Abstract
Anomalous expansion of a polymorphic tract in Ataxin-1 causes the autosomal dominant spinocerebellar ataxia type 1. In addition to polyglutamine expansion, requirements for development of pathology are phosphorylation of serine 776 in Ataxin-1 and nuclear localization of the protein. The phosphorylation state of serine 776 is also crucial for selection of the Ataxin-1 multiple partners. Here, we have used FRET for an in cell study of the interaction of Ataxin-1 with the spliceosome-associated U2AF65 and the adaptor 14-3-3 proteins. Using wild-type Ataxin-1 and Ser776 mutants to a phosphomimetic aspartate and to alanine, we show that U2AF65 binds Ataxin-1 in a Ser776 phosphorylation independent manner whereas 14-3-3 interacts with phosphorylated wild-type Ataxin-1 but not with the mutants. These results indicate that Ser776 acts as the molecular switch that discriminates between normal and aberrant function and that phosphomimetics is not a generally valid approach whose applicability should be carefully validated.
Similar content being viewed by others
Introduction
Anomalous expansion of the genetically unstable CAG codon above a threshold of 35–42 repeats in the coding region of the ATXN1 gene leads to spinocerebellar ataxia type 1 (SCA1)1. This dominant inherited neurodegenerative disease, currently incurable, is an autosomal dominant neurodegenerative pathology characterized by motor coordination deficits caused by progressive loss of Purkinje cells in the cerebellar cortex and neurons in the brain stem and spinocerebellar tracts2,3,4. As with other members of the polyQ expansion family, SCA1 is thought to be caused by aggregation and misfolding of the carrier protein promoted by the polyQ insolubility and modulated by the protein context5,6,7,8,9,10. Ataxin-1 (Atx1), the gene product of ATXN1, is a ~98 kDa mostly nuclear protein that makes of it one of the smaller components of the polyQ expansion proteins associated to disease11,12.
Despite its medical importance, the non-pathologic functional role of Atx1 in the cell and the mechanisms underlying SCA1 pathogenesis are still poorly understood. Several studies have focused on determining the Atx1 interactome and on how this could be altered in the presence of polyQ expansion. Studies addressing Atx1 association with different protein complexes that have diverse functional significance could lead to further insights into the role of the protein in the normal cellular function as well as in disease. The discovery of interactions of Atx1 with several transcriptional co-regulators such as the leucine-rich acidic nuclear protein13, polyQ-binding protein14, silencing mediator of retinoid and thyroid hormone receptors15, Capicua7, Boat8, Gfi-1/Senseless9, the nuclear hormone co-activator Tip6016 and the transcriptional co-repressor C-terminal binding protein 217 have in fact suggested a role for Atx1 in transcriptional regulation. Recently, Atx1 was also shown to interact with components of the spliceosome machinery such as RBM17 and U2AF6518,19. These studies seem to indicate that Atx1 associate with different protein complexes and that the polyQ expansion has the potential to interfere with such associations. It still remains open how the protein can discriminate among these multiple interactions at each time-point as well as in terms of its location.
A serine residue at position 776 (Ser776) of the Atx1 sequence was suggested to have a critical role in Atx1 partner selection19,20. Preliminary data seem to indicate that non-phosphorylated Atx1 binds splicing factors such as U2AF65 while Ser776-phosphorylated Atx1 binds the 14-3-3 adaptor protein19. Phosphorylated Atx1 also recognises other cellular partners such as the transcriptional repressor Capicua (CIC) and other proteins involved in transcriptional regulation, such as the MEF2-HDAC4 complex7,21. The phosphorylation state of Ser776 seems also to be important for understanding disease onset since requirements for developing the disease seem to be, in addition to polyQ expansion, nuclear localization and phosphorylation of Ser77610,20.
While the importance of Ser776 is generally unquestioned, no direct in cell studies confirm these interactions and clarify how partner selection occurs. Furthermore, some discrepancy remains between in vivo and in vitro studies. Some of the in vivo work was carried out using an aspartate mutant of Ser776 as a phosphomimetic of phosphorylated Atx122. This strategy was obviously dictated by the flexibility that the use of a genetic mutation provides in simulating a permanent phosphorylation and has increasingly been adopted23,24,25,26. However, preliminary studies using synthetic peptides spanning residues 769–777 of Atx1 have suggested that substitution of Ser776 by Asp does not efficiently recapitulate the properties of phosphorylation, shedding some doubts on the general applicability of this approach19. Other independent examples also suggest some scepticism against a generalised tout-court use of phosphomimetics27,28,29. If confirmed, this discrepancy would have important consequences in our readout of the available data and bear important consequences for our understanding of Atx1 function and its relation to disease.
To gain further insights into the effect of Ser776 phosphorylation, we have undertaken an analysis of full-length Atx1 in a cellular study. Using FRET (Förster Resonance Energy Transfer) analysis we show that U2AF65 interacts in cells with Atx1 in a phosphorylation independent manner, while 14-3-3 interacts with the wild type protein but it is unable to interact with the aspartate or alanine mutants. Our in cell study consolidates our understanding of the regulatory role of Ser776 and provides direct evidence of a role of Ser776 as a molecular switch able to discriminate between pathology and normal function of Atx1. It also provides a direct in cell evidence that, although powerful, phosphomimetics cannot be used without careful consideration.
