Abstract
We set out to reduce the chemical constitution of a living organism to 19 amino acids. A strain was constructed for reassigning the tryptophan codon UGG to histidine and eliminating tryptophan from Escherichia coli. Histidine codons in the gene for an essential enzyme were replaced with tryptophan codons and the restoration of catalytic activity by missense suppressor His-tRNA bearing a CCA anticodon was selected. We used automated cultivation to assess the stability of this genetic construct during evolution. Histidine to tryptophan mutation at codon 30 in the transketolase gene from yeast and its cognate suppressor tRNA were stably propagated in a tktAB deletant of E. coli over 2500 generations. The ratio of histidine misincorporation at tryptophan sites in the proteome increased from 0.0007 to 0.03 over 300 days of continuous culture. This result demonstrated that the genetic code can be forced to evolve by permanent metabolic selection.
Similar content being viewed by others
Introduction
To what extent can the usage of an amino acid be decreased in a living cell? Can a lineage with a smaller set of canonical amino acids evolve from an extant species? These two questions have been considered in the context of prebiotic and evolutionary reconstructions, but have rarely been addressed experimentally1,2. However, it would be of interest to investigate these issues in the framework of synthetic biology. The current approach toward minimal life considers the reduction of the total number of building blocks used for constructing simplified cells rather than the reduction of the number of categories of building blocks from which cells are assembled3. Compositional extremophiles are living species whose proteomes, transcriptomes or genomes have unusually low levels of certain amino acids, nucleotides or deoxynucleotides. Such species exist in nature, although none has been found to lack a canonical amino acid so far completely4,5,6. Artificial versions of polypeptides and polynucleotides assembled from a restricted set of monomers and endowed with functional activity have been synthesized both in vitro and in vivo, but only as single isolated biopolymer chains. Whole living organisms based on such constructs have not been obtained1,7.
Our goal is manipulate E. coli so as to obtain cells with reduced proteomes, from which one amino acid has been completely eliminated. We chose tryptophan (Trp) as our first target for elimination. This is the rarest amino acid in wild-type E. coli and, to the best of our knowledge, has not been found to be essential for the catalytic moiety of any E. coli enzyme.
Our approach is to establish a stable ambiguity in translating the codons for a "prey" amino acid (Zaa) into a "predator" amino acid (Baa). Cells carrying this ambiguity would be then cultured for many successive generations subject to tight control of the exogenous supply of the prey amino acid. It is expected that adaptive mutations will accumulate in the genome of a bacterial cell such that eventually all codons specifying Zaa in the progenitor will be read as Baa in its progeny. Adaptive mutations might change Zaa codons in the genome to amino acids less deleterious at sensitive protein sites. Also, mutations at other sites mays be selected to tolerate the occupancy of a given Zaa site by Baa. The principles which guided the design of our experimental setup can be listed as follows: (i) hyposteric replaceability; (ii) mutational irreversibility; (iii) selective enforceability.
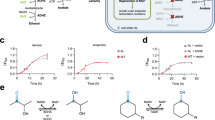
Hypostericity refers to the ability to nest Baa into the steric envelope of Zaa. This issue is likely to be the most critical constraint in reassigning the codons of a canonical amino acid to another amino acid (Fig. 1). Amino acid substitutions that cause steric clashes are known to be the most disruptive changes in protein folding and function8. However, changing isoleucine for valine (Fig. 1), valine for alanine or for cysteine, tyrosine for phenylalanine, alanine for glycine are expected to be the least disruptive.
Hyposteric amino acid substitutions.
Examples of valid and invalid hyposteric substitutions. Circles indicate steric groups absent from one amino acid, but present in the counterpart amino acid. Notice that aspartate is not hyposteric to glutamate because their common carboxylate moieties are not superimposable. In spite of the large difference in size, histidine can fit in the steric envelope of tryptophan.
Our approach requires a strong selective pressure to maintain a tRNA recognizing the codon for Zaa and incorporating Baa into the corresponding proteins (Baa-tRNAZaa). Changing a Baa codon to a Zaa codon at a critical site in an essential gene is expected to favor maintenance of such a suppressor tRNA. However, restoration of enzyme function through stable proteome miscoding using a suppressor Baa-tRNA/Zaa would be circumvented if the Zaa codon could easily mutate back to a Baa triplet. Therefore, the possible reversibility of a codon for the prey Zaa to any codon for the predator Baa through spontaneous mutations needs to be as low as possible. The most favorable cases correspond to substituting all three bases of a codon such as CUU (leucine) for AAA or AAG (lysine) and GCG (alanine) for UGC (cysteine).
