Abstract
We describe the microbiota of two hypersaline saltern ponds, one of intermediate salinity (19%) and a NaCl saturated crystallizer pond (37%) using pyrosequencing. The analyses of these metagenomes (nearly 784 Mb) reaffirmed the vast dominance of Haloquadratum walsbyi but also revealed novel, abundant and previously unsuspected microbial groups. We describe for the first time, a group of low GC Actinobacteria, related to freshwater Actinobacteria, abundant in low and intermediate salinities. Metagenomic assembly revealed three new abundant microbes: a low-GC euryarchaeon with the lowest GC content described for any euryarchaeon, a high-GC euryarchaeon and a gammaproteobacterium related to Alkalilimnicola and Nitrococcus. Multiple displacement amplification and sequencing of the genome from a single archaeal cell of the new low GC euryarchaeon suggest a photoheterotrophic and polysaccharide-degrading lifestyle and its relatedness to the recently described lineage of Nanohaloarchaea. These discoveries reveal the combined power of an unbiased metagenomic and single cell genomic approach.
Similar content being viewed by others
Introduction
Marine salterns consist of a series of shallow, interlinked ponds where seawater is gradually concentrated until sodium chloride is precipitated. In geographic regions with low average annual precipitation, the individual saltern ponds resemble a continuous bioreactor whose salinity is kept constant over long periods of time. The composition of saltern microbial communities has been found to be fairly stable over time1, even when subjected to environmental perturbations2. Furthermore, past small-scale studies have described the salterns as a system with low species richness, high density of prokaryotic cells and short food chains3,4,5,6,7,8. This simplicity makes salterns prospective model systems9, similar to, yet presumably easier to analyse than a more diverse marine environment.
The set of hypersaline ponds of the Santa Pola salterns near Alicante (Spain), in particular the crystallizer pond CR30, has been the focus of multiple studies for over 30 years. The ponds have been studied using several approaches, including classical cultivation, PCR 16S rRNA gene amplification and sequence analysis, fluorescent in-situ hybridization and metagenomic fosmid library sequence analysis3,4,5,6,7,10,11. In the NaCl-saturated crystallizer pond, the encountered microbial diversity is low. Only two microbial species are found to be abundant and both are ‘salt-in’ strategists – the hyperhalophilic square archaeon Haloquadratum walsbyi and a hyperhalophilic bacterium Salinibacter ruber3,4,6,10. Another set of extensively studied ponds corresponds to the total salt concentration of 19%, about half way through the concentration process between seawater and NaCl saturation. Pure cultures of diverse, moderate halophiles of assorted bacterial subdivisions and the Halobacteriales have been obtained from these waters11. Primary productivity has been estimated to be relatively low, probably due to the replacement of less halophilic primary producers by the species of halophilic algae Dunaliella12. Finally, studies carried out by molecular approaches6,7 indicate that the prokaryotic assemblages are composed of diverse representatives of Gammaproteobacteria, Bacteroidetes and Halobacteriaceae with 16S rRNA sequences often unrelated to any cultivated microbes. This study revisits the ponds of total salinity of 19% and 37% at a sequencing depth much higher than before.
We used the Roche 454 GS-FLX sequencing platform to carry out deep shotgun sequencing of environmental DNA from two saltern ponds of 19% and 37% salt concentrations, collecting 475 Mb and 361 Mb of genomic sequence, respectively. The 19% salinity pond (SS19, the number in the acronym indicates the approximate salinity), i.e. about 5 times seawater salinity, was characterized by precipitation of gypsum, which forms thick layers at the bottom13. This particular habitat is widespread in nature and occurs whenever seawater is concentrated by evaporation, e.g. in coastal lagoons14. The second sample was taken from a terminal crystallizer pond (SS37), with 37% total salt concentration (or about 10 times seawater salinity), a very extreme environment, characterized by extensive precipitation of sodium chloride14. Two more datasets were included in the analysis in order to be able to detect characteristic metagenomic features of the halophilic microbial communities: a metagenome from a hypersaline coastal lagoon Punta Cormoran (PC6) in the Galapagos islands (64 Mb), with a total salt concentration of 6%15 and a dataset available from Mediterranean marine waters (DCM3, 312 Mb) with a total salt concentration of 3.5%16. We first discuss the overall sequence characteristics, microbial species richness and community structure as depicted by various approaches and then identify and describe new, abundant groups in different salinity levels.
Results
GC content and Isoelectric Point Profiles
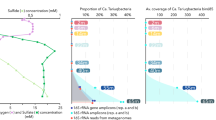
The general characteristics of the datasets are shown in Supplementary Table S1. We analyzed the sequence composition of the individual reads in each dataset by GC content (Figure 1). The DCM3 dataset (a pristine marine-salinity environment) showed a unimodal GC content distribution with a low GC peak at about 34%. This shape is typical of marine datasets, especially surface and near surface samples. In all the three hypersaline datasets, the GC content distribution plots were bimodal and had a high GC peak at ∼65%. The second, low GC peak varied between datasets and was ∼54% GC in PC6 dataset (a coastal hypersaline lagoon) and ∼47% GC in both SS19 and SS37 datasets (solar saltern ponds). In SS37, the latter was also the predominant peak and is attributed mainly to the haloarchaeon Haloquadratum walsbyi which has an average genomic GC content of 47%10.
Comparison of GC% of sequences from four metagenomic datasets of increasing salinities.
DCM3: Deep Chlorophyll Maximum (3% salinity), PC6: Punta Cormoran (6% salinity), SS19: Solar Saltern (19% salinity) and SS37: Solar Saltern (37% salinity). GC% was computed for each read and the percentage of the dataset in intervals of bin width 5 is shown.
We examined the distribution of amino acids (Supplementary Figure S1) and the predicted isoelectric points (pI) of the translated metagenomes (Figure 2, see methods), known to reflect halophilic adaptations. We found that the majority of the proteins in DCM3 dataset have a high pI (∼9.6) and only a small proportion have low pI (∼4.0). The two peaks are conserved in the hypersaline datasets, but the ratio of the low to the high peaks keeps increasing with increasing salinity getting closer to the typical pI plot of a “salt-in” strategist's proteome, i.e. “salt-out” strategists are replaced by “salt-in” ones as salinity increases.
Comparison of isoelectric point profiles of the predicted proteins in the four metagenomic datasets of increasing salinities.
