Abstract
Background
To define the optimal chemotherapy regimen for each patient we therefore used tissue from patients to identify molecular prognostic or predictive biomarkers.
Methods
Endoscopic biopsy specimens from primary lesions and surgical specimens on a phase III trial in patients with unresectable advanced or recurrent gastric cancer treated with docetaxel with cisplatin plus S-1 (DCS) or cisplatin plus S-1 (CS), were collected. We measured the mRNA expression of ERCC1 and analyzed SNPs in GSTP1 and ERCC1.
Results
Low ERCC1 expression was associated with favorable prognosis for overall survival, OS by multivariable analysis (P = 0.001). There were significant interactions between the two treatment arms of DCS and CS, and ERCC1 mRNA expression. In patients with low ERCC1 expression of a favorable prognosis, DCS therapy was inferior to CS (P = 0.046). In addition to GSTP1 rs1695 (HR 1.728), ERCC1 rs3212980, ERCC1 rs2298881, ERCC1 rs3212964 with high expression of ERCC1 mRNA were associated with significantly worse prognosis with regard to OS.
Conclusions
ERCC1 mRNA is an independent prognostic factor and predictive marker that can be used to guide the addition of docetaxel. The SNPs of ERCC1 and GSTP1 could be also prognostic or predictive factors.
Similar content being viewed by others
Introduction
Fluoropyrimidine and platinum-based combination therapies are the most commonly used and acceptable first-line therapies for patients with HER-2 negative gastric cancer worldwide [1]. The V325 study demonstrated the superiority of triplet chemotherapy using docetaxel plus cisplatin and 5-fluorouracil (5-FU) over doublet chemotherapy with cisplatin and 5-FU for patients with advanced gastric cancer [2]. This triplet regimen has not been accepted globally as a standard palliative treatment because it elicits severe neutropenia and confers a small survival advantage. JCOG 1013 trial showed that the triplet therapy with docetaxel added to cisplatin and S-1 (DCS) did not prolong overall survival (OS) and progression-free survival (PFS) in patients with unresectable advanced or recurrent gastric cancer compared with the doublet of cisplatin and S-1 (CS) [3]. Poor performance status (PS), peritoneal metastasis, liver metastasis, histological type, and disease status (unresectable advanced or recurrent) are established clinical prognostic factors for metastatic gastric cancer [4, 5]. On the other hand, perioperative triplet therapy with docetaxel, fluoropyrimidine, and oxaliplatin showed survival benefit for patients with locally advanced resectable gastric cancer [6, 7]. These mixed results show that a better understanding of biological predictive or prognostic markers of conventional cytotoxic agents is required. Armed with this knowledge, physicians then can give patients the optimal drugs to prolong their survival and improve their quality of life. This is especially important for the use of cytotoxic drugs, which are not always effective in every patient and often cause severe adverse events.
Excision repair cross-complementation group 1 (ERCC1) is an important component of the nucleotide excision repair pathway, which repairs DNA intra-strand, inter-strand, and DNA-protein crosslinks caused by cisplatin. DNA repair systems allow cells to overcome the DNA damage induced by chemotherapy [8]. In the JCOG9912 trial, low ERCC1 mRNA expression was a significant independent favorable prognostic factor in patients with metastatic gastric cancer who received first-line chemotherapy with 5-FU monotherapy, S-1 monotherapy, or cisplatin plus irinotecan [9]. Low mRNA levels of ERCC1 in primary gastric cancer have been associated with a higher overall response rate and longer survival following cisplatin treatment [9,10,11,12,13,14]. The expression of ERCC1 mRNA was suggested as a predictive and prognostic marker in resectable gastric cancer patients receiving chemotherapy. Providing complementary roles to ERCC1, X-ray repair cross-complementing group (XRCC1) is critical mediator of base excision repair and single-strand break repair [15, 16].
