Abstract
Crystallization is an essential process for understanding a molecule's aggregation behavior. It provides basic information on crystals, including their nucleation and growth processes. Deoxyribonucleic acid (DNA) has become an interesting building material because of its remarkable properties for constructing various shapes of submicron-scale DNA crystals by self-assembly. The recently developed substrate-assisted growth (SAG) method produces fully covered DNA crystals on various substrates using electrostatic interactions and provides an opportunity to observe the overall crystallization process. In this study, we investigated quantitative analysis of molecular-level DNA crystallization using the SAG method. Coverage and crystal size distribution were studied by controlling the external parameters such as monomer concentration, annealing temperature and annealing time. Rearrangement during crystallization was also discussed. We expect that our study will provide overall picture of the fabrication process of DNA crystals on the charged substrate and promote practical applications of DNA crystals in science and technology.
Similar content being viewed by others
Introduction
Molecular crystallization is a supramolecular process by which an ensemble of randomly organized molecules in the fluid state come together to form a well-ordered molecular arrangement called a crystal1,2,3,4,5. This process allows molecular-scale growth by crystals attaching themselves to nucleating structures through a specific interaction that arises from the nature of the specific constituent molecules. Thus, each solid-state crystal has distinct growth properties following geometrical and chemical characteristics of their molecules6,7,8,9. During the past few decades, DNA has stood out as a potential material for crystals because of its remarkable ability to construct complex architecture. Due to selective binding interactions following Watson-Crick complementary rules, all dimensional shapes of DNA crystal have been created by DNA molecules10,11,12,13,14,15. Additionally, simply modified DNA molecules enable DNA crystals to serve as a template for various heteromaterials to arrange within several nanometre-scale resolutions16,17,18,19,20. DNA crystals are generally fabricated by annealing into a physiological buffer solution containing divalent magnesium ions. During this process, each DNA strand forms the predesigned shape of a DNA crystal following its individual growth mechnism21,22,23. Since this information aids in understanding fundamental behaviors during the fabrication process, study on growth mechanisms are commonly accompanied by the design of a new DNA crystal.
The recently developed annealing method called SAG24,25,26 uses a different fabrication method than the conventional free solution annealing method. For SAG processes, a negatively charged substrate is immersed into physiological DNA buffer solution. Through electrostatic interaction between negatively charged DNA molecules and the substrate mediated by divalent magnesium ions, DNA crystals are formed on the substrate during the annealing process. DNA Crystals grown by SAG methods show somewhat different topologies, including the aspect ratio and dimensionality24,25,26. In addition, accurate coverage control of DNA crystals onto substrate can be achieved from 0 to 100% at relatively low DNA concentrations26. We develop a theoretical model and carry out some simulations in limited cases27, but still we are unclear that overall picture in the model is sufficient. So we have to investigate the phenomenon by detailed experiments using some important physical parameters. This article aims to provide systematic and quantitative study of the nucleation and growth mechanism of DNA crystals by SAG. During SAG, DNA molecules are generally affected by three external parameters, i.e., monomer concentration (Cm), initial annealing temperature (T) and total annealing time (t). Once these growth behaviors are well established, they will serve as an important guide in the fabrication of coverage controllable DNA crystals. To this end, we report a molecular-level DNA crystallization process with various physical conditions.
