Abstract
Transient protein-protein interactions play crucial roles in all facets of cellular physiology. Here, using an analysis on known 3-D structures of transient protein-protein complexes, their corresponding uncomplexed forms and energy calculations we seek to understand the roles of protein-protein interfacial residues in the unbound forms. We show that there are conformationally near invariant and evolutionarily conserved interfacial residues which are rigid and they account for ∼65% of the core interface. Interestingly, some of these residues contribute significantly to the stabilization of the interface structure in the uncomplexed form. Such residues have strong energetic basis to perform dual roles of stabilizing the structure of the uncomplexed form as well as the complex once formed while they maintain their rigid nature throughout. This feature is evolutionarily well conserved at both the structural and sequence levels. We believe this analysis has general bearing in the prediction of interfaces and understanding molecular recognition.
Similar content being viewed by others
Introduction
Protein-protein interactions form one of the most important components of the cellular machinery in maintaining homeostasis1,2,3,4. A myriad of biophysical techniques5, ranging from X-ray crystallography, various spectroscopic techniques, cross-linking methods, mutation studies etc, have been employed to understand the atomic picture of the protein-protein interface. Many studies use datasets of known 3-D structures of proteins in complex and/or unbound states from the Protein Data Bank6 to understand specific aspects of the interface. Several excellent reviews on the structure of protein-protein interfaces provide a comprehensive survey of the different aspects of interfaces3,7,8,9. Protein-protein interfaces are characterized by several distinguishing features with respect to the rest of surface: surface planarity10, moderately enhanced residue conservation11,12, decreased flexibility13, predominance of aromatic residues14,15, modular architecture9,16, uneven distribution of binding energy17,18 and close-packing19. Apart from enhancing our understanding of molecular recognition at protein-protein interfaces, a combination of these features has been exploited for various purposes: distinguishing biologically relevant interfaces from crystal contacts20,21, discriminating obligate from transient interactions12,15,22, prediction of protein-binding sites23,24 and improved protein-protein docking25,26.
Knowledge of these features has enabled the understanding of the interface as a whole. The first study of interface architecture proposed the delineation of the interface into the core and rim area, the former consisting largely of buried atoms and the latter formed mainly by exposed atoms27. Another viewpoint (O-ring hypothesis) proposes the existence of a hot-spot enriched region at the centre shielded from water by an outer ring of non-conserved residues18. Structural analysis of several complexes indicates a modular organization of the interface: several hot-spot enriched complementary pockets present on the two chains binding to each other28,29. Modularity of the interface region was demonstrated conclusively by mutagenesis and X-ray crystallographic studies of TEM1-Barstar16.
Though the mechanism of molecular recognition has been studied for several years, the roles of interface residues have been focused primarily on the bound forms of the complexes. These studies show that interface residues form tightly packed interfaces with decreased flexibility in comparison to other surface residues13. The availability of structures of unbound forms of the interacting proteins and the bound forms of transient protein-protein complexes30,31 provides the opportunity to study the roles and behaviour of the protein-protein interface residues in the unbound form. MD-simulations of bound and unbound forms of transient complexes have shown that the core of the interface is less flexible than the periphery32, owing to the formation of hydrogen bonds33. Further, Rajamani et al pinpointed the presence of one or few ‘anchor’ interface residues in the unbound form34. Although these studies provide a structural basis of molecular recognition starting from the unbound form for diverse set of protein-protein complexes, the preponderance in the interface and evolutionary conservation of these features is largely unexplored. In our analysis, we investigate the different kinds of interfacial residues in the unbound form. We differentiate interfacial residues as rigid and non-rigid and also show the physicochemical basis of this classification using empirical free energy computations. Using homologues of known 3D structure, we demonstrate that the rigid interfacial residues are well conserved evolutionarily, in terms of nature, sequence and side-chain orientation even in the uncomplexed form.
Results
Comparison of B-factors of interfacial residues in unbound/bound forms of proteins and identification of a subset of rigid interfacial residues
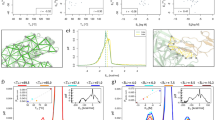
We analyzed the distribution of normalized crystallographic temperature factors (B-factors) of protein-protein interfacial residues in protein-bound and unbound forms of transient complexes (Figure 1a, also see Supplementary Table S1 online). We found that the interfacial residues in the bound form have lower B-factors as expected due to the increased burial upon protein binding (Unbound vs. bound for All-Int Paired t-test: t = 15.93; df = 1087; P = 1.65E−51; Unbound vs. bound for Core-Int Paired t-test: t = 12.39; df = 427; P = 2.48E−30) in comparison to the rest of the tertiary structural surface (Unbound vs. bound for NISurf Paired t-test: t = 3.24; df = 8204; P = 1.20E−3) (Figure 1a). Although in general, the distribution of B-factors of interfacial residues in the unbound form is not significantly different from that of solvent exposed surface residues (Unpaired t-test: t = 1.637; df = 9692; P = 1.02E−1) (Figure 1a), the trend is opposite when the core interfacial residues was compared to the solvent-exposed surface residues (Unpaired t-test: t = 6.981; df = 9027; P = 3.14E−12) (Figure 1a). While we noted interface-residues with low and high B-factors in the unbound form, interestingly, despite being exposed to solvent, a substantial proportion of interface residues in the protein-unbound form show normalized B-factors comparable to those observed for buried residues in protein tertiary structures. Using a cut-off value of 0.04, corresponding to the upper 90 percentile value of normalized B-factors for buried residues (See methods), all surface residues in the unbound form were classified into Rigid (R) and non-Rigid (NR) residues. The change in the normalized B-factors between protein-bound and free forms for the rigid interfacial residues is very small in comparison to the non-rigid interfacial residues (Figure 1b). We find that the proportion of rigid residues is highest in the core of the interface (65.42%; n = 428 sites). This proportion is less for all the interface residues considered together (54.25%; n = 1093 sites) and the rest of the surface (50.61%; n = 