Abstract
Amyotrophic lateral sclerosis (ALS) is an incurable and fatal neurodegenerative disease characterized by the loss of motor neurons. Despite substantial research, the causes of ALS remain unclear. Glycoprotein nonmetastatic melanoma protein B (GPNMB) was identified as an ALS-related factor using DNA microarray analysis with mutant superoxide dismutase (SOD1G93A) mice. GPNMB was greatly induced in the spinal cords of ALS patients and a mouse model as the disease progressed. It was especially expressed in motor neurons and astrocytes. In an NSC34 cell line, glycosylation of GPNMB was inhibited by interaction with SOD1G93A, increasing motor neuron vulnerability, whereas extracellular fragments of GPNMB secreted from activated astrocytes attenuated the neurotoxicity of SOD1G93A in neural cells. Furthermore, GPNMB expression was substantial in the sera of sporadic ALS patients than that of other diseased patients. This study suggests that GPNMB can be a target for therapeutic intervention for suppressing motor neuron degeneration in ALS.
Similar content being viewed by others
Introduction
Amyotrophic lateral sclerosis (ALS), also known as Lou Gehrig disease, is a devastating adult-onset neurodegenerative disease characterized by progressive muscular paralysis reflecting the degeneration of motor neurons and causing death within 3–5 years of diagnosis1. Approximately 10% of ALS cases are genetically inherited, whereas the remaining 90% have no clear genetic cause2. Several ALS-linked genes have been identified, including SOD1 (encoding superoxide dismutase 1), TARDBP (encoding TAR DNA binding protein-43), FUS/TLS (encoding RNA-binding protein FUS) and others3. Furthermore, new models based on these genes have been established during recent years, improving the understanding of ALS pathogenesis3,4. Despite enormous research efforts, however, a mechanistic understanding of the neurodegenerative disease processes in vivo is still largely lacking and no effective treatments to halt the progression of ALS have yet been developed.
Gene expression profiling studies using microarrays have been conducted on various tissues from rodent models for ALS5,6,7,8, cell cultures9 and postmortem ALS central nervous system tissues10,11,12,13,14,15 to identify new disease-relevant genes and targets for therapeutic intervention in ALS and many novel genes involved in the disease pathogenesis have been identified. Furthermore, these studies have highlighted many key issues pertaining to microarray analysis in ALS, such as differences in (i) animal models and human cohorts, (ii) familial and sporadic ALS (SALS), (iii) tissue collection points at the presymptomatic or symptomatic stages and (iv) cell specificity. Consequently, the results of genome-wide screening have tended not to reflect the development of a scientific and rational approach for ALS treatment owing to poor reproducibility. Indeed, only ~5% of the genome is overlap in the same direction in more than one study16.
By microarray analysis, we identified glycoprotein nonmetastatic melanoma B (GPNMB) as a novel ALS-related factor from the spinal cords of mutant superoxide dismutase (SOD1G93A) mice. GPNMB is a type I transmembrane protein that is also known as Osteoactivin, Dendritic Cell–Heparin Integrin Ligand or Hematopoietic Growth Factor Inducible Neurokinin-1 type and was initially cloned from poorly metastatic melanoma cells as a regulator of tumor growth17. GPNMB is crucial for the differentiation and functioning of osteoclasts18 and osteoblasts19, the impairment T-cell activation20, the regulation of degeneration/regeneration of extracellular matrix in skeletal muscles21, the invasion and metastasis of several cancers, including uveal melanoma22, glioma23,24, breast cancer25, hepatocellular carcinoma26 and cutaneous melanoma27. Furthermore, it was recently reported that mutant GPNMB (GPNMBR150X) in the DBA/2J mice was involved in pigmentary glaucoma28, however there was no report about the involvement of GPNMB in neurodegenerative disorders, including ALS.
Herein we describe the investigation of new pathogenic factors for ALS and attempt to use an inclusive approach to promote translational research in ALS to overcome the current challenges of microarray analysis. First, we identified GPNMB as a novel ALS-related factor. Second, we showed the expression and intracellular localization of GPNMB in the spinal cords of the mice. Importantly, the phenotypes of GPNMB differed between motor neurons and astrocytes expressing SOD1G93A: the former suppressed GPNMB glycosylation, resulting in vulnerability, whereas the latter increased GPNMB expression and promoted secretion. Moreover, high GPNMB protein levels were observed in the cerebrospinal fluid (CSF), sera and spinal cords of human patients with ALS. These results provided evidence that GPNMB contributes very broadly to ALS and perhaps to other related neurodegenerative disorders, making it an important therapeutic target for ALS.
Results
Identification of candidate genes involved in ALS pathogenesis
We initially performed a microarray analysis to identify genes differentially expressed in the spinal cords of 14-week-old SOD1G93A and wild type (WT) mice using the Agilent feature extraction software version 10.5.1.1. More than 26,000 genes from 41,000 gene probes on the array were detected in each sample and a representative scatter plot comparison of gene expression with DNA microarray between SOD1G93A and WT mice is shown in Fig. S1. These analyses identified 1,130 genes (up: 934 genes; down: 196 genes) with more than 2-fold changed expression in SOD1G93A mice. The upregulated genes are shown in Table S1 and downregulated genes in Table S2. We focused on the GPNMB gene, which showed the most dramatic change, as a candidate gene. The GPNMB messenger RNA upregulation (P = 0.0023) in the spinal cords of SOD1G93A mice was confirmed using real-time reverse transcriptase-polymerase chain reaction (Fig. 1a).
A progressive increase of glycoprotein nonmetastatic melanoma protein B (GPNMB) in the lumber spinal cords of mutant superoxide dismutase (SOD1G93A) mice.
(a) Quantitative real-time polymerase chain reaction reveals that GPNMB messenger RNA expression is enriched in the lumber spinal cords of 14-week-old SOD1G93A (G93A) mice. Values are mean ± SEM (n = 4). **P < 0.01 versus wild-type (WT) mice (Student's t-test). UM, upper marker; LM, lower marker. (b) Time-dependent increase in GPNMB in the lumber spinal cords of SOD1G93A mice. Scale bar = 50 μm. (c) Quantification of the density of GPNMB immunoreactivity in the lumber spinal cords of WT and SOD1G93A mice. Values are mean ± SEM (n = 3 to 5). **P < 0.01 versus WT mice (Student's t-test). ##P < 0.01 versus 6-week-old SOD1G93A mice (Dunnett's test). (d) Cleavage and up-regulation of GPNMB during disease progression in SOD1G93A mice determined using immunoblot analysis. (e, f) Enhanced GPNMB immunoreactivity in NeuN-positive motor neurons and glial fibrillary acidic protein-positive astrocytes, but not in ionized calcium binding adaptor molecule 1-positive microglia in the spinal cord gray (e) or white (f) matter of 14-week-old SOD1G93A mice. Scale bar = 50 μm.