Results
Analysis of coverage for molecular population
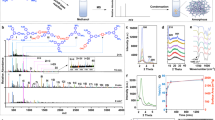
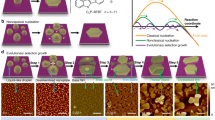
Cm is directly related to the coverage of DNA crystals on a substrate, usually mica in DNA research. To demonstrate the coverage dependence of Cm, four different types of DNA crystals were annealed at different DNA Cm values (supplementary information Figs. S1, S2 and Tables S1, S2 for details). Double-crossover (DX) crystals10 were fabricated from DX tiles consisting of two adjacent duplex DNAs connected by two crossover junctions. Eight-helix lattices (8 HL), 5-helix lattices (5 HL) and 5-helix ribbons (5 HR) were constructed from single-stranded tiles (SST)11. Schematic diagrams of the various DNA crystals are illustrated in Figures 1a–1d and corresponding atomic force microscopy (AFM) images are shown in the same column. To achieve consistency and reliability, four reference conditions were fixed during the process: substrate size (5 mm × 5 mm), total DNA sample volume (200 uL), initial annealing temperature (80°C) and total annealing time (6 hours in heat block, see DNA crystal fabrication in methods), while different Cm values were used as a control parameter. The assembly of DNA crystals began to appear at monomer threshold concentrations (Cm-th) of ~5 nM for DX, 8 HL and ~10 nM for 5 HL, 5 HR. Consequently, full coverage was reached at monomer saturation concentrations (Cm-s) of ~20 nM for DX, 8 HL and ~25 nM for 5 HL and ~40 nM for 5 HR, respectively (Fig. 1e and supplementary information Fig. S3). In the SAG system, nucleation of DNA molecules on the substrate occurred slightly below 10 nM rather than at the conventional free solution concentration of ~50 nM. This was due to the catalytic behavior of the substrate, by which the Coulomb force between the substrate and DNA molecules pays a partial entropic cost for DNA crystallization26. This force creates higher DNA molecular density close to the substrate compared to the rest of the solution, providing appropriate conditions for DNA crystallization.
Coverage dependence on monomer concentration (Cm) for each DNA crystal.
(a–d) Schematic diagrams of various DNA crystals grown by substrate-assisted growth (SAG); (a) double-crossover, (b) 8 helix lattice, (c) 5 helix lattice and (d) 5 helix ribbon crystals. AFM images of coverage at different Cm in the same column as the schematic diagrams of DNA crystals (a–d). (e) Plots of coverage controlled by Cm and CMol (inset). The yellow region encloses the coverage deviation of 2D 5 helix lattice and 1D 5 helix ribbon crystals. All scale bars are 500 nm.
For consistent comparison, the Cm was substituted with molecular concentration (CMol) to avoid confusion regarding the total molecular amount in a DNA solution (supplementary information Table S3 for details). Since each DNA crystal has different size motifs composed of different numbers of base pairs, the Cm of each DNA sample solution does not equal the molecular weight for each DNA motif. An inset plot exhibited in Fig. 1e represents the modified coverage dependence of DNA crystals. All DNA crystals tested in this research initiated crystal assembly at (5.8 ± 1) × 10−7 g/cm3 of molecular threshold concentration (CMol-th). Coverage increased until CMol reached molecular saturation concentration (CMol-s) for full coverage of DNA crystals on the substrate. CMol-s was obtained at (19 ± 2) × 10−7 g/cm3 for DX, 8 HL and 5 HL, although they have different geometries and hybridizing pathways. However, the CMol-s for 5 HR (~27 × 10−7 g/cm3) was relatively higher than those of other DNA crystals, even though it has the same structure as 5 HL. Comparative differences in CMol-s between DNA crystals may be due to different degrees of freedom for crystal growth directions. Contrary to a 1D 5 HR crystal, the DX, 8 HL and 5 HL crystals were constructed by bidirectional crystal growth. Kinetic interaction between excess molecules and DNA crystals with bilateral crystal growth directions was more likely once the crystals nucleated on the substrate. In contrast, the unidirectional crystallization of 5 HR was suppressed by crystals that had already formed on the substrate. This created a need to supersaturate excess CMol in the test tube26. Another interesting effect of CMol on 5 HR was observed when CMol passed a certain critical level of ~24 × 10−4 g/cm3, at which the randomly arranged 5 HR topology suddenly transformed to a well-oriented arrangement (supplementary information Fig. S3). This highly packed system is an effective way to minimize surface free energy while providing full coverage of 1D DNA crystals on the substrate28.