The reads that had a reliable hit to a protein sequence in the NR database were used for the analyses (see methods). The pI was computed for each translated read and is shown as a percentage of the dataset in intervals of bin width 1.
Community Structure
We used 16S rRNA reads from all the datasets to build a picture of the high level phylogenetic groups in each dataset (Figure 3, see methods). We complemented this analysis by comparing the metagenomic reads to known microbial genomes (Supplementary Figure S2, Supplementary Table S2, Supplementary Table S3 and Supplementary Table S4, see methods) and also assigned genus-level names to sequences in each dataset (Table 1, see methods). 10% of 16S rRNA reads from DCM3 (all bacterial), ∼4% reads from PC6 (nearly all bacterial), ∼8% from SS19 (nearly all bacterial) and only ∼2% from SS37 (nearly all archaeal) could not be classified to a high-level taxon using the 16S rRNA sequences. However, the results of all methods of classification were broadly in agreement and show interesting features. All datasets, except the SS37 (which was massively dominated by H. walsbyi, 64% of classified reads, Supplementary Table S4) show a surprisingly diverse assemblage of taxonomic groups. Remarkably, Euryarchaeota remain as the major group at the intermediate salinity of SS19 as well and H. walsbyi, the square archaeon, was by far the most abundant microbe (15% of classified reads, Supplementary Table S3). Some other known trends are readily visible along the salinity gradient, e.g. cyanobacteria are restricted to only DCM3 and PC6 dataset, but at 19%, they are absent. It has been known for a long time that photosynthetic eukaryotes take over the function of primary producers (e.g. Dunaliella) with increasing salinity2,8,12,17.
Apart from Euryarchaeota, other groups like Alphaproteobacteria, that were the dominant group in DCM3, especially because of the abundant presence of Candidatus Pelagibacter of the SAR11 clade16, gradually decreased in numbers as salinity increased and were totally absent at 37% salinity. Similarly, Gammaproteobacteria appear to thrive at all levels of salinity except for the crystallizer where only a very small fraction persists. Bacteroidetes were abundant members of the community at all salinity levels. Salinibacter ruber is the most halophilic bacteria known and has been shown to be a dominant member of crystallizer communities, (5–25% using fluorescence in-situ hybridization)3. In our datasets it appeared as an abundant microbe constituting nearly 8.3% of classified reads in SS19 and 4% in SS37. Moreover, nearly 14% of all reads could be assigned to the taxon Bacteroidetes in the SS19 dataset. The Salinibacter ruber genome is high GC (66% GC), but the Bacteroidetes that are more abundant, at least in the SS19 dataset, are low GC (Supplementary Figure S3). The majority of these reads were assigned to Flavobacteria (Gramella, Croceibacter, Flavobacterium and Cellulophaga), as shown in Supplementary Table S3, in contrast to the high GC Bacteroidetes reads that were assigned to Sphingobacteria (e.g Salinibacter). These low GC Flavobacteria did not appear in the crystallizer dataset SS37, where the majority of these reads were ascribed only to the high GC Salinibacter (Supplementary Table S4).
Apart from these groups, Actinobacteria were clearly present in high numbers in both the PC6 (22%) and SS19 (7%) datasets. Numerous Actinobacteria have even been isolated from salterns, e.g. Actinopolyspora halophila18, Haloactinobacterium album19, but we have very little direct information regarding actinobacterial abundance in hypersaline habitats.
At the genus level (>95% identity to a known species), we were able to assign 52.6%, 44.3%, 44.6% and 70.0% of the 16S rRNAs in DCM3, PC6, SS19 and SS37 datasets, respectively (Table 1). The taxonomic core of the SS19 community appeared to be composed of the genera Haloquadratum (16.5% of the assigned dataset), Halorubrum (16.6%), Alkalilimnicola (15.7%) and Salinibacter (7.0%). Some of these sequences had identities above 96% to described species and belonged to Haloquadratum walsbyi, Salinibacter ruber and Halorubrum lacusprofundi. Alkalilimnicola rRNA sequences shared 96% – 98% sequence identity with Alkalilimnicola ehrlichii 16S rRNA sequence. The finding of Alkalilimnicola-related sequences is not surprising even though this microbe has not yet been reported from salterns, although it was isolated from a hypersaline lake20. Representatives of several other genera not known to have halophilic representatives were found in the SS19 dataset (i.e. Clavibacter, Leifsonia, Roseobacter, Renibacterium and Nitrococcus) (Table 1). In contrast, we were unable to affiliate any rRNA sequence with moderate halophiles of the genera Halomonas and Chromohalobacter which are commonly obtained in pure culture from similar salinities21,22. rRNA sequences recovered from SS37 were affiliated to seven haloarchaeal genera and genus Salinibacter. The most abundant rRNA sequences shared >97.0% sequence identity with Haloquadratum walsbyi (79.0% of assigned dataset) and Salinibacter ruber (9.0% of assigned dataset).
The “non-halophilic” bacteria in SS19
We were interested to know whether bacterial groups presumed to be non-halophilic, identified using the rRNA approach in the SS19 dataset showed any signatures of hypersaline adaptation. Thus, we examined available genomes from cultivated representatives of genera detected in our study. In translated metagenomes (see methods), we looked for low average pI, preference for arginine instead of lysine and comparatively higher acidic amino acid percentages23. We found that cultivated non-halophilic representatives of genera detected in our study (Clavibacter, Leifsonia, Roseobacter, Renibacterium and Nitrococcus) had halophile-like acid-shifted proteomes when compared to marine (Prochlorococcus) or fresh water bacteria (Polynucleobacter) (Table 1 and Supplementary Figure S4). The patterns of amino acid composition in all these genomes showed a preference for arginine instead of lysine (Supplementary Figure S5), Gramella being the only exception to this trend. In addition, several of these genomes also showed high percentages of proteins with acidic isoelectric points, another well-known halophilic adaptation23 (Supplementary Table S5).