8601 sites). To assess if the low normalized B-factors of surface residues from unbound forms alone can distinguish the core interfacial residues from the rest of the surface residues, we measured sensitivity and specificity values at various cut-off values and represented the results in terms of ROC curve (See Supplementary methods). We show that the signal provided by temperature factors is modest with Area Under the Curve (AUC) of 0.62 (See Supplementary fig. S3 online). As expected from the fact that only a subset of interfacial residues is rigid the AUC for distinction between all the interfacial residues and non-interfacial surface residues is lower (0.53). This corroborates the previous analysis on the choice of cut-off of 0.04 closer to the optimal −0.25 (See Supplementary fig. S3 online). These AUC values re-emphasize that only a sub-set of interfacial residues correspond to low normalized B-factors. The rigid residues are more clustered in the interface than on the rest of the surface making this feature specific to the interface (Figure 1c). This clustering was obtained as the proportion of rigid surface residues surrounding any given residue with a Cα—Cα distance ≤ 9.0 Å (See methods). We found more rigid surface residues around any given rigid interface residue (6.06 ± 2.3) than non-rigid interface residue (3.00 ± 2.2) and surface residues (4.09 ± 2.8) (Figure 1c). We also found that the difference in the surface exposure for the rigid interface residues in the unbound and fictitiously unbound form, (i.e. the surface exposure of the same residue taken from complex structure without the interacting partner protein) is low (|ΔASA| = 9.677 ± 12.1 Å2) in comparison to non-interacting rigid residues (|ΔASA| = 18.25 ± 26.0 Å2) indicating that there is little or no change in the conformation to show a change in surface exposure (Figure 1d). This result is further corroborated with the observation of low local RMSD values for rigid interface residues (mean RMSD = 0.85 ± 0.62 Å) in comparison to non-rigid interface residues (mean RMSD = 1.15 ± 0.64 Å) when comparing bound and unbound forms of proteins (Supplementary fig. S1 online). However, we note that rigid and non-rigid residues at the core of the interface undergo comparable changes in RMSD upon binding (0.82 ± 0.64 Å for R and 0.90 ± 0.55 Å for NR). An example of a transient complex of β-Actin and Profilin is shown in the bound and unbound forms along with structures of homologous proteins (Figure 2). We highlight the rigid and non-rigid interface residues and their side-chain orientations in the bound, unbound and homologous proteins structures (Figure 2).
Identification and description of rigid and non-rigid interfacial residues.
(a) Distribution of normalized B-factors showing mean ± s.e.m. for various residues of bound (shaded, B) and unbound (U) forms of proteins. (b) Distribution of normalized B-factors showing mean ± s.e.m. for rigid (R) and non-rigid (NR) residues of interface (All Int) and core interface (Core Int) in bound (shaded, B) and unbound (U) forms of proteins. (c) Distribution of clustering of rigid surface residues within 9.0 Å of any given surface residue (All Int, Core Int and NISurf) in unbound form. (d) Absolute change in accessible surface area for rigid and non-rigid residues in interface and core interface between true unbound form and fictitious unbound form denoting a measure of change in conformation. * and ** denotes significance at α of 0.05 and 0.01 respectively. n.s. denotes P > 0.05.
Rigid and non-rigid interface residues of Profilin involved in β-actin-Profilin complex.
(a) The binding mode of β-actin (red) with Profilin (blue) highlighting the side-chain conformations of interfacial residues in Profilin (orange sticks) (PDB accession id: 2BTF). The Profilin molecule has been translated by 10 Å away from the interfacial axis to show the binding mode (b) All interface residues (marine blue sticks) along with core interface residues (orange sticks) in Profilin molecule are shown. (c) Rigid interacting residues (red sticks) and non-rigid interacting residues (marine blue sticks) in the unbound form of Profilin (PDB accession id: 1PNE) are shown. (d) The structural alignment of unbound form (grey) and bound form (green) of Profilin showing the lower structural variation of rigid interfacial residues (red) than non-rigid interfacial residues (blue). (e) Structural alignment of unbound form (grey) with two homologues of Profilin (teal and pale yellow; PDB accession id: 1FIL & 2VK3) showing the lower structural variation in rigid interfacial residues (red) in comparison to non-rigid interfacial residues (blue).
Physicochemical basis of rigidity
We performed empirical free energy computations to understand the basis for the demarcation of interfacial residues into rigid and non-rigid residues. We found that the free energy contribution of rigid residues towards the overall stability in the unbound form (0.013 ± 1.13 Kcal/mol) was more favourable than that of non-rigid residues (0.58 ± 0.94 Kcal/mol) (Unpaired t-test: t = 8.838; df = 1087; P = 3.85E−18) (Figure 3a). The origin of this differential contribution of free energy in the unbound form also stems from the different micro-environments of the rigid and non-rigid interfacial residues. The rigid residues are generally well packed (0.56 ± 0.06 for rigid; 0.51 ± 0.06 for non-rigid; Unpaired t-test: t = 12.77; df = 1087; P = 6.63E−35) (Figure 3b) and more buried (36.32 ± 23.13 for rigid; 52.55 ± 23.95 for non-rigid; Unpaired t-test: t = 11.37; df = 1087; P = 2.17E−28) (Figure 3c) in comparison to non-rigid interfacial residues. The rigid residues of the interface have a free energy contribution which is intermediate to that of the surface and buried residues (Figure 3a), as opposed to the non-rigid interface residues whose contribution is less than that of the surface residues (Unpaired t-test: t = 8.25; df = 9000; P = 1.75E−16) (Figure 3a). Both the rigid and non-rigid residues contribute favourably towards the stability of the complex upon binding (Bound form contribution for rigid = −0.61 ± 1.34 Kcal/mol; non-rigid = −0.09 ± 1.32 Kcal/mol) (Figure 3a). In-order to understand the origins of stability contributed by rigid residues in the unbound forms, we analyzed the enthalpy and entropy contributions of these observed free energies. Though the side-chain entropy contribution for the stability is slightly less for the rigid residues (0.49 ± 0.41 Kcal/mol) than the non-rigid residues (0.40 ± 0.38 Kcal/mol) (Unpaired t-test: t = 3.72; df = 1087; P = 2.09E−4) (Figure 4a), the enhanced enthalpy contribution (Figure 4b) more than compensates for the overall loss of conformational entropy in the unbound form (rigid = −1.87 ± 3.06 Kcal/mol; non-rigid = −1.25 ± 2.78 Kcal/mol). Upon complexation the rigid interface residues undergo minimal loss of entropy (rigid = 0.29 ± 0.64 Kcal/mol) in comparison to the non-rigid residues (non-rigid = 0.53 ± 1.00 kcal/mol), which become ordered (Figure 4a). The change in the enthalpy contribution is more or less the same for the rigid and non-rigid residues upon binding (rigid = −0.78 ± 2.33 Kcal/mol; non-rigid = −0.88 ± 3.12 Kcal/mol) (Figure 4b).
