Abstract
Most acute hepatitis C virus (HCV) infections become chronic and some progress to liver cirrhosis or hepatocellular carcinoma. Standard therapy involves an interferon (IFN)-α-based regimen and efficacy of therapy has been significantly improved by the development of protease inhibitors. However, several issues remain concerning the injectable form and the side effects of IFN. Here, we report an orally available, small-molecule type I IFN receptor agonist that directly transduces the IFN signal cascade and stimulates antiviral gene expression. Like type I IFN, the small-molecule compound induces IFN-stimulated gene (ISG) expression for antiviral activity in vitro and in vivo in mice and the ISG induction mechanism is attributed to a direct interaction between the compound and IFN-α receptor 2, a key molecule of IFN-signaling on the cell surface. Our study highlights the importance of an orally active IFN-like agent, both as a therapy for antiviral infections and as a potential IFN substitute.
Similar content being viewed by others
Introduction
Hepatitis C virus (HCV) infection affects 170 million people worldwide1 and most acute HCV infections become chronic, with some progression to liver cirrhosis or hepatocellular carcinoma. HCV has a plus-strand RNA genome that encodes both structural proteins and the nonstructural (NS) proteins 2, 3, 4A, 4B, 5A and 5B. Current standard therapy against chronic HCV infection includes the use of host factor-targeting pegylated interferon (PEG-IFN)-α and ribavirin2, which is effective in only 50% of patients chronically infected with HCV genotype 13. The main causes of this low rate of efficacy may be (i) single-nucleotide polymorphisms (SNPs) in the upstream region of the IL28B gene and (ii) low compliance with the therapy, which must be administered subcutaneously. Regarding the first cause—SNPs—the host factors that are important in the early response to therapy remain unknown. However, recent studies report that genetic variants near IL28B, which encodes IFN-λ3 (interleukin 28B), correlate with the response to treatment of chronic hepatitis C infection using IFN-α/ribavirin combination therapy4,5,6,7. Patients with an rs12979860 SNP genotype of CC are reported to have a stronger response to IFN therapy (up to an 80% sustained virological response (SVR) rate with the combined therapy) than those with TC or TT genotypes4. Regarding the lack of compliance, the current therapy using recombinant IFN is a weekly injectable formulation that is unstable, requires refrigeration and is expensive and complex to administer. Furthermore, therapy is often poorly tolerated as a result of the presence of many adverse effects, including flu-like symptoms, hematological abnormalities and adverse neuropsychiatric events, any of which may require early discontinuation8. These side effects may result in dose modifications that lead to less-than-optimal responses.
Recent trends in drug development focus on drugs targeted against viral proteins such as NS 3/4A serine protease, RNA helicase, NS4B, NS5A and NS5B RNA-dependent RNA polymerase9. Very recently, two NS3/4A protease inhibitors, telaprevir and boceprevir, have been approved as new anti-HCV agents. Adding such an inhibitor to the standard therapy in the ADVANCE10 and the SPRINT-211 trials achieved significantly higher SVR rates, but the issue still remains that using these inhibitors without injectable IFN possibly yields clinical resistance12. To overcome this problem and alleviate the low compliance outlined above, an orally available IFN would be valuable because the dosing regimen is less complex.
IFNs induce the expression of a subset of IFN-stimulated genes (ISG)13, some of which show antiviral activity or are involved in lipid metabolism, apoptosis, protein degradation and inflammation14. IFNs are not only effective against HCV infection, but are also essential for innate immunity. Broadly speaking, type I IFNs (IFN-α and -β) bind to their receptor, causing the phosphorylation and activation of JAK1 and Tyk2, which is followed by the phosphorylation of signal transducers and activators of transcription (STATs) and subsequent ISG expression. To activate the JAK/STAT pathway, IFN-α requires the IFN-α/β receptor, which consists of 2 subunits, IFN-α receptor (IFNAR) 1 and IFNAR2. These IFNAR subunits together form a heterodimer upon IFN stimulation. This association of IFNAR2 and IFNAR1 is required to mediate the antiviral, antiproliferative and apoptotic effects of type I IFNs15,16,17 because the dimerization of IFNARs induces the phosphorylation of the receptor-associated tyrosine kinases, JAK1 and Tyk2 and the phosphorylated JAK kinases then phosphorylate STAT1 and STAT2. In turn, phosphorylated STAT1 and STAT2 bind to IRF9 to form the transcriptional activator IFN-stimulating gene factor 3 (ISGF3) that induces the expression of a subset of ISGs13.
Using quantitative high-throughput screening (HTS), we identified in this study a novel small molecule that acts like IFN by directly interacting with the type I IFN receptor to drive ISG expression. Our results indicate that oral administration of the small-molecule IFN agonist stimulates ISG expression in mice and that the ISG expression from this small-molecule IFN provides antiviral activity, indicating that the compound may be a potential therapeutic IFN substitute.
Results
Identification of antiviral small-molecule IFN agonists by high-throughput chemical library screening
HCV replicon cells, which were established ten years ago18 using a cell line that expresses the HCV genotype 1b subgenomic replicon19, possess a luciferase gene as a reporter optimized for use in a robust HTS system. The HTS system provides in-depth analysis of primary screening results, including detailed information regarding potency, efficacy and structure-activity relationships. IFNs show strong inhibition using this system and have been used as a positive control in the assay. Many compounds, including HCV protease inhibitors and HCV polymerase nucleoside/non-nucleoside analogs, have been assessed and are being developed for clinical testing. Analysis of data from the combination of target/counterscreen HTS, data from other assays measuring cellular toxicity, in vitro sphingolipid biosynthesis and HCV enzymatic activity (including protease and polymerase) allowed us to select compounds with potentially novel modes of action from the primary screen.
A secondary IFN signal assay, using a luciferase reporter gene which was located downstream of the IFN-stimulated response element (ISRE), eliminated assay-related false-positive compounds. Of the remaining anti-HCV replicon compounds, one of the most active was an imidazonaphthyridine with the structural formula 8-(1, 3, 4-oxadiazol-2-yl)-2, 4-bis (trifluoromethyl) imidazo [1, 2-a] [1, 8] naphthyridine (RO4948191, hereinafter RO8191) (Fig. 1a). This compound strongly suppressed HCV replicon activity at 72 h in a dose-dependent manner (Fig. 1b, left graph) without inducing host cell toxicity, as measured by the WST-8 (Fig. 1b, right graph) and CellTiter-Glo assays (data not shown). The IC50 (50% inhibitory concentration) of the compound in an anti-HCV replicon assay was 200 nM. The compound suppressed viral replication within 24 h and showed even more effective inhibition, without cytotoxicity, after 7 days (Supplementary Fig. 1). In addition, the HCV RNA replicon levels significantly decreased after incubation with the compound for 72 h, as determined by real-time reverse transcription (RT)-polymerase chain reaction (PCR) analysis (Fig. 1c). Immunostaining showed that levels of the proteins HCV NS3 and NS4A, which are localized mainly in the perinuclear region of the replicon cells, were also reduced after RO8191 treatment for 24 h (Fig. 1d). This treatment also resulted in the disappearance of viral proteins such as NS3, NS4A/B and NS5A/B, as shown by western blot analysis (Fig. 1e). The luciferase activity of HCV subgenomic genotype 2 replicon cells (JFH1, data not shown) and, surprisingly, the HCV viral titer of JFH120 in a Huh-7/K4 cell line were also reduced by RO8191 treatment (Fig. 1f).