Selection stringency could be tuned by modifying the ratio of the amounts of the canonical and miscoding aminoacyl-tRNAs recognizing the Zaa codon. Thus, to drive the process, the cellular availability of Baa-tRNA/Zaa needs to be maximized and that of the competing canonical Zaa-tRNA/Zaa minimized. Expression of appropriate genes encoding a mutant suppressor tRNA or a suppressor aminoacyl-tRNA synthetase have enabled the implementation of ambiguous proteomes (statistical proteins)9,10,11. The formation of canonical Zaa-tRNA/Zaa can be controlled by limiting the exogenous supply of Zaa and disrupting its biosynthesis. The most difficult amino acid biosyntheses to be irreversibly disrupted in E. coli are Ala, Asp,Ser ; somewhat easier: Glu, Phe, Gly, Pro, Cys, Arg, Tyr ; quite easy: His, Ile, Lys, Met, Asn, Gln, Thr, Val, Trp12.
In addition, the substitution of the anticodons with other triplets often leads to a large decrease in charging of the amino acid by the cognate synthetase in many cases13. The most tolerant synthetases in bacteria are those for leucine, alanine and serine and the least tolerant are those for valine, isoleucine, methionine and cysteine; histidyl-tRNA synthetase shows an intermediate sensitivity to anticodon mutations when charging its tRNA13,14. Overall, the reassignment of the Trp codon UGG to His appeared to be a good tradeoff between these various constraints. Tryptophan, encoded by a single codon UGG in most organisms, is the rarest amino acid in the proteome of E. coli and most other bacteria2. Three substitutions are needed to convert UGG into either of the two histidine codons CAU and CAC. Its lengthy biosynthetic pathway can be entirely disrupted by a single deletion of the Trp operon. Histidine is smaller by four methyne groups ( = C-H) than tryptophan and can be cleanly nested into tryptophan’s steric envelope (Fig. 1). Histidine is a catalytic residue in numerous enzymes and contributes to metal coordination in active sites15, covalent acyl transfer16 and proton shuttling17.
We describe the identification of an essential histidine in an essential protein. We replaced the codon for this histidine with a tryptophan codon and transformed the cells with a construct encoding a suppressor tRNA (recognizing the Trp codon and incorpotating a His residue). This system can be used subsequently to diminish the use of tryptophan throughout the whole E. coli proteome.
Results
Construction and validation of a miscoding His-tRNA Trp in vivo
To investigate the feasibility of incorporating a His at a Trp codon, a His-tRNA allele of E. coli bearing a Trp anticodon CCA (hisT:CCA) was synthesised and inserted into a high and into a low copy vector. After transformation of these constructions in E. coli MG1655 wild type, thermosensitivity in rich and poor medium was observed for these two transformant strains; it was exacerbated in the high expression case. This toxicity already suggested a misincorporation of amino acids through erroneous codon reading18. We looked for enzymes harbouring essential histidine residues to validate the histidine misincorporation at a tryptophan codon using this missense tRNA in vivo. The activity of chloramphenicol acetyl transferase encoded by the cat gene results from a covalent catalysis at the His 193 residue. The substitution of this invariant histidine inactivates the enzyme16. We confirmed that a cat gene containing this his codon mutated to a Trp codon (H193W mutant) did not confer an E coli resistance to chloramphenicol. Co-expression of the hisT:CCA gene encoding the cognate suppressor tRNA with the H193W cat allele restored chloramphenicol resistance (Supplementary figure S1). This shows that the His-tRNACCA was charged by a histidyl-tRNA synthetase in vivo allowing the introduction of a histidine at the Trp 193 codon of the mRNA from the H193W cat allele. Expression of Histidyl-tRNA synthetase in tandem with the His-tRNACCA did not lead to any enhancement of chloramphenicol resistance expressed by the H193W cat allele. Since maintaining an antibiotic resistance in protocols of directed evolution in vivo is notoriously unreliable, we attempted to set up a stable missense selection screen based on a biosynthetic function and more specifically on an enzyme involved in high flux carbon metabolism.