Features of free energy effects and microenvironment of rigid and non-rigid interfacial residues.
(a) The free energy contribution of different interface residue types in the unbound (U) and bound (B, grey) forms are shown as box plots. The buried (Bur) and non-interacting surface residues (NISurf) residues serve as control data sets. The distributions of (b) residue packing density and (c) RSA in unbound form for different interface residue types in the unbound form are shown as box plots. Buried (Bur) and non-interacting surface residues (NISurf) residues serve as control data sets. Mean values are indicated by ‘+'. The significantly differential contributions of the rigid (R) and non-rigid (NR) interacting residues in the unbound form are indicated. ** denotes significance at α of 0.05.
Distribution of enthalpic and entropic contributions of rigid/non-rigid interface residues towards stabilization of bound/unbound forms.
(a) Distribution showing mean ± s.d. of main-chain and side-chain entropic contribution to free energy for rigid (R) and non-rigid (NR) interfacial residues in the unbound (U) and bound (B, grey) forms. (b) The distribution of mean ± s.d. of all the enthalpic (hydrophobic solvation potential (SolvH), polar solvation potential (SolvP), back-bone hydrogen bond (BkHb), side-chain hydrogen bond (ScHb) and van der Waals (vdW) contributions to free energy for rigid (R) and non-rigid (NR) interfacial residues in the unbound (U) and bound (B, grey) forms. * and ** denote significance at α of 0.05 and 0.01 respectively. n.s. denotes P > 0.05.
The hotspot residues in the protein-protein interfaces contribute highly to the stabilization of the protein-protein complex. However the subset of protein-protein interfacial residues discussed in the current analysis are rigid and they need not correspond to hotspots. However it is interesting to address the question if some of these rigid residues also correspond to hotspots. To determine, the relationship between rigid residues of the interface and interaction hotspot residues, we assessed the overlap between the list of rigid residues and the experimentally determined hotspot residues (See Supplementary methods & Supplementary table S5 online). The proportion of rigid interfacial residues among hotspot residues (29.41%; n = 34) was low as opposed to non-hotspot residues (46.33%; n = 218). Although there was no clear relationship between the B-factors and the tendency to be a hotspot residue, we found considerable overlap of these two classes (See Supplementary fig. S4 online).
Most interface residues contribute towards stabilizing the bound form. However interestingly we observed that some of the interfacial residues predominantly stabilize the self-protein i.e. the protein in which they are situated (−1.55 ± 0.87 Kcal/mol) and have a negligible contribution towards stabilization of the bound form (−0.17 ± 0.72 Kcal/mol) (Figure 5, see Supplementary methods and Supplementary fig. S2 online). Thus, though these residues are located in the protein-protein interface their main role seems to be in the stabilization of the self-protein both in the unbound and bound forms. We classified these residues as Self-protein Stabilizing Residues (SSR) (6.93%; n = 75) and the rest as Neutrally Stabilizing Residues (NSR) (42.60%; n = 461) and Complex Stabilizing Residues (CSR) (50.46%; n = 546) based on the extent of change in energy contribution towards the unbound form and the complex form. This classification of interfacial residues is independent of the rigid/non-rigid classification and is solely based on the residue level energetic contributions. It is important to note that, the proportion of rigid residues is more in SSR (73.33%; n = 55) than in NR (58.13%; n = 268) and CSR (48.90%; n = 267) sites (Figure 5). These results suggest that energetic differences in rigid and non-rigid interfacial residues play a crucial role in complexation.
The classification of protein-protein interfacial residues.
The protein-protein interfacial residues have been classified in two different ways. Using crystallographic temperature factors they are classified as rigid and non-rigid. Independently they are also classified as Self-protein stabilizing (SSR), Neutrally stabilizing residues (NSR) and Complex stabilizing residues (CSR) depending on the energetic contribution of the residue in stability of the structure of the protein which it is present and that of the protein-protein complex. The numbers of residues in various categories are indicated in the figure.