To understand better the stage at which GPNMB upregulation occurs, we further investigated GPNMB expression in the spinal cords of SOD1G93A mice. In immunohistochemical analyses, GPNMB immunoreactivity was significantly increased in 10-week-old (presymptomatic stage; P = 0.00059, Student's t-test) SOD1G93A mice and further enhanced during disease progression (Fig. 1b and c). Only a faint signal was detected in WT mice (see Fig. 1b and c).
Recently, the extracellular domain of GPNMB has been reported to be proteolytically cleaved by matrix metalloproteinases (MMPs), e.g., a disintegrin and metalloproteinase (ADAM) 10 and ADAM12; this process described as ectodomain shedding29,30. The study on ALS by Liu et al. has reported that mutant SOD1 expression in microglia enhanced the secretion of the neurotoxic cytokine tumor necrosis factor α via ADAM10 or ADAM17 activation31. Given these observations, we hypothesized that the ectodomain shedding of GPNMB might facilitate ALS pathogenesis. Practically, GPNMB extracellular fragments with molecular mass of 90 kDa were detected in 14-week-old (onset) SOD1G93A mice and were further enhanced during disease progression (Fig. 1d) and glycosylated GPNMB with a molecular mass of approximately 100 kDa was dramatically upregulated in 20-week-old (end-stage) SOD1G93A mice (see Fig. 1d). We also identified two small C-terminal fragments with molecular weights of approximately 25 and 15 kDa (see Fig. 1d). Furthermore, we performed double immunofluorescent staining to confirm the localization of GPNMB in the spinal cords (Fig. 1e and f). GPNMB colocalized with NeuN-positive motor neurons and glial fibrillary acidic protein (GFAP)-positive astrocytes but not with ionized calcium binding adaptor molecule 1 (Iba-1)-positive microglia in the spinal cord gray (E) or white matter (F) of 14-week-old SOD1G93A mice. Conversely, in WT mice, GPNMB-immunoreactive signals colocalized with NeuN-positive motor neurons but not with GFAP- or Iba-1-positive glial cells (see Fig. 1e and f).
Downregulation of GPNMB increased the motor neuron vulnerability
To investigate the role of GPNMB in each cell (i.e., motor neuron or astrocytes), we first examined whether GPNMB expression was altered by SOD1G93A in NSC34 cells. The expression of SOD1G93A was detectable within 48 h of transfection (Fig. 2a). Protein levels of glycosylated GPNMB were significantly decreased (P = 0.016) in the SOD1G93A-expressing NSC34 cells but not in WT SOD1 (SOD1wt) for 48 h (see Fig. 2a and b). The glycosylated GPNMB was specifically down-regulated in the SOD1G93A-expressing NSC34 cells, conversely increased by SOD1H46R (see Fig. S2). On the other hand, non-glycosylated GPNMB and GPNMB messenger RNA levels were unchanged by SOD1G93A in NSC34 cells, indicating that the post-translational modification of GPNMB was influenced by SOD1G93A (see Fig. 2a and c, Fig. S3).
Downregulation of glycosylated GPNMB through interaction with SOD1G93A, leading to motor neuron death.
(a-c) NSC34 motor neuron cells were lysed after transfection with Myc-tagged WT SOD1 (SOD1wt) or SOD1G93A for 48 h. (a) Expression of GPNMB was examined using immunoblotting. The protein expression levels of glycosylated (b), but not non-glycosylated (c), GPNMB were decreased in the SOD1G93A-expressing cells. Values are mean ± SEM (n = 3 or 4). *P < 0.05 versus SOD1wt (Student's t-test). (d) Lysates from NSC34 cells transfected with Myc-tagged SOD1wt or SOD1G93A for 48 h were immunoprecipitated with an antibody to GPNMB (IP: GPNMB) and analyzed using immunoblotting with antibodies to ubiquitin, GPNMB and Myc. (e) Laser scanning confocal photomicrographs of NSC34 cells showing the expression of GPNMB (green), ubiquitin (red) and Hoechst 33342 (blue). In the SOD1G93A-expressing cells, the aggregates of GPNMB in cytoplasm are partly colocalized with ubiquitin (arrowheads). Scale bar = 5 μm. (f, g) NSC34 cells were cotransfected with small interfering RNA (siRNA) against GPNMB or a nonspecific sequence (negative control) and enhanced green fluorescent protein-tagged mock, SOD1wt, or SOD1G93A for 48 h. Representative fluorescence microscopy showing nuclear staining for Hoechst 33342 (blue) and propidium iodide (red) (f). FBS, fetal bovine serum. The cell viability was reduced 48 h after the cotransfection of small interfering RNA and SOD1G93A (g). Values are mean ± SEM (n = 6). #P < 0.05 versus mock (Student's t-test). **P < 0.01 versus each negative control (Tukey's test). Scale bar = 100 μm.
Mutant SOD1, but not SOD1wt, is known to interact with several proteins—e.g., translocon-associated protein, heat shock protein 25, heat shock protein/heat shock cognate 70 and degradation in endoplasmic reticulum protein 132,33,34,35. In the present study, SOD1G93A in NSC34 cells coimmunoprecipitated with GPNMB and clearly increased the amount of ubiquitinated GPNMB (Fig. 2d). The ubiquitin-proteasome system degrades cytosolic aggregated misfolded proteins and, when dysfunctional, contributes to endoplasmic reticulum stress36. In enhanced green fluorescent protein-tagged mock or SOD1wt-expressing cells, GPNMB is mainly localized to the cytoplasm; however, the intracellular aggregates of ubiquitinated GPNMB were observed in the cytoplasm of SOD1G93A-expressing NSC34 cells (Fig. 2e). These findings suggest that the polyubiquitinated GPNMB may be degraded by the proteasome before the downregulation of glycosylated GPNMB.
We next assessed the involvement of GPNMB in SOD1G93A-induced neurotoxicity using small interfering RNA (siRNA) against GPNMB. GPNMB siRNA #2 effectively suppressed protein levels of GPNMB in both SOD1wt and SOD1G93A-expressing NSC34 cells (Fig. S4). We examined whether reducing GPNMB affected SOD1G93A-induced neurotoxicity using WST-8 assays and combination staining with fluorescent dyes—i.e., Hoechst 33342 and propidium iodide. Compared with negative control siRNA, GPNMB siRNA aggravated SOD1G93A or serum-deprivation-induced cell death and the down-regulation of cell proliferative activity (Fig. 2f and g, Fig. S5), indicating that endogenous GPNMB protects motor neurons from SOD1G93A-induced neurotoxicity. Interestingly, in the presence of SOD1wt, cell viability was significantly decreased by GPNMB depletion compared with that in the control siRNA (see Fig. 2g). These results also suggest that GPNMB plays a critical role in maintaining homeostasis in motor neurons.