Transketolase selection setup
Transketolase is a key enzyme of the phosphopentose pathway that shuttles glycoaldehydes between the phosphosugars Xylulose-5P, Ribose-5P, Sedpheptulose-7P Erythrose-4P Fructose-6P and Glyceraldehyde-3P (Fig. 2). E. coli strains lacking the two paralogous genes for transketolase, tktA and tktB, are able to grow on mineral medium with glucose if also supplied with shikimate and pyridoxine. Two catalytic activities are required for converting phosphoribulose into phophotrioses and further glycolytic processing. The catalytic site of transketolase contains several conserved histidine residues19. Substitution of the residues His30 or His481 in yeast transketolase has been reported to impair transketolase function20. H481W or H30W alleles of the yeast transketolase gene (tktY) in a p15a vector were used to transform an E coli strain G91 deleted for tktAB. These transformants were unable to grow on glucose or ribose (table 1). Co-expression of the His-tRNACCA gene in tandem with each His to Trp tktY allele (H30W or H481W) in the G91 strain resulted in marginal growth in MS glucose medium (table 1). This experimental system thus requires incorporation of a His residue at a Trp codon for the strain to grow. The more stringent condition of cultivation with ribose (2 g/L) supplied as sole carbon and energy source in mineral medium MAM was subsequently used for selecting missense mutants of transketolase suppressed by a His-tRNACCA. Converting ribose into glycolysis intermediates requires two enzymatic steps catalysed by transketolase (Fig. 2).
Evolutionary kinetics
The genetic stability of these two His to Trp tktY alleles was studied by subjecting the corresponding strains in duplicate to turbidostat selection in a self-cleaning cultivation GM3 device21,22. This set-up selects for ever-faster growing mutants in the population while counterselecting the attachment of bacterial cells to the bioreactors and thus the development of biofilms21. To accelerate the evolution of strains in the GM3, the mutT gene was deleted from the ΔtktAB strain: this increases the frequency of T:A to G:C transversions 1000-fold23. We first studied the genetic stability of the H481W tktY allele. The doubling time of strain G1857 (ΔtktAB ΔmutT p tktY H481W tRNAHis/Trp(CCA)) at the start of the experiment was 7 hours. After 30 days of cultivation in the turbidostat, the doubling times of the two cultures were 3.8 and 3.2 hours (table 2). Isolates were obtained from the two independently evolved populations after 30 days of continuous culture in the GM3. Sequencing of the plasmids in these isolates revealed that the Trp codon at position 481 (UGG) had mutated to a codon for glycine (GGG). Directed mutagenesis was used to construct a H481G allele of the yeast transketolase gene. This allele was introduced into strain G91 ΔtktAB and restored the ability to grow on glucose, albeit slowly. Therefore, a single point mutation resulting in the incorporation of a glycine at position 481 restores weak transketolase activity. Consequently, the continuous cultures of strain G1857 in GM3 were interrupted and additional mutations in the plasmids or chromosomes of the evolvants were not investigated. N-terminally His tagged enzymes corresponding to the wild type and the H481G tktY alleles were constructed in a pET vector, produced and purified on columns. The transketolase activity, estimated from kcat and Km measurements, of the H481G enzyme was three- to ten-times lower than that of the wild-type (table 3). These results are consistent with findings for of H481A and H481S mutants20. This experiment confirms that the GM3 self-cleaning turbidostat can be used for selecting enzyme variants.
As many different histidine residues are implicated in the catalytic site of the yeast transketolase, we used the same approach with the His at position 30. Substitution of His30 by most amino acids leads to enzyme inactivation in S. cerevisiae and E. coli24. Tandem coexpression of the suppressor tRNA and the tktY H30W alleles resulted in very slow growth: the generation times at the start of two parallel cultures in turbidostat devices were 25 h and 44 hours (table 2). After 300 days of cultivation corresponding to 2554 and 2674 generations under permanent selection for faster growth yielded populations with doubling times of 2.18 and 2. 48 hours respectively (table 2). The mutated codon at position 30 of tktY was unchanged in both evolved populations after 300 days. In one population an additional mutation was found, K518T through a CG to GC transversion, which appeared after 150 days of continuous culture. A double allele W30H and K518T allele did not complement ΔtktAB in strain G91 and therefore did not express transketolase activity. Therefore, a mutation at codon 30 of the yeast tkt in E. coli seems to ensure long-term maintenance of the suppressor tRNA. This model is therefore suitable for applying evolutionary pressure for His miscoding at Trp codons and thus selection for replacement of Trp residues with His.