Evolutionary aspects of interface rigidity
To explore if the physicochemical basis of rigid and non-rigid nature of residues has a selective advantage in the evolution of protein-protein interactions, we computed the conservation of these sites using information from both structures and sequences of homologues of interacting proteins (See methods, Supplementary table S2 online). We find that rigid nature of interfacial residues was well conserved (70.01% conservation; n = 474) at topologically equivalent positions in high resolution structures of unbound forms of homologues. For the same set of interfacial residues we found a higher proportion of sites with identical amino acid (75.77%; n = 513). In these sites, the rigid nature is 73.09% conserved (n = 375). For the set of substituted interfacial residues (24.22%; n = 164) we found that the rigid nature is again conserved to a higher extent (60.36%; n = 99). We also found that the rigid interfacial residues are significantly conserved better (Unpaired t-test: t = 2.987; df = 587; P = 2.93E−3) than the non-rigid interfacial residues in homologous sequences indicating a selective constraint at these interface positions (Figure 6a). Further, we observed that the variation in side-chain orientations (RMSD) of identical rigid interface residues in structures of unbound forms of homologous proteins is significantly lower (0.76 ± 0.54 Å local RMSD) than that of the non-rigid interfacial residues (0.85 ± 0.58 Å local RMSD) (Unpaired t-test: t = 3.45; df = 1770; P = 5.65E−04) (Figure 6b). It should be noted that in the unbound forms both rigid and non-rigid residues are exposed to the solvent and despite this solvent exposure rigid residues show high conservation of side-chain orientation. A small subset of rigid interfacial residues (15.5%; n = 92) showed very high specificity in side-chain orientation (i.e. local RMSD ≤ 0.50Å). We refer these residues as rigid specific residues (RS). They also retain their side-chain orientation in the bound form (0.419 ± 0.28 Å local RMSD for all interface) (Figure 6c). These sites show very high residue conservation (0.74 ± 0.86 normalized conservation score) in homologous sequences (Figure 6a) indicating enhanced physicochemical and evolutionary constraints. The proportion of residues showing high specificity in side-chain orientation in non-rigid interfacial (NRS) residues is substantially low (3.6%; n = 18).
Evolutionary conservation of rigid/non-rigid interface residues in sequence and structural homologues.
(a) The distribution of normalized conservation scores, computed as Jensen-Shannon divergence measure, for different residues types is shown as box-plots. The enhanced conservation scores for rigid specific (R-Spec) interacting (Int) and core-interacting (C-Int) residues are highlighted. Buried (Bur) and non-interacting surface residues (NISurf) residues serve as control data sets. (b) The extent of structural change in side-chains of identical residues (RMSD) in structures of unbound forms and their homologous proteins is shown using box plots for different residue types. Buried (Bur) and non-interacting surface residues (NISurf) residues serve as control data sets. (c). The extent of structural change in side-chains of identical residues (RMSD) in the structures of unbound and bound forms (grey) is shown for rigid specific (RS) and non-rigid specific (NRS) interacting residues in comparison with all rigid (R) and non-rigid (NR) interacting residues using box plots. Mean values are indicated by ‘+’. The significantly differential contributions of the rigid (R) and non-rigid (NR) interacting residues in the unbound form are indicated. ** denotes significance at α of 0.05. n.s. denotes P > 0.05.
Assessment of potential of analyzed structural features in prediction of subset of interfacial residues
We used rigidity and the above mentioned features of rigid interfacial residues to predict subset of interface residues from the structures of unbound forms alone to understand the potential of these features in contributing towards prediction of interfacial residues (See Supplementary methods online). We show more rigid interface residues can be predicted (Coverage = 30.55±19.1% of the total interface) when we use a combination of B-factors and residue sequence conservation score in comparison to a combination using B-factors and preservation of side-chain conformation (Coverage = 12.32±10.8% of the total interface) (See Supplementary table S6 online). Although the overall accuracy of the interface prediction is low, we observe that use of sequence conservation in addition to temperature factors is better in comparison to the use of conservation of side-chain conformation (See Suppelementary table S6 online). Among the interfaces predicted for 7 proteins, the interacting surface residues of Tripsinogen and Carboxypeptidase A are predicted with relatively high coverage (47.36%; n = 9 and 66.66%; n = 8 respectively) and accuracy (26.47% and 18.18%) using a combination of rigidity and conservation alone. The structural properties analyzed in this work, therefore, carry reasonable potential for prediction. These parameters are likely to be more effective in prediction of interface when used along with other parameters such as extent of their conservation, propensity of the residues to be in the interface, spatial clustering of rigid residues and consideration of geometry (flatness) of the putative interface.
Discussion
Most of the signalling and regulatory proteins, involved in a large number of cellular processes, participate in transient protein-protein interactions12,30,31. The availability of structures of stable unbound forms of the interacting proteins and bound forms of these transient protein-protein complexes provides an opportunity to study the behaviour of the residues participating in interaction before the protein complexation event. Using a high resolution dataset of unbound forms of a diverse non-redundant dataset of 67 protein structures, we observe significantly low normalized B-factors at the core of the interface in the unbound form, indicating higher rigidity (Figure 1a). This result is concurrent with a study on MD-simulations of the unbound forms of proteins corresponding to 22 protein-protein complexes, which showed that core interfacial residues exhibit lower flexibility for substantial proportion of simulation time than peripheral interface residues32. Using a non-redundant dataset of high-resolution structures of unbound forms of homologues, we also show that comparative B-factor analysis provides evidence for two different subsets of interfacial residues (rigid/non-rigid) (Figure 1b). We show that these two classes are differentially conserved in evolution using a dataset of homologous protein sequences and structures (Figure 6). The rigid interfacial residues show enhanced conservation than non-rigid residues in homologous proteins (Figure 6a). Although the rigid nature of interface residues in homologous structures is conserved better for identical residues (73.09%), it is also well conserved at substituted sites (60.36%). This indicates that rigidness is a property of the topological location of the residue at the interface and not the identity of the amino acid present at that site. Several observations substantiate this argument. Rigid interfacial residues are well packed (Figure 3b) and clustered together (Figure 1c). The tightly packed and clustered microenvironment of rigid interface residues places increased constraints on side-chain motion of these residues. Propensity of interfacial rigid and non-rigid residues is not very different for all residue types (see Supplementary Table S1 online) further reiterating that rigidness here is not the property of individual side-chains. This enormous constraint on a surface residue to be rigid might be expected to be unfavourable energetically. Most of the surface residues are flexible due to high side-chain and main-chain entropies. On the contrary rigid interfacial residues in the unbound forms are stabilized by the high enthalpic contributions (Figure 4b), owing to their well packed and slightly buried microenvironments. This feature more than compensates for the loss of conformational entropy in the unbound forms and aids in the overall stabilization.