Low GC Halophilic Actinobacteria
All existing Actinobacteria isolates from hypersaline environments have high GC genomes24. Recently, freshwater Actinobacteria have been reported to be low GC, challenging the long-held nomenclature of Actinobacteria as “High-GC Gram-Positive” microbes25. Actinobacterial reads comprised significant proportions in the PC6 and SS19 datasets (22% and 7% respectively). However, the analysis of GC content of the reads assigned to the different taxa for PC6 and SS19 showed that Actinobacteria inhabiting PC6 and SS19 had very different GC content (Supplementary Figure S3). The majority of the actinobacterial reads from the SS19 dataset were low GC, while those from PC6 were mainly high GC, although a minor low GC peak was also observed in PC6. This indicates that Actinobacteria in intermediate salinity ranges are also low GC, thus adding new members to the group of low-GC Actinobacteria. The small percentage of reads assigned to Actinobacteria in the DCM3 dataset were also low GC but the number of reads was very low. Despite low values, this finding does indicate that marine Actinobacteria (from near surface samples) might also be low GC organisms. More datasets will need to be examined to show this more convincingly.
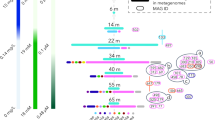
We also examined the pre-assembled scaffolds from the Punta Cormoran dataset15 to see if we could identify scaffolds that might belong to low GC Actinobacteria. As there are no reference genomes available for Actinobacteria from hypersaline habitats and metagenomic assemblies are susceptible to production of chimeric sequences, we designed a careful screening strategy to filter out scaffolds likely to be chimeric (see methods). Although most actinobacterial scaffolds were high GC, we identified a clear cluster of low GC scaffolds while examining GC% and length (Supplementary Figure S6). We did not find any low GC scaffolds with an actinobacterial 16S rRNA sequence. However, a direct search for 16S rRNA in the PC6 and SS19 unassembled metagenomic datasets yielded 208 and 77 actinobacterial rRNAs respectively. Analysis of these metagenomic reads using a comprehensive phylogenetic framework of freshwater actinobacterial sequences26 and our analysis of several 16S sequences of Actinobacteria recovered from hypersaline habitats from all around the world shows clearly that the vast majority of the actinobacterial reads from SS19 and 36% of actinobacterial reads from PC6 were actually related to the freshwater Luna1-A clade26 (Figure 4). The remaining PC6 reads could be binned into the acIII-A1 tribe (35%)26, the acSTL lineage (29%)26 and weakly into the Luna1 lineage (18%). However, nearly all actinobacterial reads in the SS19 dataset were low GC and a small fraction in PC6 are also low GC (Supplementary Figure S3). Moreover, a small fraction of the assembled scaffolds of the PC6 dataset were also low GC (Supplementary Figure S6). This indicates that the actinobacteria related to the Luna1-A clade are likely to be low GC Actinobacteria, similar to the freshwater Actinobacteria described recently25.
Phylogenetic affiliation of the 16S rRNA actinobacterial reads in the Punta Cormoran and the SS19 datasets.
The names of the actinobacterial clades are indicated to the right. Locations of the metagenomic reads from Punta Cormoran and SS19 datasets are indicated in bold with the number of reads shown within brackets. The scale bar represents 10 base substitutions per 100 nt positions.
Assembly of the 19% salinity dataset SS19
A metagenomic assembly is expected to recreate genomic fragments originating from the most abundant organisms in the sample. However, the possibility of assembling chimeric sequences is high and the results difficult to judge if reference genomes are not available. Assembly was performed using very stringent criteria and tested against the sequenced genome of H. walsbyi, the most abundant archaeaon in this dataset (see methods). From all the 88 assembled contigs, 69 could be ascribed to Euryarchaeota, 15 to Gammaproteobacteria, 2 to Actinobacteria and 1 each to Alphaproteobacteria and Nanoarchaeota. From all euryarchaeal contigs, 44 contigs belonged to low-GC Euryarchaeota, of which 30 were clearly from H. walsbyi (and largely syntenic to the reference genome, see methods). The other 14 low-GC Euryarchaeota contigs were not H. walsbyi.
In order to further assess the relatedness of these contigs we performed principal component analysis (PCA) on the normalized tetranucleotide frequencies of these contigs and observed four distinct clusters (Figure 5). All H. walsbyi contigs formed a very tight cluster, indicating a common pattern in their tetranucleotide frequencies. At least three other clusters, one of low-GC Euryarchaeota contigs (non H. walsbyi), one of high GC Euryarchaeota contigs (note proximity to reference genomes) and another of gammaproteobacterial contigs can be demarcated. The tight clustering obtained in the low-GC euryarchaeal and the gammaproteobacterial contigs suggests that these two clusters might represent two abundant and as yet unknown microbes. The low-GC Euryarchaeota contigs cluster is interesting in that it appears to be considerably different from H. walsbyi and the other reference halophiles. Moreover, the single Nanoarchaeota contig also appears within this cluster.
Principal component analysis of tetranucleotide frequencies of assembled contigs from SS19.
Reference genomes are shown as larger circles. The following types of contigs are shown, Dark Yellow: Gammaproteobacterial contigs, Blue: High GC Euryarchaeota contigs, Green: Assembled contigs assigned to Haloquadratum walsbyi, Yellow: Assembled H. walsbyi contigs with only a single gene without a best hit to H. walsbyi (but still all hits to Euryarchaeota), Light Blue: Low GC Euryarchaeota contigs. The total number of contigs for each cluster (Gammaproteobacteria, High GC Euryarchaeota, H. walsbyi and Low GC Euryarchaeota), the total length, mean length and GC% range is also indicated.
To get an estimate of how abundant these microbes might be in comparison to H. walsbyi, we compared the entire SS19 dataset to these clustered contigs and examined how many reads were recruited by each cluster. The results suggest that both the low- GC Euryarchaote and the gammaproteobacterial contigs are longer than H. walsbyi contigs and recruit even more reads from the SS19 dataset, suggesting that they are at least as abundant (Supplementary Figure S7). However, these genomic fragments do not recruit any reads in the SS37 dataset, indicating these organisms prefer a lower salinity range than found in the crystallizer (Supplementary Figure S8).
The genes in the gammaproteobacterial contigs were most similar to genes of Nitrococcus and Alkalilimnicola. No rRNA genes were found within these contigs, preventing us from further establishing finer phylogenetic affiliation. However, a 16S rRNA gene sequence distantly related to Nitrococcus mobilis was found abundant in this very same environment in a previous PCR-based study and could perhaps correspond to same organism6. We plotted the distribution of pI of the predicted proteins encoded in these contigs. The shape of this plot was similar to that of Alkalilimnicola ehrlichii (Supplementary Figure S9) and indicated that this microbe is a salt-out strategist.