Biochemical evidence of histidine misincorporation by mass spectrometry
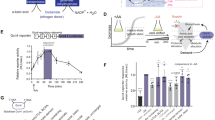
We used mass spectrometry to provide direct evidence of histidine misincorporation in suppressed strains. We designed a reporter system using the 4 oxalocrotonate tautomerase, a short monomeric enzyme that can be overexpressed in E. coli. The xylH gene from Pseudomonas putida encoding this enzyme has no Trp codon and contains only two His codons at position 7 and 5025. A synthetic gene encoding a Trp at position 7 and a His tag at the N terminus was inserted into an E. coli vector. This reporter construct was introduced into the various strains prior to (G2140) and after 300 days of directed evolution (G2141 and G2220). Protein extracts were prepared from cultures of all these transformants and the His-tagged protein was purified by affinity chromatography on NTA nickel resins and HPLC. The purified His-tagged xylH protein preparations were then digested with protease V8 that cleaves peptide bonds C-terminal to glutamic acid residues. The V8 digested samples were subjected to MALDI-TOF mass spectrum analysis. Six peptides could be identified (Fig. 3A). A peptide at m/z 2556 corresponding to the His miscoding at codon Trp 7 was detected in addition to a peptide at m/z 2605 corresponding to the same fragment but with a Trp at position 7 (Fig. 3B). The peptide at m/z 2556 resulting from histidine misincorporation was detected only in samples prepared from strains carrying both the HisT CCA gene for His miscoding tRNA and the reporter xylH UGG 7 (Fig. 3C). The amino acid sequences of peptides at m/z 2556 and at 2605 were determined by MSMS analysis (Supplementary figure S2) and confirmed the misincorporation of histidine at codon Trp7. The xylH reporter system, convenient for mass spectrometry analysis, thus clearly demonstrated His misincorporation in vivo. However, this approach is not suitable for measuring the His misincorporation into the whole proteome.
Mass spectrometry of the XylH H7W reporter protein.
(A) Sequences of predicted V8 protease digested products of XylH H7W. Amino acid composition of XylH7W (reporter protein) and amino acid composition of XylH with a His misincorporation at position7 (mistranslated reporter protein). For both proteins, the predicted peptides are shown in light blue with the corresponding mass indicated below. His misincorporation at position W7 of XylH is characterized by the appearance of a peptide of mass 2556.33 (B) MALDI-TOF mass spectrometry of V8 protease-digested XylHH7W. The protein was produced in the strain G2141. Peptides with masses of 2605 and 2556 correspond to the reporter protein and to the mistranslated reporter protein, respectively. The m/z are indicated. (C) Comparison of MALDI-TOF mass spectrometry in the m/z 2540 and 2620 region of V8 protease-digested XylHH7W purified from the various strains: (a) G2052 (no tRNA suppressor) (b) G2140 (c) G2141 (d) G2220.
His misincorporation measurement in suppressed cells
In our selection model, His misincorporation at H30W codon is necessary for cell growth but it can occur at each Trp codon of cellular proteins. To measure and compare His misincorporation into proteins of suppressed cells, we developed a new quantitative assay for proteome analysis based on mass spectrometry. E. coli proteins from each strain lysate were migrated on a SDS page gel. Five major bands from each lysate were excised from the gel, digested by trypsin and analyzed by Nano LC MSMS. For each band only the most abundant peptides were selected for MSMS analysis. His misincorporation was detected by using Sequest software to screen for a difference of 49 Daltons between otherwise identical peptides; 49 Da is the difference in mass between Trp and His. In strains G2141 and G2220 (strains isolated after forced evolution), 16 and 31 His misincorporations were identified in 12 and 20 different cellular proteins respectively (table 4). In the non-evolved G2140 strain, only two His misincorporations were detected and both were in the Cat proteins encoded by the plasmid. Peptides derived from plasmid-encoded proteins were about 100 times more abundant than peptides from cellular proteins. No His misincorporation in any peptide was found in the control strain G2052 not containing HisT UUG gene. The overall ratio of His replacement in cellular peptides was 0.0007 in G2140 but substantially higher in the evolved strains: 0.030 in G2141 and 0.032 in G2220 (table 4). The only His misincorportation detected in the Cat protein in the three strains was at the Trp16 position in the tryptic peptide ITGYTTVDISQWHR. The misincorporation ratio at this position was 0.012 in G2141, 0.032 in G2220 and 0.0006 in G2140. Therefore permanent selection for His miscoding at Trp codons over 2500 generations resulted in fifteen- to thirty-fold amplification of suppression.