Identification of a small molecule that inhibits HCV replication.
(a) The chemical structure of RO8191. (b) After treatment with various concentrations of RO8191 or 100 IU/mL IFN-α for 72 h, HCV replication levels were examined using a luciferase assay (left graph) and cell viabilities were determined using a WST-8 assay (right graph). The mean values and their SDs were recorded for treated cells as a percentage of the values for untreated cells and the values represent the means of 3 independent experiments. (c) Total RNA was extracted from HCV replicon cells cultured with the indicated concentration of RO8191 or 100 IU/mL IFN-α for 72 h; HCV RNA levels were analyzed using real-time RT-PCR. The mean values and their SDs were recorded for treated cells relative to the mRNA levels of β-actin and are shown as a percentage of untreated cells. The values represent the means of 3 independent experiments. (d) HCV replicon cells were treated with control medium (left panels) or 10 μM RO8191 (right panels) for 24 h and immunostained with Hoechst 33452 (blue), anti-NS3 antibody (green) and anti-NS4A antibody (red). The results were then merged (yellow). (e) HCV replicon cells were treated with the indicated concentrations of RO8191 or 100 IU/mL IFN-α for 72 h. Whole cell lysates were immunoblotted with antibodies specific to the indicated HCV NS proteins. (f) After infection with the HCV JFH1 strain, Huh-7/K4 cells were treated with the indicated concentrations of RO8191 for 72 h. Total RNA was extracted and the HCV RNA levels were analyzed using quantitative real-time RT-PCR.
RO8191 induces IFN signals, ISGs expression and JAK/STAT phosphorylation
To clarify whether RO8191 shows inhibitory activity against another RNA virus, we tested its action in encephalomyocarditis virus (EMCV)-infected A549 cells. RO8191 showed a cell-protective activity against EMCV infection similar to that of IFN-α (Fig. 2a). Because IFN-α is the most common host cell factor to exert its antiviral activity against HCV21,22 by inducing ISG expression13, we compared the gene expression profiles of IFN-α and RO8191 by conducting a global-scale DNA microarray analysis to identify genes, especially ISGs23, that were regulated by RO8191. As expected, RO8191 increased the expression of some ISGs (Supplementary Fig. 2 and Supplementary Table 1). DNA microarray analysis showed that RO8191 induced the expression of IP-10 (CXCL10), known as an ISG, RO8191 did not induce the genes encoding inflammatory cytokines and chemokines (Supplementary Fig. 3a). And, a reporter gene assay was performed using an NF-κB reporter gene. We transiently transfected the reporter gene to HCV-naïve HuH-7 cells and then treated them with RO8191 or TNF-α. In Huh-7 cells, NF-κB reporter gene was activated by TNF-α treatment, but not by RO8191 (Supplementary Fig. 3b). These results indicate that RO8191 specifically induces the IFN-relevant gene expressions.
RO8191 activates JAK/STAT and induces IFN-like signals.
(a) The cytopathic effect of EMCV infection was inhibited by the indicated concentrations of RO8191 or IFN-α. Viable cells were stained with crystal violet. The data shown are the mean values and SDs based on experiments performed in quadruplicate. (b) ISG expression levels were measured using real-time RT-PCR. Total RNA was extracted from HCV replicon cells cultured in the presence of 50 μM RO8191 or 100 IU/mL IFN-α for 2 or 8 h and known ISGs were analyzed. Asterisk indicates 2-h treatment with agents. The values shown are the mean fold change induction compared to the mRNA level of human β-actin and the fold change induction compared to untreated cells. (c) HCV replicon cells were treated with various concentrations of the indicated agents for 15 min. Total lysates were immunoblotted with antibodies to STATs or JAK kinases. The HCV replicon IC50 of IFN-α was 0.4 IU/mL, that of IFN-β was 3 IU/mL and that of IFN-γ was 0.3 ng/mL.
Real-time RT-PCR analysis also revealed that RO8191 induced many ISGs similar to those expressed in IFN-α-treated cells (Fig. 2b and Supplementary Table 2), suggesting that the antiviral mechanism of RO8191 depends on ISG expression. In addition to HCV replicon cells, we tested the compound in several cancer cell lines and normal human primary hepatocytes (Hc cells). Real-time RT-PCR analysis showed that RO8191 induced ISG expression in cultured cell lines and human primary hepatocytes (Supplementary Table 3 and Supplementary Fig. 4). These results suggest that RO8191 induces an IFN signal and that the application in clinical of RO8191 is not limited to suppressing HCV infection.
As mentioned earlier, type I IFNs phosphorylate JAK kinases and STAT proteins by inducing a heterodimerization of both IFNAR1 and IFNAR2 and the complexes thus formed transduce signals from IFN. Since RO8191 induces ISGs in a similar profile to IFN-α, we examined the phosphorylation of IFN signaling molecules. Immunoblotting analysis was performed to detect phosphorylated tyrosine (Tyr) and serine (Ser) residues of the STATs and JAK kinases using cell lysate that was treated with 50∼5000 times the HCV replicon IC50s of RO8191, IFN-α, IFN-β and IFN-γ (type II IFN). The degree to which both RO8191 and IFN-β phosphorylated STAT1, STAT5 and STAT6 was similar, as shown in Fig. 2c and the phosphorylation level of STAT2 by RO8191 was quite similar to that of IFN-α. Interestingly, STAT3 and JAK1 were more strongly phosphorylated by RO8191 than by IFN-α, -β, or -γ. On the other hand, Tyk2 was phosphorylated by type I IFNs, but not by IFN-γ or RO8191, indicating that Tyk2 is dispensable for RO8191 activity (Fig. 2c). Taken together, the phosphorylation profile of STAT proteins by RO8191 is generally similar to that of type I IFNs and the phosphorylation profiles of STAT1–3, 5, 6 and JAK1 in HCV replicon cells treated with RO8191 or type I IFNs suggest a common mechanism that differs from the mechanism in cells treated with type II IFN.