The isoelectric point profile of the predicted proteins in the high GC Euryarchaeota contigs was similar to salt-in strategists (H. walsbyi and S. ruber) (Supplementary Figure S9). These genes gave hits to several haloarchaea (e.g. Haloarcula, Natronomonas, Haloferax) indicating that this organism is a haloarchaeon related to cultured isolates, but unlike the others which we did not find to be very abundant (except H. walsbyi), it was present in high numbers in SS19 dataset (i.e. at levels comparable to H. walsbyi). Only a few genes were present in these two clusters of contigs (107 genes in gammaproteobacterial contigs and 117 genes in the high GC Euryarchaeota contigs) and this relatively small number of genes does not allow us to hypothesize in detail regarding the physiology of both these abundant microbes. However, now that we have some information about these microbes it may be possible to design strategies for their isolation and culture.
Single Cell Genomics Reveals an Abundant Low-GC Archaeon in 19% salinity
We collected a small water sample from the CR30 crystallizer and isolated individual microbial cells using fluorescence-activated cell-sorting (see methods). 16S rRNA amplification using archaeal and bacterial primers was used to identify potentially interesting SAGs (Single cell Amplified Genomes). From one of these SAGs, we sequenced 1/8th of a Roche 454 plate and obtained 241928 sequences, total ∼94 Mb of sequence data from a SAG (referred to as G17 henceforth). Assembly and annotation of G17 (see methods) yielded nearly 448 contigs more than 500 bases (total length ∼1.2 Mb). The GC content of the raw reads (42.11%) and the assembled contigs (42.01%) were much lower than H. walsbyi. This is lower even than the recently described low GC Nanohaloarchaeaon Candidatus Nanosalina (GC 43.5%), which was assembled from the metagenome of acidic, hypersaline Lake Tyrell in Australia27.
Direct comparison of the assembled contigs of G17 against the SS19 and SS37 metagenomes revealed that this microbe was abundant in the SS19 sample but not in SS37 (Supplementary Figure S10). We previously assembled the contigs of an abundant low-GC Euryarchaeote from the metagenome that was also more abundant in SS19 than in SS37. We wondered if the contigs from the assembled metagenome and those from the SAG came from the same microbe. The comparison of the cluster of low GC Euryarchaeota contigs assembled from the SS19 metagenome (nearly 100 kb of total sequence) revealed a near perfect alignment of ∼60kb with the assembled G17 with a %identity level of 99%, confirming they are indeed the same microbe, predicted and isolated by completely different, yet complementary methodologies. The other assembled contig sequences of this group (∼40 kb) did not find any hits in G17, but this is not surprising as we assembled only a partial genome from the SAG. Moreover, principal component analysis (PCA) of the tetranucleotide frequencies of contigs (>5 kb) from the single cell genome along with the assembled contigs from the SS19 metagenome showed a clear clustering of the contigs ascribed to low-GC euryarchaeote (Supplementary Figure S11). Furthermore, the sequences obtained from the PCR amplification of the G17 (forward and reverse) were compared to the assembled rRNA sequence in the G17 assembly and were found to be 100% identical over their entire lengths. This clearly shows that there is only a single organism whose genome has been amplified and assembled. So by remarkable coincidence we have amplified a partial genome of a novel archaeon that is abundant at 19% salinity even though our original sample was taken from 37% salinity. This is not completely unexpected as the water in the solar saltern ponds is continuously moved towards the crystallizer and microbes are carried over in significant amounts.
Single cell genomic data gives more confidence than a metagenomic assembly that the 16S rRNA sequence assembled is non-chimeric, a highly likely possibility with metagenomes, considering its high sequence identity across microbes. We detected only one rRNA sequence in the assembled contigs of the G17. This was a full-length rRNA sequence (1478 bp) long and appeared in a contig that had the 23S ribosomal rRNA gene as well. BLAST searches of this 16S sequence indicated only a distant relationship to known rRNA sequences, with the best hits being in the range of 90% sequence identity. The top hits were clearly archaeal, frequently haloarchaea and methanogenic archaea. A more detailed phylogenetic analysis, using 16S rRNA sequences from several fully sequenced genomes of Euryarchaeota, indicated that this new microbe clusters with the recently described lineage of Nanohaloarchaea27 (Supplementary Figure S12). This 16S rRNA sequence is 90% identical to Candidatus Nanosalina and 88% to Candidatus Nanosalinarum. However, neither of the previously described Nanohaloarchaea recruit well in the SS19 and SS37 datasets (Supplementary Figure S13). We have named this new microbe, G17, as Candidatus Haloredivivus (Halo: salt , redivivus: reconstructed). Examination of taxonomy of the best hits of all the genes of this microbe in the genomic fragments had revealed that the majority of the genes could be affiliated to Halobacteria (n = 286, average similarity 66.5%), followed by Methanomicrobia (n = 86, average similarity 64.6%), Methanococcus (n = 72, average similarity 64.8%), Thermococci (n = 58, average similarity 65.8%) and Methanobacteria (n = 43, average similarity 63.7%). These appear consistent with the 16S rRNA phylogeny that indicates the close relationship of Candidatus Haloredivivus to both Halobacteria and Methanogens and a likely affiliation with Nanohaloarchaea.
An examination of the functions of the predicted genes revealed that this microbe likely has a photoheterotrophic lifestyle like the other haloarchaea, indicated by the presence of a rhodopsin, similar to Cyanothece but different from other Nanohaloarchaea, (73% similar in protein sequence to Candidatus Nanosalina and 75% similar to Candidatus Nanosalinarum) (Figure 6). We also detected a photolyase suggesting interactions with light. Importantly, some genes involved in the degradation of polysaccharides (cellulase, amylase and chitinase) were found in the genome, indicating a heterotrophic metabolism. Other typically archaeal genes were also found in the genome e.g. orc1, radA and a gene coding for the S-layer protein. We also found a gene corresponding to a one Type IV flagellar protein so it might be motile as well. In addition, this microbe appears to be a salt-in strategist, like H. walsbyi, preferring to accumulate salt inside the cytoplasm, as indicated by the isoelectric point profiles of the predicted proteins (Supplementary Figure S9).
Phylogeny of the rhodopsin gene fragments detected in the metagenomic datasets.