Results
Expression of YFP-and CFP-tagged fusion proteins in COS cells
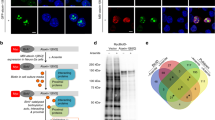
To analyse the interaction of Atx1 with the partner proteins U2AF645 and 14-3-3, we generated Atx1 expression plasmids with a YFP tag and U2AF65 and 14-3-3 with CFP tags (Figure 1). We first verified that the expression of these fusion proteins were as expected in COS cells. COS cells were individually transfected with each of these fusion-constructs. Observation of cells under fluorescent microscope showed the pattern expected for wild-type S-Atx1-YFP with several cells exhibiting nuclear Atx1 foci (Figure 2A). The pattern observed for YFP-Atx1 and mutant Atx1 expressing COS cell nuclei (Figures 2 A,B,C) is in agreement with fluorescent protein tagged wild-type Atx1 expressing cells reported earlier, with a large majority of cells forming intranuclear Atx1 foci15,30 Counter staining of the wild-type Atx1-YFP with antibody specific to Ser776 phosphorylated Atx1 also confirmed the presence of Ser776 phosphorylated Atx1 in the nuclei of these cells but not in cells transfected with either mutants (Figure S1).
Transfection of COS cells showed that CFP-U2AF65 is localized, as expected, in the nucleus whereas CFP tagged 14-3-3 (CFP-Wt-14-3-3) is mainly cytoplasmic with only around 7% of the cells showing a nuclear signal (Figures 2 D,E). This is in agreement with the literature31. Transient co-expression of CFP-Wt-14-3-3 and S-Atx1-YFP showed a similar distribution of 14-3-3 between the nuclear and cytoplasmic compartments with 2–3% of cells showing both proteins in the nucleus (data not shown). However, when D-Atx1-YFP was co-expressed with CFP-14-3-3, the already low levels of Atx1-14-3-3 co-expression in the nucleus diminished drastically. This suggests that Atx1 Ser776 phosphorylation could be important for the nuclear localisation of 14-3-3.
To study the interaction of the Atx1 mutants and 14-3-3, we generated a CFP-14-3-3 construct (CFP-14-3-3-NLS) with a nuclear localisation signal. This construct showed 14-3-3 in the nuclei of the majority of the transfected cells, as expected (Figure 2F). The expression patterns of individual fusion proteins and the co-expression patterns of Atx1 and 14-3-3 variants are summarized in the Suppl. Tables S1 and S2. Figure S2 confirms that the staining patterns observed in cells expressing fluorescent protein tagged proteins are comparable to corresponding proteins expressed without tags and stained with antibodies.
Wild-type Atx1 and its mutants interact directly with U2AF65
Interaction of Atx1 and U2AF65 was analysed by acceptor photo-bleaching FRET microscopy. We co-expressed CFP-U2AF65 with YFP-S-Atx1 to confirm that these proteins interact directly. Following photobleaching of the acceptor (YFP-Atx1) in COS cells co-expressing CFP-U2AF65 and S-Atx1-YFP, an enhancement in the donor (CFP-U2AF65) emission was observed (Figure 3C, upper panels). This de-quenching effect is indicative of an abolition of FRET due to photobleaching of the acceptor fluorophore. Since the distance between the fluorophores must be less than approximately 10 nm for FRET to occur32,33,34, we conclude that the two tagged proteins interact directly in situ. Images were acquired before (Figure 3A and B, left panels) and after (Figures 3A,B, right panels) photobleaching the acceptor (YFP-Atx1, Figure 3B). The post-bleaching images were collected immediately after and showed the dequenching effect on the donor (Figure 3C, upper panels) that results from loss of the acceptor molecule fluorescent properties (Figure 3C, lower panels). No detectable dequenching was observed in unbleached regions similar of intensity and structure to the bleached regions (compare unbleached foci to bleached foci in Figure 3). Co-expression of untagged, non-fluorescent Atx1 with S-Atx1-YFP and CFP-U2AF65 resulted in a specific, concentration dependent decrease in FRET efficiency, confirming that the direct binding of Atx1 and U2AF65 is physiologically relevant (Figure S3).
‘Rainbow’ look-up table (LUT)-encoded pseudocolour pre- and postbleach images of CFP-U2AF65 (A,D,G) and YFP-Atx1 (B,E,H: S-Atx1-YFP, A-Atx1-YFP and D-Atx1-YFP, respectively).
Magnified crops of both CFP and YFP signals in the bleach region (black circles) are depicted for pre- and post-bleach for each FRET pair (C,F,I). All scale bars are 5 μm.
We next analysed the interaction between CFP-U2AF65 and the two Atx1 mutants, A-Atx1-YFP and D-Atx1-YFP. With both mutants clear dequenching was visible in the CFP channels (Figure 3 panels D–I). Specifically, increased donor emission (CFP-U2AF65, Figures 3F and I, upper panels) was observed following photobleaching of either A-Atx1-YFP (Figure 3F, lower panels) or D-Atx1-YFP (Figure 3I, lower panels), again indicating a direct interaction between the proteins to which the fluorophores were attached. Compared to S-Atx1-YFP and A-Atx1-YFP, the FRET signal was stronger for the Asp mutant.
These results indicate that U2AF65 interacts directly with Atx1 in a Ser776 phosphorylation independent manner.