In addition to imiquimod, an IFN inducer and a Toll-like receptor (TLR) 7 agonist24, small-molecule ligands recognized by TLRs and RIG-I–like receptors are known to induce ISG expression by inducing IFN25. The chemical structure of RO8191 is similar to imiquimod so we examined whether RO8191 has a mechanism of activity like imiquimod. However, imiquimod did not affect HCV replicon cells (Supplementary Fig. 5a) and, moreover, did not stimulate STAT1 phosphorylation, whereas RO8191 caused prolonged STAT1 activation (for up to 8 h post-treatment; Supplementary Fig. 5b). Next, RO8191 is actually a small-molecule and we could not exclude the possibility that the antiviral activity might be induced through TLRs. To confirm that, we tested ISG induction using Tlr3, 4, 7 and 9 knockout (KO) mouse embryo fibroblasts (MEF)24,26,27,28. We treated the MEFs with RO8191 or murine IFN-αA for 8 h and both induced Oas1b mRNA in wild type and Tlr-KO MEFs (Supplementary Fig. 5c), demonstrating that RO8191 induces ISG expressions independently of TLRs. To exclude the possibility that RO8191 acts by inducing type I IFN, we examined its effects in Vero cells that lack the IFN gene locus29,30. Whereas imiquimod did not show IFN-like effects in Vero cells, RO8191 and IFN-α induced the ISRE activation (Supplementary Fig. 5d), indicating that RO8191 acts independently of the inducing IFN and is quite distinct from imiquimod.
RO8191 exerts antiviral activity dependent on IFNAR2/JAK1, but is independent of IFNAR1/Tyk2
IFNs require IFN receptor subunits for their activity and we hypothesized that RO8191 uses the IFN receptor to exert anti-HCV activity. To determine the contributions of IFNAR1 and IFNAR2 toward the anti-HCV replicon activity of RO8191, we suppressed the expression of these receptors using specific siRNA and treated the cells with RO8191 or IFN-α. The knockdown efficiency was determined using RT-PCR in the HCV replicon cells transfected with each siRNA, as shown in Supplementary Fig. 6. As expected, a knockdown of each receptor subunit decreased the antiviral activity of IFN-α (Fig. 3a and b, Supplementary Fig. 7a and b). IFNAR2 knockdown attenuated the antiviral activity of RO8191 (Fig. 3b and Supplementary Fig. 7b) but, interestingly, IFNAR1 knockdown did not change the antiviral activity (Fig. 3a and Supplementary Fig. 7a), suggesting that RO8191 is independent of IFNAR1. To address whether IFN-α receptor contributes to RO8191 signaling, we evaluated RO8191 using Ifnar1-KO MEF cells31. We treated Ifnar1-KO MEFs with 50 μM RO8191 or 1,000 IU/mL of murine IFN-αA for 8 h and analyzed the expression of murine Oas1b mRNA using real-time RT-PCR. Murine IFN induced murine Oas1b mRNA only in wild type MEFs, not in Ifnar1-KO MEFs, although RO8191 induced Oas1b in both wild type and Ifnar1-KO MEFs (Supplementary Fig. 8). Therefore, RO8191 induces IFN-like activity even in Ifnar1-KO MEFs.
RO8191 requires and binds IFNAR2.
(a, b) The anti-HCV replicon activity of RO8191 was attenuated by knockdown of IFNAR2 (b), but not IFNAR1 (a). Inhibition of HCV replicon replication by each agent is shown (the mean and SD from 3 experiments). The HCV replicon cells were transfected with 50 nM of the indicated siRNAs (blue, red and yellow bars). Forty-eight hours after transfection, the HCV replicon cells were treated with 1.5 μM RO8191 or 3 IU/mL IFN-α for 24 h. Twenty-four hours after treatment with each agent, the replication levels of HCV RNA were analyzed using a luciferase assay. (c, d) U5A cells that lack IFNAR2 were transfected with either an empty vector or a vector expressing the IFNAR2 gene. (c) Forty-eight hours after transfection, the cells were treated with 50 μM RO8191 (red bars) or 100 IU/mL IFN-α (yellow bars). After an additional 8 h of incubation, total RNA was extracted from the U5A cells and the OAS1 mRNA level was measured using real-time RT-PCR. The values shown are relative to the mRNA level of human β-actin. (d) Forty-eight hours after transfection, the cells were lysed and the whole cell lysates were immunoblotted with the indicated antibodies. (e) Real-time kinetic SPR analysis of the binding of RO8191 to the IFNAR2 ECD (red and blue lines). The results are consistent with 1:1 binding. PEG-IFN-α2a was also injected as a positive interacting control for IFNAR2 (black line, KD: 30 nM). (f, g) The phosphorylation of STAT1 was attenuated by a knockdown of IFNAR2 (g) but not IFNAR1 (f). The HCV replicon cells were transfected with the indicated siRNAs (10 nM). Forty-eight hours after transfection, the cells were treated for 15 min with 10 μM RO8191 or 200 IU/mL IFN-α. The total lysates were subjected to western blot analysis to analyze the phosphorylated and total protein levels of STAT1. The data were statistically analyzed using Student’s t-test.