The rigidity also manifests in terms of low structural variation of these residues upon complexation (see Supplementary fig. S1 online). This feature contrasts with the general behaviour of interface residues, which are usually the most altered set of residues upon complexation (see Supplementary fig. S1 online). We also note that the side-chain orientation is less altered in the high-resolution structures of unbound forms of homologues for rigid interfacial residues in comparison to non-rigid interfacial residues (Figure 6b). In particular, rigid interfacial residues contain a small subset of residues which are highly specific in their side-chain orientation (i.e. RMSD ≤ 0.5Å), both in unbound forms of homologues and in the bound form (Figure 6b and 6c). Their high specificity and high sequence conservation (Figure 6a) indicates that they may be crucial in complex formation. We believe these residues possibly correspond to the ‘anchor' interface residues identified by Rajamani et al. They pinpointed the presence of one or few ‘anchor’ interface residues in the unbound form, serving as pre-made recognition motifs34. The very high sequence conservation of these residues coupled with retention of side-chain orientation in unbound forms of homologues also suggests that some of these residues are similar to the set of hot-spot residues studied by Li et al., which are clustered in complemented pockets at the binding interface and are pre-organized in the unbound form29. Our analysis corroborates with Yogurctu et.al.'s study showing the overlap between these various sets of interfacial residues33.
We note that some of the interface residues (Self-protein stabilizing residues - SSR) and neutrally stabilizing residues (NSR), whose energetic contribution is greater (SSR) or moderate (NSR) towards the stabilization of the unbound protein vis-a-vis the protein complex, are enriched in rigid sites (Figure 5, see Supplementary methods, Supplementary discussion online). We note for the first time that most of the rigid interfacial sites make a more favorable energetic contribution to the free energy of the unbound form (Figure 3a) than non-rigid residues. This suggests that these rigid residues, apart from contributing to complex formation, play a more important role in the stabilization of the unbound form of the protein (Figure 3a).
The conformational side-chain entropy of a subset of interfacial residues (Rigid) is very low and that of the others (Non-rigid) is high (Figure 4a). This striking feature has two consequences for the binding process. Upon binding the conformational entropy of side-chains and main-chain atoms decreases. This decrease (in ΔS) contributes negatively for the free energy for binding. This difference (ΔS) in very different for the two sets of interface residues. The rigid residues undergo a minimal change in the entropy, enabling a favourable free energy for binding. The non-rigid residues have a large change in conformational entropy which is compensated by the enthalpy interactions across the interface. Understanding these energetic contributions of interfacial residues, calculated for a substantial number of protein-protein complexes, confirms earlier claims33,34, based on specific protein-protein complexes, that the overall reduced loss of conformational entropy (specific to side-chains) upon complexation (Figure 4a) owing to the significant proportion of rigid residues, plays a pivotal role in the energetics of complex formation. Different enthalpic energy terms, such as hydrogen bonds33, van der Waals interactions and solvation energy, appear to be important contributors to the favorable free energy contribution made by rigid residues in the unbound form, in spite of their higher entropic cost (Figure 4b). Although this entropic cost is high, the change in the entropy is low as explained above, making the overall binding process favourable. The strength of the overall binding free energy depends on the balance of the enthalpic and entropic contributions. Although the rigid interface residues have low ΔS upon binding, this might be just barely sufficient to reduce the overall free energy of binding.
In the case of hotspot residues of the interface, the overall binding free energy is very high (>2.0 kcal/mol). For a rigid residues to be a hotspot residue, the reduced entropy loss i.e. -TΔS should be at least 2.0 Kcal/mol greater than the net change in enthalpy (ΔH). This is less frequent, as we found only ∼30% overlap between the hotspot and rigid interface residues. For majority of the rigid residues (∼70%) the TΔS although is negative, is not greater than the threshold value to be identified as a hotspot residue.
The differential behaviour of rigid and non-rigid interfacial residues in the unbound forms and their varied entropic effects gives these two sets of residues varied roles in the interface prior to binding. The rigid interfacial residues provide stability to the unbound form and maintain a surface structure congenial for molecular recognition. The non-rigid residues undergo a lot of changes in side-chain orientation and help in adapting to the new micro-environment formed upon binding. This also increases the overall free-energy of binding. These contrasting features suggests that the classification of the interface residues into rigid and non-rigid has a strong energetic basis and the two sets of residues function differently as they have a differential contribution towards the energetics of complex formation.
In conclusion, rigid interfacial residues form a substantial proportion of core interfacial residues; they contribute significantly to the molecular recognition process by reducing the entropic cost on complexation by virtue of their pre-ordered conformation. We have shown for the first time that some of the rigid interfacial residues stabilize the protein-unbound conformation more than the complex despite being in the protein-protein interface. This suggests that not all the protein-protein interfacial residues have the major role of stabilizing the complex; some of these residues seem to have more significant role in the unbound form than the bound form. The rigid interfacial residues are under strong evolutionary selection to retain the transient functional interactions vital in the cellular context. This feature is highlighted by observed substantial conservation of microenvironment (rigidness), sequence (amino acid identity/similarity) and structure (specific side-chain orientation) in homologous proteins. We showed close to ∼30% coverage in the prediction of interfaces using temperature-factors and sequence conservation alone. A combination of such features along with amino-acid propensity and neighbourhood information can be instrumental in enhancing the coverage and accuracy of prediction of interfacial residues. These features can supplement the current methods employed in developing molecular docking tools. Maintenance of rigidity and side-chain orientation of some of the core interfacial residues attracts the possibility of designing small molecules to occupy these residues thus, potentially preventing protein-protein interactions. The retention of orientation of these residues in the complexed and free forms suggests that the significant conformational changes at the ligand binding site are unlikely which is an advantage in the design of small molecules.
