Abstract
The generation of induced pluripotent stem cells (iPSCs) by the forced expression of defined transcription factors in somatic cells holds great promise for the future of regenerative medicine. However, the initial reprogramming mechanism is still poorly understood. Here we show that Zscan4, expressed transiently in 2-cell embryos and embryonic stem cells (ESCs), efficiently produces iPSCs from mouse embryo fibroblasts when coexpressed with Klf4, Oct4 and Sox2. Interestingly, the forced expression of Zscan4 is required only for the first few days of iPSC formation. Microarray analysis revealed transient and early induction of preimplantation-specific genes in a Zscan4-dependent manner. Our work indicates that Zscan4 is a previously unidentified potent natural factor that facilitates the reprogramming process and reactivates early embryonic genes.
Similar content being viewed by others
Introduction
Somatic cells have been reprogrammed to the pluripotent state by nuclear transfer (NT), in which a somatic nucleus is exposed to many unknown reprogramming factors that are thought to be present in oocytes, zygotes and 2-cell embryos1,2,3,4. More recently, somatic cells have been reprogrammed to become induced pluripotent stem cells (iPSCs) by exposing a somatic nucleus to the defined set of transcription factors: Myc, Klf4, Oct4 (official gene symbol: Pou5f1) and Sox2 in the original formulation5; other factors including transcription factors, small molecules and microRNAs in the newer formulation (reviewed in6,7,8,9).
Although both procedures produce pluripotent cells that can give rise to live offspring by injection into mouse blastocysts, they seem to exhibit notable differences. For example, it has been suggested that the reprogramming of somatic cells by NT occurs within a few cell divisions10, whereas the reprogramming of somatic cells into murine iPSCs requires about 1–2 weeks of continuous application of factors for at least the first 8–10 days11. Whole-genome DNA methylation analyses have indicated that murine pluripotent stem cells made by the iPSC procedure retain an epigenetic memory of donor somatic cells, which is not apparent in pluripotent stem cells made by the NT procedure12. Furthermore, it has been reported that the genome integrity of human iPSCs seems to be often compromised with mutations and genome alterations13,14,15,16,17. Accordingly, the efficient production of high-quality iPSCs may become feasible by factors that can make the reprogramming process similar to that which occurs during the NT procedure18. As a first step, it is desirable to find a factor that can reactivate genes that are expressed in preimplantation embryos, i.e., NT environment, during iPSC generation.
Previously, we have shown that Zscan4 (zinc finger and SCAN domain containing 4), expressed specifically in 2-cell embryos and only about 5% of ESCs at a given time19, acts critically in the formation of proper blastocysts19 and in the maintenance of genome stability and telomeres in ESCs20. Accordingly, we hypothesized that Zscan4 is a factor that is present in the NT environment, but is missing in the current repertoire of iPSC factors. Here we have tested this notion and demonstrated that Zscan4 indeed functions as a potent enhancer of the reprogramming process in iPSC formation.
Results
Zscan4 is reactivated in late-stage iPSCs
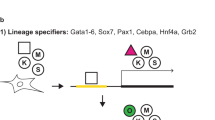
To investigate whether Zscan4 is reactivated during iPSC formation, we first generated mouse ESCs carrying an Emerald (a GFP variant) reporter driven by a 3.5 kb Zscan4 promoter, which can reproduce the expression pattern of endogenous Zscan4 in mouse ESCs20. Chimeric mice produced by injecting the ESCs (named ES-pZ-Emerald) into blastocysts were used to generate E13.5 embryos, which were subsequently used to derive mouse embryo fibroblasts (MEFs). The MEFs in which the presence of an Emerald reporter was confirmed by genotyping were named MEF-pZ-Emerald cells (Fig. 1a). Emerald fluorescence was not detectable in the MEF-pZ-Emerald cells, indicating that Zscan4 is not expressed in MEFs.
Zscan4 is not expressed during early phase of iPSC formation, but reactivated later in iPSC cells.
(a) Schematic representation of procedures to examine Zscan4 expression during iPSC formation. TA1 ESCs, F1 hybrid strain (C57BL/6J x 129S6/SvEvTac). A piggyBac transfection involves a main vector PB-TET-MKOS (shown), PB-CAG-rtTA (a tetracycline transactivator) and pCyL43 (transposase). (b) Phase-contrast microscopic images during the formation of cell colonies with authentic ES-like morphology. Day 0 is set when doxycycline (Dox) is added to the complete ES medium 24 hours after a piggyBac transfection. (c) Fluorescence images (left), fluorescence images merged with phase-contrast images (middle) and flow cytometry charts (right) of two representative cell clones established from ESC-like colonies and cultured in the absence of Dox. (d) Appearance of Emerald+ cells (represented as “+”) in the culture. Fraction of Emerald+ cells was measured by the flow cytometry on day 28.
We then transfected a piggyBac vector (PB-TET-MKOS)21,22 carrying doxycycline (Dox)-inducible Myc (M), Klf4 (K), Oct4 (O) and Sox2 (S), into the MEF-pZ-Emerald cells and then cultured the cells in ESC media supplemented with Dox. As reported, colonies with an authentic ESC-like morphology were clearly visible by day 13 (Fig. 1b). We observed the cells under fluorescence microscopes daily, but did not find any Emerald+ cells in culture. We picked 28 ESC-like colonies and passaged them into ESC culture media without Dox 11 to 14 days after the piggyBac transfection. Two clones did not survive, but the other 26 clones proliferated to form ESC-like colonies. Colonies with Emerald+ cells began to appear from day 15 and by day 28 all the colonies showed the presence of Emerald+ cells in the same pattern as typical F1 hybrid ESC lines: a small number (1.0±0.5%, S.E.M.) of Emerald+ cells (Fig. 1c, 1d). The data indicate that iPSCs, once formed, gained the expression pattern of Zscan4 similar to ESCs, suggesting a close similarity between iPSC and ESCs. Importantly, Zscan4 was not activated during the early phase of iPSC formation by the MKOS factors. Consistent with this finding, reanalysis of the published microarray data23,24,25 did not reveal the activation of Zscan4 during the early phase of iPSC formation.
