Abstract
c-Src is a non-receptor tyrosine kinase involved in numerous signal transduction pathways. The kinase, SH3 and SH2 domains of c-Src are attached to the membrane-anchoring SH4 domain through the flexible Unique domain. Here we show intra- and intermolecular interactions involving the Unique and SH3 domains suggesting the presence of a previously unrecognized additional regulation layer in c-Src. We have characterized lipid binding by the Unique and SH3 domains, their intramolecular interaction and its allosteric modulation by a SH3-binding peptide or by Calcium-loaded calmodulin binding to the Unique domain. We also show reduced lipid binding following phosphorylation at conserved sites of the Unique domain. Finally, we show that injection of full-length c-Src with mutations that abolish lipid binding by the Unique domain causes a strong in vivo phenotype distinct from that of wild-type c-Src in a Xenopus oocyte model system, confirming the functional role of the Unique domain in c-Src regulation.
Similar content being viewed by others
Introduction
C-Src is the leading member of the Src family of non-receptor tyrosine kinases (SFKs) involved in many signaling pathways1,2,3,4,5. The SH3, SH2 and kinase domains of SFK members display large sequence and structural similarity. They also have in common a myristoylated and/or palmitoylated membrane anchoring region in the N-terminus, including positively charged residues (Arg and/or Lys), known as the SH4 domain6,7. In contrast, the segment connecting the SH3 and SH4 domains, known as the Unique domain, presents unique sequences for each SFKs protein and is intrinsically disordered. With few exceptions, no clear function has been assigned to the Unique domains of SFKs8,9,10,11. However, swapping the Unique domains of c-Src and c-Yes results in a change in their specificity12,13. Previous studies indicate that, in addition to the SH4 domain, further regions in the first 111 amino acids are involved in membrane anchoring. Indeed, removal of more than 8 kDa from the amino terminus of c-Src failed to detach the protein from membranes and a c-Src mutant lacking residues 8 to 37 still binds to membranes14,15,16.
The N-terminal region of c-Src, including the SH4 and Unique domains, was found by NMR to display a partially structured section between residues 60–64 and 67–7417. Here we show that this region includes a new lipid binding site, in addition to the SH4 domain, with affinity for acidic lipids and modulated by phosphorylation of neighbor serine and threonine residues, by binding to calcium-bound calmodulin or by interaction with the adjacent SH3 domain. We further show, for the first time, that the SH3 domain of c-Src contains also a lipid binding region in the opposite side of its peptide binding site. The functional relevance of the partially structured region of the Unique domain has been demonstrated in the Xenopus laevis oocyte model system.
Together, these results reveal an important functional role for the Unique domain of c-Src and suggest the existence of a previously unrecognized regulation layer in c-Src and, possibly, other related SFKs. The complex regulation of c-Src is consistent with its role as a hub connecting many different signaling pathways.
Results
Lipid binding by the Unique domain of c-Src
A15N-labelled construct containing residues 1–85 of c-Src (USrc) was studied by NMR in the presence of bicelles containing variable proportions of dimyristoyl phosphatidyl choline (DMPC) and the acidic lipid dimyristoyl phosphatidyl glycerol (DMPG). Bicelles are small disk-like structures formed by planar bilayers of long-chain lipids closed by highly curved, micelle-like walls formed essentially by short chain lipids (dihexanoyl phosphatidylcholine, DHPC). These model membranes have been previously used to study protein-lipid interactions18,19,20. An expansion of the 1H-15N HSQC NMR spectra of USrc in the absence or presence of lipids and a plot of the lipid-induced shift changes in each residue are shown in Figures 1a and 1b , respectively. Significant shifts were observed in two regions: the N-terminal SH4 domain and residues S51, A53, A55 and 60EPKLFGGF67 of the Unique domain. The observed shifts in the Unique domain region were larger than those observed in the well known lipid binding SH4 domain. The 60–67 segment is included in the partially structured region (60–75) previously determined by NMR17 and we refer to it as the Unique Lipid Binding Region (ULBR). The SH4 domain and the ULBR showed affinity for negatively charged lipids. However, in contrast to residues in the SH4 domain, the ULBR was also substantially perturbed by neutral lipid bicelles. The specificity for different lipid classes was tested using Lipid-StripsTM. USrc binds preferentially to acidic lipids, namely phosphatidic acid (PA), cardiolipin (CL), phosphatidylserine (PS), phosphatidylinositol-4-phosphate (PtdIns(4)P) and phosphatidylinositol-3,4,5-triphosphate (PtdIns(3,4,5)P3) ( Fig. 1c ).
Lipid binding by the Unique domain of c-Src (a) Overlay of 1H-15N HSQC NMR spectra USrc alone (red) and in the presence of DMPC/DHPC bicelles (blue) and DMPG/DHPC bicelles (grey).(b) Combined 1H-15N lipid-induced chemical shift changes per residue. The color code is the same as in (a). Total lipid concentration was 8% w/v and the ratio of long chain lipids to DHPC (q) was 0.8. (c) Binding of USrc to immobilized lipids, detected by immunoblotting with anti-Strep-tag HRP (TG triglyceride, DAC diacylglyceride, PA phosphatidic acid, PS phosphatidylserine, PE phosphatidylethanolamine, PC phosphatidylcholine, PG phosphatidylglycerol, CL cardiolipin, PtdIns phosphatidylinositol). (d) Schematic representation of the structure and charge of the lipid bicelles used.