Extracellular fragments of GPNMB attenuated the neurotoxicity of SOD1G93A
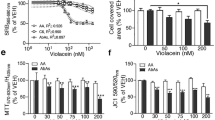
As shown in Fig. 2c, we detected extracellular fragments of GPNMB in the spinal cords of SOD1G93A mice; therefore, we investigated whether treatment with recombinant extracellular GPNMB affects SOD1G93A-induced motor neuron damage. Recombinant GPNMB at 0.025–2.5 µg/mL suppressed SOD1G93A-induced cell death in a concentration-dependent manner, with significant effects observed at 0.25 and 2.5 µg/mL (see Fig. 3a). We considered the mechanism through which GPNMB protects NSC34 cells from SOD1G93A-induced cell death. Previous work has demonstrated that treatment with a recombinant fragment of GPNMB increases extracellular signal-regulated kinase 1 and 2 (ERK1/2) signaling in fibroblasts, resulting in the upregulation of MMP329. As expected, recombinant GPNMB markedly increased ERK1/2 phosphorylation at 5–60 min (peak at approximately 30 min) and the phosphorylation levels of protein kinase B (Akt) in a time-dependent manner after treatment with 2.5 µg/mL extracellular GPNMB fragments (see Fig. 3b, Fig. S6). Furthermore, the protective effect of GPNMB was attenuated by treatment with a phosphatidylinositol 3 kinase (PI3K) inhibitor (LY294002; 30 µM) or a mitogen-activated protein kinase kinase (MEK) 1/2 inhibitor (U0126; 10 µM; Fig. 3c and d). These data suggest that GPNMB enhances survival signals such as the PI3K/Akt and MEK/ERK pathways, protecting cells from neuronal degeneration.
The extracellular fragments of GPNMB attenuated the neurotoxicity of SOD1G93A.
(a) The recombinant extracellular fragment of GPNMB at 0.25–2.5 µg/mL demonstrated a protective effect against SOD1G93A-induced cell death. Values are mean ± SEM (n = 3 to 6). #P < 0.05 versus control (Student's t-test). *P < 0.05, **P < 0.01 versus vehicle (Dunnett's test). Scale bar = 100 μm. (b) Time course of changes in phosphorylated ERK1/2 and phosphorylated Akt level after recombinant GPNMB treatment. (c, d) The protective effect of GPNMB against SOD1G93A-induced cell death was eliminated by LY294002, a PI3-kinase inhibitor, at 20 µM or by U0126, a MEK1/2 inhibitor, at 5 µM. Values are mean ± SEM (n = 6). ##P < 0.01 versus control (Student's t-test), **P < 0.01 versus SOD1G93A alone or SOD1G93A treated with GPNMB (Turkey's test). Scale bar = 100 μm. (e) Quantitative analysis of GPNMB in the conditioned media (CM) was performed using enzyme-linked immunosorbent assay. Values are mean ± SEM (n = 5). ##P < 0.01 versus mock, **P < 0.01 vs. SOD1wt (Tukey's test). (f) Expressions of GPNMB, MMP3, MMP9 and GFAP in NHA transfected with Myc-tagged SOD1wt or SOD1G93A were examined using immunoblotting. (g) CM from NHA transfected with SOD1G93A were immunoprecipitated with an antibody to GPNMB (IP: GPNMB) or control nonimmune antibody (IP: C) and added to NSC34 cells. Values are mean ± SEM (n = 5). ##P < 0.01 versus astrocyte basal medium, **P < 0.01 versus IP: C (Student's t-test). Scale bar = 100 μm.
These in vitro studies using NSC34 cells highlighted a potential role of GPNMB in motor neuron death. We found in vivo-in vitro discrepancy in the expression of GPNMB—i.e., the former was increased, whereas the latter was decreased. This discrepancy may be attributed to the use of only NSC34 cells in the in vitro study. Therefore, we next focused on astrocytes, which, like motor neurons, expressed GPNMB in the spinal cords of SOD1G93A mice (see Fig. 1e). Normal human astrocytes (NHAs) were cultured for 5 days after transfection with Myc-tagged mock, SOD1wt, or SOD1G93A and the conditioned media (CM) and corresponding cell layers were analyzed for GPNMB expression. Enzyme-linked immunosorbent assay revealed that GPNMB was secreted into the CM of the SOD1G93A-expressing NHA but not into the mock or SOD1wt NHAs (Fig. 3e). Accompanying the secretion of GPNMB, the activation of MMP3 and increase in MMP9 occurred in the SOD1G93A-expressing NHA (Fig. 3f, Fig. S7), similar to the results of previous studies that used mouse fibroblastic NIH-3T3 cells21. Because astrocytes carrying the SOD1G93A mutation have been shown specifically to induce motor neuron death37,38,39, we next investigated whether the existence of GPNMB in the CM regulates cell damage from the neurotoxicity of the SOD1G93A by eliminating GPNMB in the CM. After treatment to remove GPNMB from the CM, the number of propidium iodide-positive NSC34 cells was significantly higher than that after immunoprecipitation using normal immunoglobulin G (P = 0.000023) and astrocyte basal medium (P = 0.000013) (Fig. 3g). These findings helped to explain the in vitro/in vivo discrepancy and suggested that GPNMB secretion from SOD1G93A expression in astrocytes may confer a protective effect against motor neuron death.
GPNMB improved ALS pathogenesis in SOD1G93A mice
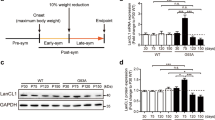
To determine the effects of GPNMB in ALS symptoms, we generated SOD1G93A/GPNMB double-transgenic (G93A/GPNMB) mice (Fig. 4a). V5-tagged GPNMB protein was detected in the spinal cords of these mice through immunoblot analysis (Fig. 4b). Disease onset was determined by the loss of motor function, which was measured using a rotarod test. The overexpression of GPNMB prolonged the decline of latency to fall off the rod (Fig. 4c) and disease onset (P = 0.0010; Fig. 4d and f). Conversely, no significant difference was found in body weight between the SOD1G93A (G93A/-) and G93A/GPNMB mice (Fig. S8). The mean disease duration (from onset to end stage) did not slow in the G93A/GPNMB mice compared to that in the G93A/- mice, however (Fig. 4g). Overall survival was extended by 8.2% (P = 0.022; G93A/GPNMB, 128.9 ± 2.0 days; G93A/-, 119.1 ± 2.4 days; Fig. 4e). Furthermore, we evaluated the effect of GPNMB overexpression on motor neuron protection in the spinal cords of G93A/GPNMB mice at 15 weeks old. The number of surviving motor neurons in G93A/GPNMB mice was significantly increased compared with that in the G93A/- mice (Fig. 4h). On the other hand, there was no significant alteration in the glial activation between G93A/- and G93A/GPNMB mice (Fig. S9). These data indicate that GPNMB significantly extended survival by delaying disease onset.