The data also indicate that, as reported previously21,22, the production of iPSC by a piggyBac vector carrying MKOS factors was robust and the majority (26 of 28) of ESC-like colonies indeed became authentic iPSCs, as confirmed by the same unique expression pattern of Zscan4 as typical ESCs. This notion was further supported by the presence of a pluripotency marker NANOG26 (Supplementary Fig. S1a, S1b), which was not used as an exogenous iPSC factor and the expression of a GFP marker under the control of the Oct4 promoter27 (Supplementary Fig. S1c) in the ESC-like colonies generated by the PB-TET-MKOS. Therefore, in subsequent experiments, we primarily used the ESC-like colony morphology combined with alkaline phosphatase staining (ALP+) to score the number of authentic iPSCs.
Zscan4 enhances the efficiency of iPSC formation
To test the effects of exogenous Zscan4 overexpression on iPSC formation, we cotransfected PB-TET-MKOS with either PB-TET-DsRed (control), PB-TET-Zscan4c, or PB-TET-Zscan4-ERT2 into wild-type MEF cells (MEF-WT) (Fig. 2a). The PB-TET-Zscan4-ERT2 was made by fusing Zscan4c to ERT2 (a mutated ligand-binding domain of the human estrogen receptor, which can be activated by the presence of tamoxifen [Tmx])28 (Fig. 2a). We found that Zscan4 increased the number of ALP+ ESC-like colonies by about 1.5-fold compared to the DsRed control (Fig. 2b, 2c; see also Supplementary Fig. S1d for the NANOG+ assay and Supplementary Fig. S1e for the Oct4-GFP assay). We also found that the human ZSCAN4 gene functions similarly to mouse Zscan4 in mouse iPSC formation (Supplementary Fig. S2a, S2b). As expected, the cotransfection of PB-TET-Zscan4-ERT2 increased the number of ALP+ ESC-like colonies only in the Tmx+ condition (Fig. 2b, 2c). When these ESC-like colonies, made with or without Zscan4 contribution, were picked and passaged onto regular ESC culture conditions (Dox- Tmx-), most (18 of 24) of them proliferated to form ESC-like colonies, which were confirmed to be authentic iPSCs by the expression of endogenous pluripotency markers (Oct4, Sox2, Nanog, Rex1 and Dax1), ALP-staining, embryoid body (EB) formation and the ability to differentiate into cells of three germlayers (Fig. 2d–2k). Taken together, Zscan4 can enhance the efficiency of MKOS-mediated iPSCs formation. The results also indicate that Zscan4-ERT2 can be used as a Tmx-controllable Zscan4 factor. Therefore, in the subsequent analysis, we used the Zscan4-ERT2.
Forced expression of exogenous Zscan4 increases the efficiency of iPSC formation from the wild-type MEFs.
(a) Schematic representation of procedures to examine the effects of exogenous Zscan4 on iPSC formation from wild-type MEFs (C57BL/6J x 129S6/SvEvTac). (b) Representative pictures of 6-well plates stained for ALP 14 days after Dox (and Tmx- or Tmx+) induction. (c) Colonies with authentic ESC morphology and ALP+ were scored. Transfections and Dox inductions were performed in triplicate. Data from two independent experiments are shown. Data are represented as mean±S.E.M. (triplicate wells). *, P < 0.01. (d) Several ESC-like colonies were picked, propagated in the Dox- condition and established as cell clones: 5 clones (MKOS factors: A2–A6) and 4 clones (MKOS factors + Zscan4). RT-PCR analysis of these clones with pluripotency gene markers: endogenous Oct4, endogenous Sox2, Nanog, Zfp42 (Rex1) and Dax1 (Nr0b1). Gapdh was used as a control. The clone B5 (MKOS factors + Zscan4) was used for the subsequent analyses (e–g). (e) A representative image of ALP staining. (f) A microscopic image showing embryoid bodies (day 4) generated from the clone. (g) Fluorescence microscopic images of the clone after in vitro differentiation from the embryoid body shown in (F) and stained with antibodies against αSMA (mesoderm), AFP (endoderm), GATA4 (endoderm) and βIII-tubulin (ectoderm). Pictures (bottom) are the same images after merging with DAPI-staining. Scale bar, 100 μm. (h–k) The same assays were performed on 5 clones (MKOS factors + Zscan4-ERT2 [Tmx-]: C1-C6) and 4 clones (MKOS factors + Zscan4-ERT2 [Tmx+]: D2–D6). The clone D3 (MKOS factors + Zscan4-ERT2 [Tmx+]) was used for the detailed analyses (i–k).
To facilitate the analysis of Zscan4-aided iPSC formation, we generated ESCs (named ES-ZERT), in which mouse Zscan4c fused to ERT2 was constantly expressed under a strong and ubiquitous CAG-promoter29 (Fig. 3a). The expression of Zscan4-ERT2 was confirmed by qRT-PCR (Supplementary Fig. S3a). The Tmx+ condition slowed down the proliferation of ES-ZERT cells, whereas the Tmx- condition did not (Supplementary Fig. S3b). Because the Tmx did not affect the proliferation of wild-type ESCs (Supplementary Fig. S3b), this result indicates that Zscan4 function can be turned on in ESCs carrying a Zscan4-ERT2 construct in the presence of Tmx+. The effect of Zscan4 on ESC proliferation was reversed within three days when Tmx was removed from the media (Supplementary Fig. S3c, S3d). Microarray analysis also showed that the effect of Zscan4 on the ESC transcriptome mostly receded within three days after Tmx withdrawal (Supplementary Fig. S3e – S3i). MEFs (MEF-ZERT) derived from ES-ZERT (Fig. 3a, 3b) also showed slower proliferation in the Tmx+ condition than in the Tmx- condition (Fig. 3c).
Zscan4 increases the efficiency of iPSC formation in Tmx-dependent manner from the Zscan4-ERT2-expressing MEFs.