Only S-Atx1, but not its mutants, interacts directly with 14-3-3
We took a similar approach to analyse the direct in situ interaction between Atx1 and 14-3-3. However, we encountered unforeseen difficulties expressing CFP-Wt-14-3-3 together with either of the A-Atx1-YFP and D-Atx1-YFP mutants. When CFP-Wt-14-3-3 was co-expressed with the Atx1 mutants, it invariably appeared in very small quantities and was localised in the cytoplasm. Since Atx1 is a nuclear protein2 and the microscopic detection of interaction between Atx1 and 14-3-3 in transfected cells and in SCA1 affected neurones have been exclusively nuclear35,36, we decided to attach a nuclear localisation signal (NLS) to 14-3-3. This allowed us to have sufficient CFP-14-3-3 signal in the nucleus to be able to detect any potential FRET signal. Nuclear localised CFP-14-3-3-NLS exhibited a clear enhancement of donor emission upon photobleaching of the S-Atx1-YFP acceptor (Figure 4A–C), indicative of a direct interaction between the two tagged proteins. There was no increase in the CFP signal following acceptor photobleaching of A-Atx1-YFP (Figure 4D–F), thus confirming that Ser776 is important for the Atx1 interaction with 14-3-3.
‘Rainbow’ pseudocolour look-up table (LUT)-encoded pre- and postbleach images of CFP-14-3-3-NLS (A,D,G) and YFP-Atx1 (B,E,H: S-Atx1-YFP, A-Atx1- YFP and D-Atx1-YFP, respectively).
Magnified crops of both CFP and YFP signals in the bleach region (black circles) are depicted for pre- and post-bleach for each FRET pair (C,F,I). All scale bars are 5 μm.
Next, we tested if D-Atx1-YFP interacted with CFP-14-3-3-NLS. If Ser776 phosphorylation could be mimicked by an Asp then co-expression of the proteins should result in positive FRET. However, as with A-Atx1-YFP, the representative cell and nuclear foci bleach showed no increase in CFP-14-3-3-NLS donor emission upon D-Atx1-YFP acceptor photobleaching (Figures 4G–I). Assuming that Atx1 Ser776 phosphorylation is necessary for interaction with 14-3-3, these results indicate that substitution of Serine for Aspartic acid is not, in this case, a true phosphoserine functional mimic. Expression pattern and localisation of D-Atx1-YFP and A-Atx1-YFP co-expressed with CFP-14-3-3-NLS can be seen on a whole-cell basis in Figure S4.
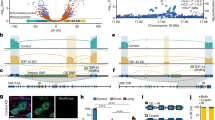
The corrected FRET efficiencies obtained with the different protein pairs and combined from different photobleaching experiments were calculated (Figure 5). For U2AF65, mean corrected FRET efficiencies of 11.6%, 10.1% and 18.2% were obtained for the interaction with S-Atx1-YFP, A-Atx1-YFP and D-Atx1-YFP, respectively, within Atx1 nuclear foci (Figure 5A). The interaction of U2AF65 with S-Atx1 and its mutants appears to be predominantly within the nuclear foci (compare f = foci vs n = non-foci, Figure 5A). An exception to this is the apparent non-foci interaction between CFP-U2AF65 and D-Atx1-YFP, which has a FRET efficiency of 10.1% ± 1.3, which is comparable to that observed for A-Atx1 within the foci. FRET efficiency is also higher for D-Atx1-YFP within the nuclear foci where a constitutively active binding is expected.
Box and whisker plots depicting population distribution of percentage corrected FRET and showing maximum, minimum, upper & lower quartiles and sample median.
Bleach regions tested were inside nuclear foci (foci, f) and outside nuclear foci (non-foci, n). (A) CFP-U2AF65 Vs YFP-Ataxin-1 (wild-type, Alanine and Aspartic acid substitution mutants). B) CFP-Wt-14-3-3 (orange) and CFP-14-3-3- NLS (blue) Vs YFP-Ataxin-1 (wild-type, Alanine and Aspartic acid substitution mutants). Means +/− standard errors, rounded to one decimal place, are shown above each boxplot. Statistical significance bars are shown and represent results of unpaired t-tests of mean difference = 0 (* = P<0.05,** = P<0.01, *** = P<0.001). N numbers (left to right) are: (A) N = 16, N = 10, N = 24, N = 22, N = 21, N = 21; (B) N = 15, N = 12, N = 22, N = 26, N = 22, N = 14, N = 19, N = 15; and represent number of individual bleach events pooled from at least 4 individual cells.
For the interaction between CFP-Wt-14-3-3 and S-Atx1-YFP, mean FRET efficiencies of 25.7% and 6.9% were observed for inside and outside Atx1 nuclear foci, respectively (Figure 5B). Whilst we observed a reduced FRET efficiency when pairing CFP-14-3-3-NLS with S-Atx1-YFP, both pairs show a strong FRET efficiency within nuclear foci but low FRET efficiency outside, indicating that the NLS-tagged fusion still retains the physiological function of its wild-type counterpart. We thus conclude that CFP-14-3-3-NLS behaves sufficiently akin to the wild-type to give reliable interaction data. Compared to S-Atx, a very low FRET efficiency of 1.4% (foci) or lower (non-foci) was observed for CFP-14-3-3-NLS vs D-Atx1-YFP (Figure 5B). Therefore, our data indicate that Atx1 Ser776 is critical for efficient binding to 14-3-3 and n.14-3-3 and that substitution of Ser with an Asp does not, in this case, effectively mimic phosphorylation.