Since IFNAR2 knockdown reduced RO8191 activity, we focused on and analyzed the IFNAR2 function using RO8191. First, U5A cells, which do not respond to IFN-α because of the lack of IFNAR2 expression32, were treated with RO8191. Although RO8191 and IFN-α did not induce OAS1 expression in the U5A cells, IFNAR2-overexpressing U5A cells were successfully complemented and this conferred susceptibility to both RO8191 and IFN-α (Fig. 3c and d). RO8191 also induced the OAS1 gene in HT1080 and 2fTGH cells, the parental cell lines of U5A cells (Supplementary Table 3, lowest and second lowest rows). Second, to elucidate whether RO8191 directly interacts with IFNAR2, we obtained a recombinant IFNAR2 extracellular domain (ECD) protein (N-terminal half of the protein, amino acids from Ile27- Lys243). The protein was subjected to surface plasmon resonance (SPR) spectroscopy using a Biacore system to directly evaluate the binding activity of the recombinant protein with its possible ligands, RO8191 and IFN-α. The IFNAR2 ECD protein was fixed on the surface of the CM7 sensor chip by amine coupling and PEG-IFN-α2a and RO8191 were injected as analytes. We comparatively analyzed 0.31 and 0.63 μM of RO8191 and both concentrations showed similar binding affinities of 480.5 and 484.5 nM, respectively (Fig. 3e), whereas a chemically derivatized compound of RO8191 that cannot inhibit HCV replication did not bind to IFNAR2 ECD (data not shown). The SPR results were consistent with 1:1 binding and showed an interaction between RO8191 and the IFNAR2 ECD protein in a dose-dependent manner, indicating that the compound and IFNAR2 may directly interact on the cell surface. In addition to the anti-replicon activity, the phosphorylation of STAT1 by RO8191 was also repressed by IFNAR2 knockdown (Fig. 3g), but the knockdown of IFNAR1 did not inhibit the STAT1 phosphorylation (Fig. 3f), indicating that RO8191 requires IFNAR2 but not IFNAR1 to achieve activity. Similarly, the knockdown of JAK1 attenuated the activity of RO8191 and IFN-α (Fig. 4a and Supplementary Fig. 9a), while, in contrast, Tyk2 and JAK2 were not required for RO8191 activity (Fig. 4b, Supplementary Fig. 9b and c). The phosphorylation of STAT1 by RO8191 was also inhibited by a knockdown of JAK1 (Supplementary Fig. 10a), but not by a knockdown of Tyk2 (Supplementary Fig. 10b), indicating JAK1 essentiality for the antiviral activity of RO8191. To confirm Tyk2-independency, we used U1A (Tyk2-deficient) cells and the parental cell, 2fTGH33 and treated them with 50 μM RO8191 or 100 IU/mL IFN-α for 8 h to analyze the expression of OAS1. As expected from previous reports34, IFN-α induced the OAS1 expression only in 2fTGH cells but not in U1A cells; however, RO8191 induced the expression in both 2fTGH and U1A cells significantly (Supplementary Fig. 11). Thus, RO8191 activates ISGs in a JAK1-dependent and Tyk2-independent manner; on the other hand, IFN-α depends on both factors. Next, we conducted a knockdown of the transcription factors related to type I IFNs activity, STAT1, STAT2 and IRF-9, to clarify whether RO8191 required these factors for antiviral activity. As expected, STAT1-siRNAs partially blocked IFN-γ activity (Supplementary Fig. 9d), which is mediated by STAT1 homodimers35. STAT1 was significantly phosphorylated by both RO8191 and IFN-α (Fig. 2c); however, the STAT1 knockdown affected neither RO8191 nor IFN-α activity (Fig. 4c and Supplementary Fig. 9d). We also analyzed the impact of STAT1 knockdown on OAS1 induction by RO8191 and found OAS1 induction by RO8191 was inhibited (Supplementary Fig. 12). STAT2 and IRF9 knockdown attenuated the inhibitory activity of both RO8191 and IFN-α (Fig. 4d and e and Supplementary Fig. 9e and f). In summary, RO8191 only binds to IFNAR2 and activates JAK1, STATs and IRF9, thereby exhibiting type I IFN-like activity (Fig. 4f).
RO8191 activates a novel IFN-like pathway.
(a–e) The inhibition of HCV replicon replications by each agent is shown (mean and SD). The HCV replicon cells were transfected with 50 nM of the indicated siRNAs (blue, red and yellow bars). (a, b) The anti-HCV replicon activity of RO8191 was reduced by knockdown of JAK1 (a) but not Tyk2 (b). Forty-eight hours after transfection, the HCV replicon cells were treated with 1.5 μM RO8191 or 3 IU/mL IFN-α. Twenty-four hours after treatment with each agent, the replication levels of HCV RNA were analyzed using a luciferase assay. (c–e) The anti-HCV replicon activity of RO8191 was reduced by knockdowns of STAT2 (d) and IRF9 (e), but not STAT1 (c). Forty-eight hours after transfection, the HCV replicon cells were treated with 1.5 μM RO8191, 3 IU/mL IFN-α, or 50 ng/mL IFN-γ. Twenty-four hours after treatment, the replication levels of HCV RNA were analyzed using a luciferase assay. (f) A schematic showing the pathways of RO8191 and IFN-α. The data were statistically analyzed using Student’s t-test and differences were considered significant at p values < 0.05.
RO8191, an IFNAR2 agonist, stimulates IFN signals in mice
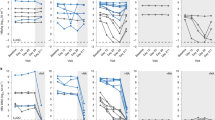
To evaluate whether RO8191 could be a clinical lead for drug development, we studied the effects of RO8191 on IFN signaling in mice. The compound or vehicle was orally administered to mice and, 24 h after treatment, the liver was removed and examined. The antiviral genes Oas1b, Mx1 and Pkr were significantly induced in the livers of mice treated with RO8191 (Table 1). As expected, gene homologs that were induced in the livers of HCV patients treated with PEG-IFN-α2b36 were also induced in mouse liver (Ifit3, Isg15, Mda5, Rig-I, Socs1 and Stat1; Table 1). In addition, genes that had previously been reported to be induced by IFN-β in mouse liver37 were also induced in the livers of RO8191-treated mice (Cxcl10, Ifit3, Isg15, Socs1 and Usp18; Table 1,). We also measured inflammatory cytokine and chemokine expressions and RO8191 did not significantly induce the expression of these genes (Supplementary Table 4). To evaluate anti-HCV activity of RO8191 in vivo, RO8191 was orally administered to HCV-infected humanized liver mice38. The results of this humanized liver mice study showed that RO8191 reduced HCV titer in vivo (Supplementary Fig. 13). These data show that RO8191 stimulates IFN signaling and is an orally available agent in mice.
Discussion
In this study, we identified a small-molecule IFN receptor agonist, RO8191, by quantitative HTS of a chemical library. This compound showed antiviral activity against both HCV and EMCV, suggesting a broad spectrum of target viruses. To learn more about the possible mechanism of action of IFN signal induction by RO8191, we investigated IFN-induced signaling and ISG induction by the small-molecule compound in vitro and in vivo. A comparison of microarray expression profiles in HCV replicon cells stimulated by IFN-α or RO8191 indicates that the IFN signal was induced not only by IFNs, but also by the small-molecule compound (Fig. 2b and Supplementary Fig. 2). Thus, this compound is an IFN-α-like small molecule, but the mechanism of the RO8191 antiviral activity remained unknown. Therefore, we examined the JAK/STAT activation pathway, which includes key players in the IFN signaling cascade.