Lipid binding by SH3 domain of c-Src
We next investigated lipid binding by the Unique domain attached to the SH3 domain ( Fig. 2a,b ) (USH3, residues 1–150) and the isolated SH3 domain ( Fig. 2c,d ) (SH3, residues 86–150). The effect of lipids in the isolated or SH3-bound Unique domain were very similar. However, large lipid-induced shifts were also observed for residues 98–102 and 114–116 of c-Src SH3 both in the USH3 construct ( Fig. 2a ) and in isolated SH3 domain ( Fig. 2d ). These residues are located in the opposite face to the one containing the canonical peptide binding site of the SH3 domain ( Fig. 2e ). Lipid binding by the isolated SH3 domain was also observed using Lipid-StripsTM and showed a similar lipid selectivity than the Unique domain ( Fig. 2c ). However, the USH3 construct, where the two domains are simultaneously present, did not bind phosphoinositides (PIPs) although it could still bind to other acidic lipids ( Fig. 2b ). The changes in lipid selectivity of the two-domain construct with respect to the isolated domains suggest the two domains are interacting.
Lipid binding by Unique-SH3 (USH3) and isolated SH3 domains.
Lipid binding by the USH3 construct (a, b) and the isolated SH3 domain (c,d,e). (a, d)) Combined 1H-15N chemical shift perturbations induced by the presence of 8% w/v of DMPG/DHPC bicelles (q = 0.8). (b,c) Immunoblots showing the lipid binding specificity (see Fig. 1c for the identity of the lipids and to compare with the isolated Unique domain). (e) Residues perturbed in presence of lipids are highlighted in yellow on the SH3 surface (PDB code: 1SHG). The polyproline binding site is located in the opposite side and indicated by the presence of a polyproline ligand.
Interaction between the Unique and SH3 domains of c-Src
In order to test for inter-domain interactions we compared the chemical shifts of equivalent residues in the USH3, USrc and SH3 constructs. Representative HSQC NMR spectra are given in Supplementary Figure 1 . Chemical shift changes between isolated and linked domains are shown in Figures 3a,b . Significant shifts were observed for H25, T37, H47, A55, N68, a region overlapping with the ULBR (Unique domain) and residues 98–103, 114–116, 134 and 145 (SH3 domain). The SH3 residues belong to the conserved RT (98–103) and nSrc (114–116) loops ( Fig. 3d ). Some of these residues (R98, E100, D102, L103, Y134) and S97 experienced small chemical shift changes when USrc was added to the isolated domain ( Supplementary Fig. 2 ) confirming that the interaction could occur, even when the two domains are not linked.
Interaction between Unique and SH3 domains (a,b) Combined absolute values 1H-15N NMR chemical shift differences between linked and isolated domains.
(a) USH3 versus USrc. (b) USH3 versus SH3. (c) Intensity ratios of NH cross-peaks from MTLS-labeled (A59C/USH3 at pH 7.0) between paramagnetic and diamagnetic (DTT reduced) forms. (d) SH3 residues perturbed by interaction with the Unique domain are highlighted in orange. The polyproline binding site is located in the opposite site and indicated by the presence of a polyproline ligand. (e) Cartoon representation of the interaction between SH3 and Unique domains observed by PRE.
The mapping of the interacting residues was confirmed by Paramagnetic Relaxation Enhancement (PRE) induced by (1-oxy-2,2,5,5-tetramethyl-D-pyrroline-3-methyl)-methanethiosulfonate (MTSL) linked to a cysteine residue introduced by site specific mutagenesis at position 59. NMR signals from nuclei that are, even transiently, close to the paramagnetic centre ( Fig. 3e ) are broader and less intense than the equivalent peaks from diamagnetic controls. Figure 3c shows the ratio of intensities of NH signals between paramagnetic and diamagnetic forms, plotted along the protein sequence.
The clusters of residues in c-Src SH3 most affected by the paramagnetic probe in residue 59 of the Unique domain matched those showing the largest chemical shift changes between SH3 and USH3 (cf. marked regions in Figs. 3b and 3c ). This result also confirmed that A59 is close to the interacting region.
The Unique-SH3 interaction is allosterically prevented by binding of a polyproline peptide to the SH3 domain
The SH3 domain of c-Src binds peptides containing the Pro-X-X-Pro motif in a polyproline II helix21. We investigated by NMR the effect of binding of a consensus high affinity unlabelled SH3 peptide ligand (Ac-VSLARRPLPPLP-OH) on the interaction between the Unique and SH3 domains in15N-labeled USH3. Figures 4a–e show expansions of NMR spectra including signals from the NH group of A55 and a non-perturbed residue (E150) in the presence of increasing amounts of peptide. In the presence of substoichiometric amounts of peptide, the A55 residue appeared duplicated and the relative intensities of the two peaks changed during the titration. This behavior is characteristic of slow exchange in the NMR time scale between peptide-bound and free USH3. The chemical shifts of NH groups in the Unique domain of peptide-bound USH3 matched those of the isolated Unique domain ( Fig. 4f ), indicating that peptide binding to the SH3 domain abolished its interaction with the Unique domain. The SH3 residues that interact with the Unique domain are located on the opposite side of the SH3 domain where polyproline ligands bind and therefore the loss of the interaction between the Unique and SH3 domain is not the result of direct competition with the peptide but an allosteric effect ( Fig. 4g ).
Allosteric effect of polyproline peptide binding on the Unique-SH3 interaction.
(a–e) Expansion of HSQC spectra showing residues A55 and E150 (acting as control) at molar rations of the Unique-SH3 construct (USH3) to Ac-VSLARRPLPPLP-OH of (a) 0, (b) 0.2, (c) 0.4, (d) 0.7 and (e) 1.0. (f) Expansion of HSQC spectrum of the isolated Unique domain (USrc) containing residue A55. All spectra were recorded at 25°C and 800 MHz. (g) Cartoon representation depicting the fact that he fast equilibrium between the open-close interaction between the Unique and SH3 domains is cancelled by the interaction of a polyproline (PP) peptide with the SH3 domain in spite of the fact that the two binding sites are separated.