Single nucleotide polymorphisms (SNPs) ERCC1 rs3212986, rs2298881, rs11615, XRCC1 rs25487, and rs1695 in glutathione S-transferase pi 1 (GSTP1; an enzyme that is involved in cytosolic platinum detoxification [16, 17]), have been suggested as prognostic markers in preclinical studies [18,19,20,21,22]. The ERCC1 genotypes had no significant association with OS in patients who received perioperative therapy with epirubicin, cisplatin, and 5-FU (ECF) in the MAGIC trial [23]. However, patients with a TYMS 2 R/2 R genotype derived a larger benefit from perioperative ECF than patients with TYMS 3 R genotypes [23]. Although low ERCC1 protein expression may be a better prognostic marker, the lack of adequate commercially available antibodies to detect the active ERCC1 subtype has limited the interpretation of immunohistochemical studies [24,25,26]. Therefore, we designed the current study to identify differences in survival and tumor regression after CS or DCS therapy. By taking a multi-omics approach, our aim was to quantify the real-world utility of these candidate molecular markers in clinical practice.
Materials and methods
Patients were randomly assigned (1:1) to receive DCS (docetaxel 40 mg/m2 and cisplatin 60 mg/m2 on day 1 intravenously, and S-1 40–60 mg twice a day orally for 2 weeks, every 4 weeks) or CS (cisplatin 60 mg/m2 intravenously on day 8, and S-1 40–60 mg orally twice a day for 3 weeks, every 5 weeks) in the JCOG1013. Written informed consent to be enrolled in JCOG1013 was obtained before registration and the opportunity to refuse to provide tumor samples was provided through web sites of the National Cancer Center and the Japan Clinical Oncology Group (JCOG) according to the Japanese Ethical Guidelines for Medical and Biological Research Involving Human Subjects. The protocol of this translational study was approved by the institutional review board of the National Center for Global Health and Medicine and each participating hospital and complied according to the criteria of REMARK (reporting recommendations for tumor marker prognostic studies [27].
For the analysis, 5 × 10-μm sections or 10 × 4- or 5-μm sections were prepared from formalin-fixed paraffin embedded tumor tissues (FFPE). The tumor cells on the sections of interest were selectively isolated by macrodissection. ERCC1 and TYMS and an internal reference gene (β-actin) were quantified with a fluorescence-based real-time detection method (LightCycler96 System and FastStart essential DNA Probes Master, Roche Diagnostics, Rotkreuz, Switzerland), both OS and PFS in the patients with lower ERCC1 mRNA and lower TYMS mRNA expression were better than those in the higher ERCC1 and higher TYMS expression in our previous study [9]. The primers and probes used have been described previously [13]. Gene expression values (relative mRNA levels) are expressed as quantification cycle (Cq) ratios (differences between Cq values) between the genes of ERCC1 or TYMS and an internal reference gene (β-actin) [28, 29].
The NCC Oncopanel test is a hybridization capture-based NGS assay designed to examine mutations, amplifications, and homozygous deletions of the entire coding region of 123 genes of clinical or preclinical relevance, along with rearrangements of 13 oncogenes included in the panel [30]. We modified the NCC Oncopanel for pharmacogenetic analysis to examine 66 SNPs in DPYD, VEGFA, ABCB1, PRKDC, MGMT, GSTP1, ACRV, TYMS, XRCC1, POLR1G, and ERCC1. We paid particular attention to the genetic changes of XRCC1 and GSTP1 that have already been reported to affect the effect of cisplatin as well as ERCC1.
Immunohistochemical staining of ERCC1 was performed using antibody 9D11 [31] and an Autostainer Link 48 device (Agilent, Santa Clara, CA). The evaluation area was limited to the region where gastric cancer cells were present in the total tissue of biopsy specimens, and in approximately three locations identified by low magnification (objective lens x4) of the surgical resection samples. The staining intensity was graded on a scale of 0–3. The expression of ERCC1 protein in cancer cells was normalized to the average ERCC1 nuclear staining intensity in intraregional vascular endothelial cells, which was set at 2. Thus, cancer cells expressed similar levels if their average staining intensity was 2, stronger expression if the value was 3, weaker if the value was 1, and were considered negative with a staining intensity of 0. The strongest intensity of ERCC1 expression of cancer cells in the region was measured.