Like type I IFNs, RO8191 significantly phosphorylates and activates STATs, in particular, STAT1 and STAT2 (Fig. 2c). Intriguingly, in HCV replicon cells, STAT1 expression knockdown did not affect the antiviral activity of RO8191 or IFN-α, although IFN-γ activity was inhibited (Fig. 4c). These data suggest that, in addition to inducing similar gene expression, RO8191 and IFN-α exhibit similar STAT phosphorylation profiles. Although RO8191- and IFN-α-mediated antiviral activity remained constant when STAT1 expression was reduced, this could be because IFN-α signaling in HuH-7 cells requires minimal amounts of STAT1 protein and STAT1 expression was not reduced below such a critical threshold by siRNA in our system. In contrast, the inhibitory activity of RO8191 was attenuated to the same extent as that of IFN-α when the expression of other components of ISGF3 (STAT2 and IRF-9) were reduced by siRNA (Fig. 4d and e). Incidentally, STAT1 siRNA did not attenuate RO8191 or IFN activity in EMCV-infected A549 cells (supplementary Fig.14), which supports the notion that STAT2 is an essential component of type I IFN signaling39. Type I IFN stimulates the formation of other STAT-containing complexes, including STAT1:STAT1, STAT3:STAT3 and STAT5:STAT5 homodimers, as well as STAT1:STAT3 and STAT2:STAT6 heterodimers40,41,42. Like IFN, RO8191 induced the phosphorylation of STAT1 and STAT2, which function as a gateway to the type I IFN signal cascade and stimulated the phosphorylation of STAT3, 5 and 6. Another possible cause for the fact that STAT1 knockdown did not show any effect on RO8191 inhibition could be compensation by these IFN signaling-stimulated STAT complexes. This finding matches the recent report by Perry et al. on the STAT dependency of IFN activity against Dengue virus, that belongs to flavivirus43. They showed that STAT2 mediates IFN antiviral signals even in STAT1 KO cells and they discussed the possibility that other STAT family proteins would compensate for STAT1 deficiency. In summary, with regards to the activation of transcription factors and ISG expression, RO8191 and IFN-α mediate the same pathway.
IFNs activate JAK kinases via IFN receptors to induce STAT phosphorylation. RO8191 robustly phosphorylated JAK1 (Fig. 2c) in comparison with IFN-α or -β and therefore we focused on IFNAR2 (a JAK1-binding subunit of the type I IFN receptor). As with IFN-α, the activity of RO8191 was inhibited by IFNAR2 knockdown (Fig. 3b). The suggestion that IFNAR2 is an essential molecule for RO8191-induced signal transduction is supported by the fact that an IFNAR2-deficient cell line, U5A, did not respond to RO8191. Furthermore, after IFNAR2 expression had been complemented in the U5A cells, ISG induction by both RO8191 and IFN-α recovered (Fig. 3c and d). The mechanism was directly explained by SPR spectroscopy, which showed an interaction between RO8191 and the IFNAR2 ECD (Fig. 3e). RO8191 strikingly phosphorylates the IFNAR2-associated kinase JAK1, when compared to other IFN-treated cell lysates (Fig. 2c). JAK1 siRNA expression inhibited RO8191 activity (Fig. 4a), indicating that JAK1 is also an essential molecule for RO8191 activity. Interestingly, RO8191 activity remains static when IFNAR1 expression is knocked down, unlike IFN-α activity (Fig. 3a). The IFNAR1-binding-kinase Tyk2 is not required for RO8191 activity (Fig. 4b) and Tyk2 was not phosphorylated (Fig. 2c) by RO8191. Also, RO8191 induced its signal even in the Ifnar1 KO MEF and the Tyk2-deficient cell line (Supplementary Fig. 8 and 11). IFN-α induces a signal via IFNAR1/Tyk2 and IFNAR2/JAK1 although RO8191 and IFN-α induce common ISGs (Fig. 2b and Supplementary Fig. 2), RO8191 activity was dependent only on IFNAR2.
We therefore propose a novel model of the induction of IFN-like signal transduction by this small molecule (Fig. 4f). So far, the IFNAR2 homodimer has been suggested to play various roles in IFN signal transduction34,44,45 and RO8191 would induce the ISG expression via such IFNAR2 homodimer. For type I IFN, both IFNAR1 and IFNAR2 cooperate and induce phosphorylation of STATs via JAK1 and Tyk2. Conversely, for RO8191, IFNAR2 alone, as a homodimer, activates JAK1 phosphorylation and subsequent STATs activation. Experiments using siRNA and deficient cells have also shown that IFNAR1 and Tyk2 were not required to induce antiviral activity in the RO8191 compound pathway. These findings suggest a novel aspect of the IFN signaling pathway that may contribute to the understanding of other molecular signaling in IFN pathways.
RO8191 is a small molecule whose oral administration is feasible and effective in a murine model (Table 1 and Supplementary Fig. 13). In the chimeric mice study, the anti-HCV effect of RO8191 and PEG-IFN was similar in that they both showed strong activity at day 1 after treatment with subsequent weak suppression of HCV replication possibly due to the immunodeficiency of chimeric mice. Further development of RO8191 by using rational chemical modifications is therefore required to produce more potent molecules for testing as an antiviral molecule which will substitute current recombinant IFN. Although RO8191 has the potential to cause IFN-like adverse effects, further development of the small-molecule agonist offers the advantages of inexpensive production cost, convenient oral administration, dose-control to reduce some adverse effects and potentially increased activity versus current recombinant IFNs.
Whereas oral NS3 protease inhibitors in monotherapy development yield resistant viruses46,47, these protease inhibitors show a significantly high rate of SVR when combined with PEG-IFN10,11 and a NS5B polymerase inhibitor also shows additive efficacy in combination with PEG-IFN48. In addition to the results of the in vivo study, we found that RO8191 induced ISGs at a level similar to IFN-α in human primary hepatocytes (Supplementary Fig. 4); we therefore expect that RO8191 will show IFN-like activity in clinical use. As an alternative strategy to protease/polymerase inhibitors with PEG-IFN, the combined use of these direct-acting antiviral agents with RO8191 in a new oral regimen may help overcome some of the delivery problems associated with current IFNs. SVR rates of individuals infected with HCV genotype 1 have increased from 5-20% with IFN monotherapy and up to 50% with a combination of IFN and ribavirin. However, the refractory patients in this therapy constitute an unmet medical need. Thus, the development of a novel IFN receptor agonist, used alone or in combination with direct-acting antiviral drugs, will add a new milestone to the treatment of chronic hepatitis C. In addition to HCV infection, type I IFNs have been approved for the treatment of multiple clinical conditions, including hairy cell leukemia, malignant melanoma, AIDS-related Kaposi’s sarcoma, multiple sclerosis and chronic hepatitis B49. Thus, R08191 shows strong potential as a lead compound for IFN substitutes.