Regulation of amyotrophic lateral sclerosis (ALS) pathogenesis by GPNMB.
(a) Non-transgenic (−/−), GPNMB (-/GPNMB), SOD1G93A (G93A/-) and SOD1G93A/GPNMB double-transgenic (G93A/GPNMB) mice at approximately 5 weeks old. (b) The protein expression levels of V5-tagged GPNMB were increased in GPNMB-transgenic spinal cord tissues compared with those of the WT. (c) Motor performance assessed using the rotarod test for G93A/- and G93A/GPNMB mice. Values are mean ± SEM (n = 10). ##P < 0.01 versus G93A/- mice (two-way repeated measure ANOVA). *P < 0.05, **P < 0.01 versus G93A/- mice (Student's t-test). (d) Age of disease onset for G93A/- (n = 10) and G93A/GPNMB (n = 10) mice. Disease onset was defined at the day when a mouse first dropped off the rotarod within 600 s. P < 0.01 using the log-rank test. (e) Survival curve for G93A/- (n = 10) and G93A/GPNMB (n = 10) mice. P < 0.05 using the log-rank test. (f, g) Mean onset (f) and mean duration of disease progression (from onset to end stage; g) for G93A/- and G93A/GPNMB mice. Values are mean ± SEM (n = 10). **P < 0.01 versus G93A/- mice (Student's t-test). (h) Cresyl violet staining in the spinal cords of non-transgenic (−/−), SOD1G93A (G93A/-) and SOD1G93A/GPNMB double-transgenic (G93A/GPNMB) mice at 15 weeks old. Values are mean ± SEM (n = 3−5). ##P < 0.01 versus non-transgenic (−/−) mice, **P <0.01 versus SOD1G93A (G93A/-) mice. Scale bar = 25 μm.
GPNMB expression in the sporadic ALS patients
To investigate the involvement of GPNMB in SALS pathogenesis, we measured GPNMB in the CSF and sera of SALS patients. Quantitative enzyme-linked immunosorbent assay revealed that GPNMB protein levels in the CSF of SALS patients were significantly increased by 1.7-fold (P = 0.0041 vs. those of non-neurological disease controls; Fig. 5a). Furthermore, the expression of GPNMB in the sera of SALS patients was significantly higher: 1.5-fold (P < 0.01) that in non-neurological disease controls and 1.3-fold that in Alzheimer disease (P < 0.01) and Parkinson disease (P < 0.01; Fig. 5b). We next performed immunohistochemistry to confirm the distribution of GPNMB in human spinal cord tissues. In the lumbar anterior horns of SALS patients and controls, GPNMB immunoreactive signals mainly localized within motor neurons (Fig. 5c). GPNMB is known to have 60% homology to the precursor of Pmel-17/gp100, a melanocyte-specific protein17,40. Pmel-17 is a pigment-cell-specific integral membrane protein that participates in the formation of melanosome-like fibrils with features of amyloid41,42. Interestingly, GPNMB is deposited extracellularly in the lumber spinal cords of SALS patients (see Fig. 5c, arrows). These results suggest that the extracellular fragments of GPNMB are partly secreted into the bloodstream and CSF and others form the deposits of GPNMB plaques in ALS pathogenesis.
Representation of GPNMB protein in cerebrospinal fluid (CSF), sera and lumber spinal cord tissues of sporadic ALS (SALS) patients.
(a, b) The amount of GPNMB secreted into CSF (a) or sera (b) in SALS patients was higher than that in controls and patients with Alzheimer disease and Parkinson disease. Values are mean ± SEM (n = 10 to 28). **P < 0.01 versus control (Mann–Whitney U-test; CSF samples). **P < 0.01 versus controls and patients with Alzheimer and Parkinson diseases (Tukey's test; sera samples). (c) Representative photographs are shown for the lumber spinal cords in the control and SALS patients. Extracellular deposition of GPNMB is observed in the lumber spinal cords of SALS (arrows). Scale bar = 100 μm. Values are mean ± SEM (n = 3). **P < 0.01 versus controls (Student's t-test).
Discussion
Given that the causes of ALS remain unclear despite substantial research, determining the original disease-related factors is important. In the present study, we identified GPNMB as a potential effector of neurodegeneration in an ALS mouse model. Our results suggested that glycosylated GPNMB in NSC34 cells was reduced by polyubiquitination, increasing cell vulnerability and that GPNMB was secreted extracellularly through ectodomain shedding from activated astrocytes to prevent motor neuron degeneration, as summarized in Fig. 6. The protective effect of GPNMB was mediated by the activation of the PI3K/Akt and MEK/ERK pathways. Similar to outcomes of in vitro studies, motor performance and survival time were improved by GPNMB overexpression in SOD1G93A mice. Finally, we detected the expression of GPNMB in the CSF, sera and spinal cords of human patients with ALS. These findings suggested that GPNMB could contribute to ALS pathogenesis. To our knowledge, this study is the first to report the association between GPNMB and neurodegenerative diseases represented by ALS.
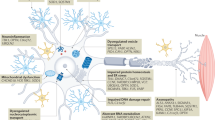
Hypothesized mechanisms for GPNMB regulation of motor neuron degeneration in ALS.
In motor neurons, glycosylation of GPNMB is inhibited by the interaction with SOD1G93A and GPNMB polyubiquitination. The downregulation of glycosylated GPNMB increases motor neuron vulnerability, ultimately triggering motor neuron death. Activated astrocytes secrete the extracellular fragments of GPNMB. The secretion of GPNMB is mediated by metalloproteinases such as a disintegrin and metalloproteinases and the fragments attenuate the neurotoxicity of SOD1G93A in motor neurons. Promoting the release of GPNMB extracellular fragments may rescue the motor neurons.
In the present study, we showed that the extracellular domain of GPNMB had protective effects against mutant SOD1-induced neurotoxicity via ERK1/2 and Akt pathways. Two previous reports have linked GPNMB to tumor cell apoptosis. GPNMB-dependent augmentation of tumor growth has been attributed to decreased apoptosis and increased angiogenesis in GPNMB-expressing tumors30. Recently, Rose et al.43 have also reported that treatment with CDX-011, a GPNMB-specific antibody, elevates apoptosis in GPNMB-expressing breast cancer cells. Furthermore, the recombinant fragment of GPNMB increases MMP3 expression through the ERK1/2 pathway in fibroblasts29. These findings suggest that GPNMB may activate survival signals via the MEK/ERK and PI3K/Akt pathways.
