Abstract
Conditions within the endoplasmic reticulum (ER) influence most secretory proteins that pass through the ER. Therefore, eukaryotic cells must strike a balance between the ER stress response, which changes the conditions in the ER and other considerations associated with protein secretion. Here, an interaction between the ER stress and defence responses in rice is described. Expression of OsWRKY45, which encodes a transcription factor that plays a central role in defence mediated by salicylic acid (SA), is induced by ER stress. Additionally, expression of some genes encoding pathogenesis-related (PR) secretory proteins is reduced by the ER stress response mediated by the stress sensor IRE1. Concomitant activation of the SA and ER stress responses suppresses the induction of ER stress-responsive genes, with the exception of OsWRKY45 and the reduction of PR gene expression. These findings demonstrate a functional integration between the defence and ER stress responses in plants.
Similar content being viewed by others
Introduction
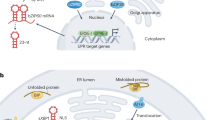
The ER stress response is triggered by the accumulation of unfolded proteins in the ER lumen. ER stress activates intracellular signal transduction pathways, such as the unfolded protein response, that contribute to the relief of the stress1. In plants, the signal transduction pathways involved in the ER stress response have been identified2. However, the relationships between the ER stress response and other plant cellular responses remain poorly understood. In particular, because plants are simultaneously subjected to biotic and abiotic stresses in their natural environments, understanding interference among multiple stress responses represents a major challenge in plant science.
In many eukaryotes including plants, ER stress is detected by the transmembrane protein IRE1. The N-terminal portion of IRE1 resides in the ER lumen; the C-terminal portion resides in the cytosol and contains a serine/threonine kinase domain and an endonuclease domain1,3,4. When activated by ER stress, IRE1 mediates an unconventional splicing of the mRNA encoding a key bZIP transcription factor, HAC1 (in yeast), XBP1 (animals), AtbZIP60 (Arabidopsis) or OsbZIP50 (rice); the spliced forms of these mRNAs are translated as active forms5,6,7,8,9. In animals, ER stress is also sensed by ATF6, a type II transmembrane protein activated by proteolysis by the site 1 and 2 proteases. Cleavage of ATF6 liberates its cytosolic domain, which contains a bZIP transcription factor that, like XBP1/AtZIP60/OsZIP50, activates transcription of ER stress response genes10,11. In plants, AtbZIP17 and AtbZIP28 (Arabidopsis) and OsbZIP39 and OsbZIP60 (rice) are the counterparts of ATF69,12,13,14,15. In the case of rice, the activated forms of OsbZIP39, OsbZIP60 and OsbZIP50 induce the expression of genes that encode ER quality control (ERQC)-related factors such as the ER chaperone BiP9,15.
In the life cycle of higher plants, massive amounts of seed storage proteins are produced during seed maturation according to a developmental program, resulting in induction of the ER stress response in response to the large load of secretory protein16,17. Defects in secretion of storage proteins are associated with expression of ERQC-related genes in maturing seeds. Furthermore, when plants are attacked by pathogens, multiple types of proteins including pathogenesis-related (PR) proteins are secreted through the ER as part of the defence response. Consistent with this, defects in ERQC disturb the expression of secretory proteins required for defence responses18,19. Furthermore, in Arabidopsis, some ER chaperones transiently induced by transcription factor NPR1, which plays a central role in the SA response, promote efficient secretion of PR proteins20. These findings suggest that ER function and defence in plants are interrelated.
The mechanisms of the defence response differ between rice and Arabidopsis. For example, the level of SA, a phytohormone that plays a central role in the defence response, is several orders of magnitude higher in rice than in Arabidopsis21. Additionally, the SA response in rice involves both an ortholog of Arabidopsis NPR1 as well as an SA-regulated transcription factor, OsWRKY45, which is absent from Arabidopsis22. Although overexpression of the OsWRKY45 gene alone is not sufficient to activate the defence response fully, the resulting accumulation of OsWRKY45 contributes to efficient activation of the defence response genes when the plant is attacked by pathogens22. Therefore, the biological activities of OsWRKY45 are predicted to enhance disease resistance in rice22. Understanding the specific mechanisms of the rice defence response is an important challenge in agriculture.