Reference sequences have been introduced to provide a framework and represent all major types of microbial rhodopsins described (identified by the name of the microbe and accession number in GenBank). The metagenomic reads are all identified by the name of the dataset from which they were identified except for the one retrieved from the Candidatus Haloredivivus genome. The numbers of identical sequences are indicated in brackets after each read identifier.
Discussion
Aquatic, hypersaline environments have been studied using various molecular approaches3,4,5,6,7,10,11. In this study, we collected vast amounts of sequence data (784 Mb) using a metagenomics approach which provides a PCR and cloning bias-free perspective on community structure. Inclusion of other datasets, e.g. the marine DCM3 dataset and the hypersaline coastal lagoon of Punta Cormoran (PC6) provided an opportunity to examine taxonomic trends with increasing salinity, hypersaline adaptations and to identify novel microbes in these environments.
The isoelectric point of microbial proteins is indicative of the survival strategy employed in hypersaline conditions. There are two principal strategies that microbes employ to keep the cellular proteins stable in the presence of high salt concentrations. The proteins of “salt-in” strategists remain soluble in the presence of high cytoplasmic salt concentrations due to the presence of acidic amino acid residues on the protein surface. Consequently, the pI values of halophilic proteins are more acidic compared to their non-halophilic counterparts23,28. Some halophilic and halotolerant organisms deal with high osmotic pressure by synthesizing compatible solutes, a “salt-out” strategy. Two recent studies addressed the question of amino acid-coding bias on an environmental scale. In communities inhabiting hypersaline mat layers, Kunin et al.29 observed that the average isoelectric point is conspicuously acid-shifted when compared to most bacteria and microbiomes that are non-halophilic. Furthermore, a metagenomic study on a series of environments representing a range of salinities showed that the proportion of encoded arginine increases with salinity30. Both of these phenomena were observed in all three hypersaline datasets analyzed, but also in the genomes of the majority of the non-halophilic relatives of the species found in SS19 dataset. In concordance with the above authors we conclude that the described trends can be considered as indications of hypersaline adaptation. So it appears that these organisms, which have not been suspected to have close halophilic representatives previously, might indeed be capable of survival in hypersaline waters at least up to 19% salinity.
The analysis of the SS19 dataset revealed considerable novelty. We found very little evidence of the presence of species of Halomonas, Chromohalobacter or Salinivibrio, which are commonly isolated from this environment21,22. We were further surprised to find abundant metagenomic reads affiliated with Haloquadratum walsbyi and Salinibacter ruber, at NaCl concentration nearly below their reported range of laboratory growth31. Besides, we found a large number of sequences of presumably non-halophilic bacterial genera that, although targeted, were not recovered in previous PCR-based studies6. In addition, the recently described low GC Actinobacteria25, appear not to be restricted to freshwater alone, but also appear as abundant community members in saline systems up to at least 19%. This is the first report of low GC Actinobacteria in hypersaline habitats and it appears that these low GC Actinobacteria belong to the Luna1-A clade.
However, the most interesting discovery was the partial sequence assembly of a novel and quite abundant low-GC Euryarchaeote collected from the CR30 crystallizer, using a combination of metagenomic and single cell genome sequencing. This single cell amplified genome was found to be identical to the assembled contigs from the 19% salinity dataset. This microbe was found to be abundant in the 19% dataset but was apparently absent from the 37% dataset. Similar microbes have been found to be abundant in the acidic hypersaline Lake Tyrrell in Australia, but the microbes found here are only similar at the level expected to different genera of the same family and actually seem to thrive at different salinities i.e. 19% in this case versus saturation for NaCl in the Lake Tyrrell metagenomic contigs.
We show that by utilizing carefully planned assembly strategies, novel microbes can be detected in metagenomic data. The new group of low GC Actinobacteria, the novel Euryarchaeotes (both low and high GC) and a new abundant Gammaproteobacteria are examples of the utility of this approach. These studies also bring out the wide gap between our perception of the microbial world, blinkered by what is cultivable and what is as yet uncultured and similar studies, carried out across varied habitats will provide us with a more realistic view of the microbial world.
Methods
Two samples were collected from ponds of Bras del Port salterns, located near Alicante, Spain (38° 12' N, 0° 36' W). The crystallizer pond sample (SS37) was taken on June 26, 2008 and the concentrator pond sample (SS19) on July 21, 2008. The salinity was measured using a hand refractometer. Each 50 litre sample was sequentially filtered through a 5.0 μm and 0.22 μm-pore size polycarbonate filters using a peristaltic pump. The environmental DNA was extracted as described before32 and 5 μg of each sample was sent separately for sequencing (Roche 454 GS-FLX system, Titanium chemistry, by GATC, Konstanz, Germany). The amount sequenced corresponded to one full FLX plate for SS19 and half a plate for SS37. The average read length obtained was 417 bp in SS37 and 361 bp in SS19 dataset. Low quality regions were completely clipped using sff_extract (by Jose Blanca). The metagenomic reads were also annotated using the MG-RAST server33. Translated metagenome was obtained by conducting a BLASTX search of the metagenomic reads against the NCBI NR database and extracting the translated query sequences from the alignment (e-value 1e-5). Isoelectric points were computed using the program iep in the EMBOSS package34.
For the selection of actinobacterial scaffolds from the pre-assembled from Punta Cormoran we predicted all the genes in all scaffolds35 and used sequence similarity against the NCBI non-redundant protein database to identify the best hits for each gene. We retained only those scaffolds in which all the genes gave best hits only to Actinobacteria. In addition, to remove, small and spurious scaffolds, we retained only those that were larger than 5 kb or had less than three genes. This left us with 255 scaffolds that could be confidently ascribed as actinobacterial.
