Abstract
Left-right asymmetry is subtle but pervasive in the human central nervous system. This asymmetry is initiated early during development, but its mechanisms are poorly known. Forebrains and midbrains were dissected from six human embryos at Carnegie stages 15 or 16, one of which was female. The structures were divided into left and right sides, and RNA was isolated. RNA was sequenced with 100 base-pair paired ends using Illumina Hiseq 4000. After quality control, five paired brain sides were available for midbrain and forebrain. A paired analysis between left- and right sides of a given brain structure across the embryos identified left-right differences. The dataset, consisting of Fastq files and a read count table, can be further used to study early development of the human brain.
Design Type(s) | transcription profiling by high throughput sequencing design • organism part comparison design |
Measurement Type(s) | transcription profiling assay |
Technology Type(s) | RNA sequencing |
Factor Type(s) | regional part of brain • Cerebral Hemisphere |
Sample Characteristic(s) | Homo sapiens • forebrain • midbrain |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
In addition to the anterior-posterior axis and the dorsal-ventral axis, the body has a left-right axis. Left-right asymmetry of the internal organs such as the heart and liver is obvious, but the central nervous system is asymmetric as well. The asymmetry of the central nervous system is at least partly uncoupled from the asymmetry of the internal organs, as evidenced from observations of people with reversed organ placement1,2.
The earliest observed evidence for structural brain asymmetry in development is a small difference in average volume of the left and right sides of the choroid plexus in human foetuses at 11 post conception weeks (PCW)3. From 8 PCW, behavioural asymmetry has been observed: embryos more often moved their right arms than their left arms4,5. In a previous study, we showed that human spinal cord and hindbrain have left-right differences in gene expression at stages between 4 and 8 PCW6. These observations indicate a well-regulated genetic programme to manage early asymmetrical brain development. However, whether the human midbrain and forebrain also show transcriptional asymmetry at such early stages was unknown prior to generation of the current dataset (previous studies had analyzed foetuses at later stages). In fish, molecular studies have revealed mechanisms and genes that are involved in brain laterality7, but it is unknown how much of this is conserved in humans.
Our previous studies6 suggested a difference in maturation rate between left and right sides of spinal cord and hindbrain as one mechanism to arrive at left-right differences. Also, we observed that the left-right pattern in the hindbrain was a mirror-image of that in the spinal cord. To further elucidate the left-right differences in the developing human brain, we now dissected midbrains and forebrains of six human embryos, at Carnegie stages 15 or 16, into left and right. The embryos were from social pregnancy terminations. RNA was isolated from each brain side separately. Barcoded cDNA fragments were sequenced on an Illumina HiSeq 4000 sequencer, using paired-end sequencing with a read length of 100 bases. After filtering for quality, the median size was 5.65 Gb per library. Data are made available as Fastq-files and a processed data table containing gene counts. Our analysis of this dataset will be published elsewhere.
Methods
Collection, library preparation and sequencing
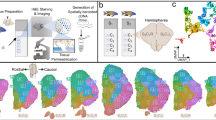
These methods are expanded versions of descriptions in our related work titled “Subtle left-right asymmetry of gene expression profiles in embryonic and foetal human brains”8. A schematic overview of this data collection can be found in Fig. 1.
Six human embryos at Carnegie stages 15 or 16. Forebrain and midbrain were dissected and divided into left and right. RNA was isolated from the complete tissue sample. Paired end sequencing (100 bp) was done at BGI, China. Alignment and read counts were performed against GrCh38 with RefSeq gene definitions. Adapted from Version 8.25 from the Textbook OpenStax Anatomy and Physiology Published May 18, 2016, licensed under the Creative Commons Attribution 4.0 International license.