Discussion
The transketolase active site, with its network of conserved His residues provides a potent selection screen for miscoding His in response to the Trp codon UGG. Maintaining this metabolic selection under conditions of permanent growth in a turbidostat allowed us to validate the genetic stability of the Trp30 residue over 2500 generations. By contrast, Trp substitution at the conserved site His481 was reproducibly followed by recovery of activity through point mutation, to Gly481, within 160 generations. Similarly, an inactivating His263 to Trp substitution reverted to a functional Arg263 transketolase variant within 262 generations in the turbidostat (data not shown). The GM3 turbidostat technology constitutes a powerful approach to modeling protein evolution and to studying amino acid plasticity at particular positions in protein sequences.
After 2500 generations (300 days), prolonged selection for Trp30 suppression led to a 30-fold increase in His misincorporation. This result is highly significant as it provides experimental evidence for the feasibility of genetic code reassignment through natural selection as an adaptive response to pressure for miscoding enforced metabolically. It would be interesting to elucidate the molecular mechanisms underlying this enhancement of miscoding by genomic and enzymologic studies.
Our next step will be to reduce tryptophan availability in our model system, by deleting the Trp operon and progressively eliminating tryptophan from the medium. This will provide pressure to eliminate this amino acid from the E coli proteome through reassignment of UGG to histidine. Direct analysis of the proteome by mass spectrometry, as developed here, will allow quantification of the His misincorporation rate in the evolving strains. The medium-swap pulse-feed regime, as previously used for replacing thymine with chlorouracil in E. coli, could be applied for this purpose22.
Methods
Strains and growth conditions
The strains used and constructed in this study are all derivatives of the wild type Escherichia coli K12 strain MG1655 (Supplementary Table S2). Bacteria were routinely grown in rich LB medium or in mineral medium (MS) containing 2 g of D-glucose or D-ribose per liter. Liquid and solid cultures were incubated at 30°C. Strain G1851 was obtained by deletion of the mutT allele from the G91 strain (ΔtktAB)26 as previously described27. The mutT was replaced by an excisable kanamycin resistance cassette. Strains G1857 and G2140 were cultivated in continuous culture in a GM3 device22 at 30°C in MAM medium (15 μM Na2HPO4, 10 mM KH2PO4, 10 mM (NH4)2SO4, 1 mM MgCl2, 100 μM nitrilotriacetic acid, 3 mM FeCl3, 1 μM MnCl2, 1 μM ZnCl2, 0.3 μM (CrCl3, H3BO3, CoCl2, CuCl2, NiCl2, Na2MoO2, Na2SeO3) containing 0.02% D-ribose. After 300 days of continuous culture in two independent GM3 devices, G2140 evolved strains were named G2141 and G2220.
Plasmid constructions
The following synthetic oligonucleotide (5’GGAATTCGAGCTCGGTACCCGGGGATCCTCTAGAGGCCCGTTGGTCAAGCGGTTAAGACACCGCCCTTTCACGGCGGTAACACGGGTTCGAATCCCGTACGGGTCACCAGCATGCCTAGGTTTAAACTAAGGAGGTTAATTAA 3’) was ligated between EcoRI and PacI sites in pVDM18 vector26 to give a p15a vector with two multiple cloning sites under the control of the same lacZ promoter but downstream from two different ribosome binding sites. The first MCS contains the following restriction sites EcoRI, PstI, SacI, KpnI, SmaI, BamHI, XbaI, SphI, BlnI and PmeI and the second MCS contains PacI, NotI and HindIII restriction sites. This plasmid was named pEVL649. The EcoRI/HindIII DNA fragment containing these two consecutive MCS was also inserted into a pUC18 vector to give pEVL683. The DNA sequence corresponding to the His tRNA from E. coli containing the Trp anticodon CCA: (5’GGATCCGGTGGCTATAGCTCAGTTGGTAGAGCCCTGGATTCCAATTCCAGTTGTCGTGGGTTCGAATCCCATTAGCCACCCCAGCATGC 3’) was synthesized in vitro and inserted between the BamHI and SpHI restriction sites in pEVL649. The yeast transketolase genes carrying mutations were individually ligated between the PacI and Not I restrictions sites in the second MCS of both the plasmids with and without the gene for the His tRNA CCA. pEVL683 derivatives with the gene for the His tRNA CCA between the BamHI and SpHI sites, with and without the histidyl tRNA synthetase gene between the PacI and NotI sites were constructed. The chloramphenicol acetyl transferase gene carrying a H193W mutation was inserted into pEVL550 (itself constructed by ligation of a DNA fragment containing the PacI and NotI restriction between EcoRI and HindII sites of pSU38). The synthetic xylH encoding the 4 oxalocrotonate tautomerase with a His to Trp substitution at position 7 and 6 His tag at the N-terminus was inserted between the XbaI/HindIII sites of pUC18 and the resulting plasmid was named pGEN695. The tkt gene from S. cerevisiae carried in pUC18 was subjected to site-directed mutagenesis. Correct construction of sequences encoding the H30W and H481W mutants and the absence of other mutations were verified by sequencing. The mutated genes were then subcloned independently into pEVL649 without the His/Trp tRNA.