Modulation of lipid binding by phosphorylation
USrc was monophosphorylated at S17 using PKA or diphosphorylated at T37 and S75 positions with GST-Cdk5 activated with GST-p25 as previously described17. Figure 5 compares the lipid-induced NMR chemical shift perturbations in phosphorylated and unphosphorylated forms of USrc. Phosphorylation at S17 almost abolished the interaction with lipids of the SH4 region, as seen by the significant decrease in lipid-induced NMR shifts but had only a moderate effect on the ULBR ( Fig. 5a ). Conversely, phosphorylation of S37/T75 caused a significant reduction in lipid binding by the ULBR with minor effects on the lipid interaction by the SH4 domain ( Fig. 5b ). The effects observed in residues distant from the phosphorylation sites suggest the two lipid binding regions show some degree of binding cooperativity. In previous work we had demonstrated that phosphorylation of S17, S37 and T75 has only local effects and does no induce structural perturbations in isolated USrc17. Therefore, we suggest that phosphorylation destabilizes electrostatically the interaction of the Unique domain of c-Src with acidic lipids.
Calmodulin modulates lipid binding by the Unique domain of c-Src
The interaction of some disordered proteins with lipids is modulated by calcium levels through binding to calmodulin22,23. The effect of adding calcium-loaded calmodulin (CaM) to USrc is shown in Figure 6 . Remarkably, chemical shift changes and/or broadening were observed for a number of residues in the Unique domain. The largest changes were observed for residues A44, S51, A55, F64 and G65, partially overlapping with the ULBR ( Fig. 6c ).
Calmodulin binding by the Unique domain of c-Src.
Two different expansions of 1H-15N HSQC NMR spectra obtained during the titration of USrc with Ca2+-calmodulin (CaM) are shown in panels a and b. (c) Combined absolute values of 1H-15N chemical shift changes induced by 2-fold excess of Ca2+-calmodulin in USrc residues.
No changes were observed when apo-calmodulin and calcium were added independently or when CaM was added to the isolated SH3 domain (results not shown). Instead, after addition of CaM to the Unique-SH3 construct, chemical shifts changes were observed for SH3 residues 97SRTETDL103 (in the RT-loop) and 133GYI135, which correspond to a subset of those perturbed by the interaction with the Unique domain ( Supplementary Fig. 3a,b and d ). The effect of adding CaM to the USH3 construct was to return the chemical shifts of these residues to values typical of the free SH3 domain. We conclude that calmodulin binds to the Unique domain of c-Src and this interaction competes with the intra-molecular interaction with the SH3 domain. However, the chemical shifts of SH3 residues 113IVNN116 in the nSrc loop, which also interact with the Unique domain, remained unaffected in presence of CaM, indicating that this region of the SH3 domain is still interacting with the Unique domain, even when it is bound to calmodulin.
The effect of CaM on lipid binding by the USH3 construct was also tested using Lipid-StripsTM ( Supplementary Fig. 3c ). The addition of a five-fold excess of CaM drastically reduced the capacity of USH3 to bind lipids. This effect is mediated by the Unique domain as CaM had no effect on the lipid interactions of the isolated SH3 domain.
In vivo effects of mutations in the Lipid Binding Region of the Unique domain
Xenopus laevis oocytes were used as model system to test the effect of mutations of the ULBR in the context of full-length c-Src. The ULBR and phosphorylation sites shown to be relevant for Unique domain interactions are highly conserved even in phylogenetically distant species ( Fig. 7 ). A charge-conserving mutant was generated by replacing residues 63–65 (LFG) by 63AAA65 (mutant AAA). A second mutant had residues 63–68 of c-Src (LFGGFN) replaced by 63AAAEAE68 (mutant EAE), where the additional mutations and the extra negative charges were expected to completely abolish lipid binding. Indeed, addition of lipids did not perturb the NMR chemical shifts of Unique domain residues of the EAE mutant and had only a small effect on Unique domain of the AAA mutant. Lipid binding by the SH4 domain was still observed in both mutants ( Supplementary Fig. 4 ).
Sequence alignment of the Unique domain of Src.
Sequence alignment of the Unique domains of human, mouse, chicken and Xenopus laevis Src is shown.Green boxes highlight residues S17, T37 and S75 that are known to be phosphorylated in vitro. Orange box shows the position of the 6 aminoacids of the ULBR that were mutated in the oocytes maturation experiment (see Fig. 8).
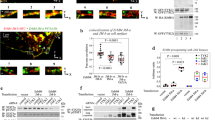
Progesterone induced maturation of Xenopus oocytes has been previously shown to be accelerated by the expression of constitutively active viral or Xenopus Src24. Oocytes were injected with in vitro transcribed mRNAs encoding constitutively active human c-Src protein, with either wild-type, AAA, or EAE Unique domains. Figure 8a shows the percentage of oocytes that completed maturation at different times after the addition of progesterone. Maturation was assessed by the appearance of a white spot at the animal pole of the oocytes, which indicates germinal vesicle breakdown (GVBD) and meiosis I entry. Oocytes injected with mRNA encoding wild-type c-Src started maturation around 2 h before control oocytes, either untreated or injected with H2O. This observation agrees with previous reports25 and confirms that Xenopus oocytes are an appropriate model to functionally characterize human c-Src. Interestingly, oocytes injected with mRNAs encoding the AAA or EAE Unique domain mutants also showed accelerated maturation in response to progesterone but only around 70%–80% completed the process ( Figure 8a , inset). In addition, 2 h after the appearance of the white spot, about half of the matured oocytes that expressed AAA or EAE mutants showed progressive depigmentation and started to die ( Fig. 8b ). These effects were not a consequence of different expression levels of the wild type and mutant forms of c-Src as all three proteins were expressed at similar levels in the injected oocytes ( Fig. 8c ).
