actuation system. Smart Mater. Struct. 18, 115017 (2009).
Berg, H. C. The rotary motor of bacterial flagella. Biochemistry 72, 19–54 (2003).
Behkam, B. & Sitti, M. Effect of quantity and configuration of attached bacteria on bacterial propulsion of microbeads. Appl. Phys. Lett. 93, 223901 (2008).
Kim, D., Liu, A., Diller, E. & Sitti, M. Chemotactic steering of bacteria propelled microbeads. Biomed. Microdevices 14, 1009–1017 (2012).
Martel, S., Mohammadi, M., Felfoul, O., Lu, Z. & Pouponneau, P. Flagellated Magnetotactic Bacteria as Controlled MRI-trackable Propulsion and Steering Systems for Medical Nanorobots Operating in the Human Microvasculature. Int. J. Robot Res. 28, 571–582 (2009).
Park, S. J. et al. Motility enhancement of bacteria actuated microstructures using selective bacteria adhesion. Lab Chip 10, 1706–1711 (2010).
Lefevre, C. T., Bernadac, A., Yu-Zhang, K., Pradel, N. & Wu, L. F. Isolation and characterization of a magnetotactic bacterial culture from the Mediterranean Sea. Environ. Microbiol. 11, 1646–1657 (2009).
Min, J. J., Nguyen, V. H., Kim, H. J., Hong, Y. J. & Choy, H. E. Quantitative bioluminescence imaging of tumor-targeting bacteria in living animals. Nat. Protoc. 3, 629–636 (2008).
Forbes, N. S. Engineering the perfect (bacterial) cancer therapy. Nat. Rev. Cancer 10, 784–793 (2010).
Yoo, J.-W., Irvine, D. J., Discher, D. E. & Mitragotri, S. Bio-inspired, bioengineered and biomimetic drug delivery carriers. Nat. Rev. Drug Discov. 10, 521–535 (2011).
Le, U. N. et al. Engineering and visualization of bacteria for targeting infarcted myocardium. Mol. Ther. 19, 951–959 (2011).
Nguyen, V. H. et al. Genetically engineered Salmonella typhimurium as an imageable therapeutic probe for cancer. Cancer Res 70, 18–23 (2010).
Jiang, S.-N. et al. Inhibition of Tumor Growth and Metastasis by a Combination of Escherichia coli–mediated Cytolytic Therapy and Radiotherapy. Mol. Ther. 18, 635–642 (2010).
McEldowney, S. & Fletcher, M. Variability of the influence of physicochemical factors affecting bacterial adhesion to polystyrene substrata. Appl. Environ. Microbiol. 52, 460–465 (1986).
Cho, S., Park, S. J., Ko, S. Y., Park, J. O. & Park, S. Development of bacteria-based microrobot using biocompatible poly(ethylene glycol). Biomed. Microdevices 14, 1019–1025 (2012).
Lin, W., Li, J. & Pan, Y. Newly isolated but uncultivated magnetotactic bacterium of the phylum Nitrospirae from Beijing, China. Appl. Environ. Microbiol. 78, 668–675 (2012).
Bren, A. & Eisenbach, M. How signals are heard during bacterial chemotaxis: protein-protein interactions in sensory signal propagation. J. Bacteriol. 182, 6865–6873 (2000).
Jiang, S.-N. et al. Engineering of bacteria for the visualization of targeted delivery of a cytolytic anticancer agent. Mol. Ther. 10.1038/mt.2013.183 (2013).
Min, J.-J. et al. Noninvasive real-time imaging of tumors and metastases using tumor-targeting light-emitting Escherichia coli. Mol. Imaging Biol. 10, 54–61 (2008).
Hejazi, A. & Falkiner, F. Serratia marcescens. J. Med. Microbiol. 46, 903–912 (1997).
Wolfe, A. J. & Berg, H. C. Migration of bacteria in semisolid agar. Proc. Natl. Acad. Sci. USA. 86, 6973–6977 (1989).
Ahmed, T., Shimizu, T. S. & Stocker, R. Bacterial chemotaxis in linear and nonlinear steady microfluidic gradients. Nano Lett. 10, 3379–3385 (2010).
Ahmed, T. & Stocker, R. Experimental verification of the behavioral foundation of bacterial transport parameters using microfluidics. Biophys. J. 95, 4481–4493 (2008).
Bainer, R., Park, H. & Cluzel, P. A high-throughput capillary assay for bacterial chemotaxis. J. Microbiol. Methods. 55, 315–319 (2003).
Yu, H. S. & Alam, M. An agarose-in-plug bridge method to study chemotaxis in the Archaeon Halobacterium salinarum. FEMS Microbiol. Lett. 156, 265–269 (1997).
Bradburne, J., Godfrey, P., Choi, J. H. & Mathis, J. In vivo labeling of Escherichia coli cell envelope proteins with N-hydroxysuccinimide esters of biotin. Appl. Environ. Microbiol. 59, 663–668 (1993).
Xia, Y. & Whitesides, G. M. Soft lithography. Annu. Rev. Mater. Sci. 28, 153–184 (1998).
Acknowledgements
This research was supported by Pioneer Research Center Program through the National Research Foundation of Korea, funded by the Ministry of Science, ICT & Future Planning (2009-0082954 and 2009-0082935). J.-J.M. was supported by the National Research Foundation of Korea (NRF) (No. 2012-0006072). H.E.C. was supported by the Intelligent Synthetic Biology Center of Global Frontier Project funded by MEST (2011-0031958).
Author information
Authors and Affiliations
Contributions
S.J.P. and S.-H.P. are co-first authors; J.-J.M., S.P. and J.-O.P. are co-last authors. J.-J.M., S.P. designed all experiments. J.-J.M., H.E.C., Y.H., S.P., J.-O.P. and S.Y.K. supervised experiments and interpreted results. S.J.P. and S.-H.P. executed most of experiments and analyzed data. S.C., D.-M.K. and Y.L. assisted with mouse experiments. J.-J.M., S.P., S.J.P. and S.-H.P. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Supplementary Movie 1. Bacteriobot
Supplementary Information
Supplementary Movie 2. PS microbead
Supplementary Information
Supplementary Movie 2. NIH lysate
Supplementary Information
Supplementary Movie 2. NIH 3D
Supplementary Information
Supplementary Movie 2. CT-26 lysate
Supplementary Information
Supplementary Movie 2. CT-26 3D
Supplementary Information
Supplementary Movie 2. 4T1 lysate
Supplementary Information
Supplementary Movie 2. 4T1 3D
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Park, S., Park, SH., Cho, S. et al. New paradigm for tumor theranostic methodology using bacteria-based microrobot. Sci Rep 3, 3394 (2013). https://doi.org/10.1038/srep03394
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03394
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.