Methods
Cell culture, mice and reagents
The #Huh7/3-1 cell line, which expresses HCV replicons, was a kind gift from F. Hoffmann-La Roche. The cells were cultured in 0.5 mg/mL G418-containing Dulbecco’s modified Eagle’s medium (DMEM, GIBCO) supplemented with 10% fetal bovine serum (FBS, HyClone). The replicon construct was derived from pFK-I377neo/NS3-3′/WT, as previously reported19. Hc cells (DS Pharma Biomedical) were cultured in CSC Complete Defined Serum-Free Medium (Cell Systems Corporation) supplemented with SF4ZR-500-D Rocket Fuel. Tlr KO MEFs were purchased from OrientalBioService, Inc. Ifnar1 KO MEF was kindly gifted by Prof. Takaoka. 2fTGH and U1A and U5A cells were kindly gifted by Prof. Stark. Culture conditions for the other cell lines are shown in Supplementary Table 5. Six-week-old C57BL/6J mice were obtained from Charles River Laboratories. Chimeric mice harboring a functional human liver cell xenograft were purchased from PhoenixBio. The protocol was reviewed by the Institutional Animal Care and Use Committee of Chugai Pharmaceutical Co., Ltd. and all mouse experiments were performed in accordance with the Guidelines for the Accommodation and Care of Laboratory Animals promulgated in Chugai Pharmaceutical Co., Ltd. Recombinant human IFN-α2a was a kind gift from F. Hoffmann-La Roche. Recombinant murine IFN-αA and human IFN-β1a were purchased from PBL Interferon Source. Recombinant TNF-α and IFN-γ were purchased from R&D Systems. Imiquimod was purchased from LKT Laboratories. JAK inhibitor I was purchased from Merck.
Luciferase assay
Luciferase activity was quantified using the Steady-Glo Luciferase assay system (Promega) and the EnVision 2013 Multilabel Reader (PerkinElmer).
WST-8 assay
The viability of drug-treated Huh-7 cells was determined using a WST-8 cell counting kit (Dojin Laboratories).
Real-time RT-PCR
Total RNA was extracted using Rneasy (Qiagen) and cDNA was synthesized using a Transcriptor First Strand cDNA Synthesis Kit (Roche Applied Science). Gene expression was measured using the LightCycler 480 System and LightCycler 480 Probes Master (Roche Applied Science). The amplification used 50 cycles of: 95°C for 5 min, 95°C for 10 s and 60°C for 30 s. Human β-actin or rodent glyceraldehyde 3-phosphate dehydrogenase (GAPDH; Applied Biosystems) expression was used as the endogenous reference for each sample. Primers and TaqMan probes for genes were designed using the Universal Probe Library Assay Design Center (Roche Applied Science; Supplementary Table 6). The probes used were from the Roche Universal Probe Library (Roche Applied Science). The samples were run in triplicate for each target gene and each reference gene was used as an internal control.
Western blotting and immunostaining
Cells were lysed in CelLytic M Cell Lysis Reagent (Sigma-Aldrich) containing Protease Inhibitor Cocktail (Sigma-Aldrich) and PhosSTOP (Roche Applied Science). Rabbit polyclonal antibodies against STAT1, STAT3, STAT6, pY701-STAT1, pY690-STAT2, pY705-STAT3, pS727-STAT3, pY694-STAT5, pY641-STAT6, pY1022/1023-JAK1 and pY1054/1055-Tyk2 were purchased from Cell Signaling Technology. Rabbit polyclonal antibodies against actin, STAT2 and STAT5 were purchased from Santa Cruz Biotechnology. Anti-Tyk2 rabbit polyclonal antibody was purchased from Upstate. Anti-IFNAR1 mouse monoclonal (MAB245) and anti-IFNAR2 sheep polyclonal antibodies were purchased from R&D Systems. Anti-NS3, anti-NS5A and anti-NS5B rabbit polyclonal antibodies were a kind gift from F. Hoffmann-La Roche. Anti-NS4A and anti-NS4B mouse monoclonal antibodies were a kind gift from the Tokyo Metropolitan Institute of Medical Science. Proteins were detected using the Odyssey Infrared Imaging System (LI-COR). For immunostaining analysis, the cells were fixed on a 35-mm glass-based dish (Iwaki) with 4% paraformaldehyde, blocked using 5% fetal bovine serum in phosphate-buffered saline and then incubated with anti-NS3 and anti-NS4A antibodies. The cells were then washed and incubated with Alexa488-labeled anti-rabbit IgG and Alexa568-labeled anti-mouse IgG (Molecular Probes) and analyzed using confocal laser microscopy.
JFH-1 antiviral assay
A cured K4 cell line derived from HuH-7 HCV replicon cells was maintained in DMEM supplemented with 10% fetal calf serum (FCS), high-glucose nonessential amino acids and HEPES (Invitrogen). The JFH-1/K4 cell line, which was persistently infected with the HCV JFH-1 strain, was maintained under the same conditions as the cured K4 cell line. For the anti-HCV assay of JFH-1/K4 cells persistently infected with the JFH-1 strain, JFH-1/K4 cells were seeded in a 24-well tissue culture plate containing DMEM supplemented with 10% FCS, high-glucose nonessential amino acids and HEPES (Invitrogen). After overnight incubation, serial dilutions of reagent in growth medium were added. After 72 h, total RNA was purified from the JFH-1/K4 cells using Isogene (Nippon Gene). HCV-RNA was quantified by real-time PCR as previously reported50.
EMCV cytopathic effect assay
This assay was performed on A549 cells seeded in a 96-well tissue culture plate containing DMEM supplemented with 10% FBS. After overnight incubation, the indicated concentrations of each reagent were added to the growth medium. After 12 h, 100 TCID50/mL EMCV was added and after another 48 h, viable cells were stained with 0.5% crystal violet. RNAi experiment using EMCV was also performed on A549 cells seeded in a 96-well tissue culture plate containing DMEM supplemented with 10% FBS. We transfected STAT1- or STAT2-siRNA to A549 cells and after 72 h we infected EMCV to the cells and treated them with 1 μM RO8191 or 2 IU/mL IFN. After additional 48 h incubation, we evaluated the cell viability by staining with crystal violet.
GeneChip and data analysis
Total RNA was extracted from 107 HCV replicon cells cultured for 8 h in the presence of 2 μM RO8191 or 4 IU/mL IFN-α with TRIzol Reagent (Invitrogen). Reverse transcription, RNA labeling (5 μg of total RNA), hybridization to Human Genome U133 Plus 2.0 Arrays (Affymetrix) and scanning were performed according to the manufacturer’s instructions (Affymetrix, http://www.affymetrix.com). GC-RMA (GeneChip Robust Microarray Analysis) algorithms were used to generate scaled gene expression values. The fold change compared to untreated cells was calculated and probe sets were selected for genes that were at least 2.0-fold upregulated in RO8191- and IFN-α-treated cells relative to the control cells.
RNA interference
For all double-stranded RNAs, ON-TARGET Plus siRNA reagents (Dharmacon) were used (Supplementary Table 7). The siRNAs were transiently transfected using Lipofectamine RNAiMAX Transfection reagent (Invitrogen) according to the manufacturer’s protocols for reverse transfection.