Effect of Src mutants on Xenopus oocyte maturation.
The effect of the injection of oocytes with mRNAs encoding wild-type c-Src (green), AAA mutant (blue), EAE mutant (magenta), or water (orange) is compared with that of non-injected oocytes (yellow).(a) Percentage of oocytes that underwent maturation, as determined germinal vesicle breakdown (GVBD), at different times after the addition of progesterone. The inset shows the percentages of morphologically normal oocytes at the end of the experiment, when 100% of the control oocytes treated with progesterone reached GVBD. (b) Oocyte appearance 2 and 4 h after GVBD. About half of the matured oocytes that express AAA or EAE mutants show progressive depigmentation starting 2 h after GVBD. (c) Analysis of the expression levels of wild-type and mutants forms of human c-Src and endogenous Xenopus c-Src. Oocyte lysates were separated by SDS-PAGE and analyzed by immunoblotting with anti-human c-Src (top panel), anti-Xenopus c-Src (middle panel) and anti-MPK1 (bottom panel) as a loading control.
Discussion
The regulation of c-Src has been extensively studied. Known mechanisms implicate the kinase (SH1), SH2, SH3 and SH4 domains and the C-terminal regulatory tail. The regulatory processes involve, among others, the phosphorylation of tyrosine residues in the C-tail and kinase domain and the recognition of peptide motifs by SH2 and SH3 domains. Palmitoylation and/or myristoylation of the SH4 domain mediate localization and anchoring to membrane surfaces that also modulates the activity of other kinases of the Src family. c-Src has only a single myristoyl group. The role of the intrinsically disordered Unique domain connecting the SH4 and SH3 domains is poorly understood, although some earlier reports have suggested that it may participate in c-Src function and regulation11,12,13. In this article, we show that the Unique domain and the SH3 domains of c-Src participate in a number of intra- and intermolecular interactions suggesting the existence of a new regulation layer in c-Src and possibly in other members of the c-Src family.
We demonstrate, for the first time, that human c-Src has at least two additional lipid binding regions in addition to the SH4 domain that show specificity for acidic lipids, including phosphoinositides. One of them is located in the intrinsically disordered Unique domain and includes residues 51, 53, 55 and 60 to 67 (the Unique Lipid Binding Region - ULBR). The second one includes residues in the RT and nSrc loops of the SH3 domain.
Previous examples of lipid binding by SH3 domains are restricted to helical extended specialized SH3 domains, (hSH3), reported to bind acidic phospholipids26,27. These specialized hSH3 domains display a strongly positively charged surface and lipid binding requires an extra helical domain docked on the domain core. In contrast, lipid binding by c-Src SH3 takes place in the absence of an extended helical region. Previously, PtdIns(3,4,5)P3 binding by the c-Src SH2 domain had been reported28.
Lipid binding outside the SH4 domain suggests a more complex interplay of c-Src with the membranes to which is anchored through the myristoylated domain. Importantly, lipid binding through residues 60-75 of the flexible Unique domain would result in a significant decrease of the distance the folded core of the protein can reach away from the membrane surface and, therefore, a change in the ligands or substrates that are accessible to these domains. The possible relevance of such a “positional regulation” mechanism is stressed by the discovery of, at least, two additional regulatory mechanisms that modulate the interaction of the Unique domain with lipids. The first one is phosphorylation of T37/S75 and S17 in the Unique domain, that decreases affinity for acidic phospholipids. The same phosphorylation events have previously been associated to physiological processes. Phosphorylation of S17 is required for cAMP activation of Rap1, inhibition of extracellular signal-regulated kinases and inhibition of cell growth, although no mechanism was proposed29. T34, T46 and S72 in chicken c-Src (corresponding to T37 and S75 in human c-Src; T46 is not conserved) are phosphorylated by Cyclin dependent-kinase 1 (Cdk1) during mitosis, whereas Cdk5 is responsible for mitosis independent phosphorylation of S75 in human Y79 retinoblastoma cells30.
A second modulation mechanism of the Unique-lipid and Unique-SH3 interactions involves binding of calcium-loaded calmodulin to the Unique domain, which suppresses lipid binding by the isolated Unique domain as well as by the USH3 construct and modulates the interaction between the SH3 and Unique domains. However, apo-calmodulin showed no interactions with either domain. Therefore, interactions involving the Unique domain of c-Src may be modulated by calcium levels through calmodulin.
A similar modulation of phosphoinositides binding by CaM has been suggested for another class of myristoylated, intrinsically disordered proteins: the myristoylated alanine-rich C-kinase substrates (MARCKS) involved in binding to PtdIns(4,5)P223. In response to increased calcium levels, CaM binding induces partial detachment of MARCKS from the plasma membrane and release of PtdIns(4,5)P2 molecules sequestered by a basic lipid binding region of MARCKS, thus increasing the levels of free inositides. The protein may remain attached to the membrane through the myristoyl group. The fact that the Unique domain of c-Src is also myristoylated, disordered and capable of phosphoinositide binding, suggests that a similar mechanism may be operative in this case.