Assembly of the metagenomic reads (only reads >100bp) was performed using a stringent criteria of overlap of at least 80 bp of the read and 99% identity and at most a single gap in the alignment (using Geneious Pro 5.4). Assembled contigs that were less than 5 kb in length and those with less than three predicted genes were discarded. We retained only those contigs that gave consistent hits to only a single high level taxon (e.g. Alphaproteobacteria, Euryarchaeota, Bacteroidetes, Actinobacteria). The strict assembly requirements combined with a taxonomic uniformity condition imposed on the assembled sequence resulted in a total of 88 contigs that were more than 5kb in length and had a consistent phylogenetic profile and were hence more likely to originate from a single organism. To test if the assembly strategy produced contigs that were “real”, we manually identified all contigs that belonged to H. walsbyi, well known to be abundant in the salterns. The criterion for assigning a contig to H. walsbyi was that all genes must give best hits to this genome. We identified 27 contigs in which all genes that gave the best hit to H. walsbyi and 3 contigs in which all genes but one gave hits to H. walsbyi (but still all hits were to Euryarchaeota, the high-level taxon). Direct nucleotide comparison using BLASTN, showed that the vast majority of these contigs were also syntenic to the reference genome with nucleotide identities ranging from 98% to 100% (Supplementary Figure S14). Tetranucleotide frequencies of the assembled contigs were computed using the wordfreq program in the EMBOSS package34 and principal component analysis was performed using the R package FactoMineR36.
The sequence data from the single cell amplified genome were processed using sff_extract (by Jose Blanca) and assembled using MIRA assembler. Gene prediction on the assembled contigs was done using MGA35. The predicted protein sequences obtained were compared using BLASTP to the NCBI nr protein database.
16S Ribosomal RNA genes were identified by comparing the datasets against the RDP database37. All reads that matched an rRNA sequence with an alignment length of more than 100 bases and an e-value of 0.001 against the database were extracted. The best named hit (that was not described as uncultured, unknown, or unidentified) was considered a reasonable closest attempt to classify the rRNA sequences. When possible, the sequences were further assigned to genus if they shared ≥95% rRNA sequence identity with a known species.
Phylogenetic analysis of rhodopsin genes was performed by comparing the datasets to the rhodopsin subset of GenBank databases using BLASTX. The reads found to match to our rhodopsin sequences at a similarity level of >70% and alignment length over 100 amino acids were analyzed. The sequence alignment was generated by MUSCLE38 and quality checked by CORE (http://tcoffee.vital-it.ch/cgi-bin/Tcoffee/tcoffee_cgi/index.cgi). Ambiguously aligned positions were removed and the tree was built using PhyML39 with substitution model WAG, a Γ law and 1000 bootstraps. Phylogenetic analysis of the rRNA genes was performed using PhyML39 after multiple sequence alignment using Tcoffee40.
Phylogenetic analysis of the actinobacterial reads was conducted by maximum likelihood (RAxML) with near full length (>1300 nt) reference 16S rRNA gene sequences from a manually curated alignment and highly variable positions masked. The tree contains all major lineages of freshwater Actinobacteria described to date26 and in addition, actinobacterial 16S sequences from several hypersaline sites were also added to create a comprehensive phylogenetic framework for binning the metagenomic reads. Metagenomic reads were added without altering tree topology using maximum parsimony criterion and a 50% base frequency filter in the ARB software package41. Bootstrap values are indicated above nodes with greater than 50% support and the scale bar represents 10 base substitutions per 100 nt positions.
Single Cell Genomics: Water sample was incubated for 10–60 min with SYTO-9 (5 µM final concentration; Invitrogen) and high nucleic acid content prokaryote cells were sorted with a MoFlo™ (Beckman Coulter) flow cytometer using a 488 nm argon laser for excitation, a 70 µm nozzle orifice and a CyClone™ robotic arm for droplet deposition into microplates. The cytometer was triggered on side scatter. The “single 1 drop” mode was used for maximal sort purity, which ensures the absence of non-target particles within the target cell drop and the adjacent drops. Under these sorting conditions, sorted drops contain a few 10's of pL of sample surrounding the target cell42, resulting in low or absent non-target DNA. The accuracy of 10 µm fluorescent bead deposition into the 384-well plates was verified by microscopically examining the presence of beads in the plate wells. Of the 2–3 plates examined each sort day, <2% wells were found to not contain a bead and only <0.5% wells were found to contain more than one bead, indicating very high purity of single cells. In addition, we verified the lack of DNA contamination in the sheath fluid and in sheath fluid lines by performing real-time multiple displacement amplification with the processed sheath fluid as the template.
Single bacterial cells were deposited into 384-well plates containing 0.6 µL per well of TE buffer. Plates were stored at −80°C until further processing. Of the 384 wells, 315 were dedicated for single cells, 66 were used as negative controls (no droplet deposition) and 3 received 10 cells each (positive controls). The cells were lysed and their DNA was denatured using cold KOH43. Genomic DNA from the lysed cells was amplified using multiple displacement amplification (MDA)43,44 in 10 µL final volume. The MDA reactions contained 2 U/uL Repliphi polymerase (Epicentre), 1x reaction buffer (Epicentre), 0.4 mM each dNTP (Epicentre), 2 mM DTT (Epicentre), 50 mM phosphorylated random hexamers (IDT) and 1 µM SYTO-9 (Invitrogen) (all final concentration). The MDA reactions were run at 30°C for 12–16 h and then inactivated by 15 min incubation at 65°C. The amplified genomic DNA was stored at −80°C until further processing. We refer to the MDA products originating from individual cells as single amplified genomes (SAGs).
The instruments and the reagents were decontaminated for DNA prior to sorting and MDA setup, as previously described45. High molecular weight DNA contaminants in all MDA reagents were cross-linked by a UV treatment in a Stratalinker (Stratagene). An empirical optimization of the UV exposure was performed to remove all detectable contaminants without inactivating the reaction. Cell sorting and MDA setup were performed in a HEPA-filtered environment. As a quality control, the kinetics of all MDA reactions was monitored by measuring the SYTO-9 fluorescence using FLUOstar Omega (BMG). The critical point (Cp) was determined for each MDA reaction as the time required to produce half of the maximal fluorescence. The Cp is inversely correlated to the amount of DNA template46. The Cp values were significantly lower in 1-cell wells compared to 0-cell wells (p<0.05; Wilcoxon Two Sample Test) in each well. The MDA products were diluted 50-fold in sterile TE buffer. Then 0.5 µL aliquots of the dilute MDA products served as templates in 5 µL real-time PCR screens targeting the SSU rRNA gene using bacterial primers 27F47 and 907R48 and archaeal primers Arch_344 and Arch_915R48. Forward (5′−GTAAAACGACGGCCAGT−3′) or reverse (5′−CAGGAAACAGCTATGACC−3′) M13 sequencing primer was appended to the 5′ end of each PCR primer to aid direct sequencing of the PCR products. All PCRs were performed using LightCycler 480 SYBR Green I Master mix (Roche) in a LightCycler® 480 II real time thermal cycler (Roche). The real-time PCR kinetics and the amplicon melting curves served as proxies detecting successful SAG target gene amplification. New, 20 µL PCR reactions were set up for the PCR-positive SAGs and the amplicons were sequenced from both ends using M13 targets and Sanger technology by Beckman Coulter Genomics. To obtain sufficient quantity of genomic DNA for shotgun sequencing, the original MDA products of the SAG AAA188-G17 were re-amplified using similar MDA conditions as above: four, replicate 150 µL reactions were performed and then pooled together, resulting in 414 µg/ng genomic dsDNA. Single cell sorting, whole genome amplification and PCR were performed at the Bigelow Laboratory Single Cell Genomics Center (www.bigelow.org/scgc). Our previous studies and other recent publications using our single cell sequencing technique demonstrate the reliability of our methodology with high purity of single cell MDA products45,49,50,51,52,53,54,55.