Plasmids and transfection
ISRE and NF-κB reporter gene were purchased from Clontech. IFNAR2 was cloned into a pCOS2 vector51 harboring the EF1α promoter. Plasmids were transfected using FuGENE HD (Roche Applied Science) according to the manufacturer’s instructions.
SPR measurements
SPR binding studies were performed using a Biacore T100. Recombinant IFNAR2 ECD protein was purchased from R&D Systems. The protein (1 mg/mL) was diluted 1:20 with 10 mM sodium acetate buffer (pH 5.0) and mixed with 2 µM RO8191 for stabilization of the binding site. The mixture was immobilized on a Series S sensor chip CM7 using amine coupling. RO8191 and PEG-IFN-α2a (Chugai Pharmaceutical) were injected onto the sensor chip at a flow rate of 0.03 mL/min. Response curves were generated by subtraction of the background signal generated simultaneously on a control flow cell. Kinetic parameters were obtained by global fitting of the sensorgrams to a 1:1 model using Biacore T100 Evaluation Software, version 2.0.1.
Humanized liver mice study
The chimeric mice were generated by transplanting human primary hepatocytes into severe combined immunodeficient (SCID) mice carrying the urokinase plasminogen activator transgene controlled by an albumin promoter38. The chimeric mice used in this study were applied from Inoue et al.52 and had a high substitution rate of human hepatocytes. Six weeks after hepatocyte transplantation, patient serum containing 106 copies of HCV genotype 1b was intravenously injected into each mouse. HCV titer reached approximately 108 copies/mL and was stable after 4 weeks of HCV injection and persistently infected for 12 weeks. Here, we used mice after 5 weeks post infection and tested for 2 weeks. The mice were treated for 14 days with RO8191 30 mg/kg/day orally or PEG-IFN-α2a 30 μg/kg subcutaneously twice weekly. HCV RNA in serum was extracted using the acid guanidinium-phenol-chloroform method. Quantification of HCV RNA was performed using real-time RT-PCR based on TaqMan chemistry, as described50. HCV inoculations, drug administration, blood collection and killing were performed under ether anesthesia. Blood samples were taken from the orbital vein and sera were immediately isolated. The protocols for animal experiments were approved by the local ethics committee. The animals received humane care according to NIH guidelines. Patients gave written informed consent before sampling.
References
Lavanchy, D. The global burden of hepatitis. Liver Int. 29, 74–81 (2009).
Ghany, M. G., Strader, D. B., Thomas, D. L. & Seeff, L. B. Diagnosis, management and treatment of hepatitis C: an update. Hepatology. 49, 1335–74 (2009).
Zeuzem, S. Interferon-based therapy for chronic hepatitis C: current and future perspectives. Nat Clin Pract Gastroenterol Hepatol. 5, 610–622 (2008).
Ge, D. et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature. 461, 399–401 (2009).
Suppiah, V. et al. IL28B is associated with response to chronic hepatitis C interferon-alpha and ribavirin therapy. Nat Genet. 41, 1100–4 (2009).
Tanaka, Y. et al. Genome-wide association of IL28B with response to pegylated interferon-alpha and ribavirin therapy for chronic hepatitis C. . Nat Genet.41, 1105–9 (2009).
Thomas, D. L. et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature. 461, 798–801 (2009).
Czepiel, J., Czepiel, J., Biesiada, G. & Mach, T. Viral hepatitis C. .Pol Arch Med Wewn. 118, 734–740 (2008).
Webster, D. P., Klenerman, P., Collier, J. & Jeffery, K. J. Development of novel treatments for hepatitis C. .Lancet Infect Dis. 9, 108–117 (2009).
Jacobson, I. M. et al. Telaprevir for previously untreated chronic hepatitis C virus infection. N Engl J Med. 364, 2405–16 (2011).
Poordad, F. et al. Boceprevir for untreated chronic HCV genotype 1 infection. N Engl J Med. 364, 1195–206 (2011).
Kwong, A. D., McNair, L., Jacobson, I. & George, S. Recent progress in the development of selected hepatitis C virus NS3.4A protease and NS5B polymerase inhibitors. Curr Opin Pharmacol. 8, 522–531 (2008).
Stark, G. R. et al. How cells respond to interferons. Annu Rev Biochem. 67, 227–264 (1998).
de Veer, M. J. et al. Functional classification of interferon-stimulated genes identified using microarrays. J Leukoc Biol. 69, 912–20 (2001).
Pestka, S., Langer, J. A., Zoon, K. C. & Samuel, C. E. Interferons and their actions. Annu Rev Biochem. 56, 727–777 (1987).
Uze, G., Lutfalla, G. & Gresser, I. Genetic transfer of a functional human interferon alpha receptor into mouse cells: cloning and expression of its cDNA. Cell. 60, 225–234 (1990).
Novick, D., Cohen, B. & Rubinstein, M. The human interferon alpha/beta receptor: characterization and molecular cloning. Cell. 77, 391–400 (1994).
Lohmann, V. et al. Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science. 285, 110–113 (1999).
Sakamoto, H. et al. Host sphingolipid biosynthesis as a target for hepatitis C virus therapy. Nat Chem Biol. 1, 333–337 (2005).
Wakita, T. et al. Production of infectious hepatitis C virus in tissue culture from a cloned viral genome. Nat Med. 11, 791–796 (2005).
Kneteman, N. M. et al. Anti-HCV therapies in chimeric scid-Alb/uPA mice parallel outcomes in human clinical application. Hepatology. 43, 1346–1353 (2006).
Dash, S. et al. Interferons alpha, beta, gamma each inhibit hepatitis C virus replication at the level of internal ribosome entry site-mediated translation. Liver Int. 25, 580–594 (2005).
Der, S. D., Zhou, A., Williams, B. R. & Silverman, R. H. Identification of genes differentially regulated by interferon alpha, beta, or gamma using oligonucleotide arrays. Proc Natl Acad Sci U S A. 95, 15623–15628 (1998).
Hemmi, H. et al. Small anti-viral compounds activate immune cells via the TLR7 MyD88-dependent signaling pathway. Nat Immunol. 3, 196–200 (2002).
Kawai, T. & Akira, S. Toll-like receptor and RIG-I-like receptor signaling. Ann N Y Acad Sci. 1143, 1–20 (2008).
Yamamoto, M. et al. Role of adaptor TRIF in the MyD88-independent toll-like receptor signaling pathway. Science. 301, 640–3 (2003).
Hoshino, K. et al. Cutting edge: Toll-like receptor 4 (TLR4)-deficient mice are hyporesponsive to lipopolysaccharide: evidence for TLR4 as the Lps gene product. J Immunol. 162, 3749–52 (1999).