Phosphoinositide phosphates play a fundamental role in controlling membrane-cytosol interfaces and constitute signals that help define organelle identity31. PtdIns(4)P and PtdIns(3,4,5)P3 are concentrated in the plasma membrane and possibly enriched in raft-like structures. PtdIns(3,4,5)P3 is present in negligible amounts in resting cells but increases dramatically in response to growth factor stimulation and mediates cell proliferation, migration, differentiation and metabolic changes32,33. PtdIns(3,4,5)P3 recruits proteins involved in the regulation of the actin cytoskeleton and mediates the effect of growth factors on the formation of peripheral ruffles implicated in cell migration34.
In addition to interacting with lipids, the Unique and SH3 domains of c-Src interact with each other, as seen by NMR. The protein-protein interaction regions overlap with the lipid-binding regions in both domains. The SH3 region interacting with the Unique domain includes the RT and nSrc loops located opposite to the classical peptide binding site. Peptide binding outside the polyproline binding cleft had been previously observed in other SH3 domains35. Also, the interaction of non-canonical peptide motifs in intrinsically disordered regions with SH3 domains was predicted36 and has been recently observed37 and simultaneous binding of peptides at two sites of an SH3 domain has been reported38. However, we have found that binding of a poly-proline peptide to the SH3 domain of c-Src allosterically inhibits its interaction with the Unique domain. Previous NMR studies have shown that peptide binding to the c-Src SH3 domain triggered a cascade of perturbations along a chain of hydrogen bonds connecting the peptide binding region with the RT and nSrc loops located on the opposite face of the domain39. Interestingly, in the inactive basal state of c-Src the SH3 domain interacts with the polyproline linker region connecting the SH2 and kinase domains. This interaction is released when c-Src is activated40,41. Since the lipid and protein binding regions in both SH3 and Unique domains overlap, our results suggest that the activation state of c-Src may affect lipid binding by the Unique/SH3 domains and provides a connection between a “classical” regulation mechanism (polyproline recognition by the SH3 domain) and a new regulatory layer, involving lipid binding by the Unique and SH3 domains.
We have demonstrated the biological relevance of the interactions involving the ULBR in the context of full length c-Src, by comparing the effects of proteins with wild type or mutated ULBR sequences in the maturation of Xenopus laevis oocytes, used as a model system. The motif present in the ULBR formed by the sequence FGG(F/V) followed by a stretch of small polar residues (N/D/S/T) is highly conserved in c-Src of a range of species including Xenopus laevis. The ULBR motif is also present in the Unique domain of c-Fyn and c-Yes, the two kinases most similar to c-Src. Injection of mRNA encoding a constitutively active form of human c-Src caused an acceleration of progesterone-induced maturation of oocytes, which is very similar to that induced by constitutively active Xenopus c-Src or v-Src24,25. This result validates the heterologous model. However, injection of in vitro transcribed mRNAs encoding for mutants in the ULBR caused that a significant portion (20%–25%) of the oocytes failed to mature after treatment with progesterone. In addition, about half of the oocytes that matured, showed depigmentation and possible apoptotic symptoms. Thus, while wild-type and mutant forms of human c-Src are similarly active in the initiation of oocyte maturation, the failure to complete maturation and the lethal phenotype observed only in matured oocytes expressing mutated c-Src demonstrates that the ULBR plays an essential role in the regulation of c-Src.
These results constitute a proof of concept of the functional importance of the ULBR and support the idea that the observed interactions are indeed part of a new level of regulation for c-Src.
Given the large number of interactions potentially involving the Unique domain of c-Src, much work is still needed to elucidate the details of the regulation mechanisms in which it may be involved. Considering that c-Src is attached to membrane surfaces by its myristoylated SH4 domain, an intriguing possibility would be that secondary lipid binding by the Unique and SH3 domains limits the sites accessible to the kinase and SH2 domains, to those that are located closer to the maximum separation from the membrane surface allowed by the Unique domain. Thus, the effective length spanned by the flexible Unique domain may introduce a certain degree of compartimentalization in the direction perpendicular to the plane of the membrane that could prevent the interaction between c-Src and other proteins, even if they are attached next to each other on the membrane. Modulation of the lipid-ULBR interaction could provide a mechanism to change the effective length of the Unique domain that connects the folded domains of c-Src to the membrane and therefore, the specificity of c-Src with respect to upstream and downstream signaling partners. Our group is actively investigating this and alternative models of c-Src regulation involving the Unique domain. The presence of this additional regulatory layer may help to understand how c-Src effectively connects a wide variety of upstream and downstream effectors.
Methods
Cloning and protein expression
The cDNA encoding (1–85) human c-Src region with a Strep-tag in C-terminal position for purification purposes was cloned into a pET-14b vector (Novagen, UK). Human USH3 protein (SH4, Unique and SH3 domains of c-Src protein, 1–150) or SH3 c-Src domain (86–150) were expressed as His-GST fusion proteins using the pETM-30 vector (EMBL). Plasmids were transformed in Escherichia coli RosettaTM (DE3)pLysS cells (Novagen, UK) and cells were grown in M9 minimal medium supplemented with either [15N]NH4Cl, or [15N]NH4Cl and D-[U-13C]-glucose (Cambridge Isotope Laboratories, UK).
USrc protein was isolated using Strep-tactin Sepharose (IBA, Göttingen) and size exclusion chromatography (Superdex 75 26/60, GE Healthcare, Spain). His-GST proteins were purified using a Ni-NTA resin (Qiagen), cleaved from their fusion partner with TEV protease (S219V mutant with an N-terminal polyhistidine tag) and re-purified with Ni-NTA resin and size exclusion chromatography. Mutations were introduced using the QuickChange site-directed mutagenesis kit (Stratagene).