From a therapeutic perspective, we propose that a successful treatment strategy for ALS will be found by characterizing the sheddases specific for GPNMB and by promoting ectodomain shedding as early as possible by activating these enzymes. Recently, non-neuronal cells such as astrocytes and microglia have been reported to contribute to the disease process of ALS and as a consequence, motor neuron death in ALS is considered a “non-cell autonomous” process44,45,46. Our findings showed that GPNMB expression was increased in the motor neurons and astrocytes in the spinal cords of SOD1G93A mice and the activated astrocytes secreted GPNMB extracellularly. Astrocytes have many important functions relevant to motor neuron physiology, including the release of several neurotrophic factors to maintain neuronal health47,48. Glycosylation of GPNMB was in fact inhibited through interaction with SOD1G93A, increasing motor neuron vulnerability. Furthermore, activated astrocytes secreted extracellular fragments of GPNMB, attenuating the neurotoxicity of SOD1G93A in motor neurons (see Fig. 6). These results suggest that the ectodomain shedding of GPNMB may be performed to maintain surrounding cells, such as motor neurons. In our in vivo study, the overexpression of GPNMB in SOD1G93A mice significantly extended survival by delaying disease onset; however, the effect was less than that expected. One of the reasons for this result was the difference in of time points between motor neuron degeneration and astrogliosis. Previously, we have shown that motor neuron loss is observed at the presymptomatic stage (10-week-old) in SOD1G93A mice, whereas astrocytes are dramatically activated at the end stage49, a phenomenon similar to GPNMB upregulation in the spinal cords of SOD1G93A mice. These results suggest that promoting ectodomain shedding of GPNMB at an earlier stage is a key for ALS treatment. Thus, much stronger effects of GPNMB were expected in in vivo models. Further studies will be required to identify a GPNMB-specific sheddase as well as GPNMB receptors in the central nervous system.
In conclusion, we demonstrated that GPNMB inhibits motor neuron death and plays a critical role in motor neuron survival. Accordingly, GPNMB may be a potential new therapeutic target for ALS.
Methods
An expanded Methods section is available in the section of SUPPLEMENTARY MATERIAL.
Expression plasmids, cell culture and transfection
Human WT SOD1 (SOD1wt), mutant SOD1G93A, SOD1G37R, SOD1H46R and SOD1G85R cDNAs containing the entire coding region were transferred from Nagoya University Graduate School of Medicine50. NSC34 cells, a hybrid neuroblastoma x spinal cord cell line, were purchased from Cosmo Bio Co., Ltd. (Tokyo, Japan) and maintained in Dulbecco's modified Eagle's medium containing with 10% fetal bovine serum (Valeant, Costa Mesa, CA, USA), 100 U/mL penicillin (Meiji Co., Ltd., Tokyo, Japan) and 100 µg/mL streptomycin (Meiji Co., Ltd.). Normal human astrocytes (NHAs) were maintained in Astrocyte Basal Medium (ABM; CC-3187; Lonza Group Ltd., Basel, Switzerland) supplemented with the growth medium (CC-4123; Lonza), according to the manufacturer's instructions. Transfections were performed using Lipofectamin 2000 (Invitrogen, Carlsbad, CA, USA) according to manufacture's recommendations.
siRNA knockdown of GPNMB
NSC34 cells were transfected with 100 nM of mouse GPNMB-specific RNAi oligo and control RNAi oligo (Nippon EGT Co., Ltd., Toyama, Japan) using Lipofectamine 2000 (Invitrogen) according to manufacturer's recommendations. Knockdown was analyzed by immunoblot (IB) with antibody to GPNMB. Sequences were as follows:
#1, 5′-CCAUCUUGCUGUACAAAAAdTdT-3′ (sense) and 5′-UUUUUGUACAGCAAGAUGGdTdT-3′ (antisense);
#2, 5′-GCACGGGUUUCUAUAAACAdTdT-3′ (sense) and 5′-UGUUUAUAGAAACCCGUGCdTdT-3′ (antisense);
#3, 5′-GAAGCUUUUUGUUUGGAAAdTdT-3′ (sense) and 5′-UUUCCAAACAAAAAGCUUCdTdT-3′ (antisense); and universal negative control small interfering RNA (siRNA).
RNA isolation
Spinal cord biopsy specimens were cut into 0.5 cm cubes or smaller and stored in RNA later (Ambion, Austin, TX, USA) overnight at 4 ± 3°C, then stored at –80°C until RNA extraction. Total RNA was extracted using TRIzol reagent (Invitrogen) according to the manufacturer's instructions. Total RNA was further purified using the Qiagen RNeasy Mini Kit (QIAGEN, Valencia, CA, USA) according to the manufacturer's instructions. In NSC34 cells, total RNA was isolated using NucleoSpin RNA II (Takara Bio Inc., Shiga, Japan). RNA quantity and quality were determined using a Nanodrop ND-1000 spectrophotometer (Thermo Fisher Scientific, Inc., Waltham, MA, USA), an Agilent Bioanalyzer (Agilent Technologies, Palo Alto, CA, USA), or NanoVue Plus (GE Healthcare Japan, Tokyo, Japan), as recommended.
cRNA amplification and labeling
Total RNA from spinal cords was amplified and labeled with Cyanine 3 (Cy3) using Agilent Quick Amp Labeling kit, one-color (Agilent Technologies) following the manufacturer's instructions. Briefly, 500 ng of total RNA was reversed transcribed to double-strand cDNA using a poly dT-T7 promoter primer. Primer, template RNA and quality-control transcripts of known concentration and quality were first denatured at 65°C for 10 min and incubated for 2 h at 40°C with 5X first strand Buffer, 0.1 M DTT, 10 mM dNTP mix, MMLV RT and RNase-out. The MMLV RT enzyme was inactivated at 65°C for 15 min. cDNA products were then used as templates for in vitro transcription to generate fluorescent cRNA. cDNA products were mixed with a transcription master mix in the presence of T7 RNA polymerase and Cy3 labeled-CTP and incubated at 40°C for 2 h. Labeled cRNAs were purified using QIAGEN's RNeasy mini spin columns and eluted in 30 μL of nuclease-free water. After amplification and labeling, cRNA quantity and cyanine incorporation were determined using a Nanodrop ND-1000 spectrophotometer and an agilent bioanalyzer.