Six embryos were collected by the MRC/Wellcome-Trust funded Human Developmental Biology Resource (HDBR – www.hdbr.org) (United Kingdom) at either CS15 or CS16, therefore estimated between 5 and 5.5 weeks post conception. The embryos were obtained anonymously from voluntary medical terminations (a combination of mifepristone and misoprostol) or physical termination (according to the mother’s choice), following appropriate informed consent by the donors, and with ethical approval from the Newcastle and North Tyneside NHS Health Authority Joint Ethics Committee. Donors to HDBR are asked to give written consent for the embryonic material to be collected, and are only approached once a decision to terminate their pregnancy has been made. The abortions were not because of observed congenital malformations or suspected genetic disorders. Karyotypes were normal.
The development of the embryos was assessed and designated to the relevant Carnegie stage (CS)9, using a practical staging guide devised to enable staging to a particular CS and using the external morphology of a single sample10. Forebrain and midbrain were separated and then dissected into left and right sides. The dissections were performed by multiple people from a small 4 member group following the same dissection procedure.
RNA was extracted at HDBR Newcastle. Tissue samples were divided into sub-samples each weighing thirty milligrams, which was the maximum loading capacity of the columns of the RNA purification kits). The sub-samples were homogenised using a Precellys 24 bead mill homogeniser (Bertin Corp. Rockville, MD, USA) using ceramic 1.4 mm beads for soft tissue homogenising (CK14) with 600ul of RTL plus Buffer with 10 μl/ml of β-Mercaptoethanol and 5 μl/ml of reagent DX (Qiagen, Venlo, the Netherlands). RNA and DNA was extracted from the tissue with a QIAcube using an AllPrep DNA/RNA Mini Kit (Qiagen) following the manufacturer’s recommended protocol. RNA was then pooled from all 30 mg sub-samples belonging to a given embryo’s left or right midbrain, or left or right forebrain. RNA quality was assessed using an Agilent 2100 Bioanalyzer (Applied Biosystems, Santa Clara, CA, USA), and then RNA was shipped on dry ice to Beijing Genomics Institute (BGI) Shenzhen/HongKong, China (www.genomics.cn). The embryos were five males and one female. The female was from a physical pregnancy termination, the males from chemical pregnancy terminations.
At BGI, the RNA was treated with DNAse and quality was determined again on an Agilent 2100 Bioanalyzer. All samples passed the quality filters of: ≥ 4 μg RNA; concentration ≥ 80 ng/μL; RIN ≥ 7.0; 28 S/18 S ≥ 1.0; smooth baseline and normal 5 S peak. 200 ng of RNA was treated with Oligo-dT beads to enrich for mRNA out of the pool of total RNA. Afterwards, the purified RNA was broken into short segments in Fragment Buffer (New England Biolabs). After fragmentation, cDNA was generated with random hexamer primers, using First Strand Master Mix with Superscript II (Invitrogen) reverse transcriptase, and next the Second Strand Master Mix. End-repair Mix was added, and the repaired fragments were purified with AMPure(R) XP beads (Agencourt(R) ). A-tailing mix was added, then cDNA fragments were connected with adapters, following standard Illumina(R) procedures. Next followed again a round of purification with AMPure XP beads. To enrich the library further, several rounds of PCR were used, followed by a new round of purification. The final library was quantitated in two ways: the average molecule length was determined using the Agilent 2100 bioanalyzer instrument (Agilent DNA 1000 Reagents), and the library was quantified by real-time quantitative PCR (QPCR) (TaqMan Probe, Thermo Fisher Scientific). The libraries were now firstly amplified within the flow cell on the cBot instrument for cluster generation (HiSeq® 4000 PE Cluster Kit, Illumina). Then, the clustered flowcell was loaded onto the HiSeq 4000 Sequencer for paired-end sequencing (HiSeq® 4000 SBS Kit, Illumina) with read lengths of 100 bases.
Data processing
At BGI, raw reads were filtered to exclude reads with more than 5% unknown bases, reads which contained more than 20% bases with quality score below 15, and reads with adapters. After filtering, the median size was 5.65 Gb per library (range 3.66–7.26 Gb). RNA sequencing data were produced as fastq-files by BGI. These files comprise the bulk of the current data release (Data Citation 1).