(a) A pCAG-Zscan4cERT2 vector was transfected into V6.5 ESCs (C57BL/6 x 129/Sv) to make Zscan4-ERT2 ESCs (ES-ZERT). ES-ZERT cells were microinjected into blastocysts from ICR mice to generate male chimeric mice, which were subsequently mated with ICR female mice. E13.5 embryos were used to generate mouse embryo fibroblasts (MEFs). MEFs were subjected to genotyping and quantitative RT-PCR. MEFs that carried pCAG-Zscan4cERT2 DNA and expressed exogenous Zscan4c were designated as MEF-ZERT and MEFs that did not were designated as MEF-WT (wild type). (b) Zscan4 expression in MEF-ZERT cell lines compared to that in ESCs. Expression levels of Zscan4 in two different series of MEFs (A1–A13 and B1–B11) were examined by qRT-PCR. Data in triplicate were represented as mean±S.E.M. MEF lines marked with * were used in this work: MEF-ZERT (#A2, #A7, #B5); MEF-WT (#A3). (c) Growth curves of MEF-WT and MEF-ZERT cultured and passaged in Tmx+ and Tmx- conditions. (d) A piggyBac vector (PB-TET-MKOS) carrying doxycycline (Dox)-inducible Myc (M), Klf4 (K), Oct4 (O) and Sox2 (S), was transfected into MEF-ZERT and MEF-WT, respectively. The cells were cultured under the Dox+ Tmx- or Dox+ Tmx+ condition for 14 days, fixed and stained for alkaline phosphatase (ALP). (e) Representative pictures of 6-well plates stained for ALP. (f) ALP+ ESC-like colonies were scored (mean±S.E.M.). *, P < 0.05. (g) A representative phase-contrast image of an iPSC clone derived from the MEF-ZERT (Tmx+) condition. (h) A representative image of the clone stained for ALP. (i) Fluorescence-microscopic images of the clone after staining with antibodies against SSEA-1 and NANOG. Pictures (right) are the same images after merging with DAPI-staining. (j) Karyotype analysis of iPSCs generated with MKOS (Tmx-: 5 independent clones) and ZMKOS (Tmx+: 8 independent clones) conditions. For each clone, the fraction of cells with a normal karyotype (% euploid) was calculated based on more than 40 metaphase spreads. Then, mean + S.E.M. was calculated for each group (P < 0.05). (k) A representative picture of live E13.5 embryos entirely generated from iPSC clone made with the ZMKOS.
After transfecting a PB-TET-MKOS vector, the MEF-ZERT cells and control MEF-WT cells were cultured with or without Tmx in standard iPSC generation conditions (Dox+) (Fig. 3d). Scoring the number of ALP+ ESC-like colonies after 14 days revealed that the Tmx+ condition increased the number of iPSC colonies by 1.5- to 2-fold in the MEF-ZERT cells, whereas the Tmx+ condition did not affect the efficiency of iPSC formation in the MEF-WT cells (Fig. 3e, 3f). These data further confirmed that the Tmx-controlled Zscan4-ERT2 system works during iPSC formation and that Zscan4 enhances iPSC formation by MKOS factors.
When these ESC-like colonies were picked and transferred to the Dox- Tmx- condition, most of the clones (9 of 12) made by both ZMKOS factors (Dox+ Tmx+ conditions) and MKOS factors (Dox+ Tmx- conditions) continued to proliferate without these exogenous factors. Furthermore, the ZMKOS clones did not show any difference from the MKOS clones in the standard tests for the formation of authentic iPSCs, i.e., ALP-staining of colonies (Fig. 3g, 3h), immunohistochemistry of pluripotency markers (Fig. 3i), RT-PCR analysis of pluripotency genes (Supplementary Fig. S4a), DNA methylation patterns (Supplementary Fig. S4b), embryoid body formation (Supplementary Fig. S4c) and ability to differentiate into three germ layers (Supplementary Fig. S4d). However, karyotype analysis of >40 metaphase spreads each from 5 independent MKOS clones and 8 independent ZMKOS clones suggested that iPSCs generated with ZMKOS seem to have better karyotypes than those made with MKOS (Fig. 3j). Although an extensive comparison was not done, we also found that the injection of iPSCs (ZMKOS) into tetraploid (4N) blastocysts formed an entire mouse embryo (Fig. 3k). As the embryos are entirely derived from iPSCs in the tetraploid complementation system30, the results demonstrated that iPSCs generated with ZMKOS factors were fully pluripotent.
Zscan4 can replace Myc for the iPSC formation
Previous studies have shown that oncogene Myc, which increases cell proliferation and suppresses genome stability, is required for efficient iPSC formation (reviewed in31). To test whether Zscan4 can enhance iPSC formation without Myc, we transfected MEF-WT and MEF-ZERT cells with a PB-TET-KOS vector carrying Dox-inducible KOS factors and cultured the cells in Dox+ Tmx+ or Dox+ Tmx- conditions for 2 weeks (Fig. 4a). As expected, MEF-WT cells produced only a small number of ALP+ ESC-like colonies (Fig. 4b). However, MEF-ZERT cells produced 40- to 70-fold more ALP+ ESC-like colonies in Dox+ Tmx+ (i.e., ZKOS) conditions than in the control Dox+ Tmx- (i.e., KOS) conditions (Fig. 4b). The result was confirmed in independent experiments (Supplementary Fig. S5a, S5b). When the ESC-like colonies were picked from the ZKOS condition (i.e., Dox+ Tmx+) and cultured without the exogenous factors (i.e., in the Dox- Tmx- condition), most of them (9 of 11) proliferated to form authentic iPSCs judged by the RT-PCR analysis of pluripotency genes (Supplementary Fig. S5c), the ALP-staining of colonies (Supplementary Fig. S5d), embryoid body formation (Supplementary Fig. S5e), ability to differentiate into three germ layers (Supplementary Fig. S5f) and 4N complementation assay (Supplementary Fig. S5g). The number of iPSC colonies generated by ZKOS factors (Fig. 4b) was comparable to that generated by MKOS factors (Fig. 2c, 3f). Thus, in combination with KOS factors, Zscan4 can replace Myc for iPSC formation.
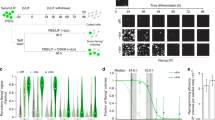
Zscan4 enhances iPSC formation without Myc and is required only for the first few days of induction.