Analysis of crystal size distribution by three external parameters
DNA crystallization has a significant and complicated energy correlation with several external parameters, including Cm, T, t for molecular population, thermodynamic energy and reaction time, respectively. The crystal size distribution was analyzed to demonstrate the effects of the aforementioned parameters on DNA crystallization. For Cm, different concentrations of DX solution with a mica substrate were annealed by cooling in 5°C decrements for 30 min. from 80°C to room temperature (25°C) (T = 80°C, t = 6 h) using a heat block (for details, see DNA crystal fabrication in methods). Fig. 2a displays AFM images corresponding to the DX crystal fabricated at each crystallization condition. Average crystal size (Savg) analyzed by Cm showed that, with conditions of T = 80°C and t = 6 h, crystal size increased until near the end of the process at 20 nM. As mentioned above, the assembly of DX crystals began at the Cm-th of 5 nM with Savg of ~3.8 × 105 nm2 (each single crystal domain is enclosed by yellow dotted lines in Fig. 2a). Savg then proportionally increased to ~12.6 × 105 nm2 until Cm-s of 20 nM, at which full coverage was achieved (Fig. 2b, supplementary information Fig. S4 and Table S4). No distinct crystal size increases were observed for Cm larger than 20 nM.
Analysis of crystal size distribution depending on monomer concentration (Cm), initial annealing temperature (T) and total annealing time (t).
(a) AFM images of double-crossover (DX) crystals grown by substrate-assisted growth (SAG) at each crystallization condition. As control parameters increased under specific crystallization conditions (black arrows), average crystal size increased gradually. Single DNA crystal domains are enclosed in yellow dotted lines. (b–d) Plots of average crystal size distribution versus (b) Cm with fixed T = 80°C and t = 6 h, (c) T with fixed Cm = 50 nM and t = 1 h, (d) t with fixed Cm = 50 nM and T = 80°C. Red dotted lines show quadratic exponential fitted plots for growth rate and coefficient values are exhibited in (e). All scale bars in AFM images are 500 nm.
A similar trend was also observed when T and t were used as control parameters under conditions of Cm = 50 nM, t = 1 h and Cm = 50 nM, T = 80°C, respectively. To ensure 100% coverage of DX crystals on a given substrate, the sample was prepared with a DX concentration of 50 nM, about 2 times larger than the Cm-s. For T = 40°C (5°C/20 min.), a series of AFM images (Fig. 2a) shows that the substrate was fully covered by small DX crystals with Savg of ~0.2 × 105 nm2, even though the temperature was slightly higher than crystal annealing temperature of ~37°C for DX crystal formation10,22. Additionally, DNA crystals were fabricated in a relatively short period of about 1 hour rather than the normal annealing time of over 6 hours. As T increased to 60°C (5°C/~9 min.) and 80°C (5°C/~6 min.), the Savg observed by AFM showed proportional increases in crystal size of ~1.4 × 105 nm2 and ~3.5 × 105 nm2, respectively. However, the Savg did not reach the Savg (~12.6 × 105 nm2) obtained under Cm = 20 nM, T = 80°C and t = 6 h. This might be due to shorter reaction time for hybridization (Figs. 2a, 2c and supplementary information Fig. S4, Table S5). The effect of t was also investigated by annealing DNA solution under conditions of Cm = 50 nM and T = 80°C. A significant size increase was observed by ~3.5 × 105 nm2, ~5.1 × 105 nm2 and ~12. 6 × 105 nm2 following t = 1 h (5°C/~6 min.), 3 h (5°C/~15 min.) and 6 h (5°C/~30 min.), respectively (Figs. 2a, 2d and supplementary information Fig. S4, Table S6). No further increases were observed when the parameters were further extended over Cm = 20 nM, T = 80°C and t = 6 h.
Discussion
Experimental observation suggests a simple model for the Savg growth rate that predicts the average DNA crystal size controlled by Cm, T and t. Since the individual Savg growth rate for DNA crystals showed increases proportional to Cm, T and t (Fig. 2), each growth rate is independent of other constant fixed parameters. If so, the growth rate of DNA crystals can be expressed as follows,

where α(Cm), β(T) and γ(t) denote independent growth rates depending on each control parameter under specific conditions. Additionally, the fitted plot displayed in Figs. 2b–2d (red dotted line) show that the growth rates nearly follow quadratic exponential functions such as  . The coefficient values for each growth condition are exhibited in Fig. 2e. These coefficients are intrinsic parameters of DX crystals and determine the growth tendency at specific conditions. This expression can roughly predict the crystal growth rate at certain points.
. The coefficient values for each growth condition are exhibited in Fig. 2e. These coefficients are intrinsic parameters of DX crystals and determine the growth tendency at specific conditions. This expression can roughly predict the crystal growth rate at certain points.