In this study, in order to understand the relationship between the ER stress response and other cellular responses, we measured the expression patterns of many ER stress–responsive genes that are closely associated with other responses. From data obtained by DNA microarray analysis of ER-stressed rice plants, we found that expression of OsWRKY45 is also induced by ER stress. Additionally, we investigated the interactions between the ER stress and SA responses. We close by discussing the implications of the ER stress–responsive induction of OsWRKY45.
Results
Expression of OsWRKY45 induced by ER stress response in an OsbZIP50-dependent manner
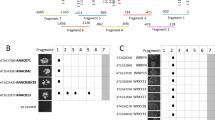
Our first DNA microarray screen for ER stress-responsive genes revealed that the expression of the SA-responsive gene OsWRKY45 is also markedly induced by ER stress. Quantitative reverse transcription-PCR (RT-PCR) analysis confirmed that expression of OsWRKY45 was induced by tunicamycin (Tm) treatment, which induces ER stress by inhibiting N-linked glycosylation. The induction by Tm was suppressed by 4-phenylbutyric acid (PBA), which acts as a chemical chaperone by masking unfolded proteins (Figure 1a). Additionally, the induction of OsWRKY45 expression by ER stress was markedly decreased in OsbZIP50 knockdown (KD) lines, indicating that the induction depended strongly on OsbZIP50 (Figure 1b). To determine whether OsbZIP50 interacts directly with the promoter region of OsWRKY45, we performed chromatin immunoprecipitation (ChIP) assays. Quantitative PCR analysis of ChIP output material demonstrated that DNA fragments derived from upstream regions of OsWRKY45 gene were specifically precipitated with anti-OsbZIP50 antibodies9 (Figure 1c). This result indicates that OsbZIP50 is directly involved in the induction of OsWRKY45 expression in response to ER stress. By contrast, the induction of OsWRKY45 expression by SA was similar between wild type and OsbZIP50 KD, indicating that induction by SA does not depend on OsbZIP50 (Figure 1d).
Expression of OsWRKY45 is induced by the ER stress response in an OsbZIP50-dependent manner.
(a, b, d) Relative mRNA levels of OsWRKY45. Wild type plants after treatment for 4 h with DMSO, tunicamycin (Tm) or Tm and 4-phenylbutyric acid (Tm+PBA) (a). Wild type (open bars) and OsbZIP50 knockdown (KD) plants (filled bars) after treatment with Tm (b) or SA (d). Quantitative RT-PCR analyses. Data are expressed as means ± s.d of three or four biological replicates. (c) ChIP-PCR analysis of DNA fragments precipitated with anti-OsbZIP50 antibodies. A short region upstream of the transcription initiation site of OsWRKY45 gene was analysed by quantitative PCR. DNA samples before ChIP were used as INPUT. IgG that does not react with OsbZIP50 was used as a control. Similar results were observed in two independent experiments (c).
ER stress-induced gene expression is generally suppressed by SA
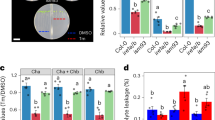
When Tm was co-administered with SA, the effects on OsWRKY45 expression were additive (Figure 2). By contrast, high-level expression of many ER stress–inducible genes by Tm was significantly suppressed by SA treatment, although induction of OsWRKY45 itself was not suppressed (Figure 2). This suppression was also observed upon addition of benzothiadiazole (BTH), a functional analogue of SA that activates the SA signalling pathway (Supplementary Figure S1), indicating that the suppression is a result of SA signalling. Furthermore, the same effect was observed when DTT was used in place of Tm (Supplementary Figure S2), indicating that the SA pathway suppresses expression of genes that are responsive to ER stress per se, rather than to some specific consequence of Tm treatment. Inversely, in order to investigate whether expression of SA-inducible genes is suppressed by ER stress, we investigated expression levels of several SA-inducible genes (Os07g0418500 encoding P450 and Os07g0129200 encoding PR1a) using the same samples. However, induction of these genes by SA was not suppressed by Tm treatment (Figure 2). The induction of Os07g0418500 expression by the SA response depends on OsWRKY4522. However, in plants treated with Tm alone, although transcripts of OsWRKY45 accumulated to high levels, the expression level of Os07g0418500 was not elevated (Figure 2). This data suggests that OsWRKY45 induced by ER stress confers a “priming” effect, in which the OsWRYK45 target genes are potentiated for expression but must await activation of the SA response in order to be expressed at high levels.