(a) Schematic representation of experimental procedures for iPSC generation. (b) Representative pictures of 6-well plates stained for ALP 20 days after the Dox induction. ALP+ ESC-like colonies were counted (mean±S.E.M.). *, P < 0.05. (c, d) Efficiency of iPSC formation was examined after different Tmx treatments. ALP+ ESC-like colonies were counted 20 days after induction (mean±S.E.M.). Different letters denote statistically significant differences between groups (P < 0.05).
Zscan4 is required only for the first few days during iPSC formation
To further clarify the role of Zscan4 in reprogramming, we examined when and how long Zscan4 is required. It has been shown that efficient iPSC formation requires the ectopic overexpression of MKOS factors for at least 8–10 days11,24. Using a Tmx-inducible system of Zscan4 expression in MEF-ZERT cells, which was independent of the Dox-inducible expression of KOS factors, we could vary the time of exposure to Zscan4 (Fig. 4a). The absence of Tmx for the first 4 or 7 days significantly reduced the number of iPSC colonies, whereas the presence of Tmx for only the first 4 or 7 days was sufficient to produce a number of ALP+ ESC-like colonies comparable to that obtained by the presence of Tmx for 20 days (Fig. 4c). Further experiments with progressive shortening of exposure established that Tmx is required only for the initial day of iPSC formation (Fig. 4d). As we showed above, the addition of Tmx turned on Zscan4 effects within a day, whereas the removal of Tmx turned off Zscan4 effects within 3 days. Therefore, these data indicate that the forced expression of Zscan4 is required only for the first few days and can thus influence the early phase of reprogramming.
Secondary MEF system further confirms that Zscan4 plays a distinct role among iPSC factors
It has been shown that the secondary MEF system facilitates the analysis of early events leading to iPSC formation, because every cell carries inducible iPSC factors11,18,23. To generate secondary MEF cells, we first cotransfected MEF-WT cells with PB-TET-KOS and PB-TET-Zscan4-ERT2 vectors (Fig. 5a). After culturing these cells in Dox+ Tmx+ condition for 2 weeks, ESC-like colonies were picked and propagated in the Dox- Tmx- condition. Clones #2 and #4 were analyzed further and confirmed as authentic iPSC cells based on the RT-PCR analysis of pluripotency genes (Fig. 5b), ALP-staining of colonies (Fig. 5c, 5d), immunohistochemistry of pluripotency markers (Fig. 5e), the formation of the embryoid body (Fig. 5f), the ability to differentiate into three germ layers (Fig. 5g) and the production of live embryos by 4N complementation assay (Fig. 5h). These iPSCs also showed >80% normal karyotype and global gene expression profiles indistinguishable from ESCs (Fig. 5i). The E13.5 embryos generated from the iPSC (ZKOS#2) by the 4N complementation assay were subsequently used to derive the secondary MEFs (named MEF-KOS-ZERT2nd) (Fig. 5a).
Secondary MEFs derived from iPSCs can generate iPSCs without transfection in Zscan4-dependent manner.
(a) iPSC colonies were generated by transfecting MEF-WT (C57BL/6Jx129S6/SvEvTac) with piggyBac vectors (PB-TET-KOS and PB-TET-Zscan4ERT2-IRES-HisDsRed) and culturing the cells for two weeks under the Dox+ Tmx+ condition. Under fluorescence microscope, Zscan4-ERT2+ colonies could be identified by red-fluorescence. The two red colonies were picked from the wells and propagated in the ES cell culture condition on feeder cells, resulting in the establishment of two cell clones (#2, #4). (b) RT-PCR analysis of these clones and control MEF-WT cells with pluripotency gene markers. (c) A phase-contrast image of the clone (ZKOS#2). (d) A phase-contrast image of the clone (ZKOS#2) after staining with ALP. (e) Fluorescence microscopic images of the clone (ZKOS#2) after staining with antibodies against SSEA-1 and NANOG. (f) A microscopic image showing embryoid bodies (day 4) generated from the iPSC clone (ZKOS#2). (g) Fluorescence images of the clone (ZKOS#2) after in vitro differentiation from the embryoid body shown in (f) and stained with antibodies against αSMA, AFP, GATA4 and βIII-tubulin. Scale bar, 100 μm. (h) E13.5 embryos derived from the clone (ZKOS#2) by the 4N complementation. These embryos were used to generate secondary MEFs (MEF-KOS-ZERT2nd) as described in (a). (i) Global expression profiles of cells were generated by using DNA microarrays. (Upper Panel) A scatter-plot showing pair-wise comparison between the clone (ZKOS#2) and MEF-WT. (Lower Panel) A scatter-plot showing pair-wide comparison between the clone (ZKOS#2) and V6.5 ESC. Spots in color represent genes whose expression show statistically significant differences between samples (FDR≤0.05, fold-change≥2)45. (j) Morphologies of the MEF-KOS-ZERT2nd cells during the first 6 days of Dox and Tmx treatments. Pictures of cells after ALP-staining on day 17 are also shown. (k) qRT-PCR analysis of the MEF-KOS-ZERT2nd cells. Expression levels of genes are presented as fold-difference compared to those in ESCs. Results using primer pairs that recognize 3′-UTR of genes represent the expression levels of endogenous genes, whereas results using primer pairs that recognize the open reading frame (ORF) of genes represent the combined expression levels of endogenous and exogenous (from the piggyBac vectors) genes.
As expected, without ZKOS induction (i.e., Dox- Tmx- and Dox- Tmx+ conditions), ALP+ ESC-like colonies were not formed from the MEF-KOS-ZERT2nd cells by day 17. By contrast, the forced expression of ZKOS factors (Dox+ Tmx+ condition) produced a large number (∼400) of ALP+ ESC-like colonies, whereas the KOS factor only (Dox+ Tmx- condition) produced a much smaller number (∼20) of ALP+ ESC-like colonies (Fig. 5j). The Dox-dependent expression of exogenous Zscan4-ERT2, Klf4, Oct4 and Sox2 were confirmed by qRT-PCR (Fig. 5k). The upregulation of an endogenous Myc, which was not included as an exogenous iPSC factor, was not observed (Fig. 5k). The secondary MEF system also showed that Zscan4 is required only for the first few days of iPSC formation (Supplementary Fig. S6a). By contrast, KOS factors were required for at least the first 6 days (Supplementary Fig. S6b). These results further provide independent confirmation of our aforementioned observation that Zscan4 acts in the early phases of reprogramming and plays a distinct role among iPSC factors.