Sequence reads were then aligned to the Human reference GRCh38 from UCSC (http://genome-euro.ucsc.edu) using Hisat2 (v2.0.4). Using the same reference with RefSeq gene annotations, reads were then counted per gene using RSEM (v1.3.09). Both packages used bowtie211. In R (version 3.3.2), expression data were normalized and transformed into log2 cpm (counts per million). From these processes we produced a file containing per gene log2 transformed read counts per sample, with genes indicated by Entrez IDs, which is also included in the current release (Data Citation 1). The report from BGI with details about the sequencing and mapping results is included as a supplement (Supplementary File 1). Figures 6 and 7 of this Supplementary File 1 show a satisfactory coverage of transcripts by the reads.
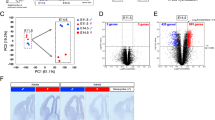
Finally, multidimensional scaling (MDS) analysis in R showed that forebrain and midbrain separated into two distinct clusters on the basis of overall gene expression similarity. However, the right side of one of the forebrain samples fell into the midbrain cluster. Data for the forebrain were therefore discarded for this embryo, and are excluded from the current release. In addition, both sides of one midbrain sample clustered with the forebrain group, and the midbrain samples for this embryo were discarded as well, and excluded from the current release. We are therefore releasing data from four males and one female for each structure, though not exactly the same embryos for forebrain and midbrain (See Table 1 for an overview of the samples).
Note that for the purposes of MDS analysis, and our other downstream analysis of this dataset which we report elsewhere, genes were additionally filtered to retain only those for which at least three libraries had at least five reads per gene, separately for forebrain and midbrain. However, this step has not been applied to the gene count table which we are releasing.
Code availability
Codes that were used for processing the data are available as supplement (Supplementary File 2).
Data Records
FASTQ sequencing files for six embryos times two brain structures times two sides, minus one forebrain and one midbrain = 20 libraries (2 files per library because of the paired reads), have been deposited to the Gene Expression Omnibus with series number GSE99302 (Data Citation 1). Individual accession numbers for each biological sample are also provided in Table 2 and with more details in Supplementary File 3 (Excel).
For users not interested in re-processing the data, a processed file of per gene log2 transformed read counts per sample, with genes indicated by Entrez IDs, is also included ‘GSE99302_BGI_2Tissues_expression_log2.txt.gz’ at NCBI Gene Expression Omnibus GSE99302. Per sample, text files are available with the following columns: gene_id (Entrez gene id), expected count (number of reads), FPKM (Fragments Per Kilobase Million), Symbol (Gene Symbol).
Technical Validation
Descriptive analysis with the FastQC software (v0.11.5, Babraham Bioinformatics, Cambridge, USA) showed that the quality was high without overrepresented sequences or adapters, and phred base quality score mostly > 37 (Fig. 2). GC content varied between 48 and 50%.
A separate pipeline using HiSat (v0.1.6 beta, for mapping against hg19) and GATK (v3.4.0 (ref. 12)) was used at BGI to create genotype calls for single nucleotide polymorphisms (SNPs) from the RNA sequencing data. The SNP data were then used in plink (v1.0713) to confirm that left and right pairs of matched samples came from the same individual, and to confirm the sexes of the samples. A second confirmation for sex was found by looking at the expression data for the X-chromosomal gene XIST and the Y-chromosomal genes EIF1AY and KDM5D. XIST should be expressed in the female, while EIF1AY and KDM5D should not. Sex was confirmed, as well as proper matching of RNA from a given embryo.
For our analysis of this dataset which we report elsewhere, MDS analysis was repeated using the expression data of the remaining five embryos per brain structure (after the aforementioned exclusions). The forebrain and midbrain clustered separately, and also the female was separated from the males, while left and right sides from the same embryo tended to cluster together (Fig. 3).
Comparison with published studies
Expression levels per gene, for the forebrain and the midbrain, were compared to those from a publically available dataset in which left and right had not been divided: E-MTAB-484014 in the ArrayExpress database (Data Citation 2). From this latter dataset, data for embryos in the ages 4 to 9 weeks were used. Data were available as fastQ files and were processed in the same way as described above. Across all genes, correlations of average expression per gene for our embryos (5–5.5 weeks old) with those in dataset E-MTAB-4840 (4–9 weeks old) were r = 0.84 for forebrain and r = 0.80 for midbrain.