Gene expression induced by ER stress is generally suppressed by SA.
mRNA levels, determined by Quantitative RT-PCR, of representative SA-responsive and ER stress–responsive genes in wild type seedlings after treatment for 4 h with DMSO (−), tunicamycin (Tm), Tm and SA (Tm+SA) or SA and DMSO (SA). ND, not detected. Data are expressed as means ± s.d. of four biological replicates.
ER stress response reduces expression of some PR genes in an OsIRE1-dependent manner
In order to examine in more detail the relationship between the ER stress response and the defence response against pathogens, we screened for defence-related genes in DNA microarray data obtained from OsIRE1 KD plants9. The first such screen revealed that after treatment with DTT, expression levels of some genes were higher in OsIRE1 KD plants than in wild type. Many of these genes encoded pathogenesis-related (PR) proteins (e.g., Os03g0663400, a thaumatin-like protein; Os050477900, a lipid-transfer protein; Os06g0726100, endochitinase; and Os11g0645400, a disease-resistance response protein) that are putative secretory proteins expressed in root tissue even under non-stressed conditions. Quantitative RT-PCR analysis confirmed that expression levels of these PR genes were severely reduced upon DTT treatment and that some of these effects depended on expression of OsIRE1 (Figures 3a and b). Decreased expression levels of these genes were also observed when DTT was replaced with Tm, although the effects of Tm at the doses we used were slightly milder than those of DTT (Figure 3c). These results strongly suggest that these effects are caused by the ER stress response. Furthermore, the reduction in expression of PR genes by ER stress was suppressed by SA treatment, although expression of the PR genes was not induced by SA alone (Figure 3c).
ER stress response reduces expression of some PR genes in an OsIRE1-dependent manner.
mRNA levels, determined by Quantitative RT-PCR, of representative PR protein–encoding genes that exhibit ER stress-responsive changes in expression. Data are expressed as means ± s.d. of three or four biological replicates. Wild type plants after treatment for 2 h with or without DTT (a). Wild type plants (WT) and OsIRE1 knockdown (IRE1 KD) plants after treatment for 2 h with DTT (b). Wild type plants after treatment for 4 h with DMSO (−), tunicamycin (Tm), Tm and SA (Tm+SA) or SA and DMSO (SA) (c). Asterisks indicate significant differences relative to treatment with Tm (*, P<0.05; **, P<0.01; ***P<0.001; t-test).
SA does not relieve or mask ER stress
As described above, the SA response suppressed multiple effects of the ER stress response, raising the possibility that the SA response relieves or masks ER stress. However, ER stress–responsive ERQC-related factors such as BiPs, which contribute to relief or masking of ER stress, were not induced in SA-treated plants (Figure 2). Additionally, SA treatment slightly induced expression of OsbZIP50 and OsIRE1-mediated splicing of OsbZIP50 mRNA, but the induction of gene expression was suppressed by a chemical chaperone (Supplementary Figure S3). These data suggest that SA acts to induce, rather than relieve, ER stress. These observations strongly suggest that the SA response suppresses ER stress-inducible gene expression by affecting the functions of the signalling components of the ER stress response.
In summary, our results demonstrate that ER stress up-regulated the expression of genes encoding ERQC-related chaperones (such as BiPs) and OsWRKY45, but down-regulated expression of a subset of PR genes. This regulation of gene expression by ER stress response was widely suppressed by the SA response. Additionally, it is possible that the SA response utilises the OsWRKY45 protein accumulated during ER stress response to activate defence responses.
Discussion
Comparison of the gene expression patterns induced by distinct signalling pathways can provide new insight into the relationship between the responses. In this study, we found that OsWRKY45 was induced by both the ER stress and SA responses. Consequently, we investigated the possibility that interference between the ER stress and defence responses might influence the regulation of gene expression. Our findings reveal a functional integration between these two stress responses and suggest that the ER stress response has a close relationship with the defence systems that are unique to plants.