Sequence data have been deposited in the INSDC Sequence Read Archive under the accession SRP007685. The Whole Genome Shotgun project has been deposited at DDBJ/EMBL/GenBank under the accession AGNT00000000. The version described in this paper is the first version, AGNT01000000.
References
Rodriguez-Brito, B. et al. Viral and microbial community dynamics in four aquatic environments. ISME J 4, 739–751 (2010).
Gasol, J. M. et al. Control of heterotrophic prokaryotic abundance and growth rate in hypersaline planktonic environments. Aquatic Microbial Ecology 34, 193–206 (2004).
Anton, J., Llobet-Brossa, E., Rodriguez-Valera, F. & Amann, R. Fluorescence in situ hybridization analysis of the prokaryotic community inhabiting crystallizer ponds. Environ Microbiol 1, 517–523 (1999).
Anton, J., Rossello-Mora, R., Rodriguez-Valera, F. & Amann, R. Extremely halophilic bacteria in crystallizer ponds from solar salterns. Appl Environ Microbiol 66, 3052–3057 (2000).
Benlloch, S. et al. Archaeal Biodiversity in Crystallizer Ponds from a Solar Saltern: Culture versus PCR. Microb Ecol 41, 12–19 (2001).
Benlloch, S. et al. Prokaryotic genetic diversity throughout the salinity gradient of a coastal solar saltern. Environ Microbiol 4, 349–360 (2002).
Casamayor, E. O. et al. Changes in archaeal, bacterial and eukaryal assemblages along a salinity gradient by comparison of genetic fingerprinting methods in a multipond solar saltern. Environ Microbiol 4, 338–348 (2002).
Pedros-Alio, C. et al. The microbial food web along salinity gradients. FEMS Microbiol Ecol 32, 143–155 (2000).
Rodriguez-Valera, F. et al. Explaining microbial population genomics through phage predation. Nat Rev Microbiol 7, 828–836 (2009).
Legault, B. A. et al. Environmental genomics of "Haloquadratum walsbyi" in a saltern crystallizer indicates a large pool of accessory genes in an otherwise coherent species. BMC Genomics 7, 171 (2006).
Rodriguez-Valera, F., Ventosa, A., Juez, G. & Imhoff, J. F. Variation of environmental features and microbial populations with salt concentrations in a multi-pond saltern. Microbial Ecology 11, 107–115 (1985).
Estrada, M., Henriksen, P., Gasol, J. M., Casamayor, E. O. & Pedros-Alio, C. Diversity of planktonic photoautotrophic microorganisms along a salinity gradient as depicted by microscopy, flow cytometry, pigment analysis and DNA-based methods. FEMS Microbiol Ecol 49, 281–293 (2004).
Javor, B. Hypersaline environments - Microbiology and Biogeochemistry. (Springer-Verlag, 1989).
Warren, J. K. Evaporites, sediments, resources and hydrocarbon., (Springer, 2006).
Rusch, D. B. et al. The Sorcerer II Global Ocean Sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol 5, e77 (2007).
Ghai, R. et al. Metagenome of the Mediterranean deep chlorophyll maximum studied by direct and fosmid library 454 pyrosequencing. The ISME Journal 4, 1154–1166 (2010).
He, Q. et al. Cloning and characterization of a plastidic glycerol 3-phosphate dehydrogenase cDNA from Dunaliella salina. J Plant Physiol 164, 214–220 (2007).
Gochnauer, M. B. et al. Isolation and characterization of Actinopolyspora halophila, gen. et sp. nov., an extremely halophilic actinomycete. Can J Microbiol 21, 1500–1511 (1975).
Tang, S. K. et al. Haloactinobacterium album gen. nov., sp. nov., a halophilic actinobacterium and proposal of Ruaniaceae fam. nov. Int J Syst Evol Microbiol 60, 2113–2119 (2010).
Hoeft, S. E. et al. Alkalilimnicola ehrlichii sp. nov., a novel, arsenite-oxidizing haloalkaliphilic gammaproteobacterium capable of chemoautotrophic or heterotrophic growth with nitrate or oxygen as the electron acceptor. Int J Syst Evol Microbiol 57, 504–512 (2007).
Arahal, D. R. et al. Chromohalobacter salexigens sp. nov., a moderately halophilic species that includes Halomonas elongata DSM 3043 and ATCC 33174. Int J Syst Evol Microbiol 51, 1457–1462 (2001).
Arenas, M. et al. Halomonas ilicicola sp. nov., a moderately halophilic bacterium isolated from a saltern. Int J Syst Evol Microbiol 59, 578–582 (2009).
Paul, S., Bag, S. K., Das, S., Harvill, E. T. & Dutta, C. Molecular signature of hypersaline adaptation: insights from genome and proteome composition of halophilic prokaryotes. Genome Biol 9, R70 (2008).
de la Haba, R., Sanchez-Porro, C., Marquez, M. C. & Ventosa, A. in Extremophiles Handbook (ed Horikoshi K.) (Springer, 2011).
Ghai, R., McMahon, K. D. & Rodriguez-Valera, F. Breaking a paradigm: cosmopolitan and abundant freshwater actinobacteria are low GC. Environ Microbiol Reports (2011).
Newton, R. J., Jones, S. E., Eiler, A., McMahon, K. D. & Bertilsson, S. A guide to the natural history of freshwater lake bacteria. Microbiol Mol Biol Rev 75, 14–49 (2011).