Discussion
Atx1 is a human protein of medical interest because it is associated with the rare but dominant SCA1 pathology4,37,38. An important step for understanding the mechanisms that trigger SCA1 is that of determining the physiologic function of Atx1 through studying its interactome. Using this approach, increasing evidence suggests a role of Atx1 as a transcription regulator able to interact with multiple partners6,7,14,19,21,39,40,41,42. A crucial importance in complex formation and in partner selection has been demonstrated for Ser7767,18,19. Phosphorylation of this residue also seems to discriminate between normal and pathological function since, together with polyQ expansion and nuclear localization, Ser776 phosphorylation is necessary for development of SCA1 in animal models20. It is therefore essential to establish beyond any doubt whether phosphomimetics of Ser776 could be used as a valid research tool in Atx1 studies and how far we can rely on this approach for dissecting the physiologic function. Different in vitro studies, mostly using synthetic peptides, have in fact already shed some doubts about the general validity of this concept27,28,29,43,44,45. Our previous in vitro data have also suggested opposite binding behaviour for phosphorylated versus Asp mutated Atx1 synthetic peptides19.
As a read-out for direct protein interaction in situ, we have undertaken FRET analysis of Atx1 and its binding partners. This method provides a more accurate picture than co-localization or co-IP approaches that, although useful for detecting the association of proteins, do not lead to the conclusive confirmation of a one-to-one interaction. Our FRET data confirm that there is no requirement of Ser776 phosphorylation for U2AF65 recognition. Wild-type Atx1 as well as the Asp and Ala mutants bind U2AF65, indicating that Asp substitution does not prevent U2AF65 from binding to Atx1 and even increases FRET efficiency. A possible explanation for the behaviour of D-Atx1-YFP is that this mutant could have higher affinity for U2AF65 and is thus able to interact also outside the natural protein complex. Our in vitro data using Atx1 peptides do not, however, support this hypothesis19: the affinity of the complex of U2AF65 with a peptide spanning Atx1 residues 769–779 with an aspartate in position 776 is fully comparable with those of peptides with either serine or alanine. Alternatively, a third binding partner with strong affinity could bring U2AF65 and Atx1 into a conformation that allows direct interaction.
A very different behaviour was observed for 14-3-3: we could detect direct interaction when wild-type Atx1 and 14-3-3 were co-expressed in a form that led to their co-localization in the nucleus. Even though predominantly a cytoplasmic protein with no obvious NLS signal, various studies have shown the presence of 14-3-3 in the nucleus, as well as the co-localisation of Atx1 and 14-3-3 in the nucleus35,36. Since it is not possible to express constitutively phosphorylated proteins, Atx1 is expected to be present in the cell as a mixed population of unphosphorylated as well as phosphorylated forms (as shown in Figure S1). We observed a FRET signal between wild-type Atx1 and 14-3-3 indicating a direct interaction of the two proteins in the nucleus. Conversely, we could not detect an interaction with either the alanine or the aspartate mutants even though the proteins still co-localise in the nucleus. These observations are in excellent agreement with our previous in vitro data19 and confirm the absolute requirement for Ser776 phosphorylation for 14-3-3 recognition. As a corollary, they also indicate that the aspartate mutant cannot, in this case, mimic a phosphoserine.
Glutamate and aspartate have been widely used as phosphomimetics to substitute phosphotyrosines and phosphoserines23,24,25,26, The substitution is particularly useful because it allows a flexible way to introduce mutations in animal or cellular models to test the effect of phosphorylation. This allows for insight into the effect of the phosphorylation in cells as well as in animal models and hence the approach has far reaching possibilities in translational medicine including in the design of therapeutic strategies. In many instances, the phosphorylation effect may be mostly electrostatic in which case an aspartic, for instance, can be expected to be sufficiently close to be an effective substitute of phosphorylation. However, this may not necessarily hold true for all the interactions: in at least some instances, positional effects and conformational constraints may come into play in addition to or in spite of the charge effect. Previous crystallographic studies may explain the lack of binding of the aspartate mutant to 14-3-346,47. A model built on the crystal structure of the complex of 14-3-3- with a peptide (accession name 1qjb) shows that the phospho-group faces directly three charged residues (K49, R56 and R127) from a helical bundle of 14-3-3 with which it can hydrogen bond (Figure 6). The phospho-group is also close to the hydroxyl group of the aromatic ring of Y128. While still retaining the negative charge present in a phosphoserine, the aspartate substitution has several very basic flaws: The aspartate side chain is too short to establish the same interactions with the environment since any overall movement of the hairpin in 14-3-3 would introduce main distortions in the structure. This implies that some if not all of the hydrogen bonds formed by the phospho-protein will not be established or will be weaker in the mutant (see a comparison of the distances in Supplementary Table S3). The aspartate has only two of the three oxygens in the phosphoserine, resulting in a completely different geometry. Even more importantly, the charge distribution and density will certainly be very different in the two negatively charged groups. Finally, like most physiological 14-3-3 binding proteins, Atx1 contains a proline residue at a position two residues C-terminal to the phosphorylated serine. A sharp alteration of the peptide chain direction at this position could change the conformational landscape of the chain and affect the kink exerted by the proline residue C-terminal to the phosphoserine. These considerations will need to be kept in mind in further usage of phosphomimics.
Structural determinants of the interaction between 14-3-3 andphosphopeptides.
A model of the complex of Atx1 with 14-3-3 was built using the accession name 1qjb as a template by replacing the Atx1 residues using phosphorylated serine (left) or an aspartate (right). The backbone of 14-3-3 is shown as green ribbons. The side chains of residues that surround the peptide are explicitly shown in yellow. The peptide is shown in green. The position of the phospho-serine or aspartate is indicated with a red circle. The substitution leads to a different geometry and to longer distances between the charged groups.