Microarray analysis
Total RNA was amplified and labeled with 1.65 μg of Cy3 for the test sample using Agilent's Low RNA Input Linear Amplification Kit (Agilent Technologies, Palo Alto, CA, USA) following the detailed protocol described in the kit manual (version 5.7). Briefly, 500 ng of total RNA was reversed transcribed to doublestrand, cDNA using a poly dT-T7 promoter primer. Primer, template RNA and quality-control transcripts of known concentration and quality were first denatured at 65°C for 10 min and incubated for 2 h at 40°C with 5x first strand Buffer, 0.1 M DTT, 10 mM dNTP, MMLV RT and RNase-out. The MMLVRT enzyme was inactivated at 70°C for 15 min. cDNA products were then used as templates for in vitro transcription to generate fluorescent cRNA. cDNA products were mixed with a transcription master mix in the presence of T7 RNA polymerase and Cy3 labeled-CTP and incubated at 40°C for 2 h. Labeled cRNAs were purified using Qiagen's RNeasy mini spin columns and eluted in 30 mL of nuclease-free water. After amplification and labeling, cRNA quantity and cyanine incorporation were determined using a NanoDrop ND. Microarray expression experiments were performed on Agilent Mouse GE 4x44K v1 Microarray (Design ID: 014868) according to the manufacturer's instructions. Images of the arrays were acquired using a microarray scanner G2565BA (Agilent Technologies) and image analysis was performed using Feature Extraction software version 9.5 (Agilent Technologies).
Data analysis of microarray
Intensity values of each scanned feature were quantified using Agilent feature extraction software version 10.5.1.1, which performs background subtractions. We only used features which were flagged as no errors (present flags) and excluded features which were not positive, not significant, not uniform, not above background, saturated and population outliers (marginal and absent flags). Normalization was performed using Agilent GeneSpring GX version 10.0.1 and normalized by setting all measurements, 0.01 to 0.01, normalizing each chip to the 75 percentile of all measurements taken for that chip and normalizing each gene to the median measurement for that gene across all chips.
Real-time PCR
Single-stranded cDNA was synthesized using a PrimeScript RT Master Mix (Takara). Species-specific primer set designed to detect mouse GPNMB mRNA was 5′-TCTGAACCGAGCCCTGACATC-3′ (sense) and 5′-AGCAGTAGCGGCCATGTGAAG-3′ (antisense). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) mRNA was used as a loading control employing 5′-TGTGTCCGTCGTGGATCTGA-3′ (sense) and 5′-TTGCTGTTGAAGTCGCAGGAG-3′ (antisense). Relative quantitative real-time PCR was performed using the Thermal Cycler Dice Real Time System TP800 with SYBR Premix Ex Taq™ II (Takara), according to manufacture's recommendations. The thermal cycler conditions were as follows: 5 sec at 95°C and then 30 sec at 60°C, followed by two-step PCR for 60 cycles consisting of 95°C for 15 sec followed by 60°C for 1 min. For each PCR, we obtained the slope value, R2 value and linear range of a standard curve of serial dilutions. The results were expressed relative to the GAPDH internal control.
Immunoprecipitation
NSC34 cells and NHA were transfected with Myc-tagged Mock, SOD1wt, or SOD1G93A for 48 h or 5 days, respectively. GPNMB was immunoprecipitated from NSC-34 cell lysates or conditioned media (CM) clutured NHA using Pierce Classic IP kit (Thermo), according to manufacture's recommendations. The cell lysates or CM were incubated with anti-GPNMB antibody (S-24; Santa Cruz Biotechnology, Santa Cruz, CA, USA) or normal rabbit IgG (PeproTech Inc. Rocky Hill, NJ, USA). For immunoblotting, immunoprecipitates were released by incubation in 8% (w/v) sodium dodecyl sulfate (SDS) sample buffer solution (Wako Pure Chemicals, Osaka, Japan).
Immunoblot analysis
Cells, spinal cord tissues and serum proteins were lysed using a buffer (RIPA buffer R0278; Sigma-Aldrich, St. Louis, MO, USA) with protease (P8340; Sigma-Aldrich) and phosphatase inhibitor cocktails (P2850 and P5726; Sigma-Aldrich). The protein concentration was measured by comparison with a known concentration of bovine serum albumin using a BCA Protein Assay kit (Thermo). Equal amounts of protein in sample buffer with 10% 2-mercaptoethanol was subjected to SDS-polyacrylamide gels (PAGE) using 5–20% gradient gels (SuperSep Ace; Wako) and the separated proteins were transferred onto a polyvinylidene difluoride membrane (Immobilon-P; Millipore Corporation, Bedford, MA, USA). After blocking with Blocking One-P (Nacalai Tesque, Kyoto, Japan), membranes were incubated with the primary antibodies as follows: goat anti-GPNMB (AF2330; R&D Systems Inc., Minneapolis, MN, USA), mouse anti-ubiquitin (Cell Signaling Technology, Danvers, MA, USA), rabbit anti-extracellular signal-regulated kinase 1 and 2 (ERK1/2) (Cell Signaling), rabbit anti-p-ERK1/2 (Cell Signaling), rabbit anti-protein kinase B (Akt) (Cell Signaling), rabbit anti-p-Akt (Cell Signaling), rabbit anti- matrix metalloproteinase (MMP) 3 (Sigma-Aldrich), rabbit anti-MMP9 (Millipore), mouse anti-glial fibrillary acidic protein (GFAP) (Millipore), mouse anti-V5 antibody (Invitrogen), mouse anti-Myc-Tag (Cell Signaling) and mouse anti-β-actin (Sigma-Aldrich). The band intensities were measured using an ImmunoStar LD (Wako).
Cell death assay
NSC34 cells were seeded at 7 × 103 cells per well into a collagen-coated 96-well plate and then incubated at 37°C in a humidified atmosphere of 95% air and 5% CO2. Twenty four hours after plating cells were transfected with plasmids coding for the various proteins, which were enhanced green fluorescent protein (GFP)-tagged Mock, SOD1wt, or SOD1G93A and in some cases, replaced with the serum–free media for 24 h. In case of GPNMB knockdown, the cells were transfected them simultaneously with the plasmid coding for the various proteins. In case of using the recombinant GPNMB (2550-AC; R&D Systems), we treated the recombinant proteins just after replaceing with the serum–free media. In other experiments, NSC34 cells were replaced with the CM from NHA immunoprecipitated of GPNMB. Assessment of cell viability was performed using two methods. The first method was a single-cell digital imaging-based method employing fluorescent staining of nuclei. Cell death was assessed on the basis of combination staining with fluorescent dyes [namely, Hoechst 33342 and propidium iodide (Invitrogen)], observations being made using an OLYMPUS IX70 inverted epifluorescence microscope (Olympus, Tokyo, Japan). In a blind manner, a total of at least 200 cells per condition were counted using image-processing software (Image-J ver. 1.33f; National Institutes of Health, USA). As the second method for measuring cell viability, cell metabolic activity was quantitatively assessed. Cell viability was assessed following immersion in 10% WST-8 solution (Cell Counting Kit-8; Dojin Kagaku, Kumamoto, Japan) for 2 h at 37°C and absorbance was recorded at 450 nm. This absorbance is expressed as a percentage of that in control cells, after subtraction of background absorbance.