Additional information
How to cite this article: de Kovel, C. G. F. et al. Transcriptomic analysis of left-right differences in human embryonic forebrain and midbrain. Sci. Data 5:180164 doi: 10.1038/sdata.2018.164 (2018).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
McManus, I. C., Martin, N., Stubbings, G. F., Chung, E. M. & Mitchison, H. M. Handedness and situs inversus in primary ciliary dyskinesia. Proc Biol Sci 271, 2579–2582 (2004).
Tanaka, S., Kanzaki, R., Yoshibayashi, M., Kamiya, T. & Sugishita, M. Dichotic listening in patients with situs inversus: brain asymmetry and situs asymmetry. Neuropsychologia 37, 869–874 (1999).
Abu-Rustum, R. S., Ziade, M. F. & Abu-Rustum, S. E. Reference Values for the Right and Left Fetal Choroid Plexus at 11 to 13 Weeks An Early Sign of “Developmental” Laterality? J Ultras Med 32, 1623–1629 (2013).
Hepper, P. G. The developmental origins of laterality: Fetal handedness. Developmental Psychobiology 55, 588–595 (2013).
Hepper, P. G., Shahidullah, S. & White, R. Handedness in the human fetus. Neuropsychologia 29, 1107–1111 (1991).
de Kovel, C. G. F. et al. Left-Right Asymmetry of Maturation Rates in Human Embryonic Neural Development. Biol Psychiatry 82, 204–212 (2017).
Roussigne, M., Blader, P. & Wilson, S. W. Breaking symmetry: the zebrafish as a model for understanding left-right asymmetry in the developing brain. Dev Neurobiol 72, 269–281 (2012).
de Kovel, C. G. F., Lisgo, S. N., Fisher, S. E. & Francks, C. Subtle left-right asymmetry of gene expression profiles in embryonic and foetal human brains. Sci Rep 8, 10.1038/s41598-018-29496-2 (2018).
O’Rahilly, R. & Muller, F. Developmental stages in human embryos: revised and new measurements. Cells Tissues Organs 192, 73–84 (2010).
Bullen, P., Wilson, D. in Molecular Genetics of Early Human Development (eds Strachan, T., Lindsay, S. & Wilson, D. I. ) (Garland Science, 1997).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359 (2012).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome research 20, 1297–1303 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. American journal of human genetics 81, 559–575 (2007).
Lindsay, S. J. et al. HDBR Expression: A Unique Resource for Global and Individual Gene Expression Studies during Early Human Brain Development. Front Neuroanat 10, 86 (2016).
Data Citations
Gene Expression Omnibus GSE99302 (2017)
ArrayExpress E-MTAB-4840 (2017)
Acknowledgements
We thank the women who donated the embryos and foetuses to the Human Developmental Biology Resource (UK). C.G.F.de.K was supported by an Open Programme grant (824.14.005) to C.F. from the Netherlands Organization for Scientific Research (NWO).
Author information
Authors and Affiliations
Contributions
Clyde Francks designed the project, acquired the funding and did the supervision. Steven Lisgo was in charge of collecting and dissecting the embryos, extracting the RNA and performing the quality controls. Carolien de Kovel performed the bioinformatics, such as data cleaning and alignment of the Fastq files, generating gene counts etc. Clyde Francks and Carolien de Kovel wrote the manuscript, and Steven Lisgo critically read the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
ISA-Tab metadata
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
de Kovel, C., Lisgo, S. & Francks, C. Transcriptomic analysis of left-right differences in human embryonic forebrain and midbrain. Sci Data 5, 180164 (2018). https://doi.org/10.1038/sdata.2018.164
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2018.164
This article is cited by
-
Subtle left-right asymmetry of gene expression profiles in embryonic and foetal human brains
Scientific Reports (2018)