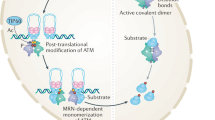
An even more important take-home message from our work concerns the consequences of our results for understanding both the normal function of Atx1 and its role in SCA1 pathology. Both U2AF65 and 14-3-3 have been shown to bind the same short region of Atx1, termed the Atx1 ULM19. This competition calls for a mechanism of regulation to determine which protein may bind Atx1 at a given time. Our results clearly indicate that phosphorylation acts as the molecular switch that causes Atx1 to interact with a substantially different network of nuclear proteins than non-phosphorylated Atx1, thus determining its localization and function. Nonphosphorylated expanded Atx1, which does not manifest a SCA1 phenotype20, recognizes UHM-containing nuclear factors but is unable to bind 14-3-3. Participation of Atx1 in the interaction with the large spliceosome machinery could play a protective role against self-association of the protein induced by polyQ expansion thus preventing aggregation. Conversely, when Atx1 is Ser776-phosphorylated and recruited by the smaller 14-3-3, the interaction with the spliceosome complex is hampered making expanded Atx1 susceptible to self association through regions such as the polyQ stretches and/or the AXH domain6. This hypothesis proposes a scenario in which normal function and aberrant aggregation are competing pathways. A similar concept was already proposed for another polyQ protein, ataxin-3 whose aggregation and misfolding is mediated by the same exposed surfaces that are involved in the recognition of the physiologic partners48. Such a competition could be a general mechanism able to explain the causes of several if not all misfolding diseases. Further studies are needed to test the general validity of the model proposed.
Methods
Plasmid constructs
Yellow Fluorescent protein (YFP)-tagged Wild-type Atx1 (30Q) was generated by PCR amplification and cloning into pEYFP-N1 vector (Clontech). This construct was used as a template to obtain Ser776 to Ala or Asp mutated YFP-Atx1 constructs. Serine to Ala mutation was achieved by PCR using the primers 5′-CGAGGAAGAGGAGGTGGGCCGCGCCAGAGAGCCGC-3′ and 5′-GCGGCTCTCTGGCGCGGCCCACCTCCTCTTCCTCG-3′. Serine to Aspartic acid mutation was generated by using primers 5′-CGAGGAAGAGGAGGTGGGACGCGCCAGAGAGCCGC-3′ and 5′-GCGGCTCTCTGGCGCGTCCCACCTCCTCTTCCTCG-3′. CFP tagged U2AF65 and 14-3-3 were obtained by PCR amplification and cloning into the pECFP-C1 vector (Clontech). Green Fluorescent protein (GFP)-U2AF65 (a gift from Juan Valcarcel, Barcelona) was used as a template for U2AF65 and GST-14-3-3ζ (provided by Katrin Rittinger, London), for 14-3-3. SV40 Nuclear localisation signal (NLS) was amplified from pEF/nuc vector (Invitrogen) and ligated in frame to the 3′ end of the Cyan Fluorescent Protein (CFP)-14-3-3 construct to generate 14-3-3 fusion protein targeted to the nucleus (CFP-14-3-3-NLS). The following primers were used: 5′-TGCAGGTCGACCTCGAG ATCAAA CGG-3′ and 5′-ATATAGGATCCTTTTTGTTCTGCGGCCGTATCTAC-3′.
Cell culture and transfections
COS cells were cultured as monolayers in Dulbecco's modified Eagle medium supplemented with 10% fetal bovine serum and 100 U/mL penicillin-streptomycin (Invitrogen Life Technologies). For the expression and microscopic analysis of fluorescent fusion proteins, cells were grown on chamber slides and transfected with appropriate plasmid DNA using GeneCellin tranfection reagent (BioCellChallenge) according to the manufacturer's instructions. Cells were fixed using 4% paraformaldehyde 36–48 h post-transfection and slides were mounted using CitiFluor (Agar Scientific).
FRET microscopy
To detect FRET by the acceptor photobleaching method32,33, cells transfected with appropriate expression constructs were fixed with 4% paraformaldehyde and mounted as described. Samples for FRET were imaged on a Zeiss LSM 510 confocal microscope using a 63× 1.4NA Plan NeoFluar oil immersion objective. CFP was excited with the 458 nm line of an Argon-Krypton ion laser and YFP was excited using the 514 nm line. The Argon laser was set to a tube current of 7.3 A. A 545 nm dichroic mirror was used to split the two emission channels, followed by a band-pass 475–525 nm filter for the CFP channel and a long pass 530 nm filter for the YFP channel. In each case the pinhole was set such that the optical slice was indicated as 5 μm by the LSM 510 AIM software. Each image line was scanned four times at scan speed 8 using line-by-line unidirectional multi-tracking mode and digitised at 70 nm/pixel by using a scan zoom of 4×. Separate photomultiplier tube detectors were used for each channel and images were digitised to 12-bit. The coverslip was tilescanned at 10x and cells showing strong signals in both CFP and YFP channels were marked and their XY positions stored for viewing at 63x. Preliminary scans of cells were cropped with a 4x zoom then a rectangular acquisition Region of Interest (ROI) was drawn around the nuclear and perinuclear region. Focus was positioned where the maximum number of nuclear foci were clearly visible. For nuclear-foci bleaches, the gain for the CFP channel was set to approximately 75% of dynamic range (12-bit, 4096 grey levels), whilst the YFP channel was set to the nearest greyvalue to maximum without actually saturating any pixels. For the non-foci bleaches, the gain for the CFP channel was again set to approximately 75% of the dynamic range whilst that of the YFP channel was set such that the non-foci regions had a sufficient dynamic range for mathematical operations; typically between 70–99% of the dynamic range. Since the majority of material tended to be found in the nuclear foci, this resulted in saturated pixels within the dense nuclear foci regions. Any saturated pixels were automatically excluded from FRET calculations and so did not adversely affect the FRET results. For all channels and all bleach experiments, offsets were set such that backgrounds were at least 1 grey level in value. The timelapse mode was used to collect one prebleach image for each channel before bleaching. YFP was photobleached using the 514 nm line of the 30 mW Argon-Krypton laser set to a tube current of 8.1 A. A transmission of 100% was scanned across the bleach ROI at scan speed 9 using this wavelength and power and the bleach process was repeated for 200 iterations. A second post-bleach image was then collected for each channel. Bleach regions were defined by the size of nuclear foci in each particular cell and then a similar-sized bleach ROI was used for all non-foci bleaches on that same cell. Pre- and post-bleach CFP and YFP images were imported into Mathematica 7.0 for processing as previously described49. Briefly, images were smoothed using a 3x3 box mean filter, background subtracted and post-bleach images fade compensated.