Previous studies on the secondary MEFs using MKOS factors on a piggyBac vector have shown that early morphological changes of the MEFs, i.e., the mesenchymal-epithelial transition (MET), are the important first step for iPSC formation23. Interestingly, in our system, both the KOS factors and the ZKOS factors caused dramatic changes in MEF morphology by day 1: the cells became flatter and lost the typical spindle-shape of MEFs (Fig. 5j). However, only the ZKOS condition (i.e., Dox+ Tmx+ condition) produced ESC-like colony structures by day 6 and resulted in a large number of iPSC colonies by day 17 (Fig. 5j). These data thus indicate that the MET is a feature that can be caused by KOS factors alone and does not necessarily lead to iPSC formation, suggesting that the MET is not sufficient for eventual iPSC formation.
Global expression profiles reveal the transient reactivation of preimplantation-specific genes in a Zscan4-dependent manner
Consistent with a dramatic morphological change, the forced expression of KOS alone (Dox+ Tmx-) altered the transcriptomes of MEF-KOS-ZERT2nd significantly by day 1: upregulation of 1730 genes and downregulation of 947 genes (Fig. 6a, first row). By day 3 and 6, the number of expression-altered genes further increased. However, considering that the KOS condition (Dox+ Tmx-) produced only a small number of iPSCs after 17 days in culture (Fig. 5j), these transcriptome alterations were not sufficient enough to convert MEFs to iPSCs.
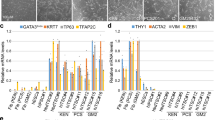
Microarray analysis reveals Zscan4-dependent activation of preimplantation- and gonads-specific genes during the early phase of iPSC formation from the MEF-KOS-ZERT 2nd cells.
(a) Scatter-plots showing pair-wise comparison between Dox+ Tmx- (KOS factors) and Dox- Tmx- (No factor); Dox+ Tmx+ (ZKOS factors) and Dox- Tmx+ (No factor); Dox- Tmx+ (No factor) and Dox- Tmx- (No factor); and Dox+ Tmx+ (ZKOS factors) and Dox+ Tmx- (KOS factors) conditions. Cells were harvested on day 1, 3 and 6 after beginning the Dox or Tmx treatment. Figures in each scatter plot represent the number of genes that showed statistically significant differences between the conditions (FDR≤0.05, fold-change≥2). A list of non-redundant 231 genes were obtained by combining 12 (day 1), 90 (day 3) and 178 (day 6) genes that were more highly expressed in Dox+ Tmx+ (ZKOS) condition than in Dox+ Tmx- (KOS) condition. (b) A heatmap showing the fold-difference of expression levels of 231 genes between Tmx+ and Tmx- conditions. The fold difference for each gene was calculated by dividing the expression level (Tmx+) by the expression level (Tmx-). Among 231 genes, Pramel6 showed the highest fold-difference: 10.2-fold on day 3 (see the Supplementary Table S1). Results obtained by searching the EST database for 231 genes are shown as symbols after gene names: Red circle, genes expressed predominantly in oocytes; blue circle, genes expressed predominantly in preimplantation embryos (1-cell to blastocysts); pink square, genes expressed predominantly in testes or ovaries.
By contrast, a comparison between the iPSC-producing ZKOS condition (Dox+ Tmx+) and the KOS condition (Dox+ Tmx-) revealed only a minor transcriptome difference (Fig. 6a, fourth row): 28 genes by day 1, 162 genes by day 3 and 237 genes by day 6. That such a small difference in transcriptome yielded such a great difference in the number of iPSC colonies was astounding, suggesting the significance of these genes for Zscan4-mediated iPSC formation. After combining lists of these genes, we obtained 231 distinct genes that were more highly expressed in the ZKOS condition than in the KOS condition (with the statistical significance of FDR≤0.05, fold-change≥2) (Fig. 6a). For these 231 genes, the fold-differences between Tmx+ (i.e., Zscan4+) and Tmx- (i.e., Zscan4-) were calculated, subjected to hierarchical clustering and presented as a heatmap (Fig. 6b, Supplementary Table S1).
We noticed that some of these genes were originally identified from only preimplantation embryos through large-scale cDNA sequencing projects32, e.g., Pramel6, D5Ertd577e, D13Ertd608e, Tcstv133, Trim43a34 and Trim43b34. Accordingly, we searched the public Expressed Sequence Tags (ESTs) database (NCBI/NIH) and scored the number of EST hits for each gene. A significant fraction of the 231 genes was indeed expressed predominantly in early embryos and closely related gonads (testis and ovary): 27 genes in the preimplantation stage (1-cell – blastocysts); 14 genes in oocytes; and 37 genes in testis/ovary (Fig. 6b, Supplementary Table S2). The preimplantation- and gonad-specific expression of these genes was also confirmed by the expression patterns in the NIA Gene Expression Atlas35 (Fig. 7a; Supplementary Fig. S6c) and the GNF database36 (Fig. 7b; Supplementary Fig. S6d). Interestingly, most of these genes showed little or no expression in MEFs, ESCs and iPSCs (Fig. 7a; Supplementary Fig. S6c), indicating that these genes are activated transiently in the early phase of iPSC formation, but then downregulated once iPSCs are fully formed. These genes may thus represent the earliest markers for Zscan4-mediated iPSC formation from MEFs, because they were fully activated as early as day 1, 3, or 6, when typical pluripotency markers such as Nanog, Zfp42 (also known as Rex1) and Dppa5a (also known as Esg1) were still silent (Supplementary Fig. S7a).
Zscan4-dependent activation of preimplantation – and/or gonad-specific genes is mostly transient.