Methods
Dataset of 3D-structures of bound & unbound forms of transient protein-protein complexes
A curated dataset of structures of proteins involved in transient interactions, solved in both unbound and bound forms, were taken from Benchmark4 dataset35. Out of the 176 transient protein-protein complexes available, only those structures of unbound forms solved at a resolution better than 2Å were considered for the analysis. Further, only entries containing single chain in asymmetric unit and biological unit, with no other macromolecular ligand bound were considered, to ensure that there was no bias due to crystal contacts and ligand-binding. This dataset was further pruned by removing entries belonging to the class of antigen-antibody interactions owing to their specialized nature of interaction. The remaining entries were clustered at 25% sequence identity using BLASTCLUST algorithm (http://www.csc.fi/english/research/sciences/bioscience/programs/blast/blastclust) to remove redundant sequence information. Finally, a non-redundant dataset of 67 structures of unbound forms solved at high resolution was obtained. This dataset consists of proteins performing diverse functions, ranging from enzyme-substrates/inhibitors, signalling proteins and other proteins involved in cellular processes. Interacting proteins of each binary complex of dataset are non-redundant at the level of their SCOP families36. The PDB accession codes for the high-resolution unbound forms, the corresponding bound forms and the interacting partner in the bound form are provided in Supplementary Table S1 online.
Dataset of 3D-structures of homologous proteins in unbound form
A repository of high-resolution crystal structures of single-chain protein entries was generated by mining the PDB6 using the following criteria: presence of only a single polypeptide chain in the asymmetric and biological unit and crystallographic resolution ≤ 2Å. Each of the 67 entries in the main dataset was queried against the repository of high resolution single chain protein structures using BLAST37 at an E-value cut-off of 10−6 with the low complexity regions masked. We ensured that a non-redundant set of homologues structures were picked up for each query by filtering the hits using the following criteria: sequence identity range 40–70%; query and hit coverage ≥ 80%. We identified high-resolution structures of homologous proteins solved in unbound form for only 24 entries. The homologous sequences were further clustered at a sequence identity of 70% using BLASTCLUST to generate a non-redundant set. As there was a variation in the number of homologous proteins identified for each of the 24 entries, we used an upper cut-off of 15 homologous entries per protein to reduce sampling bias. The final dataset had 115 pair-wise combinations of high resolution structures of unbound forms with their homologues. PDB accession codes of these pairwise combinations are provided in Supplementary Table S2 online.
Identification of buried, surface and interacting residues in bound and unbound forms
The residues were classified into the following categories: buried, surface, interacting, core-interacting, non-interacting surface. Buried and Surface residues were identified in both forms by computing accessibility using NACCESS38,39 algorithm. Residues with accessibility ≤ 5% and ≥ 10% were considered buried and surface residues respectively. These cut-offs were previously optimized40 and used to define buried residues in monomeric proteins. Interfacial residues in a complex were identified by considering the inter-atomic distances between proteins of the complex. Residues across the interface with distances below a cut-off have been considered as interacting. The cut-off distance is computed as the sum of van der Waal’s radii of interacting atoms plus 0.5Å41. The van der Waal’s radii for the protein atoms were taken from Chothia (1975)42. The interacting residues which undergo a large change in the accessibility upon complexation are considered as ‘core-interacting residues’. These residues are identified on the basis of their accessibility values: residues with accessibility ≥ 10% in the unbound form and accessibility ≤ 7% in the bound are considered to be the ‘core interacting residues'43. The core-interacting residues form a subset of the interacting residues of a complex. Surface residues other than the interacting residues, i.e. non-interacting surface residues were also identified.
B-factor analysis and identification of rigid/non-rigid nature of a residue
The B-factor (atomic displacement factor) of an atom reflects the degree of isotropic smearing of electron density around its center44. The measure of flexibility/rigidity of a particular residue in a structure is provided by its normalized backbone B-factor45. Normalization with respect to all the other residues provides an idea of increase/decrease in flexibility on a common scale. Only surface residues (i.e. accessibility ≥ 10%) were considered for the normalization. The three most N-terminal and C-terminal surface residues were excluded since their B-factors are usually high and can affect the ‘mean’ of the values. The normalized B-factor per residue (Bi,N) was computed as  where Bi is the B-factor of residue i, <Bi> is the mean B-factor of the protein surface residues and σ Bi is the s.d. for the same. Buried residues have the lowest Bi,N values for a given protein structure. We used an upper 90 percentile value (0.04) corresponding to the distribution of Bi,N values of buried residues to demarcate rigid from non-rigid surface residues. All interface, core-interface and non-interacting surface residues with a Bi,N ≤ 0.04 were considered rigid and otherwise non-rigid in this analysis. The same was also extended to the dataset of homologous protein structures.
where Bi is the B-factor of residue i, <Bi> is the mean B-factor of the protein surface residues and σ Bi is the s.d. for the same. Buried residues have the lowest Bi,N values for a given protein structure. We used an upper 90 percentile value (0.04) corresponding to the distribution of Bi,N values of buried residues to demarcate rigid from non-rigid surface residues. All interface, core-interface and non-interacting surface residues with a Bi,N ≤ 0.04 were considered rigid and otherwise non-rigid in this analysis. The same was also extended to the dataset of homologous protein structures.
Computation of free energy for bound/unbound protein forms
We used an empirical effective energy function, FoldX46,47, to compute the free energies of proteins in bound and unbound states. We obtained residue level contributions to the overall free energies from computations on 67 sets of high resolution 3D structures of bound, unbound and fictitiously unbound forms. We also obtained energies corresponding to different energy terms, including all enthalpic and entropic contributors, as FoldX is an additive function of all these terms.
Residue microenvironment
The microenvironment of a residue has been studied both in terms of its extent of exposure to solvent molecules and local packing density. The solvent accessibility of a molecule is computed using NACCESS38,39. Both absolute accessible surface area (ASA) and relative surface accessibility (RSA) have been computed. The local packing density for each residue is calculated using Voronoia program suite48. It employs Voronoi tessellation49 to assess the spatial proximity of atoms in 3Dspace. The packing density for an atom was computed as the fraction of the voronoi polyhedron volume occupied by the van der Waal's volume of an atom. The density of rigid surface residues around a given residue was computed using the following formula:

Where, FiR is the fraction of rigid surface residues within a Cα―Cα distance of 9.0 Å from the residue i.