FRET efficiencies were then extracted from pixels falling inside the bleach region and plotted against the bleach efficiency on a pixel-by-pixel basis. FRET efficiency showed a linear correlation with bleach efficiency enabling determination of FRET efficiency at 100% bleach efficiency by extrapolation. Data from images were used only if YFP bleaching efficiency was greater than 50%. Finally, the FRET efficiency was converted in to the inter-fluorophore radius using:

where Ro is the Foerster radius for CFP and YFP, which is 4.95 nm.
References
Orr, H. T., Chung, M. Y., Banfi, S., Kwiatkowski, T. J. Jr., Servadio, A., Beaudet, A. L., McCall, A. E., Duvick, L. A., Ranum, L. P. & Zoghbi, H. Y. Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nat. Genet. 4, 221–226 (1993).
Zoghbi, H. Y. & Orr, H. T. Spinocerebellar ataxia type 1. Semin. Cell Biol. 6, 29–35 (1995).
Cummings, C. J., Orr, H. T. & Zoghbi, H. Y. Progress in pathogenesis studies of spinocerebellar ataxia type 1. Philos. Trans. R Soc. Lond. B Biol. Sci. 354, 1079–1081 (1999).
Matilla-Dueñas, A., Goold, R. & Giunti, P. Clinical, genetic, molecular and pathophysiological insights into spinocerebellar ataxia type 1. Cerebellum 7, 106–114 (2008).
Cummings, C. J., Mancini, M. A., Antalffy, B., DeFranco, D. B., Orr, H. T. & Zoghbi, H. Y. Chaperone suppression of aggregation and altered subcellular proteasome localization imply protein misfolding in SCA1. Nat. Genet. 19, 148–154 (1998).
de Chiara, C., Menon, R. P., Dal Piaz, F., Calder, L. & Pastore, A. Polyglutamine is not all: the functional role of the AXH domain in the ataxin-1 protein. J. Mol. Biol. 354, 883–893 (2005).
Lam, Y. C., Bowman, A. B., Jafar-Nejad, P., Lim, J., Richman, R., Fryer, J. D., Hyun, E. D., Duvick, L. A., Orr, H. T., Botas, J. & Zoghbi, H. Y. ATAXIN-1 interacts with the repressor Capicua in its native complex to cause SCA1 neuropathology. Cell 127, 1335–1347 (2006).
Mizutani, A., Wang, L., Rajan, H., Vig, P. J., Alaynick, W. A., Thaler, J. P. & Tsai, C. C. Boat, an AXH domain protein, suppresses the cytotoxicity of mutant ataxin-1. EMBO J. 24, 3339–3351 (2005).
Tsuda, H., Jafar-Nejad, H., Patel, A. J., Sun, Y., Chen, H. K., Rose, M. F., Venken, K. J., Botas, J., Orr, H. T., Bellen, H. J. & Zoghbi, H. Y. The AXH domain of Ataxin-1 mediates neurodegeneration through its interaction with Gfi-1/Senseless proteins. Cell 122, 633–644 (2005).
Klement, I. A., Skinner, P. J., Kaytor, M. D., Yi, H., Hersch, S. M., Clark, H. B., Zoghbi, H. Y. & Orr, H. T. Ataxin-1 nuclear localization and aggregation: role in polyglutamine-induced disease in SCA1 transgenic mice. Cell 95, 41–53 (1998).
Servadio, A., Koshy, B., Armstrong, D., Antalffy, B., Orr, H. T. & Zoghbi, H. Y. Expression analysis of ataxin 1 protein in tissues from normal and sca 1 affected individuals. Nat. Genet. 10, 94–98 (1995).
Skinner, P. J., Koshy, B. T., Cummings, C. J., Klement, I. A., Helin, K., Servadio, A., Zoghbi, H. Y. & Orr, H. T. Ataxin-1 with an expanded glutamine tract alters nuclear matrix-associated structures. Nature 389, 971–974 (1997).
Matilla, A., Koshy, B. T., Cummings, C. J., Isobe, T., Orr, H. T. & Zoghbi, H. Y. The cerebellar leucine-rich acidic nuclear protein interacts with ataxin-1. Nature 389, 974–978 (1997).