(a) A heatmap showing expression patterns of 201 genes (a subset of 231 genes) found in the NIA Gene Expression Atlas35 (22 different adult organs/tissues and cultured cells, from left to right: brain, cerebellum, eyes, skeletal muscle, heart, bone, liver, kidney, bladder, skin, visceral fat, lung, small intestine, large intestine, stomach, placenta, ovary, oocyte, testis, MEF cells, ESCs and iPSCs). A magnified picture can be found in Supplementary Fig. S6c. Bar graphs show the gene expression levels of two representative genes (Patl2 and D13Ertd608e) among these tissues. (b) A heatmap showing the expression patterns of 99 genes (a subset of 231 genes) found in the GNF database36 (62 different organs/tissues). A magnified picture can be found in Supplementary Fig. S6d. Bar graphs show the gene expression levels of two representative genes (Pramel6 and D5Ertd577e) among these tissues. (c) A summary diagram showing events occurring during Zscan4-mediated iPSC formation.
The early activation of preimplantation-specific genes seems to be unique to Zscan4-mediated iPSC formation. Indeed, previous microarray studies of the secondary MEF system using piggyBac MKOS factors have not identified the activation of preimplantation-specific genes as the critical features of iPSC-forming transcriptome changes23. For example, most of the 231 critical genes that we identified were not activated transiently during the early phase of MKOS-based iPSC formation (Supplementary Fig. S7b, S7c). These data suggest that Zscan4-mediated iPSC formation takes a different path from standard MKOS-mediated iPSC formation: the former path is more strongly associated with the genetic program occurring in the preimplantation embryos than the latter path.
Discussion
Previously, we have shown that Zscan4, which is not expressed in blastocysts, is activated in a subpopulation of cells by day 6 during the derivation of ES cells from blastocysts in vitro19. We have also shown that Zscan4 is transiently expressed in late 2-cell embryos19, blastomeres of which have been shown to have potent reprogramming activity in NT cloning experiments4. In this paper, we now show that Zscan4 is not expressed in the early phase of standard MKOS-mediated iPSC formation, but begins to be expressed in a subpopulation (1–5%) of iPSCs later, once they become mature iPSCs. The results led to a hypothesis that Zscan4 can make the reprogramming process similar to that which occurs during the NT procedure and prompted us to examine the effects of forced expression of Zscan4 on the process of iPSC formation. Our work has established that Zscan4 is a potent early embryonic factor with distinct properties that can help to reprogram MEFs to iPSCs.
Our work reveals a number of remarkable features of Zscan4 as a reprogramming factor (Fig. 7c). First, Zscan4 action is fast, as it is required only for the initial few days during the long process of iPSC formation. It has been well established that the continuous presence of iPSC factors for at least 8–10 days or for at least 5 days for Myc24 is required for iPSC formation. Our own results by shortening Dox exposure (i.e., overexpression of KOS factors) have also confirmed these earlier results. Furthermore, unlike other iPSC factors, which are expressed highly and rather homogeneously in ESCs, Zscan4 is expressed in only a small fraction (1%–5%) of ESCs19. Our earlier analysis of systematically perturbing the transcriptomes of mouse ESCs by overexpressing specific transcription factors showed that Zscan4 belongs to regulatory modules distinct from Oct4, Sox2, Nanog and other core pluripotency TFs37. It is also striking that the overexpression of Zscan4 causes very minor changes to the transcriptome during the early phase of reprogramming, but major differences to the outcome of iPSC formation. This is another indication that the mode of Zscan4 action is quite different from other iPSC factors. One could speculate that Zscan4 may be involved in epigenetic regulation or chromatin remodeling: the overexpression of such factors does not alter the transcriptome dramatically compared to regular transcription factors37.
Second, Zscan4-mediated reprogramming activates 231 genes in MEFs in a Zscan4-dependent manner, which are rather specifically expressed in preimplantation embryos, oocytes and gonads, suggesting that Zscan4 is a potent inducer of the preimplantation genetic program, including reprogramming activity. The data also hint at the presence of a common mechanism between reprogramming mediated by NT and reprogramming mediated by ZKOS for iPSC. While this manuscript was being reviewed, a paper reporting that Glis1, which is expressed in MetaII oocytes, could be used as a reprogramming factor that can replace Myc, was published38. We checked our microarray data and found that Glis1 was transiently upregulated on day3 in both Dox+ Tmx- and Dox+ Tmx+ conditions, thus not included in our 231 genes. On the other hand, Zscan4 does not seem to be activated during Glis1-mediated iPSC formation, at least based on the reported microarray data on day 538. Interestingly, ZSCAN4 was listed in their Supplementary Table as one of 18 factors that can replace KLF4 for iPSC formation after screening 1437 human transcription factors; however, further characterization of ZSCAN4 was not conducted38. Taken together, Zscan4 is the first reprogramming factor that can activate the expression of many preimplantation-specific genes during the early phase of iPSC formation.
One of the critical issues for the application of iPSCs to regenerative medicine is the potential cancer-risk due to the compromised genome integrity associated with iPSC formation39. In this sense, it is conceivable that Zscan4 can help reprogram cells while preserving genome stability. Zscan4 replaces the oncogene Myc in enhancing the efficiency of iPSC formation with KOS factors. Interestingly, unlike Myc and other iPSC-promoting conditions (e.g., repression of P53 (reviewed in40)), which stimulate the proliferation of MEFs and destabilize genome integrity, Zscan4 represses the proliferation of MEFs and ESCs. Furthermore, we have shown previously that Zscan4 promotes genome stability and the maintenance of a normal karyotype in ESCs20. Indeed, iPSCs generated with ZMKOS factors showed better karyotypes than iPSCs generated with MKOS factors. Although more extensive analysis, including the comprehensive genome sequencing approach13,14,15,16,17, is required to draw any firm conclusion, considering other anecdotal findings about the quality of iPSCs generated with the aid of Zscan4, such as the successful tetraploid complementation assay, these initial findings are promising. Future analyses of large numbers of iPSCs generated with the aid of Zscan4 for their genome stability and possible tumorigenicity in chimeric animals will provide the definitive answers to these important issues.