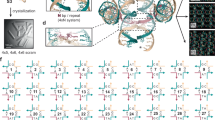
Crystallization includes nucleation and crystal growth. However, it is difficult to identify the latter since the system rapidly undergoes nucleation and crystal growth during SAG. A new experimental scheme called the direct annealing process has to be developed to achieve in situ AFM imaging while DNA crystallizes on the substrate (see Direct annealing process and sequential in situ AFM imaging in methods). Fig. 3a shows sequential in situ AFM images at a constant temperature of 40°C obtained by the direct annealing process. In this figure, Δt indicates time elapsed since collecting the AFM images after adding the last DX strand (DX1-4 shown in supplementary information Figs. S2 and S5) to the buffer solution on the preheated substrate. At Δt = 0 sec, following introduction of the DNA crystal assembly, small multimeric DX crystals began nucleation close to the substrate and consequently adsorbed leaving large sections uncovered by crystals. A distinct morphological change was observed between 0 and 550 sec, when the surface became densely populated with crystals. Full coverage was reached at Δt = ~550 sec. A magnified AFM image shows that individual DNA crystals have an Savg of 0.7 × 104 nm2 and a noise filtered reconstruction image collected by fast Fourier transformation shows clear crystal periodicity (Fig. 3a and supplementary information Fig. S5, Table S7). After Δt > 550 sec, no further distinct crystal transformation was observed until Δt near 3000 sec. In contrast to the sample crystallized at 40°C requiring time, immediate full coverage was achieved at both 50°C and 60°C. In addition, the Savg obtained at a constant temperature of 60°C corresponded to relatively large-scale DNA crystals almost 10 times larger than the Savg measured at 40°C, even though this temperature is well above the critical temperature for hybridization of DX crystals (Fig. 3b and supplementary information Fig. S6, Table S8)10,22. Crystal formation under such unfavorable conditions may be assisted by strong electrostatic interaction. This provides thermodynamic energy as a driving force for crystal formation rather than denaturation by paying some of the energy cost for DNA crystallization.
DNA crystallization process.
(a) Series of sequential in situ AFM height images showing the time-dependent covering process of DX crystals followed by nucleating and adsorbing processes of individual DX single strands on substrate at 40°C. (b) Plot of average crystal size distribution depending on the fixed substrate temperature. (c) Crystal growth followed by rearrangement of DNA tiles on the heated substrate. (d) Schematic diagram of DNA crystallization close to charged substrate. All scale bars are 500 nm unless otherwise noted.
The impetus behind crystal growth is a molecule's rearrangement behavior1. Crystal rearrangement growth represents changes in the shape of polycrystalline crystals that correspond to macroscopic changes in the orientations of adjacent ones. Similar morphological changes were observed when the temperature of fully covered polycrystalline DNA crystals fabricated at 40°C was intentionally increased to 60°C. As shown in Fig. 3c, large DX crystals with clear crystal boundaries began appearing. This suggests that small DX crystals combine neighbors together through molecular rearrangement24. Temperature might provide some activation energy for rearrangement and integration through short-range molecular mobility on the charged substrate. The molecule's rearrangement behavior on a substrate is supported by experimental data. First, the topological changes of DNA crystals depend on Cm. For 5 HR, the AFM image at low Cm shows initiation of crystallization through a random attaching process. However, the topology changed to a highly-packed 1D crystal arrangement with a certain orientation when Cm passed Cm-th (supplementary information Fig. S3). This behavior was also observed with DX crystals fabricated at different annealing times of t = 1 min. and 6 hours under the same conditions of Cm = 50 nM and T = 80°C. For instantaneous annealing time of t = 1 min., DNA strands formed small multimeric DNA crystals close to the charged substrate and randomly adsorbed onto the surface, whereas DNA crystals fabricated for 6 hours showed large DNA crystals with Savg of 12.6 × 105 nm2. Secondly, no crystals were observed on the substrate missing one DX sticky-end strand (DX1-4) in the DNA solution. However, when this strand was added to the solution after several minutes, crystal formation occurred even though electrostatic interaction suppressed further hybridization. The experimental results show that the DNA crystallization process can be roughly estimated, as shown in Fig. 3d. Due to strong electrostatic interactions between two charged materials, the appropriate molecular concentration for DNA crystallization was achieved close to the substrate. Consequently, each DNA strand began forming DNA motifs and small multimeric DNA crystals. The crystals were randomly adsorbed onto the surface by sufficient electrostatic interactions. Randomly arranged DNA crystals then further interacted with adjacent crystals through thermodynamic energy while annealing to form large DNA crystals. Simulation can be done only in limited cases, so we have less chances to verify that the theoretical model is correct. With our experimental results, we can provide reasonably good numerical values for model optimization.