Enzyme-linked immunosorbent assay
The culture media of NHA transfected with Myc-tagged Mock, SOD1wt, or SOD1G93A for 5 days or human serum samples were subjected to enzyme-linked immunosorbent assay (ELISA). ELISA for human osteoactivin/GPNMB (DY2550; R&D Systems) was carried out according to the manufacture's recommendations. The absorbance was measured at 450 nm using VARIOSKAN FLASH (Thermo).
References
Wijesekera, L. C. & Leigh, P. N. Amyotrophic lateral sclerosis. Orphanet J Rare Dis. 4, 3 (2009).
Dion, P. A., Daoud, H. & Rouleau, G. A. Genetics of motor neuron disorders: new insights into pathogenic mechanisms. Nat Rev Genet. 10, 769–782 (2009).
Vande Velde, C., Dion, P. A. & Rouleau, G. A. Amyotrophic lateral sclerosis: new genes, new models and new mechanisms. F1000 Biol Rep. 3, 18 (2011).
Jackson, M., Ganel, R. & Rothstein, J. D. Models of amyotrophic lateral sclerosis. Curr Protoc Neurosci. Chapter 9, Unit 9 13 (2002).
Ferraiuolo, L. et al. Microarray analysis of the cellular pathways involved in the adaptation to and progression of motor neuron injury in the SOD1 G93A mouse model of familial ALS. J Neurosci. 27, 9201–9219 (2007).
Olsen, M. K. et al. Disease mechanisms revealed by transcription profiling in SOD1-G93A transgenic mouse spinal cord. Ann Neurol. 50, 730–740 (2001).
Yoshihara, T. et al. Differential expression of inflammation- and apoptosis-related genes in spinal cords of a mutant SOD1 transgenic mouse model of familial amyotrophic lateral sclerosis. J Neurochem. 80, 158–167 (2002).
Lobsiger, C. S., Boillee, S. & Cleveland, D. W. Toxicity from different SOD1 mutants dysregulates the complement system and the neuronal regenerative response in ALS motor neurons. Proc Natl Acad Sci U S A. 104, 7319–7326 (2007).
Vargas, M. R., Pehar, M., Diaz-Amarilla, P. J., Beckman, J. S. & Barbeito, L. Transcriptional profile of primary astrocytes expressing ALS-linked mutant SOD1. J Neurosci Res. 86, 3515–3525 (2008).
Dangond, F. et al. Molecular signature of late-stage human ALS revealed by expression profiling of postmortem spinal cord gray matter. Physiol Genomics. 16, 229–239 (2004).
Ishigaki, S. et al. Differentially expressed genes in sporadic amyotrophic lateral sclerosis spinal cords--screening by molecular indexing and subsequent cDNA microarray analysis. FEBS Lett. 531, 354–358 (2002).
Jiang, Y. M. et al. Gene expression profile of spinal motor neurons in sporadic amyotrophic lateral sclerosis. Ann Neurol. 57, 236–251 (2005).
Malaspina, A., Kaushik, N. & de Belleroche, J. Differential expression of 14 genes in amyotrophic lateral sclerosis spinal cord detected using gridded cDNA arrays. J Neurochem. 77, 132–145 (2001).
Offen, D. et al. Spinal cord mRNA profile in patients with ALS: comparison with transgenic mice expressing the human SOD-1 mutant. J Mol Neurosci. 38, 85–93 (2009).
Wang, X. S., Simmons, Z., Liu, W., Boyer, P. J. & Connor, J. R. Differential expression of genes in amyotrophic lateral sclerosis revealed by profiling the post mortem cortex. Amyotroph Lateral Scler. 7, 201–210 (2006).
Kudo, L. C. et al. Integrative gene-tissue microarray-based approach for identification of human disease biomarkers: application to amyotrophic lateral sclerosis. Hum Mol Genet. 19, 3233–3253 (2010).
Weterman, M. A. et al. nmb, a novel gene, is expressed in low-metastatic human melanoma cell lines and xenografts. Int J Cancer. 60, 73–81 (1995).
Ripoll, V. M. et al. Microphthalmia transcription factor regulates the expression of the novel osteoclast factor GPNMB. Gene. 413, 32–41 (2008).
Abdelmagid, S. M. et al. Osteoactivin, an anabolic factor that regulates osteoblast differentiation and function. Exp Cell Res. 314, 2334–2351 (2008).
Chung, J. S., Dougherty, I., Cruz, P. D., Jr & Ariizumi, K. Syndecan-4 mediates the coinhibitory function of DC-HIL on T cell activation. J Immunol. 179, 5778–5784 (2007).
Ogawa, T. et al. Osteoactivin upregulates expression of MMP-3 and MMP-9 in fibroblasts infiltrated into denervated skeletal muscle in mice. Am J Physiol Cell Physiol. 289, C697–707 (2005).
Williams, M. D. et al. GPNMB expression in uveal melanoma: a potential for targeted therapy. Melanoma Res. 20, 184–190 (2010).
Kuan, C. T. et al. Glycoprotein nonmetastatic melanoma protein B, a potential molecular therapeutic target in patients with glioblastoma multiforme. Clin Cancer Res. 12, 1970–1982 (2006).
Rich, J. N. et al. Bone-related genes expressed in advanced malignancies induce invasion and metastasis in a genetically defined human cancer model. J Biol Chem. 278, 15951–15957 (2003).
Rose, A. A. et al. Osteoactivin promotes breast cancer metastasis to bone. Mol Cancer Res. 5, 1001–1014 (2007).
Onaga, M. et al. Osteoactivin expressed during cirrhosis development in rats fed a choline-deficient, L-amino acid-defined diet, accelerates motility of hepatoma cells. J Hepatol. 39, 779–785 (2003).
Tse, K. F. et al. CR011, a fully human monoclonal antibody-auristatin E conjugate, for the treatment of melanoma. Clin Cancer Res. 12, 1373–1382 (2006).
Anderson, M. G. et al. Mutations in genes encoding melanosomal proteins cause pigmentary glaucoma in DBA/2J mice. Nat Genet. 30, 81–85 (2002).