Preparation of bicelles
12.5% (w/w) bicellar dispersions were prepared by mixing long-chain (DMPC or DMPG) and short-chain (DHPC) phospholipids (Avanti Polar Lipids, Alabaster, AL) in chloroform or chloroform/methanol, evaporating the solvent and rehydrating the resulting lipid film in 50 mM phosphate buffer, pH 7.0. The mixture was subjected to five freeze-thaw cycles with pipetting and vortexing until the lipid solution was clear and transparent. The molar ratio of lipids in bicelle samples was 1.0 DHPC:0.4 DMPC:0.1 DMPG (q = 0.5, 13.3% DMPG molar ratio), 1.0 DHPC:0.4 DMPC:0.4 DMPG (q = 0.8, 22.2% DMPG molar ratio), 1.0 DHPC:0.8 DMPG (q = 0.8, 44.4% DMPG molar ratio) and 1.0 DHPC:0.8 DMPC (q = 0.8, 44.4% DMPC molar ratio). DMPC+DMPG/DHPC (q ratios) of 0.5–0.8 provide isotropic fast tumbling bicelles42,43. Concentrated protein solutions were added to the bicelles to a final concentration of 0.2 mM protein and 8% (w/v) of lipids. NMR samples contained 10% D2O and 0.05% sodium azide and were measured at 298 K. Bicelle integrity was checked by 31P NMR and transmission electron microscopy.
Protein-phospholipid assays
Lipid binding specificity was assessed with protein-lipid overlay assays using commercially available lipid strips blotted with 100 pmol of biologically relevant lipids (Echelon Biosciences), following manufacturer's instructions. Echelon Lipids StripsTM were blocked with 3% fatty acid-free BSA (A7030, Sigma) in TBS-Tween for 1 h at room temperature and then incubated with 1 μM USrc, HisGST-USH3 or HisGST-SH3 (10 μg/ml, 47 μg/ml and 34 μg/ml, respectively) in TBS-Tween for 1 h at room temperature. The membranes were washed with TBS-Tween and incubated with a 1:3,000 dilution of anti-Streptag HRP antibody (Novagen) or a 1:5,000 dilution of anti-Histag antibody (GE Healthcare) in TBS-Tween for 1 h at room temperature. After washing, the membranes probed with anti-His tag antibody were incubated with a 1:5,000 dilution of horseradish peroxidase (HRP)-coupled anti-mouse IgG (GE Healthcare) for 1 h at room temperature. After washing, protein-lipid interactions were detected by enhanced chemiluminiscence (ECLTM detection, GE Healthcare).
Xenopus laevis oocytes
Xenopus laevis ovaries were surgically removed from full-grown females and treated with collagenase and dispase. Stage VI oocytes were then selected and cultured in Barth's medium.
Wild type human c-Src cDNA or mutated variants in the Unique domain were cloned in the vector FTX6. Constitutively active forms were obtained replacing tyrosine 530 by phenylalanine. The different constructs were then used to prepare mRNAs by in vitro transcription with the mMessage mMachine kit (Ambion). Oocytes were microinjected with 50 nl of mRNAs (250 ng) and maintained overnight in modified Barth's medium at 18°C before progesterone stimulation (5 μg/μl, Sigma) to induce maturation. For the preparation of lysates, oocytes were homogenized in 10 μl per oocyte of ice-cold H1K buffer (80 mM β-glycerophosphate, pH 7.5, 20 mM EGTA, 15 mM MgCl2, 1 mM DTT, 1 mM AEBSF, 2.5 mM Benzamidine and 10 μg/ml each of Aprotinin and Leupeptin). Lysates were centrifuged at 10,000 × g for 10 min and the cleared supernatants were used for western blotting. Protocols are detailed in the review by Perdiguero and Nebreda44. The following antibodies were used for western blotting: anti-Src [clone 327] (ab16885, Abcam), anti c-Src [SRC 2] (sc-18, Santa Cruz biotechnology), anti-MPK1 (rabbit serum).
NMR Spectroscopy
NMR experiments were recorded in Bruker Avance 800 and 600 MHz spectrometers equipped with TCI cryo-probes at 0.2 mM protein concentration in 50 mM phosphate buffer pH 7.0 at 298 K or 278 K. The lower temperature allowed the observation of some NH signals from the Unique domain that exchanged rapidly. Reported chemical shift differences are from spectra obtained under identical conditions. Possible aggregation phenomena were ruled out by comparing spectra recorded between 0.1 mM and 0.4 mM ( Supplementary Fig. 5 ). USrc assignments had been previously reported17 USH3 backbone assignments were based on literature values (BMRB entries 3433 and 4888)45,46 measured at pH 6.5 and confirmed with 3D triple-resonance proton-based experiments CBCANH, CBCACONH and HNCO in a 0.6 mM 15N, 13C-double labelled USH3 sample prepared in 50 mM sodium phosphate, 100 mM NaCl buffer (pH 6.5) containing 10% D2O. Assignments were transferred to other experimental conditions by recording series of HSQC spectra at intermediate conditions.
Combined NH chemical shift differences were computed as in equation (1)

where ΔδH and ΔδN are the changes in chemical shift for 1H and 15N, respectively.
For PRE experiments the spin label (1-oxy-2,2,5,5-tetramethyl-D-pyrroline-3-methyl)-methanethiosulfonate, MTSL, (Toronto Research Chemicals) was attached as previously described to a cysteine residue introduced by mutation at position 5917. Diamagnetic samples were measured after the paramagnetic sample by reducing the paramagnetic tag with the addition of 5 mM DTT. PRE effects were determined as the ratio of 1H-15N HSQC cross peak intensity, in the paramagnetic (Iox) and diamagnetic (Ired) samples.