Based on our data, we propose a model (Figure 4) of the relationship between the ER stress and defence responses, in particular the SA response (Figure 4). Some PR genes are expressed even under normal conditions. For example, according to the Rice XPro microarray expression database (http://ricexpro.dna.affrc.go.jp), a thaumatin-encoding gene (Os03g0663400) analysed in this study (Figure 3) is highly expressed in root tissue. These PR proteins, which confer antibiotic resistance, were secreted through the ER (Figure 4a). When ER stress arose, ERQC-related genes were induced by activation of the OsIRE1-OsbZIP50, OsbZIP39 and OsbZIP60 pathways in order to relieve the stress (Figure 4b). In addition, ER stress down-regulated the expression of some PR genes in an OsIRE1-dependent manner (Figures 3 and 4b). This decrease in gene expression levels reduced secretion of the PR proteins, lowering the secretory burden on the ER and thereby preventing the stress condition from worsening further. Concomitantly, the ER stress response induced expression of OsWRKY45 in an OsbZIP50-dependent manner (Figures 1 and 4b). Resistance to some diseases is improved by overexpression of OsWRKY4522. Therefore, the OsWRKY45 protein accumulated in response to ER stress might offset the risk associated with decreased expression of PR genes (Figure 4b). When the SA response was activated at the same time as the ER stress response, most of the aforementioned effects of the ER stress response were suppressed, although OsWRKY45 was still induced (Figures 2, 3c and 4c). As a result of this suppression, the ER stress response refrains from consuming energy that might be used to mount the SA response and defence-related secretory proteins are permitted to pass through the ER. Additionally, accumulated OsWRKY45 protein can be subsequently activated by the SA response, resulting in rapid induction of SA-responsive gene expression (Figure 4c). Although the molecular mechanism underlying the suppression of the ER stress response by the SA response remains unknown, this finding is nonetheless important to the understanding of the environment adaptability of plants.
Model of the relationship between ER stress and defence responses.
Under normal conditions (a), some PR proteins are synthesised and secreted through the ER. When ER stress conditions arise (b), the OsIRE1-OsbZIP50, OsbZIP39 and OsbZIP60 pathways induce genes encoding ERQC-related factors (such as BiPs) in order to relieve the stress. Furthermore, the ER stress response reduces expression of some PR genes in an OsIRE-1-dependent manner and induces expression of OsWRKY45 in an OsbZIP50-dependent manner. ER stress-induced OsWRKY45 is not fully activated. When the SA response is activated concomitantly with the ER stress response (c), most of the aforementioned effects of the ER stress response are suppressed. Accumulated OsWRKY45 are subsequently activated by the SA response and contribute to SA-responsive gene expression.
Our results showed that OsIRE1 is involved in down-regulation of some PR genes under conditions of ER stress (Figure 3a). Additionally, our previous studies showed that expression levels of some seed storage proteins, secretory proteins that are produced at high levels during seed maturation, are significantly down-regulated in transgenic rice seeds under severe ER stress conditions17,23. Thus, ER stress may be responsible for regulating expression of secretory proteins other than PR proteins. In animals, IRE1 directly degrades mRNAs encoding secretory proteins24; some of the reduction in PR gene expression mediated by OsIRE1 may occur by a similar mechanism. Further study will be required to elucidate the involvement of OsIRE1 in such phenomena.
In our first DNA microarray screen for ER stress-responsive genes, we observed that OsWRKY45 was induced to a greater extent than other WRKY family members, suggesting that the transcription factor OsWRKY45 plays a specialised role. Induction of OsWRKY45 expression during ER stress is dependent on the OsIRE1-OsbZIP50 pathway (Figure 1). Taken together, these observations indicate that OsIRE1 is involved not only in expression of various ERQC-related genes, but also in expression of defence-related genes (OsWRKY45 and some PR genes), as observed in this study. Thus, over the course of evolution, IRE1 can be incorporated into species-specific pathways such as the plant defence response.
In Arabidopsis, IRE1 is also involved in immunity25. Additionally, Arabidopsis WRKY33 is induced by ER stress26, implying that the relationship between ER stress and SA responses in rice may also exist in to Arabidopsis to some extent. However, because the SA responses of the two species differ in specific detail, individual analysis will be necessary to evaluate the role of IRE1 in the Arabidopsis response.
The plant defence response is very complicated, with several distinct phytohormones participating in regulation of defence response genes. Therefore, the results presented here cannot fully explain all aspects of the defence response; nonetheless, our findings contribute significantly to our understanding of these pathways. Further studies of the relationships between ER stress and other phenomena will be required in order to elucidate plant stress responses in full detail.
Methods
Oligonucleotides
The oligonucleotides used in this study are listed in Supplementary Table S1.