Furochi, H. et al. Osteoactivin fragments produced by ectodomain shedding induce MMP-3 expression via ERK pathway in mouse NIH-3T3 fibroblasts. FEBS Lett. 581, 5743–5750 (2007).
Rose, A. A. et al. ADAM10 releases a soluble form of the GPNMB/Osteoactivin extracellular domain with angiogenic properties. PLoS One. 5, e12093 (2010).
Liu, Y., Hao, W., Dawson, A., Liu, S. & Fassbender, K. Expression of amyotrophic lateral sclerosis-linked SOD1 mutant increases the neurotoxic potential of microglia via TLR2. J Biol Chem. 284, 3691–3699 (2009).
Shinder, G. A., Lacourse, M. C., Minotti, S. & Durham, H. D. Mutant Cu/Zn-superoxide dismutase proteins have altered solubility and interact with heat shock/stress proteins in models of amyotrophic lateral sclerosis. J Biol Chem. 276, 12791–12796 (2001).
Wang, J. et al. Copper-binding-site-null SOD1 causes ALS in transgenic mice: aggregates of non-native SOD1 delineate a common feature. Hum Mol Genet. 12, 2753–2764 (2003).
Urushitani, M. et al. CHIP promotes proteasomal degradation of familial ALS-linked mutant SOD1 by ubiquitinating Hsp/Hsc70. J Neurochem. 90, 231–244 (2004).
Kunst, C. B., Mezey, E., Brownstein, M. J. & Patterson, D. Mutations in SOD1 associated with amyotrophic lateral sclerosis cause novel protein interactions. Nat Genet. 15, 91–94 (1997).
Ciechanover, A. & Brundin, P. The ubiquitin proteasome system in neurodegenerative diseases: sometimes the chicken, sometimes the egg. Neuron. 40, 427–446 (2003).
Vargas, M. R., Pehar, M., Cassina, P., Beckman, J. S. & Barbeito, L. Increased glutathione biosynthesis by Nrf2 activation in astrocytes prevents p75NTR-dependent motor neuron apoptosis. J Neurochem. 97, 687–696 (2006).
Di Giorgio, F. P., Carrasco, M. A., Siao, M. C., Maniatis, T. & Eggan, K. Non-cell autonomous effect of glia on motor neurons in an embryonic stem cell-based ALS model. Nat Neurosci. 10, 608–614 (2007).
Nagai, M. et al. Astrocytes expressing ALS-linked mutated SOD1 release factors selectively toxic to motor neurons. Nat Neurosci. 10, 615–622 (2007).
Adema, G. J., Bakker, A. B., de Boer, A. J., Hohenstein, P. & Figdor, C. G. pMel17 is recognised by monoclonal antibodies NKI-beteb, HMB-45 and HMB-50 and by anti-melanoma CTL. Br J Cancer. 73, 1044–1048 (1996).
Berson, J. F., Harper, D. C., Tenza, D., Raposo, G. & Marks, M. S. Pmel17 initiates premelanosome morphogenesis within multivesicular bodies. Mol Biol Cell. 12, 3451–3464 (2001).
Fowler, D. M. et al. Functional amyloid formation within mammalian tissue. PLoS Biol. 4, e6 (2006).
Rose, A. A. et al. Glycoprotein nonmetastatic B is an independent prognostic indicator of recurrence and a novel therapeutic target in breast cancer. Clin Cancer Res. 16, 2147–2156 (2010).
Lasiene, J. & Yamanaka, K. Glial cells in amyotrophic lateral sclerosis. Neurol Res Int. 2011, 718987 (2011).
Yamanaka, K. et al. Astrocytes as determinants of disease progression in inherited amyotrophic lateral sclerosis. Nat Neurosci. 11, 251–253 (2008).
Ilieva, H., Polymenidou, M. & Cleveland, D. W. Non-cell autonomous toxicity in neurodegenerative disorders: ALS and beyond. J Cell Biol. 187, 761–772 (2009).
Ekestern, E. Neurotrophic factors and amyotrophic lateral sclerosis. Neurodegener Dis. 1, 88–100 (2004).
Dewil, M. et al. Vascular endothelial growth factor counteracts the loss of phospho-Akt preceding motor neurone degeneration in amyotrophic lateral sclerosis. Neuropathol Appl Neurobiol. 33, 499–509 (2007).
Tanaka, H. et al. Apoptosis-inducing factor and cyclophilin a cotranslocate to the motor neuronal nuclei in amyotrophic lateral sclerosis model mice. CNS Neurosci Ther. 17, 294–304 (2011).
Niwa, J. et al. Dorfin ubiquitylates mutant SOD1 and prevents mutant SOD1-mediated neurotoxicity. J Biol Chem. 277, 36793–36798 (2002).
Acknowledgements
The authors deeply appreciate the patients and families that participated in this study. We also thank Dr. Takashi Uehara (Department of Medicinal Pharmacology, Graduate School of Medicine, Dentistry and Pharmaceutical Sciences, Okayama University) for the useful advice about this study. This study was supported by grants from the Japan Amyotrophic Lateral Sclerosis Association (2011), the Adaptable and Seamless Technology Transfer Program through Target-Driven R&D (2011, Subject No. AS231Z00098F) and the Japan Society for the Promotion of Science (2011, Subject No. 23-8471).
Author information
Authors and Affiliations
Contributions
H.T. (Hirotaka Tanaka), M.S., K.T. and H.H. designed the experiments. H.T. (Hirotaka Tanaka), M.K. and M.T. performed the experiments. H.T. (Hirotaka Tanaka) performed the analysis and wrote the paper. J.N., Y.I. and G.S. contributed several SOD1 plasmid vectors. T.N. contributed the GPNMB transgenic mice. M.Y., H.T. (Hitoshi Takahashi), I.H. and T.I. contributed the clinical samples. All authors contributed to the editing of the paper and to scientific discussions.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
SUPPLEMENTARY INFORMATIONS
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Tanaka, H., Shimazawa, M., Kimura, M. et al. The potential of GPNMB as novel neuroprotective factor in amyotrophic lateral sclerosis. Sci Rep 2, 573 (2012). https://doi.org/10.1038/srep00573
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00573
This article is cited by
-
Progranulin and GPNMB: interactions in endo-lysosome function and inflammation in neurodegenerative disease
Journal of Neuroinflammation (2023)
-
GPNMB: a potent inducer of immunosuppression in cancer
Oncogene (2022)
-
Epigenetic inactivation of the autophagy–lysosomal system in appendix in Parkinson’s disease
Nature Communications (2021)
-
Neonatal hypoxia-ischemia in rat elicits a region-specific neurotrophic response in SVZ microglia
Journal of Neuroinflammation (2020)
-
GPNMB is expressed in human epidermal keratinocytes but disappears in the vitiligo lesional skin
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.