Protein purification and in vitro assays
The transketolase genes (encoding the H481G mutant and the wild type) on peT47b plasmids were used to transform strain BL21. Cultures were grown in LB medium and at an OD (600 nm) of 0.6 gene expression was induced by incubation with 0.05 mM IPTG for 16 h at 20°C. Cells were pelleted and incubated at −80°C for 30 min. The cells in the pellets were then lyzed in the following buffer (50 mM NaH2PO4 pH 8, 300 mM NaCl, 10% glycerol, 0.5 mM TPP, 2 mM MgCl2) with lyzonase for 20 min at 30°C and subjected to sonication. The lysate was centrifuged at 10000 g for 30 min and the supernatant was applied to Protino Ni-TED columns. The eluted protein was concentrated on Amicon Centricon (3 kDA).
Determination of kinetic parameters of transketolase proteins
The assay for transketolase activity was based on a coupled system with glyceraldehyde-3-P dehydrogenase and NAD+ and was as described previously20. Kinetic steady state parameters were determined in 50 mM TrisHCl buffer at pH 7.5. For substrate parameter determinations, the MgCl2 concentration was 1 mM, the concentration of the cosubstrate was 5–10 times its Km value and the concentration of the investigated substrate was varied from 23 μM to 3 mM. All kinetics measurements were done at 25°C in duplicate.
Mass spectrometry analysis on XylH protein
Strains G2052, G2140, G2141 and G2220 were transformed with pGEN695. Cultures of 200 ml of each transformed strain were grown overnight at 30°C. Cells were lysed and sonicated as described for the transketolase enzymes. The His-tagged XylH protein was purified on Protino Ni-TED columns and concentrated on Amicon Ultra4 (Millipore). The eluted proteins were injected into a Waters 2795 HPLC apparatus. A 208MS5415 C8 column with a precolumn packed with the same phase (Grace Vydac) was used for purification. Gradient elution was performed with (A) water/0.05% trifluoroacetic acid and (B) acetonitrile/0.05% trifluoroacetic acid as mobile phases. After 5 min of isocratic elution in A, the gradient was run from A to 50% B in 20 min and then to 100% B in 5 min. The flow rate was 1 ml/min and the injection volume was 25 μl. The eluted products were detected in an UV detector (Waters 996 Photodiode Array detector) and the main peak was collected and concentrated with a SpeedVac (Eppendorf) to a protein concentration about 2 mg/mL. XylH (15 μg) was digested with 0.75 μg of Endoproteinase GluC from S. aureus V8 (sigma) overnight at 37°C in 5 μl of 100 mM NH4HCO3, pH 7.8. The matrix solution was prepared at a concentration of 10 mg/ml in 50/50 acetonitrile/water (0.1% TFA). Aliquots of 0.5 μl of a 10/1 mixture of matrix and digest solutions were spotted on a sample plate, allowed to dry in air and then inserted into the mass spectrometer (“dried droplet method”). MALDI-TOF MS and MALDI-TOF/TOF MS/MS analyses were performed using a 4800 MALDI-TOF/TOF Analyser mass spectrometer (AB Sciex, Les Ulis, France). The instrument was equipped with an Nd:YAG laser (operating at 355 nm wavelength, 500 ps pulse and 200 Hz repetition rate). Acquisitions were performed in the positive ion mode. For MS/MS experiments, precursor ions were accelerated at 8 keV and the MS/MS spectra were acquired using 1 keV collision energy with CID gas (air) at a pressure of 3.5 × 10−6 Torr. MS and MS/MS data were processed using DataExplorer 4.4 (AB Sciex).