We conclude that DNA crystallization on substrates is deeply associated to Cm, T and t. Interesting results were obtained by controlling each parameter to show different coverages of 1D and 2D DNA crystals. Gradual, though sometimes spontaneous, DNA crystallization onto the charged mica substrate was experimentally demonstrated. Some questions concerning the detailed pathway of DNA crystallization still remain. However, our results show deep insight concerning correlations of each external parameter. These correlations are essential resources for DNA crystallization catalyzed by electrostatic interactions under the SAG system. If any parameter is not sufficient for crystallization, no well-made or large crystals will form on the substrate even with sufficiently strong electrostatic interactions. Accurate size control and high-yield fabrication of large-scale crystals are critical issues for applying the DNA crystals constructed by SAG to current physical, chemical and biological devices and sensors that require precise regular arrangements of heteromaterials. Further studies are underway to expand the applicability of this analysis beyond fabricating an entirely single crystalline lattice on a given substrate. In addition, research is being performed on the theoretical basis of the crystallization pathway. If successful, this crystallization mechanism would greatly promote the fabrication of DNA crystals in micro- and even sub-millimetre scales and promote practical usage in physical and chemical application fields.
Methods
DNA crystal fabrication
Synthetic oligonucleotides purified by high performance liquid chromatography (HPLC) were purchased from Integrated DNA Technologies (IDT, Coralville, IA). Complexes were formed by mixing a stoichiometric quantity of each strand in physiological buffer, 1 × TAE/Mg2+. The mica substrate and DNA strands were then inserted into an AXYGEN-tube together. For annealing, the test tube was placed in a heat block and cooled stepwise from initial annealing temperature T to room temperature 25°C with time intervals of  at every 5°C. Here t is total anneal time.
at every 5°C. Here t is total anneal time.
AFM imaging
For AFM imaging, a SAG sample was placed on a metal puck using instant glue. Then, 30 μL 1 × TAE/Mg2+ buffer was pipetted onto the substrate and another 30 μL 1 × TAE/Mg2+ buffer was dispensed into the silicon nitride AFM tip (Veeco Inc.). AFM images were obtained by Multimode Nanoscope (Veeco Inc.) in liquid tapping mode.
Direct annealing process and sequential in situ AFM imaging
50 μL of physiological 1 × TAE/Mg2+ buffer solution was deposited on the substrate. The substrate had been preheated by placing it on a temperature applications controller (TAC-2, Veeco Inc.) with fixed temperature, implying a constant temperature while imaging. Then, 1 μL of each of the eight 10 μM DX strand solutions were added stepwise to buffer solution with a time interval of several tens of seconds. After that, the surface was immediately characterized by AFM under liquid tapping mode.
References
Humphrey, F. J. & Hatherly, M. Recrystallization and Related Annealing Phenomena. (Pergamon, 2004).
Mann, S. et al. Crystallization at inorganic-organic interfaces: biominerals and biomimetic synthesis. Science 261, 1286–1292 (1993).
Mann, S. Molecular recognition in biomineralization. Nature 332, 119–124 (1988).
Cashman, K. V. & Marsh, B. D. Crystal size distribution (CSD) in rocks and the kinetics and dynamics of crystallization. Crontrib. Mineral. Petrol. 99, 292–305 (1988).
Chung, S., Shin, S. H., Bertozzi, C. R. & De Yoreo, J. J. Self-catalyzed growth of S layers via an amorphous-to-crystalline transition limited by folding kinetics. Proc. Natl. Acad. Sci. U.S.A. 107, 16536–16541 (2010).
Judge, R. A., Forsythe, E. L. & Pusey, M. L. Growth rate dispersion in protein crystal growth. Cryst. Growth Des. 10, 3164–3168 (2010).