Statistical analysis
The gene expression levels of ERCC1 and TYMS were categorized into low and high groups by the median or an optimal cutoff value based on a SNP analysis to assess the associations between gene expression levels and OS, and PFS, and response rate. Categorical data were compared using Fisher’s exact test. Survival function was estimated with the Kaplan–Meier method, and differences between survival functions were compared with the log-rank test. Hazard ratios (HRs) with 95% confidence intervals (CIs) based on a Cox proportional hazards model were used to provide quantitative summaries of the gene expression data. Variables for the multivariable analysis included the genes with expression levels (high or low) that showed associations in the univariable analyses in this study, as well as the patient’s background, such as Eastern Cooperative Oncology Group (ECOG) PS, age, sex, number of metastatic sites, previous gastrectomy, presence or absence of target lesions according to RECIST version 1.0, histological classification (differentiated/undifferentiated) [32], and presence or absence of peritoneal metastasis. All reported P-values are two-sided, and the level of statistical significance was set at P < 0.05. All analyses were performed using R version 4.2.3 (R Foundation for Statistical Computing, Vienna, Austria) and SAS version 9.4 (SAS Institute Inc., Cary, NC).
Results
Relationship between ERCC1 and TYMS expression and survival
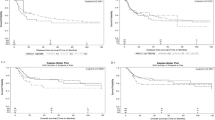
Tissue samples for this study were collected from 523 endoscopic biopsy specimens and 136 surgical specimens taken before the treatment of 741 randomized patients in JCOG1013 (Supplementary Fig. 1). The baseline characteristics were equally distributed among the subsets for ERCC1 and TYMS. A univariable analysis of the whole study population showed that both OS (HR, 0.861; 95% CI: 0.703–1.054; P = 0.147) and PFS (HR, 0.882; 95% CI: 0.726–1.071; P = 0.205) in the low ERCC1 mRNA groups were generally better than those in the high ERCC1 mRNA groups. There were significant interactions between the two treatment arms of DCS and CS, and ERCC1 mRNA expression (Table 1). High ERCC1 mRNA expression was significantly associated with worse prognosis, and DCS was inferior to CS in patients with low ERCC1 mRNA expression who had a better prognosis (Fig. 1). The response rates of CS and DCS were similar: 43% (47/109) and 36% (37/104) in the ERCC1-mRNA high group, and 29% (30/103) and 37% (40/109) in the ERCC1-mRNA low group. There were no significant differences in OS or PFS according to the expression of TYMS. Multivariable analyses for survival with ERCC1 mRNA expression and clinical characteristics showed that independent prognostic factors were ERCC1 mRNA and ECOG PS for OS, and ERCC1 mRNA and peritoneal metastasis for PFS (Table 2).
Relationship between ERCC1 mRNA and protein expression
There was no statistically significant correlation between ERCC1 Cq ratio and protein expression. The protein staining intensities (scale 0–3) in the ERCC1 mRNA-high group were 3 in 32/142 (23%), 2 in 84/142 (46%), 1 in 20/142 (38%), and 0 in 6/142 (4%). Staining intensities in the ERCC1-mRNA low group were 3 in 51/196 (26%), 2 in 98/196 (54%), 1 in 33/196 (17%), and 0 in 14/196 (7%). ERCC1 expression had no predictive and prognostic significance with regard to OS, PFS or tumor shrinkage.