In gel digest of total extract of proteins
E. coli strains (G2052, G2140, G2141 and G2220) were grown in LB overnight at 30°C. Pellets were lyzed by lyzonase 30 minutes at 30°C and centrifuged at 15000 rpm for 30 minutes at 4°C. Twenty microgrammes of proteins collected in the supernatant were loaded on a SDS PAGE gel. Five bands between 5 and 60 KDa of a SDS PAGE gel were excised for each strain of E coli. Gel pieces were thoroughly washed, reduced with DTT and alkylated with iodoacetamide. Proteins were then digested with trypsin (Promega) and the resulting peptides were extracted with a solution of acetonitrile, water and formic acid (60/38/2) and dried under vacuum.
Identification of proteins by NanoLC-MS/MS mass spectrometry and measurement of substitution rates
Peptide mixtures were injected into a nanoLC apparatus and subjected to electrospray tandem mass spectrometry. A U3000 Dionex nanoflow system connected to a LTQ Orbitrap mass spectrometer equipped with a nanoelectrospray source (Thermo-Fischer, Bremen, Germany) was used for peptide analysis. A C18 pepmap 100 column (75 μm ID, 25 cm length, 3 μm 100A Dionex) was used for chromatographic separation. Peptide mixtures were injected onto a pre-concentration column with a flow rate of 20 μl/min of 0.1% TFA in water . After a three-minute wash with the same solvent, the peptides were eluted and separated on the analytical column with at flow of 300 nl/min and a 70 min gradient from 2% to 60% acetonitrile in 0.1% formic acid. The mass spectrometer was operated in the data-dependent mode to switch automatically between orbitrap MS and MS2 in the linear trap. Survey full scan MS spectra from 500 to 2000 Da were acquired in the orbitrap with resolution R = 60000 at m/z 400, after accumulation of 500,000 charges on the linear ion trap. The most ions with the most intense signals (up to six, depending on signal intensity) were sequentially isolated for fragmentation in the linear ion trap using CID at a target value of 100,000 charges. The resulting fragments were recorded in the linear trap. Proteins were identified by high accuracy MS and MSMS Data using Sequest (Thermo-Fischer). The mass accuracy for MS and MS/MS Data was set to 5 ppm and 0.8 uma respectively. A bank of sequences from E. coli and Yeast extracted from the Uniprot database was used for data mining. The substitution of tryptophan residues by histidine residues was identified by Sequest software which looked for a loss of 49.0204 Da corresponding to the difference between the masses of the two residues. The area of each parent on the MS1 signal was calculated by proteome discoverer 1.2. (Thermo-Fischer). The substitution rate was determined by calculation of the ratio between the areas of the pairs of peaks when a substitution was found.
References
Walter, K. U., Vamvaca, K. & Hilvert, D. An active enzyme constructed from a 9-amino acid alphabet. J. Biol. Chem. 280, 37742–37746 (2005).
Wong, J. T. Evolution of the genetic code. Microbiol. Sci. 5, 174–181 (1988).
Glass, J. I. et al. Essential genes of a minimal bacterium. Proc. Natl. Acad. Sci. U.S.A. 103, 425–430 (2006).
Baudouin-Cornu, P., Surdin-Kerjan, Y., Marlière, P. & Thomas, D. Molecular evolution of protein atomic composition. Science 293, 297–300 (2001).
Elser, J. J., Acquisti, C. & Kumar, S. Stoichiogenomics: the evolutionary ecology of macromolecular elemental composition. Trends Ecol. Evol. (Amst.) 26, 38–44 (2011).
Mazel, D. & Marlière, P. Adaptive eradication of methionine and cysteine from cyanobacterial light-harvesting proteins. Nature 341, 245–248 (1989).
Reader, J. S. & Joyce, G. F. A ribozyme composed of only two different nucleotides. Nature 420, 841–844 (2002).
Nene, V. & Glass, R. E. Genetic studies on the beta subunit of Escherichia coli RNA polymerase. I. The effect of known, single amino acid substitutions in an essential protein. Mol. Gen. Genet. 188, 399–404 (1982).