Methods
Plasmid vector construction
To construct a pCAG-Zscan4-ERT2 plasmid, an entire open reading frame (ORF: 506 a.a.) of a mouse Zscan4c gene19 was fused with ERT2 (a mutated ligand-binding domain of the human estrogen receptor28; 314 a.a.) and cloned into the XhoI/NotI sites of a plasmid (pPyCAGBstXI-IP; a gift from Dr. Hitoshi Niwa). The resultant plasmid vector (pCAG-Zscan4-ERT2) expresses a Zscan4c-ERT2 fusion protein and a Puromycin-resistant protein driven by a strong CAG promoter29 (Fig. 1a). PiggyBac vectors (PB-TET-IRES- βgeo21,22, PB-TET-MKOS21,22 and PB-CAG-rtTA21,22) were purchased from Addgene.
Generation of ES-ZERT cells and chimeric mice
All experiments were performed in accordance with the guidelines of the NIA/NIH Animal Care and Use Committee (Approved Animal Study Proposal 311-LG-2012). V6.5 ESCs41 derived from an F1 hybrid strain (C57BL/6 x 129/Sv) were purchased from Thermo Scientific Open Biosystems. ESCs were cultured at 37°C in 5% CO2 in a complete ES medium: DMEM, 15% FBS, 1000 U/ml LIF (ESGRO, Chemicon), 1 mM sodium pyruvate, 0.1 mM non-essential amino acids, 2 mM GlutaMAX, 0.1 mM β-mercaptoethanol, penicillin and streptomycin. V6.5 ESCs (5×105 cells) in suspension were cotransfected with 2 μg of a linearized pCAG-Zscan4-ERT2 vector and 0.4 μg of a PL452 vector42 (a neomycin-resistant gene driven by a PGK promoter) using Effectene (QIAGEN) according to the manufacturer's protocol and plated in 100 mm cell culture dishes. After selecting with G418 for 8 days, resulting ESC colonies were picked, expanded and frozen. Subsequently, an ES-ZERT cell clone was selected based on the results of genotyping, qPCR and puromycin-resistance. ES-ZERT cells (10-15 cells) were injected into 2N blastocysts and then transferred to E2.5 recipient females. After genotyping the pups, ZERT chimeric mice carrying pCAG-Zscan4-ERT2 DNA were established.
MEF isolation
MEF-ZERT cells and MEF-WT cells were isolated from E13.5 embryos, which were obtained by crossing male ZERT mice to female ICR mice (Figure 1a). MEF-WT (C57BL/6J x 129S6/SvEvTac) cells were isolated from E13.5 embryos, which were obtained by crossing between male 129SvEvTac mice and female C57BL/6J mice. These cells were plated into a 10-cm plates in DMEM supplemented with 10% FBS, 1 mM sodium pyruvate, 0.1 mM non-essential amino acids, 2 mM GlutaMAX, 0.1 mM β-mercaptoethanol, penicillin and streptomycin.
PiggyBac-transfection and iPSC generation
iPSCs were generated as previously reported21,22 with some modifications. Briefly, MEFs were plated on gelatin-coated 6-well plates at a density of 1×105 cells/well in complete ES medium. After a 24 h incubation period, MEFs were transfected with 1.6 μg of each plasmid, which were pCyL4343 (Sanger institute), PB-CAG-rtTA and PB-TET-MKOS or -KOS with or without PB-TET-Zscan4c or PB-TET-Zscan4cERT2, using Xfect (Clontech). After 24 h, cells were fed with the complete ES medium with Doxycycline (Dox: 1.5 μg/ml) and with or without 200 nM 4-hydroxytamoxifen (Tmx). Colonies were picked after 12 days post-Dox induction and propagated as iPSCs in the complete ES medium with Dox (1.5 μg/ml), until Dox-independency was observed in replicate wells. Culture medium was changed every day.
Microarray Data Analysis
Expression profiling was carried out as described37 using whole-genome 60-mer oligonucleotide microarrays44 (Agilent). Data analysis and visualization were carried out by NIA Array Analysis Software45. Heatmaps were generated with MultiExperiment Viewer (Mev) v4.246. Heatmaps were produced by the HCL (Hierarchical clustering) module of MeV4.2. All the microarray data have been submitted to the public database GEO (accession number GSE28436, http://www.ncbi.nlm.nih.gov/geo/).
Bisulfite sequencing
Sequencing data were analyzed using QUMA47.
References
Gurdon, J. B., Elsdale, T. R. & Fischberg, M. Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature 182, 64–65 (1958).
Wakayama, T., Perry, A. C., Zuccotti, M., Johnson, K. R. & Yanagimachi, R. Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 394, 369–374 (1998).
Wilmut, I., Schnieke, A. E., McWhir, J., Kind, A. J. & Campbell, K. H. Viable offspring derived from fetal and adult mammalian cells. Nature 385, 810–813 (1997).
Egli, D., Sandler, V. M., Shinohara, M. L., Cantor, H. & Eggan, K. Reprogramming after chromosome transfer into mouse blastomeres. Curr Biol 19, 1403–1409 (2009).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Gonzalez, F., Boue, S. & Belmonte, J. C. Methods for making induced pluripotent stem cells: reprogramming a la carte. Nat Rev Genet 12, 231–242 (2011).
Hanna, J. H., Saha, K. & Jaenisch, R. Pluripotency and cellular reprogramming: facts, hypotheses, unresolved issues. Cell 143, 508–525 (2010).
Plath, K. & Lowry, W. E. Progress in understanding reprogramming to the induced pluripotent state. Nat Rev Genet 12, 253–265 (2011).
Stadtfeld, M. & Hochedlinger, K. Induced pluripotency: history, mechanisms and applications. Genes Dev 24, 2239–2263 (2010).
Egli, D. et al. Reprogramming within hours following nuclear transfer into mouse but not human zygotes. Nat Commun 2, 488 (2011).
Wernig, M. et al. A drug-inducible transgenic system for direct reprogramming of multiple somatic cell types. Nat Biotechnol 26, 916–924 (2008).
Kim, K. et al. Epigenetic memory in induced pluripotent stem cells. Nature 467, 285–290 (2011).
Hussein, S. M. et al. Copy number variation and selection during reprogramming to pluripotency. Nature 471, 58–62 (2011).
Gore, A. et al. Somatic coding mutations in human induced pluripotent stem cells. Nature 471, 63–67 (2011).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011).