Quantification of residue conservation in sequences
The degree of conservation for all the sites in a protein family was calculated using the Jensen-Shannon divergence measure50. This metric operates on the premise that most sites in a protein family are not under any evolutionary pressure and hence have a distribution similar to background amino acid distribution. Sites under evolutionary pressure, such as those contributing to function and/or stability, show amino acid distribution significantly different from the background distribution. Homologous sequences for every protein in our PPC dataset were identified by a search employing PSI-BLAST37 against the UNIREF9051 database at an E-value cut-off of 10−4 for 3 iterations. Further, only sequences with ≥ 30% identity with the query sequence and ≥ 70% length coverage with the query and hit sequences were retained to avoid false positives. Only proteins with ≥ 10 homologous sequences were considered for further analysis, resulting in a final dataset of 37 proteins. A multiple sequence alignment (MSA) of the query sequence with the homologous sequences was generated using CLUSTALW52. The conservation scores for every position in the MSA was calculated using Jensen-Shannon divergence measure50. The conservation scores range from 0–1.
Quantification of conservation of structural features
In order to estimate the extent of variation in side-chain orientation in the bound form as well as in the structures of unbound forms of homologues, pairwise structural alignments of unbound-bound forms and unbound form in main dataset – unbound forms of homologues were performed using DALI53,54. The extent of structural change for the topologically equivalent and identical residues was computed using all-atom local root mean square deviation (RMSD). The extent of conservation of rigid/non-rigid nature in structures of unbound forms of homologous proteins was computed for topologically equivalent positions of the interface.
Statistical analysis
All variables compared were tested for normality using Kolmogorov-Smirnov test. We used paired and unpaired student's t-tests for comparing the distributions. The complete details of various parameter comparisons and features are listed in Supplementary table S4 online.
References
Janin, J. & Wodak, S. J. Protein modules and protein-protein interaction. Introduction. Adv Protein Chem 61, 1–8 (2002).
Levy, E. D. & Pereira-Leal, J. B. Evolution and dynamics of protein interactions and networks. Curr Opin Struct Biol 18, 349–57 (2008).
Reichmann, D., Rahat, O., Cohen, M., Neuvirth, H. & Schreiber, G. The molecular architecture of protein-protein binding sites. Curr Opin Struct Biol 17, 67–76 (2007).
Vidal, M., Cusick, M. E. & Barabasi, A. L. Interactome networks and human disease. Cell 144, 986–98 (2011).
Lakey, J. H. & Raggett, E. M. Measuring protein-protein interactions. Curr Opin Struct Biol 8, 119–23 (1998).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res 28, 235–42 (2000).
Nooren, I. M. & Thornton, J. M. Diversity of protein-protein interactions. Embo J 22, 3486–92 (2003).
Janin, J., Bahadur, R. P. & Chakrabarti, P. Protein-protein interaction and quaternary structure. Q Rev Biophys 41, 133–80 (2008).
Keskin, O., Gursoy, A., Ma, B. & Nussinov, R. Principles of protein-protein interactions: what are the preferred ways for proteins to interact? Chem Rev 108, 1225–44 (2008).
Jones, S. & Thornton, J. M. Principles of protein-protein interactions. Proc Natl Acad Sci U S A 93, 13–20 (1996).
Choi, Y. S., Yang, J. S., Choi, Y., Ryu, S. H. & Kim, S. Evolutionary conservation in multiple faces of protein interaction. Proteins 77, 14–25 (2009).
Mintseris, J. & Weng, Z. Structure, function and evolution of transient and obligate protein-protein interactions. Proc Natl Acad Sci U S A 102, 10930–5 (2005).
Jones, S. & Thornton, J. M. Protein-protein interactions: a review of protein dimer structures. Prog Biophys Mol Biol 63, 31–65 (1995).
Lo Conte, L., Chothia, C. & Janin, J. The atomic structure of protein-protein recognition sites. J Mol Biol 285, 2177–98 (1999).
De, S., Krishnadev, O., Srinivasan, N. & Rekha, N. Interaction preferences across protein-protein interfaces of obligatory and non-obligatory components are different. BMC Struct Biol 5, 15 (2005).
Reichmann, D. et al. The modular architecture of protein-protein binding interfaces. Proc Natl Acad Sci U S A 102, 57–62 (2005).
Clackson, T. & Wells, J. A. A hot spot of binding energy in a hormone-receptor interface. Science 267, 383–6 (1995).
Bogan, A. A. & Thorn, K. S. Anatomy of hot spots in protein interfaces. J Mol Biol 280, 1–9 (1998).
Sonavane, S. & Chakrabarti, P. Cavities and atomic packing in protein structures and interfaces. PLoS Comput Biol 4, e1000188 (2008).
Bahadur, R. P., Chakrabarti, P., Rodier, F. & Janin, J. A dissection of specific and non-specific protein-protein interfaces. J Mol Biol 336, 943–55 (2004).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J Mol Biol 372, 774–97 (2007).
Zhu, H., Domingues, F. S., Sommer, I. & Lengauer, T. NOXclass: prediction of protein-protein interaction types. BMC Bioinformatics 7, 27 (2006).
Ezkurdia, I. et al. Progress and challenges in predicting protein-protein interaction sites. Brief Bioinform 10, 233–46 (2009).
Tuncbag, N., Kar, G., Keskin, O., Gursoy, A. & Nussinov, R. A survey of available tools and web servers for analysis of protein-protein interactions and interfaces. Brief Bioinform 10, 217–32 (2009).
Janin, J. Protein-protein docking tested in blind predictions: the CAPRI experiment. Mol Biosyst 6, 2351–62 (2010).
Vajda, S. & Kozakov, D. Convergence and combination of methods in protein-protein docking. Curr Opin Struct Biol 19, 164–70 (2009).