Tsai, C. J., Chen, M., Lu, H. Y., Chang, W. C. & Chen, C. H. Crystal growth rate and master curves of poly (ethylene succinate) and its copolyesters using a nonisothermal method. J. Polym. Sci. Part B: Polym. Phys. 48, 932–939 (2010).
Sanamaría-Holek, I., Gadomski, A. & Rubí, J. M. Controlling protein crystal growth rate by means of temperature. J. Phys.: Condens. Matter 23, 235101 (2011).
Korkmaz, N., Ostermann, K. & Rödel, G. Calcium dependent formation of tubular assemblies by recombinant S-layer proteins in vivo and in vitro. Nanotechnology 22, 095601 (2011).
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Yin, P. et al. Programming DNA tube circumferences. Science 321, 824–826 (2008).
Zheng, J. et al. From molecular to macroscopic via the rational design of a self-aseembled 3D DNA crystal. Nature 461, 74–77 (2009).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Lee, J. et al. Size-conrollable DNA rings with copper-ion modification. Small 8, 374–377 (2012).
Lee, J. et al. Fabrication of zigzag and folded DNA nanostructures by an angle control scheme. Soft Matter 8, 44–47 (2012).
Park, S. H. et al. Finite-size, fully addressable DNA tile lattices formed by hierarchical assembly procedures. Angew. Chem. 118, 749–753 (2006).
Maune, H. T. et al. Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates. Nature Nanotechnol. 5, 61–66 (2010).
Sharma, J. et al. Control of self-assembly of DNA tubules through integration of gold nanoparticles. Science 323, 112–116 (2009).
Yan, H. et al. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301,1882–1884 (2003).
Le, J. D. et al. DNA-templated self-assembly of metallic nanocomponent arrays on a surface. Nano Lett. 4, 2343–2347 (2004).
LaBean, T. H. et al. Construction, analysis, ligation and self-assembly of DNA triple crossover complexes. J. Am. Chem. Soc. 122, 1848–1860 (2000).
Rothemund, P. W. K., Papadakis, N. & Winfree, E. Algorithmic self-assembly of DNA sierpinski triangles. PLoS Biol. 2, 2041–2053 (2004).
Zhao, Z., Yan, H. & Liu, Y. A route to scale up DNA origami using DNA tiles as folding staples. Angew. Chem Int. Ed. 49, 1414–1417 (2010).
Hamada, S. & Murata, S. Substrate-assisted assembly of interconnected single-duplex DNA nanostructures. Angew. Chem. Int. Ed. 48, 6820–6823 (2009).
Sun, X., Ko, S. H., Zhang, C. Z., Ribbe, A. E. & Mao, C. Surface-mediated DNA self-assembly. J. Am. Chem. Soc. 131, 13248–13249 (2009).
Lee, J., Kim, S., Kim, J., Lee, C. W., Noh, Y. & Park, S. H. Coverage control of DNA crystals grown by silica assistance. Angew. Chem. Int. Ed. 50, 9145–9149 (2011).
Hamada, S. & Murata, S. Theoretical model of substrate-assisted self-assembly of DNA nanostructures. RSC Adv., 2, 7406–7412 (2012).
Erbil, H. Y. Surface chemistry of solid and liquid interfaces. (Blackwell publishing, 2006).
Acknowledgements
This paper was supported by Faculty Research Fund, Sungkyunkwan University, 2011 and by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MEST) (2012-0002000 & 2012-005985) to SHP and by Grant-in-aid for Scientific Research (Nos. 22220001 & 24104005) from Japanese government (MEXT) to SM.
Author information
Authors and Affiliations
Contributions
J.L. conceived and initiated this study. J.L. and S.U.H. prepared the samples, took AFM images and collected data. All authors contributed to the data analysis. J.L. wrote the manuscript with contributions from all authors. S.H.P. and S.M. supervised the research.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
DNA-SAG-SI
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Lee, J., Hamada, S., Hwang, S. et al. Quantitative analysis of molecular-level DNA crystal growth on a 2D surface. Sci Rep 3, 2115 (2013). https://doi.org/10.1038/srep02115
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02115
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.