Plant material and growth conditions
Oryza sativa L. cv. Kita-ake plants were grown on MS medium [1× Murashige and Skoog salt mix, 3% sucrose, B5 vitamin, 2.5 mM MES (pH 5.8) and 0.25% Gelrite] at 25°C using a 16 h light/8 h dark cycle. Seedlings (8 days old) were incubated in liquid MS medium containing 2 mM DTT, 5 µg/ml tunicamycin, 0.5 mM SA (salicylic acid, sodium salt), 0.2 mM BTH or 2 mM PBA, unless otherwise indicated. As negative controls for DTT, SA and PBA, equal volumes of water were added. As negative controls for tunicamycin and BTH, equal volumes of DMSO (final concentration of 0.1%) were added.
RNA extraction and gene expression analysis
Total RNA was extracted from root tissues using the RNeasy Plant Mini Kit (QIAGEN). For RT-PCR analysis, first-strand cDNA was synthesised from 0.8 µg of total RNA using the SuperScript III First-Strand Synthesis SuperMix for qRT-PCR (Invitrogen), which includes both oligo(dT) and random hexamers. Quantitative RT-PCR was performed using 1/80 of the prepared cDNA, specific primers (Supplementary Table S1) and SYBR® Premix Ex Taq™ (Takara). The ubiquitin-encoding gene (Os06g0681400) was used as an internal reference.
ChIP-PCR
ChIP analysis was performed as described by Haring et al. (2007) with modifications27. Chromatin samples were prepared from root tissues of wild type seedlings after treatment with DTT for 2 h and fixed by submerging the root tissues in 1% (w/v) formaldehyde solution under vacuum for 10 minutes. The fragmentation of chromatin was performed by sonication (TOMY UR-20P, output setting 8, for about 7 minutes). For immunoprecipitation, anti-OsbZIP50 antibody9 bound to protein A–Sepharose 4 Fast Flow (GE Healthcare Bio-Sciences) was pre-incubated with salmon sperm DNA and BSA. For a control IgG that does not react with OsbZIP50, we used purified anti-OsBiP4&5 antibody28. Quantitative PCR analysis of immunoprecipitated DNA fragments was performed using SYBR® Premix Ex Taq™ (Takara).
References
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Iwata, Y. & Koizumi, N. Plant transducers of the endoplasmic reticulum unfolded protein response. Trends in Plant Science (2012). doi:10.1016/j.tplants.2012.06.014
Hetz, C. & Glimcher, L. H. Fine-tuning of the unfolded protein response: Assembling the IRE1alpha interactome. Mol. Cell 35, 551–561 (2009).
Koizumi, N. et al. Molecular characterization of two Arabidopsis Ire1 homologs, endoplasmic reticulum-located transmembrane protein kinases. Plant Physiol. 127, 949–962 (2001).
Yoshida, H., Matsui, T., Yamamoto, A., Okada, T. & Mori, K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 107, 881–891 (2001).
Mori, K., Ogawa, N., Kawahara, T., Yanagi, H. & Yura, T. mRNA splicing-mediated C-terminal replacement of transcription factor Hac1p is required for efficient activation of the unfolded protein response. Proc. Natl. Acad. Sci. U.S.A. 97, 4660–4665 (2000).
Deng, Y. et al. Heat induces the splicing by IRE1 of a mRNA encoding a transcription factor involved in the unfolded protein response in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 108, 7247–7252 (2011).
Nagashima, Y. et al. Arabidopsis IRE1 catalyses unconventional splicing of bZIP60 mRNA to produce the active transcription factor. Sci. Rep. 1, 29 (2011).
Hayashi, S., Wakasa, Y., Takahashi, H., Kawakatsu, T. & Takaiwa, F. Signal transduction by IRE1-mediated splicing of bZIP50 and other stress sensors in the endoplasmic reticulum stress response of rice. Plant J. 69, 946–956 (2012).
Haze, K., Yoshida, H., Yanagi, H., Yura, T. & Mori, K. Mammalian transcription factor ATF6 is synthesized as a transmembrane protein and activated by proteolysis in response to endoplasmic reticulum stress. Mol. Biol. Cell 10, 3787–3799 (1999).
Brown, M. S., Ye, J., Rawson, R. B. & Goldstein, J. L. Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell 100, 391–398 (2000).
Liu, J.-X., Srivastava, R., Che, P. & Howell, S. H. Salt stress responses in Arabidopsis utilize a signal transduction pathway related to endoplasmic reticulum stress signaling. Plant J. 51, 897–909 (2007).