Mayshar, Y. et al. Identification and classification of chromosomal aberrations in human induced pluripotent stem cells. Cell Stem Cell 7, 521–531 (2010).
Laurent, L. C. et al. Dynamic changes in the copy number of pluripotency and cell proliferation genes in human ESCs and iPSCs during reprogramming and time in culture. Cell Stem Cell 8, 106–118 (2011).
Hanna, J. et al. Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462, 595–601 (2009).
Falco, G. et al. Zscan4: a novel gene expressed exclusively in late 2-cell embryos and embryonic stem cells. Dev Biol 307, 539–550 (2007).
Zalzman, M. et al. Zscan4 regulates telomere elongation and genomic stability in ES cells. Nature 464, 858–863 (2010).
Kaji, K. et al. Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 458, 771–775 (2009).
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009).
Samavarchi-Tehrani, P. et al. Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 7, 64–77 (2010).
Sridharan, R. et al. Role of the murine reprogramming factors in the induction of pluripotency. Cell 136, 364–377 (2009).
Koche, R. P. et al. Reprogramming factor expression initiates widespread targeted chromatin remodeling. Cell Stem Cell 8, 96–105 (2011).
Silva, J. et al. Nanog is the gateway to the pluripotent ground state. Cell 138, 722–737 (2009).
Brambrink, T. et al. Sequential expression of pluripotency markers during direct reprogramming of mouse somatic cells. Cell Stem Cell 2, 151–159 (2008).
Feil, R. et al. Ligand-activated site-specific recombination in mice. Proc Natl Acad Sci U S A 93, 10887–10890 (1996).
Niwa, H., Yamamura, K. & Miyazaki, J. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108, 193–199 (1991).
Nagy, A. et al. Embryonic stem cells alone are able to support fetal development in the mouse. Development 110, 815–821 (1990).
Nakagawa, M., Takizawa, N., Narita, M., Ichisaka, T. & Yamanaka, S. Promotion of direct reprogramming by transformation-deficient Myc. Proc Natl Acad Sci U S A 107, 14152–14157 (2010).
Ko, M. S. et al. Large-scale cDNA analysis reveals phased gene expression patterns during preimplantation mouse development. Development 127, 1737–1749 (2000).
Struwe, M. & Solter, D. (NCBI/NIH, 1998).
Stanghellini, I., Falco, G., Lee, S. L., Monti, M. & Ko, M. S. Trim43a, Trim43b and Trim43c: Novel mouse genes expressed specifically in mouse preimplantation embryos. Gene Expr Patterns 9, 595–602 (2009).
Sharov, A. A. et al. Responsiveness of genes to manipulation of transcription factors in ES cells is associated with histone modifications and tissue specificity. BMC Genomics 12, 102 (2010).
Su, A. I. et al. Large-scale analysis of the human and mouse transcriptomes. Proc Natl Acad Sci U S A 99, 4465–4470 (2002).
Nishiyama, A. et al. Uncovering early response of gene regulatory networks in ESCs by systematic induction of transcription factors. Cell Stem Cell 5, 420–433 (2009).
Maekawa, M. et al. Direct reprogramming of somatic cells is promoted by maternal transcription factor Glis1. Nature 474, 225–229 (2011).
Pera, M. F. Stem cells: The dark side of induced pluripotency. Nature 471, 46–47.
Tapia, N. & Scholer, H. R. p53 connects tumorigenesis and reprogramming to pluripotency. J Exp Med 207, 2045–2048 (2010).
Eggan, K. et al. Hybrid vigor, fetal overgrowth and viability of mice derived by nuclear cloning and tetraploid embryo complementation. Proc Natl Acad Sci U S A 98, 6209–6214 (2001).
Liu, P., Jenkins, N. A. & Copeland, N. G. A highly efficient recombineering-based method for generating conditional knockout mutations. Genome Res 13, 476–484 (2003).
Wang, W. et al. Chromosomal transposition of PiggyBac in mouse embryonic stem cells. Proc Natl Acad Sci U S A 105, 9290–9295 (2008).
Carter, M. G. et al. Transcript copy number estimation using a mouse whole-genome oligonucleotide microarray. Genome Biol 6, R61 (2005).
Sharov, A. A., Dudekula, D. B. & Ko, M. S. A web-based tool for principal component and significance analysis of microarray data. Bioinformatics 21, 2548–2549 (2005).
Saeed, A. I. et al. TM4 microarray software suite. Methods Enzymol 411, 134–193 (2006).
Kumaki, Y., Oda, M. & Okano, M. QUMA: quantification tool for methylation analysis. Nucleic Acids Res 36, W170–175 (2008).
Acknowledgements
We thank Hong Yu, Carole Stagg, Yong Qian and Dawood Dudekula for technical assistance. This research was supported entirely by the Intramural Research Program of the NIH, National Institute on Aging.
Author information
Authors and Affiliations
Contributions
M.S.H.K. conceived the project; T.H., T.A. and M.S.H.K. designed and interpreted experiments; T.H. and M.A. performed cell culture and molecular biology experiments; T.A. performed mouse experiments; Y.P. performed microarray and qRT-PCR experiments; H.G.H. performed karyotype analysis; Y.N. contributed expert advice about vector constructions; T.H. and M.S.H.K. prepared figures; and M.S.H.K. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Table S1
Supplementary Information
Table S2
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Hirata, T., Amano, T., Nakatake, Y. et al. Zscan4 transiently reactivates early embryonic genes during the generation of induced pluripotent stem cells. Sci Rep 2, 208 (2012). https://doi.org/10.1038/srep00208
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00208
This article is cited by
-
Zfp296 knockout enhances chromatin accessibility and induces a unique state of pluripotency in embryonic stem cells
Communications Biology (2023)
-
CTCF is a barrier for 2C-like reprogramming
Nature Communications (2021)
-
ZSCAN4 facilitates chromatin remodeling and promotes the cancer stem cell phenotype
Oncogene (2020)
-
The role of telomere-binding modulators in pluripotent stem cells
Protein & Cell (2020)
-
A molecular roadmap for the emergence of early-embryonic-like cells in culture
Nature Genetics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.