Okazawa, H., Rich, T., Chang, A., Lin, X., Waragai, M., Kajikawa, M., Enokido, Y., Komuro, A., Kato, S., Shibata, M., Hatanaka, H., Mouradian, M. M., Sudol, M. & Kanazawa, I. Interaction between mutant ataxin-1 and PQBP-1 affects transcription and cell death. Neuron 34, 701–713 (2002).
Tsai, C. C., Kao, H. Y., Mitzutani, A., Banayo, E., Rajan, H., McKeown, M. & Evans, R. M. Ataxin1, a SCA1 neurodegenerative disorder protein, is functionally linked to the silencing mediator of retinoid and thyroid hormone receptors. Proc. Natl. Acad. Sci. USA 10, 4047–4052 (2004).
Serra, H. G., Duvick, L., Zu, T., Carlson, K., Stevens, S., Jorgensen, N., Lysholm, A., Burright, E., Zoghbi, H. Y., Clark, H. B., Andresen, J. M. & Orr, H. T. RORalpha-mediated Purkinje cell development determines disease severity in adult SCA1 mice. Cell 127, 697–708 (2006).
Lee, S., Hong, S., Kim, S. & Kang, S. Ataxin-1 occupies the promoter region of E-cadherin in vivo and activates CtBP2-repressed promoter. Biochim Biophys Acta 1813, 713–722 (2011).
Lim, J., Jafar-Nejad, P. & Richman, R. Opposing effects of polyglutamine expansion on native protein complexes contribute to SCA1. Nature 452, 713–718 (2008).
de Chiara, C., Menon, R. P., Strom, M., Gibson, T. J. & Pastore, A. Phosphorylation of S776 and 14-3-3 binding modulate ataxin-1 interaction with splicing factors. PLoS One 4, e8372 (2009).
Emamian, E. S., Duvick, L. A. & Tousey, S. K. Serine 776 of ataxin-1 is critical for polyglutamine-induced disease in SCA1 transgenic mice. Neuron 38, 375–387 (2003).
Bolger, T. A., Zhao, X., Cohen, T. J., Tsai, C. C. & Yao, T. P. The neurodegenerative disease protein ataxin-1 antagonizes the neuronal survival function of myocyte enhancer factor-2. J. Biol. Chem. 282, 29186–29192 (2007).
Duvick, L., Barnes, J., Ebner, B., Agrawal, S., Andresen, M., Lim, J., Giesler, G. J., Zoghbi, H. Y. & Orr, H. T. SCA1-like disease in mice expressing wild-type ataxin-1 with a serine to aspartic acid replacement at residue 776. Neuron 67, 929–935 (2010).
Maciejewski, P. M., Peterson, F. C., Anderson, P. J. & Brooks, C. L. Mutation of serine 90 to glutamic acid mimics phosphorylation of bovine prolactin. J Biol Chem. 270, 27661–27665 (1995).
Madeo, F., Schlauer, J., Zischka, H., Mecke, D. & Fröhlich, K. U. Tyrosine phosphorylation regulates cell cycle-dependent nuclear localization of Cdc48p. Mol Biol Cell. 9, 131–141 (1998).
Li, Y. M., Wen, Y., Zhou, B. P., Kuo, H. P., Ding, Q. & Hung, M. C. Enhancement of Bik antitumor effect by Bik mutants. Cancer Res. 63, 7630–7633 (2003).
Kassenbrock, C. K. & Anderson, S. M. Regulation of ubiquitin protein ligase activity in c-Cbl by phosphorylation-induced conformational change and constitutiveactivation by tyrosine to glutamate point mutations. J Biol Chem. 279, 28017–28027 (2004).
Capasso, H., Palermo, C., Wan, S., Rao, H., John, U. P., O'Connell, M. J. & Walworth, N. C. Phosphorylation activates Chk1 and is required for checkpoint-mediated cell cycle arrest. J Cell Sci. 115, 4555–4564 (2002).
Paleologou, K. E., Schmid, A. W., Rospigliosi, C. C., Kim, H. Y., Lamberto, G. R., Fredenburg, R. A., Lansbury, P. T. Jr., Fernandez, C. O., Eliezer, D., Zweckstetter, M. & Lashuel, H. A. Phosphorylation at Ser-129 but Not the Phosphomimics S129E/D Inhibits the Fibrillation of α-Synuclein. J. Biol. Chem. 283, 16895–16905 (2008).
Anthis, N. J., Haling, J. R., Oxley, C. L., Memo, M., Wegener, K. L., Lim, C. J., Ginsberg, M. H. & Campbell, I. D. Beta integrin tyrosine phosphorylation is a conserved mechanism for regulating talin-induced integrin activation. J. Biol. Chem. 284, 36700–36710 (2009).
Parfitt, D. A., Michael, G. J., Vermeulen, E. G., Prodromou, N. V., Webb, T. R., Gallo, J. M., Cheetham, M. E., Nicoll, W. S., Blatch, G. L. & Chapple, J. P. The ataxia protein sacsin is a functional co-chaperone that protects against polyglutamine-expanded ataxin-1. Hum. Mol. Genet. 18, 1556–1565 (2009).
Mhawech, P. 14-3-3 proteins-an update. Cell Research 15, 228–236 (2005).