References
Brown, M. T. & Cooper, J. A. Regulation, substrates and functions of Src. Biochim. Biophys. Acta 1287, 121–149 (1996).
Thomas, S. M. & Brugge, J. S. Cellular functions regulated by Src family kinases. Annu. Rev. Cell Dev. Biol. 13, 513–609 (1997).
Martin, G. S. The hunting of the Src. Nature Rev. 2, 467–475 (2001).
Parsons, S. J. & Parsons, J. T. Src family kinases, key regulators of signal transduction. Oncogene 23, 7906–7909 (2004).
Yeatman, T. J. A renaissance for SRC. Nature Rev. 4, 470–480 (2004).
Pellman, D., Garber, E. A., Cross, F. R. & Hanafusa, H. Fine structural mapping of a critical NH2-terminal region of p60src. Proc. Natl. Acad. Sci. USA 82, 1623–1267 (1985).
Sigal, C. T., Zhou, W., Buser, C. A., McLaughlin, S. & Resh, M. D. Amino-terminal basic residues of Src mediate membrane binding through electrostatic interaction with acidic phospholipids. Proc. Natl. Acad. Sci. USA 91, 12253–12257 (1994).
Kim, P. W., Sun, Z. Y., Blacklow, S. C., Wagner, G. & Eck, M. J. A zinc clasp structure tethers Lck to T cell coreceptors CD4 and CD8. Science 301, 1725–1728 (2003).
Davis, A. M. & Berg, J. M. Homodimerization and Heterodimerization of Minimal Zinc(II)-Binding-Domain Peptides of T-Cell Proteins CD4, CD8α and Lck. J. Am. Chem. Soc. 131, 11492–11497 (2009).
Adachi, T., Pazdrak, K., Stafford, S. & Alam, R. The Mapping of the Lyn Kinase Binding Site of the Common ß Subunit of IL-3/Granulocyte-Macrophage Colony- Stimulating Factor/IL-5 Receptor. J. Immunol. 162, 1496–1501 (1999).
Gingrich, J. R. et al. Unique domain anchoring of Src to synaptic NMDA receptors via the mitochondrial protein NADH dehydrogenase subunit 2. Proc. Natl. Acad. Sci. USA. 101, 6237–42 (2004).
Hoey, J. G., Summy, J. & Flynn, D. C. Chimeric constructs containing the SH4/Unique domains of cYes can restrict the ability of Src(527F) to upregulate heme oxygenase-1 expression efficiently. Cell Signal. 12, 691–701 (2000).
Summy, J. M. et al. The SH4-Unique-SH3-SH2 domains dictate specificity in signaling that differentiate c-Yes from c-Src. J. Cell. Sci. 116, 2585–2598 (2003).
Levinson, A. D., Courtneidge, S. A. & Bishop, J. M. Structural and functional domains of the Rous sarcoma virus transforming protein (pp60src). Proc. Natl. Acad. Sci. USA 78, 1624–1628 (1981).
Kaplan, J. M., Varmus, H. E. & Bishop, J. M. The Src protein contains multiple domains for specific attachment to membranes. Mol. Cell Biol. 10, 1000–1009 (1990).
Kaplan, J. M., Mardon, G., Bishop, J. M. & Varmus, H. E. The first seven amino acids encoded by the v-src oncogene act as a myristoylation signal: lysine 7 is a critical determinant. Mol. Cell Biol. 8, 2435–2441 (1988).
Pérez, Y., Gairí, M., Pons, M. & Bernadó, P. Structural characterization of the natively unfolded N-terminal domain of human c-Src kinase: insights into the role of phosphorylation of the unique domain. J. Mol Biol. 391, 136–148 (2009).
Whiles, J. A., Deems, R., Vold, R. R. & Dennis, E. A. Bicelles in structure-function studies of membrane-associated proteins, Bioorg. Chem. 30, 431–42 (2002).
Andersson, A., Gräslund, A. & Mäler, L. NMR Solution Structure and Membrane Interaction of the N-Terminal Sequence (1−30) of the Bovine Prion Protein. Henrik Biverståhl. Biochemistry 43, 14940–14947 (2004).
Andersson, A., Almqvist, J., Hagn, F. & Maler, L. Diffusion and dynamics of penetratin in different membrane mimicking media. Biochim. Biophys. Acta 1661, 18–25 (2004).
Kuriyan, J. & Cowburn, D. Structures of SH2 and SH3 domains. Curr. Opin. Struct. Biol. 3, 828–837 (1993).
Arbuzova, A., Murray, D. & McLaughlin, S. MARCKS, membranes and calmodulin: kinetics of their interaction. Biochim. Biophys. Acta 1376, 369–379 (1998).
McLaughlin, S. & Murray, D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature 438, 605–611 (2005).
Unger, T. F. & Steele, R. E. Biochemical and cytological changes associated with expression of deregulated pp60src in Xenopus oocytes. Mol. Cel. Biol. 12, 5485–5498 (1992).
Tokmakov, A. et al. Regulation of Src kinase activity during Xenopus oocyte maturation. Dev. Biol. 278, 289–300 (2005).
Heuer, K., Arbuzova, A., Strauss, H., Kofler, M. & Freund, C. The helically extended SH3 domain of the T cell adaptor protein ADAP is a novel lipid interaction domain, J. Mol. Biol. 348, 1025–1035 (2005).
Heuer, K. et al. Lipid-binding hSH3 domains in immune cell adapter proteins. J. Mol. Biol. 361, 94–104 (2006).