Chakrabarti, P. & Janin, J. Dissecting protein-protein recognition sites. Proteins 47, 334–43 (2002).
Keskin, O., Ma, B. & Nussinov, R. Hot regions in protein--protein interactions: the organization and contribution of structurally conserved hot spot residues. J Mol Biol 345, 1281–94 (2005).
Li, X., Keskin, O., Ma, B., Nussinov, R. & Liang, J. Protein-protein interactions: hot spots and structurally conserved residues often locate in complemented pockets that pre-organized in the unbound states: implications for docking. J Mol Biol 344, 781–95 (2004).
Ansari, S. & Helms, V. Statistical analysis of predominantly transient protein-protein interfaces. Proteins 61, 344–55 (2005).
Perkins, J. R., Diboun, I., Dessailly, B. H., Lees, J. G. & Orengo, C. Transient protein-protein interactions: structural, functional and network properties. Structure 18, 1233–43 (2010).
Smith, G. R., Sternberg, M. J. & Bates, P. A. The relationship between the flexibility of proteins and their conformational states on forming protein-protein complexes with an application to protein-protein docking. J Mol Biol 347, 1077–101 (2005).
Yogurtcu, O. N., Erdemli, S. B., Nussinov, R., Turkay, M. & Keskin, O. Restricted mobility of conserved residues in protein-protein interfaces in molecular simulations. Biophys J 94, 3475–85 (2008).
Rajamani, D., Thiel, S., Vajda, S. & Camacho, C. J. Anchor residues in protein-protein interactions. Proc Natl Acad Sci U S A 101, 11287–92 (2004).
Hwang, H., Vreven, T., Janin, J. & Weng, Z. Protein-protein docking benchmark version 4.0. Proteins 78, 3111–4 (2010).
Murzin, A. G., Brenner, S. E., Hubbard, T. & Chothia, C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J Mol Biol 247, 536–40 (1995).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–402 (1997).
Lee, B. & Richards, F. M. The interpretation of protein structures: estimation of static accessibility. J Mol Biol 55, 379–400 (1971).
Hubbard, S. J., Campbell, S. F. & Thornton, J. M. Molecular recognition. Conformational analysis of limited proteolytic sites and serine proteinase protein inhibitors. J Mol Biol 220, 507–30 (1991).
Miller, S., Lesk, A. M., Janin, J. & Chothia, C. The accessible surface area and stability of oligomeric proteins. Nature 328, 834–6 (1987).
Keskin, O., Tsai, C. J., Wolfson, H. & Nussinov, R. A new, structurally nonredundant, diverse data set of protein-protein interfaces and its implications. Protein Sci 13, 1043–55 (2004).
Chothia, C. Structural invariants in protein folding. Nature 254, 304–8 (1975).
Rekha, N., Machado, S. M., Narayanan, C., Krupa, A. & Srinivasan, N. Interaction interfaces of protein domains are not topologically equivalent across families within superfamilies: Implications for metabolic and signaling pathways. Proteins 58, 339–53 (2005).
Parthasarathy, S. & Murthy, M. R. Analysis of temperature factor distribution in high-resolution protein structures. Protein Sci 6, 2561–7 (1997).
Yuan, Z., Zhao, J. & Wang, Z. X. Flexibility analysis of enzyme active sites by crystallographic temperature factors. Protein Eng 16, 109–14 (2003).
Guerois, R., Nielsen, J. E. & Serrano, L. Predicting changes in the stability of proteins and protein complexes: a study of more than 1000 mutations. J Mol Biol 320, 369–87 (2002).
Schymkowitz, J. et al. The FoldX web server: an online force field. Nucleic Acids Res 33, W382–8 (2005).
Rother, K., Hildebrand, P. W., Goede, A., Gruening, B. & Preissner, R. Voronoia: analyzing packing in protein structures. Nucleic acids research 37, D393–5 (2009).
Goede, A., Preissner, R. & Frömmel, C. Voronoi cell: New method for allocation of space among atoms: Elimination of avoidable errors in calculation of atomic volume and density. J Comput Chem 18, 1113–1123 (1997).
Capra, J. A. & Singh, M. Predicting functionally important residues from sequence conservation. Bioinformatics 23, 1875–82 (2007).
Suzek, B. E., Huang, H., McGarvey, P., Mazumder, R. & Wu, C. H. UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinformatics 23, 1282–8 (2007).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22, 4673–80 (1994).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J Mol Biol 233, 123–38 (1993).
Holm, L. & Park, J. DaliLite workbench for protein structure comparison. Bioinformatics 16, 566–7 (2000).
Acknowledgements
We thank Ms. Smita Mohanty for discussions and suggestions. L.S.S is supported by Indo-French CEFIPRA grant. R.M.B is supported by a fellowship from Council of Scientific and Industrial Research, New Delhi. This research is supported by the Department of Biotechnology, New Delhi and also by the Mathematical Biology program sponsored by Department of Science and Technology, New Delhi.
Author information
Authors and Affiliations
Contributions
LSS, RMB and NS wrote the main manuscript text and LSS, RMB and JS prepared figures 1–6. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supporting Online Material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Swapna, L., Bhaskara, R., Sharma, J. et al. Roles of residues in the interface of transient protein-protein complexes before complexation. Sci Rep 2, 334 (2012). https://doi.org/10.1038/srep00334
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00334
This article is cited by
-
A multitier virtual screening of antagonists targeting PD-1/PD-L1 interface for the management of triple-negative breast cancer
Medical Oncology (2023)
-
Interface residues of transient protein-protein complexes have extensive intra-protein interactions apart from inter-protein interactions
Biology Direct (2019)
-
Learning protein binding affinity using privileged information
BMC Bioinformatics (2018)
-
Computational and biochemical characterization of two partially overlapping interfaces and multiple weak-affinity K-Ras dimers
Scientific Reports (2017)
-
A novel index of protein-protein interface propensity improves interface residue recognition
BMC Systems Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.