Trinkle-Mulcahy, L., Chusainow, J., Lam, Y. W., Swift, S. & Lamond, A. Visualization of intracellular PP1 targeting through transiently and stably expressed fluorescent protein fusions. Methods Mol. Biol. 365, 133–154 (2007).
Hayward, R. D., Goguen, J. D. & Leong, J. M. No better time to FRET: shedding light on host pathogen interactions. Journal of Biology 9,12 (2010)
Llères, D., James, J., Swift, S., Norman, D. G. & Lamond, A. I. Quantitative analysis of chromatin compaction in living cells using FLIM–FRET. J. Cell. Biol. 187, 481–496 (2009).
Chen, H. K., Fernandez-Funez, P., Acevedo, S. F., Lam, Y. C., Kaytor, M. D., Fernandez, M. H., Aitken, A., Skoulakis, E. M., Orr, H. T., Botas, J. & Zoghbi, H. Y. Interaction of Akt-Phosphorylated Ataxin-1 with 14-3-3 Mediates Neurodegeneration in Spinocerebellar Ataxia Type 1. Cell 113, 457–468 (2003).
Umahara, T. & Uchihara, T. 14-3-3 Proteins and Spinocerebellar Ataxia Type 1: from Molecular Interaction to Human Neuropathology. Cerebellum 9, 183–189 (2010).
Zoghbi, H. Y. & Orr, H. T. Pathogenic mechanisms of a polyglutamine-mediated neurodegenerative disease, spinocerebellar ataxia type 1. J. Biol. Chem. 284, 7425–7429 (2009).
Donato, S. D., Mariotti, C. & Taroni, F. Spinocerebellar ataxia type 1. Handb. Clin. Neurol. 103, 399–421 (2012).
Goold, R., Hubank, M., Hunt, A., Holton, J., Menon, R. P. & Revesz, T. Down-regulation of the dopamine receptor D2 in mice lacking ataxin 1. Hum. Mol. Gen. 16, 2122–2134 (2007).
Crespo-Barreto, J., Fryer, J. D., Shaw, C. A., Orr, H. T. & Zoghbi, H. Partial loss of ataxin-1 function contributes to transcriptional dysregulation in spinocerebellar ataxia type 1 pathogenesis. PLoS Genet. 6, e1001021 (2010).
Tong, X., Jin, F. & Lin, P. Ataxin-1 and Brother of ataxin-1 are components of the Notch signalling pathway. EMBO Rep. 12, 428–435 (2011).
Cvetanovic, M., Patel, J. M., Marti, H. H., Kini, A. R. & Opal, P. Vascular endothelial growth factor ameliorates the ataxic phenotype in a mouse model of spinocerebellar ataxia type 1. Nat. Med. 17, 1445–1447 (2011).
McDowell, J. H., Robinson, P. R., Miller, R. L., Brannock, M. T., Arendt, A., Smith, W. C. & Hargrave, P. A. Activation of Arrestin: Requirement of Phosphorylation as the Negative Charge on Residues in SyntheticPeptides from the Carboxyl-Terminal Region of Rhodopsin. Invest. Ophthalmol. Vis. Sci. 42, 1439–1443 (2001).
Chong, H., Lee, J. & Guan, K. L. Positive and negative regulation of Raf kinase activity and function by phosphorylation. EMBO J 20, 3716–3727 (2001).
Tanaka, S., Tak, Y. S. & Araki, H. The role of CDK in the initiation step of DNA replication in eukaryotes. Cell. Div. 2, 16 (2007).
Yaffe, M. B., Rittinger, K., Volinia, S., Caron, P. R., Aitken, A., Leffers, H., Gamblin, S. J., Smerdon, S. J. & Catley, L. C. The structural basis for 14-3-3:phosphopeptide binding specificity. Cell 91, 961–971 (1997).
Rittinger, K., Budman, J., Xu, J., Volinia, S., Cantley, L. C., Smerdon, S. J., Gamblin, S. J. & Yaffe, M. B. Structural analysis of 14-3-3 phosphopeptide complexes identifies a dual role for the nuclear export signal of 14-3-3 in ligand binding. Mol. Cell 4, 153–166 (1999).
Masino, L., Nicastro, G., Calder, L., Vendruscolo, M. & Pastore, A. Functional interactions as a survival strategy against abnormal aggregation. FASEB J. 25, 45–54 (2011).
Matthews, H. K., Marchant, L., Carmona-Fontaine, C., Kuriyama, S., Larraín, J., Holt, M. R., Parsons, M. & Mayor, R. Directional migration of neural crest cells in vivo is regulated by Syndecan-4/Rac1 and non-canonical Wnt signaling/RhoA. Development 135, 1771–1780 (2008).
Acknowledgements
We gladly acknowledge financial support from an International JOINT PROJECT of the Royal Society (NIMR: U.1175.03.002.00001.01), from Ataxia UK and from the British Heart Foundation. Ataxin 1 antibodies were obtained from UC/Davis/NINDS/NIMH NeuroMab Facility, maintained by the Dept of Pharmacology, School of Medicine, University of California, Davis, CA 95616
Author information
Authors and Affiliations
Contributions
RM and DS have performed the experiments, MH and NA provided technical support for FRET experiments, CD carried out structural analysis and RM and AP written the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
SupplMaterial 8 figures and 3 Tables
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Menon, R., Soong, D., de Chiara, C. et al. The importance of serine 776 in Ataxin-1 partner selection: A FRET Analysis. Sci Rep 2, 919 (2012). https://doi.org/10.1038/srep00919
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00919
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.