Rameh, L. E., Chen, C. S. & Cantley, L. C. Phosphatidylinositol (3,4,5)P3 interacts with SH2 domains and modulates PI 3-kinase association with tyrosine-phosphorylated proteins. Cell 83, 821–830 (1995).
Mulgrew-Nesbitt, A. et al. The role of electrostatics in protein-membrane interactions. Biochim. Biophys. Acta 1761, 812–826 (2006).
Kato, G. & Maeda, S. Neuron-specific Cdk5 kinase is responsible for mitosis-independent phosphorylation oc c-Src at Ser75 in human Y79 retinoblastoma cells. J. Biochem. 126, 957–961 (1999).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
Cantley, L. C. The phosphoinositide 3-kinase pathway. Science 296, 1655–1657 (2002).
Czech, M. P. Dynamics of phosphoinositides in membrane retrieval and insertion. Annu. Rev. Physiol. 65, 791–815 (2003).
Oikawa, T. et al. PtIns(2,4,5)P3 binding is necessary for WAVE-2 induced formation of lamellipodia. Nature Cell Biol. 6, 420–426 (2004).
Kami, K., Takeya, R., Sumimoto, H., Kohda, D. Diverse recognition of non-PxxP peptide ligands by the SH3 domains from p67phox, Grb2 and Pex13p. EMBO J. 21, 4268–4276 (2002).
Beltrao, P. & Serrano, L. Comparative genomics and disorder prediction identify biologically relevant SH3 protein interactions. PLoS Comp. Biol. 1(3), e26 (2005).
Feuerstein, S. et al. Transient structure and SH3 interaction sites in an intrinsically disordered fragment of the Hepatitis C virus protein NS5A. J. Mol. Biol. 420, 310–323 (2012).
Barnett, P., Bottger, G., Klein, A. T. J., Tabak, H. F. & Distel, B. The peroxisomal membrane protein Pex13p shows a nobel mode of SH3 interaction. EMBO J. 19, 6382–6391 (2000).
Cordier, F., Wang, C., Grzesiek, S. & Nicholson, L. K. Ligand-induced strain in hydrogen bonds on the c-Src SH3 domain detected by NMR. J. Mol. Biol. 304, 497–505 (2000).
Xu, W., Doshi, A., Lei, M. Eck, M. J. & Harrison, S. C. Crystal structures of c-Src reveal features of its autoinhibitory mechanism Mol. Cell 3, 629–638 (1999).
Bernado, P., Perez, Y., Svergun, D. I. & Pons, M. Structural Characterization of the Active and Inactive States of Src Kinase in Solution by Small-Angle X-ray Scattering. J. Mol. Biol. 376, 492–505 (2008).
Glover, K. J. et al. Structural evaluation of phospholipid bicelles for solution-state studies of membrane-associated biomolecules. Biophys J. 81, 2163–2171 (2001).
Vold, R. R., Prosser, R. S. & Deese, A. J. Isotropic solutions of phospholipid bicelles: a new membrane mimetic for high-resolution NMR studies of polypeptides. J. Biomol. NMR 3, 329–35 (1997).
Perdiguero, E. & Nebreda, A. R. Use of Xenopus oocytes and early embryos to study MAPK signaling. Methods Mol Biol 250, 299–314 (2004).
Wang, C., Pawley, N. H. & Nicholson, L. K. The Role of Backbone Motions in Ligand Binding to the c-Src SH3 Domain. J. Mol. Biol. 313, 873–887 (2001).
Yu, H., Rosen, M. K. & Schreiber, S. L. 1H and 15N assignments and secondary structure of the Src SH3 domain. FEBS Lett. 324 1, 87–92 (1993).
Acknowledgements
The clones used for the expression of full-length c-Src constructs were derived from those kindly provided by M. Resh (Memorial Sloan Kettering Cancer Center). PB was supported by a Ramón y Cajal contract co-financed by the MICINN and IRB Barcelona. This work was supported by funds from the “Marato de TV3” on cardiovascular diseases, the Spanish MICINN-FEDER (BIO2010-15683 and BFU2010-17850), the “Generalitat de Catalunya” (2009SGR1352) and EU 7th FP BioNMR (Contract 261863). The facilities of the “ICTS Laboratorio de RMN de Barcelona” have been used in this work. A.R.N. acknowledges support by the Fundación BBVA.
Author information
Authors and Affiliations
Contributions
Y.P., A.R.N., P.B., M.P. designed experiments. Y.P., M.M., A.I., I.A., M.G. performed experiments. Y.P., M.M., A.I., I.A., A.R.N., P.B., M.P. analyzed data. M.P. wrote the paper with contributions from all coauthors.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Pérez, Y., Maffei, M., Igea, A. et al. Lipid binding by the Unique and SH3 domains of c-Src suggests a new regulatory mechanism. Sci Rep 3, 1295 (2013). https://doi.org/10.1038/srep01295
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01295
This article is cited by
-
Regulation of Src tumor activity by its N-terminal intrinsically disordered region
Oncogene (2022)
-
Src family kinases, adaptor proteins and the actin cytoskeleton in epithelial-to-mesenchymal transition
Cell Communication and Signaling (2021)
-
Full structural ensembles of intrinsically disordered proteins from unbiased molecular dynamics simulations
Communications Biology (2021)
-
Intrinsic disorder in the regulatory N-terminal domain of diacylglycerol acyltransferase 1 from Brassica napus
Scientific Reports (2018)
-
A head-to-tail view of L-selectin and its impact on neutrophil behaviour
Cell and Tissue Research (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.