Liu, J.-X., Srivastava, R., Che, P. & Howell, S. H. An endoplasmic reticulum stress response in Arabidopsis is mediated by proteolytic processing and nuclear relocation of a membrane-associated transcription factor, bZIP28. Plant Cell 19, 4111–4119 (2007).
Che, P. et al. Signaling from the endoplasmic reticulum activates brassinosteroid signaling and promotes acclimation to stress in Arabidopsis. Sci. Signal. 3, ra69 (2010).
Takahashi, H., Kawakatsu, T., Wakasa, Y., Hayashi, S. & Takaiwa, F. A rice transmembrane bZIP transcription factor, OsbZIP39, regulates the endoplasmic reticulum stress response. Plant Cell Physiol. 53, 144–153 (2012).
Vitale, A. & Ceriotti, A. Protein quality control mechanisms and protein storage in the endoplasmic reticulum. A conflict of interests? Plant Physiol. 136, 3420–3426 (2004).
Wakasa, Y. et al. Expression of ER quality control-related genes in response to changes in BiP1 levels in developing rice endosperm. Plant J. 65, 675–689 (2011).
Nekrasov, V. et al. Control of the pattern-recognition receptor EFR by an ER protein complex in plant immunity. EMBO J. 28, 3428–3438 (2009).
Saijo, Y. et al. Receptor quality control in the endoplasmic reticulum for plant innate immunity. EMBO J. 28, 3439–3449 (2009).
Wang, D., Weaver, N. D., Kesarwani, M. & Dong, X. Induction of protein secretory pathway is required for systemic acquired resistance. Science 308, 1036–1040 (2005).
Yang, Y., Qi, M. & Mei, C. Endogenous salicylic acid protects rice plants from oxidative damage caused by aging as well as biotic and abiotic stress. Plant J. 40, 909–919 (2004).
Shimono, M. et al. Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 19, 2064–2076 (2007).
Oono, Y. et al. Analysis of ER stress in developing rice endosperm accumulating beta-amyloid peptide. Plant Biotechnol. J. 8, 691–718 (2010).
Hollien, J. & Weissman, J. S. Decay of endoplasmic reticulum-localized mRNAs during the unfolded protein response. Science 313, 104–107 (2006).
Moreno, A. A. et al. IRE1/bZIP60-mediated unfolded protein response plays distinct roles in plant immunity and abiotic stress responses. PLoS ONE 7, e31944 (2012).
Martínez, I. M. & Chrispeels, M. J. Genomic analysis of the unfolded protein response in Arabidopsis shows its connection to important cellular processes. Plant Cell 15, 561–576 (2003).
Haring, M. et al. Chromatin immunoprecipitation: optimization, quantitative analysis and data normalization. Plant Methods 3, 11 (2007).
Wakasa, Y., Hayashi, S. & Takaiwa F. Expression of OsBiP4 and OsBiP5 is highly correlated with the endoplasmic reticulum stress response in rice. Planta (in press).
Acknowledgements
We thank Y. Ikemoto, K. Miyashita, Y. Suzuki, M. Utsuno and Y. Yajima, for technical assistance. This work was supported by Genomics for Agricultural Innovation (GMC0003) and NIAS Strategic Research Fund from the Ministry of Agriculture Forestry and Fisheries of Japan to F.T.
Author information
Authors and Affiliations
Contributions
SH, YW and FT designed the experiments. SH and YW executed the experiments. SH and YW analysed the data. SH and FT wrote the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Hayashi, S., Wakasa, Y. & Takaiwa, F. Functional integration between defence and IRE1-mediated ER stress response in rice. Sci Rep 2, 670 (2012). https://doi.org/10.1038/srep00670
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00670
This article is cited by
-
Endoplasmic reticulum stress-responsive microRNAs are involved in the regulation of abiotic stresses in wheat
Plant Cell Reports (2023)
-
Probing natural variation of IRE1 expression and endoplasmic reticulum stress responses in Arabidopsis accessions
Scientific Reports (2020)
-
Molecular dissection of a rice microtubule-associated RING finger protein and its potential role in salt tolerance in Arabidopsis
Plant Molecular Biology (2015)
-
RNA sequencing-mediated transcriptome analysis of rice plants in endoplasmic reticulum stress conditions
BMC Plant Biology (2014)
-
OsWRKY30 is a transcription activator that enhances rice resistance to the Xanthomonas oryzae pathovar oryzae
Journal of Plant Biology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.