Pezo, V. et al. Artificially ambiguous genetic code confers growth yield advantage. Proc. Natl. Acad. Sci. U.S.A. 101, 8593–8597 (2004).
Brustad, E., Bushey, M. L., Brock, A., Chittuluru, J. & Schultz, P. G. A promiscuous aminoacyl-tRNA synthetase that incorporates cysteine, methionine and alanine homologs into proteins. Bioorg. Med. Chem. Lett. 18, 6004–6006 (2008).
Schimmel, P. Mistranslation and its control by tRNA synthetases. Philos. Trans. R. Soc. Lond., B, Biol. Sci. 366, 2965–2971 (2011).
Gottschalk, G. Bacterial Metabolism. (Springer: 1986).
Beuning, P. J. & Musier-Forsyth, K. Transfer RNA recognition by aminoacyl-tRNA synthetases. Biopolymers 52, 1–28 (1999).
Vasil’eva, I. A. & Moor, N. A. Interaction of aminoacyl-tRNA synthetases with tRNA: general principles and distinguishing characteristics of the high-molecular-weight substrate recognition. Biochemistry Mosc. 72, 247–263 (2007).
Bertrand, T. et al. Crystal structure of a four-copper laccase complexed with an arylamine: insights into substrate recognition and correlation with kinetics. Biochemistry 41, 7325–7333 (2002).
Lewendon, A., Murray, I. A., Shaw, W. V., Gibbs, M. R. & Leslie, A. G. Replacement of catalytic histidine-195 of chloramphenicol acetyltransferase: evidence for a general base role for glutamate. Biochemistry 33, 1944–1950 (1994).
Polgár, L. & Bender, M. L. The nature of general base-general acid catalysis in serine proteases. Proc. Natl. Acad. Sci. U.S.A. 64, 1335–1342 (1969).
Döring, V. & Marlière, P. Reassigning cysteine in the genetic code of Escherichia coli. Genetics 150, 543–551 (1998).
Schneider, G. & Lindqvist, Y. Crystallography and mutagenesis of transketolase: mechanistic implications for enzymatic thiamin catalysis. Biochim. Biophys. Acta 1385, 387–398 (1998).
Wikner, C. et al. Identification of catalytically important residues in yeast transketolase. Biochemistry 36, 15643–15649 (1997).
Marlière, P. & Mutzel, R. Patent W0 00/34433. (2000).
Marlière, P. et al. Chemical evolution of a bacterium’s genome. Angew. Chem. Int. Ed. Engl. 50, 7109–7114 (2011).
Cox, E. C. Mutator gene studies in Escherichia coli: the mutT gene. Genetics 73, Suppl 73, 67–80 (1973).
Hibbert, E. G. et al. Directed evolution of transketolase activity on non-phosphorylated substrates. J. Biotechnol. 131, 425–432 (2007).
Chen, L. H. et al. 4-Oxalocrotonate tautomerase, an enzyme composed of 62 amino acid residues per monomer. J. Biol. Chem. 267, 17716–17721 (1992).
Simon, G. et al. Amino acid precursors for the detection of transketolase activity in Escherichia coli auxotrophs. Bioorg. Med. Chem. Lett. 19, 3767–3770 (2009).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. U.S.A. 97, 6640–6645 (2000).
Acknowledgements
We thank the sequencing department of Genoscope. We also thank Susan Cure, Valerie de Crecy-Lagard and Bruno Sargueil for improving the manuscript and Lan N’guyen and Isabelle Boko for technical assistance.
Author information
Authors and Affiliations
Contributions
V. Pezo and P. Marliere wrote the main manuscript text and figures 1 and 2. D. Louis prepared figure 3. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
A Metabolic Prototype for Eliminating Tryptophan From The Genetic Code V Pezo, D Louis, V Guerineau, JP Le Caer, L Gaillon, R Mutzel and P Marliere
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Pezo, V., Louis, D., Guérineau, V. et al. A Metabolic Prototype for Eliminating Tryptophan From The Genetic Code. Sci Rep 3, 1359 (2013). https://doi.org/10.1038/srep01359
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01359
This article is cited by
-
Experimental Evolution of a Green Fluorescent Protein Composed of 19 Unique Amino Acids without Tryptophan
Origins of Life and Evolution of Biospheres (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.