Hemmi, H. et al. A Toll-like receptor recognizes bacterial DNA. Nature. 408, 740–5 (2000).
Mosca, J. D. & Pitha, P. M. Transcriptional and posttranscriptional regulation of exogenous human beta interferon gene in simian cells defective in interferon synthesis. Mol Cell Biol. 6, 2279–83 (1986).
Diaz, M. O. et al. Homozygous deletion of the alpha- and beta 1-interferon genes in human leukemia and derived cell lines. Proc Natl Acad Sci U S A. 85, 5259–63 (1988).
Chen, H. M. et al. Critical role for constitutive type I interferon signaling in the prevention of cellular transformation. Cancer Sci. 100, 449–56 (2009).
Lutfalla, G. et al. Mutant U5A cells are complemented by an interferon-alpha beta receptor subunit generated by alternative processing of a new member of a cytokine receptor gene cluster. EMBO J. 14, 5100–5108 (1995).
Velazquez, L. Fellous, M. Stark, G. R. & Pellegrini, S. A protein tyrosine kinase in the interferon alpha/beta signaling pathway. Cell. 70, 313–22 (1992).
Lewerenz, M. Mogensen, K. E. & Uzé, G. Shared receptor components but distinct complexes for alpha and beta interferons. J Mol Biol. 282, 585–99 (1998).
Shuai, K. et al. Interferon activation of the transcription factor Stat91 involves dimerization through SH2-phosphotyrosyl peptide interactions. Cell. 76, 821–828 (1994).
Sarasin-Filipowicz, M. et al. Interferon signaling and treatment outcome in chronic hepatitis C. .Proc Natl Acad Sci U S A. 105, 7034–9 (2008).
Farnsworth, A. et al. Acetaminophen modulates the transcriptional response to recombinant interferon-beta. PloS One. 5, e11031 (2010).
Mercer, D. F. et al. Hepatitis C virus replication in mice with chimeric human livers. Nat Med. 7, 927–33 (2001).
Leung, S. et al. Role of STAT2 in the alpha interferon signaling pathway. Mol Cell Biol. 15, 1312–1317 (1995).
Li, X. et al. Formation of STAT1-STAT2 heterodimers and their role in the activation of IRF-1 gene transcription by interferon-alpha. J Biol Chem. 271, 5790–4 (1996).
Ghislain, J. J. & Fish, E. N. Application of genomic DNA affinity chromatography identifies multiple interferon-alpha-regulated Stat2 complexes. J Biol Chem. 271, 12408–13 (1996).
Gupta, S. Jiang, M. & Pernis, A. B. IFN-alpha activates Stat6 and leads to the formation of Stat2:Stat6 complexes in B cells. J Immunol. 163, 3834–41 (1999).
Perry, S. T., Buck, M. D., Lada, S. M., Schindler, C. & Shresta, S. STAT2 Mediates Innate Immunity to Dengue Virus in the Absence of STAT1 via the Type I Interferon Receptor. PloS Pathogens. 7, e1001297 (2011).
Pattyn, E. et al. Dimerization of the interferon type I receptor IFNaR2-2 is sufficient for induction of interferon effector genes but not for full antiviral activity. J Biol Chem. 274, 34838–45 (1999).
Platis, D. & Foster, G. R. Activity of hybrid type I interferons in cells lacking Tyk2: a common region of IFN-alpha 8 induces a response, but IFN-alpha2/8 hybrids can behave like IFN-beta. J Interferon Cytokine Res. 23, 655–66 (2003).
McHutchison, J. G. et al. Telaprevir with peginterferon and ribavirin for chronic HCV genotype 1 infection. N Engl J Med. 360, 1827–38 (2009).
Hézode, C. et al. Telaprevir and peginterferon with or without ribavirin for chronic HCV infection. N Engl J Med. 360, 1839–50 (2009).
Kneteman, N. M. et al. HCV796: A selective nonstructural protein 5B polymerase inhibitor with potent anti-hepatitis C virus activity in vitro, in mice with chimeric human livers and in humans infected with hepatitis C virus. Hepatology. 49, 745–52 (2009).
Gutterman, J. U. Cytokine therapeutics: lessons from interferon alpha. Proc Natl Acad Sci U S A. 91, 1198–1205 (1994).
Takeuchi, T. et al. Real-time detection system for quantification of hepatitis C virus genome. Gastroenterology. 116, 636–642 (1999).
Yabuta, N. et al. Method for screening ligand having biological activity. PCT Int. Appl. WO0206838 (2002).
Inoue, K. et al. Evaluation of a cyclophilin inhibitor in hepatitis C virus-infected chimeric mice in vivo. Hepatology. 45, 921–8 (2007).
Acknowledgements
This study was supported financially by Chugai Pharmaceutical Co., Ltd. H.K., K.O., Y.O., H.Y., H.O., M.A., A.O., H.S., N.H., A.K., K.M., T.T., N.S., Y.A., M.A. and M.S. are employees of Chugai Pharmaceutical Co., Ltd. We are grateful to George Stark for providing us the 2fTGH, U1A and U5A cell lines and Akinori Takaoka for mouse Ifnar1-knockout MEFs. We also thank Isamu Kusanagi and Chiaki Tanaka for technical assistance, AVSS Co., Ltd. for technical assistance on EMCV and Editing Services at Chugai Pharmaceutical Co., Ltd. for editorial assistance.
Author information
Authors and Affiliations
Contributions
H.K., K.O., Y.O., H.Y., Y.H., A.O. and N.H. performed the experiments; H.K., K.O., H.O. and M.A. analyzed the data; G.F. and W.A. provided experimental materials and input into the data analysis; H.S., A.K., M.K., T.T., N.S., G.F., Y.A., M.A. and M.S. provided expert information; and H.K. and M.S. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
An orally available, small-molecule interferon inhibits viral replication
Supplementary Information
An orally available, small-molecule interferon inhibits viral replication
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Konishi, H., Okamoto, K., Ohmori, Y. et al. An orally available, small-molecule interferon inhibits viral replication. Sci Rep 2, 259 (2012). https://doi.org/10.1038/srep00259
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00259
This article is cited by
-
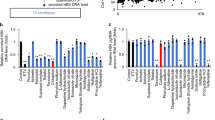
A small molecule iCDM-34 identified by in silico screening suppresses HBV DNA through activation of aryl hydrocarbon receptor
Cell Death Discovery (2023)
-
Epigenetic reprogramming promotes the antiviral action of IFNα in HBV-infected cells
Cell Death Discovery (2021)
-
Identification of a small molecule that primes the type I interferon response to cytosolic DNA
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.