Narasingarao, P. et al. De novo metagenomic assembly reveals abundant novel major lineage of Archaea in hypersaline microbial communities. ISME J (2011).
Soppa, J. From genomes to function: haloarchaea as model organisms. Microbiology 152, 585–590 (2006).
Kunin, V. et al. A bacterial metapopulation adapts locally to phage predation despite global dispersal. Genome Res 18, 293–297 (2008).
Rhodes, M. E., Fitz-Gibbon, S. T., Oren, A. & House, C. H. Amino acid signatures of salinity on an environmental scale with a focus on the Dead Sea. Environ Microbiol 12, 2613–2623 (2010).
Anton, J. et al. Salinibacter ruber gen. nov., sp. nov., a novel, extremely halophilic member of the Bacteria from saltern crystallizer ponds. Int J Syst Evol Microbiol 52, 485–491 (2002).
Martin-Cuadrado, A. B. et al. Metagenomics of the deep Mediterranean, a warm bathypelagic habitat. PLoS One 2, e914 (2007).
Meyer, F. et al. The metagenomics RAST server – a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9, 386 (2008).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 16, 276–277 (2000).
Noguchi, H., Taniguchi, T. & Itoh, T. MetaGeneAnnotator: detecting species-specific patterns of ribosomal binding site for precise gene prediction in anonymous prokaryotic and phage genomes. DNA Res 15, 387–396 (2008).
Lê, S., Josse, J. & Husson, F. FactoMineR: An R Package for Multivariate Analysis. Journal of Statistical Software 25, 1–18 (2008).
Cole, J. R. et al. The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37, D141–145 (2009).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32, 1792–1797 (2004).
Guindon, S. & Gascuel, O. A simple, fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52, 696–704 (2003).
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J Mol Biol 302, 205–217 (2000).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res 32, 1363–1371 (2004).
Sieracki, M. E., Poulton, N. J. & Crosbie, N. Automated isolation techniques for microalgae. (Elsevier Academic, New York, 2005).
Raghunathan, A. et al. Genomic DNA amplification from a single bacterium. Appl Environ Microbiol 71, 3342–3347 (2005).
Dean, F. B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc Natl Acad Sci U S A 99, 5261–5266 (2002).
Stepanauskas, R. & Sieracki, M. E. Matching phylogeny and metabolism in the uncultured marine bacteria, one cell at a time. Proc Natl Acad Sci U S A 104, 9052–9057 (2007).
Zhang, K. et al. Sequencing genomes from single cells by polymerase cloning. Nat Biotechnol 24, 680–686 (2006).
Page, K. A., Connon, S. A. & Giovannoni, S. J. Representative freshwater bacterioplankton isolated from Crater Lake, Oregon. Appl Environ Microbiol 70, 6542–6550 (2004).
Casamayor, E. O., Schafer, H., Baneras, L., Pedros-Alio, C. & Muyzer, G. Identification of and spatio-temporal differences between microbial assemblages from two neighboring sulfurous lakes: comparison by microscopy and denaturing gradient gel electrophoresis. Appl Environ Microbiol 66, 499–508 (2000).
Fleming, E. J. et al. What's new is old: resolving the identity of Leptothrix ochracea using single cell genomics, pyrosequencing and FISH. PLoS One 6, e17769 (2011).
Hess, M. et al. Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331, 463–467 (2011).
Heywood, J. L., Sieracki, M. E., Bellows, W., Poulton, N. J. & Stepanauskas, R. Capturing diversity of marine heterotrophic protists: one cell at a time. ISME J 5, 674–684 (2011).
Martinez-Garcia, M. et al. High-throughput single-cell sequencing identifies photoheterotrophs and chemoautotrophs in freshwater bacterioplankton. ISME J (2011).
Swan, B. K. et al. Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the dark ocean. Science in press (2011).
Woyke, T. et al. Assembling the marine metagenome, one cell at a time. PLoS One 4, e5299 (2009).
Yoon, H. S. et al. Single-cell genomics reveals organismal interactions in uncultivated marine protists. Science 332, 714–717 (2011).
Acknowledgements
This work was supported by projects MAGYK (BIO2008-02444), MICROGEN (Programa CONSOLIDER-INGENIO 2010 CSD2009-00006), METAGHALO (BIO2009-10138) and HALOMET (CGL2010-19303) from the Spanish Ministerio de Ciencia e Innovación, DIMEGEN (PROMETEO/2010/089) from the Generalitat Valenciana and P10-CVI-6226 from the Junta de Andalucía and NSF Grant DEB-919290. FEDER funds supported this project. R.G. and A-B.M-C were supported by a Juan de la Cierva scholarship, from the Spanish Ministerio de Ciencia e Innovación. A.B. F. was supported by a fellowship from the Junta de Andalucía. L.P. was supported by CONSOLIDER-INGENIO 2010 and the Slovenian Research Agency programme P1-0198. RS was supported by the U.S. National Science Foundation grants DEB-841933 and OCE-821374.
Author information
Authors and Affiliations
Contributions
This work was conceived by F R-V and AV. Laboratory work was done by ABF, LP and ABMC. Single cell work was conceived and carried out by RTP and RS. Analysis was carried out by RG, LP, ABF, ABMC, CMM, KDM, BRB, FRV and CSP. The manuscript was written by FR-V, RG and LP. All authors read and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Tables and Figures
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Ghai, R., Pašić, L., Fernández, A. et al. New Abundant Microbial Groups in Aquatic Hypersaline Environments. Sci Rep 1, 135 (2011). https://doi.org/10.1038/srep00135
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00135
This article is cited by
-
Metagenomic analysis of the effects of salinity on microbial community and functional gene diversity in glacial meltwater estuary, Ny-Alesund, Arctic
Brazilian Journal of Microbiology (2024)
-
Genome analysis of haloalkaline isolates from the soda saline crater lake of Isabel Island; comparative genomics and potential metabolic analysis within the genus Halomonas
BMC Genomics (2023)
-
Microbial life in 25-m-deep boreholes in ancient permafrost illuminated by metagenomics
Environmental Microbiome (2023)
-
Functional differentiation determines the molecular basis of the symbiotic lifestyle of Ca. Nanohaloarchaeota
Microbiome (2022)
-
Microbial diversity in extreme environments
Nature Reviews Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.