ERCC1, XRCC1, and GSTP1 SNPs as prognostic factors
Genomic analysis using the NCC Oncopanel was performed on 111 surgical specimens and 13 endoscopic biopsy samples; most patients were postoperative recurrent cases. DCS was superior to CS for patients with recurrent gastric cancer after gastrectomy in terms of OS (21.9 months [95%CI, 17.9–26.3] vs 15.9 months [12.9–19.0], HR = 0.64 [0.45–0.90], P = 0.0095), but not superior in patients with unresectable advanced gastric cancer (Supplementary Fig. 2 MST, 13.0 months [11.9–14.3] vs 15.0 months [14.1–16.1], HR = 1.14 [0.963–1.36], P = 0.127). There were significant differences between the baseline patient characteristics of unresectable advanced and recurrent gastric cancer regarding PS 1 (37% vs. 25%), only one metastatic site (38% vs. 66%), liver metastasis (31% vs. 21%), and bone metastasis (5% vs. 10%). Thus, patients with recurrent gastric cancer had more favorable prognostic factors than those with unresectable advanced gastric cancer. Among the 124 patients for whom data were available, DCS was superior to CS (P < 0.01). There were no differences in the distribution of ERCC1 mRNA expression between patients with unresectable advanced and those with recurrent gastric cancer. The prognostic values of ERCC1, XRCC1, and GSTP1 in patients treated with DCS or CS are shown in Table 3. ERCC1 rs3212964 (HR 1.533), ERCC1 rs2298881 (HR 1.525), and GSTP1 rs1695 (HR 2.336) were significant prognostic factors with regard to PFS. The ERCC1 rs3212980 TT, rs3212964 TT, rs11615 AA, rs3212948 GG, and rs2298881 AA alleles tended to have higher mean values of ERCC1 mRNA expression when compared with the reference alleles. Other remarkable HRs of DCS vs CS in terms of OS were 0.259 in DPYD rs2297595 TC (vs 0.514 in TT), 1.777 in ABCB1 rs7787082 AA (vs 0.435 in GA and 0.426 in GG), 0.237 in XRCC2 rs1799782 AA (vs 0.499 GA and 0.678 in GG) (Supplementary Table 2).
ERCC1, XRCC1, and GSTP1 SNPs as predictive factors
ERCC1, XRCC1, GSTP1 SNPs were predictive of the benefits of DCS or CS in terms of OS and PFS (Table 4). When compared with each reference allele, the ERCC1 rs3212980 TT, rs3212964 TT, rs11615 AA, rs3212948 GG and rs2298881 AA alleles tended to have larger HR for both OS and PFS in the DCS-treated patients versus those treated with CS. Patients with GSTP1 GG who were treated with DCS had a shorter OS. The response rates in the CS and DCS groups were 20% vs. 27% in ERCC1 rs3212980 TT, 6% vs. 24% in rs3212964 TT, 0% vs. 29% in rs11615 AA, 0% vs. 29% in rs3212948 GG, and 6% vs. 25% in rs2298881 AA.
Intra-tumoral gene mutation and outcomes
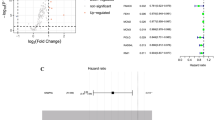
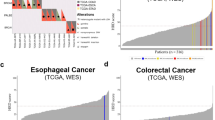
TP53 mutation was observed in 43% of cases, ARID1A in 12%, PIK3CA in 8.1%, RHOA in 7.3%, APC in 6.5%, BRCA2 in 4.8%, KRAS in 4.8%, SMAD4 in 4.8%, MSH6 in 2.4%, MLH1 in 0.8%, and MSH2 in 0.8% (Fig. 2). There was no difference of OS between mutant and wild-type of TP53 (HR 1.02; 95%CI: 0.69 to 1.51; P = 0.91). There was a tendency of poorer OS in patients with RHOA, SMAD4, or microsatellite instability- high (MSH6/MLH1/MSH2) genetic alterations.
a TP53 mutation was most commonly observed mutation in gastric cancer. b TP53 status had no impact on OS following treatment with either CS or DCS (HR, 1.02; 95% confidence interval, 0.69–1.51). c Patients with wild type TP53 had better PFS than those with TP53 mutant (HR, 1.33; 95% confidence interval, 0.91–1.93). d OS of patients segregated based on mutations of representative genes.
Discussion
DNA repair capacity is a major determinant of cisplatin resistance, with ERCC1 protein playing an essential role in nucleotide excision repair. Here, we found that patients with high ERCC1 expression had a poorer overall survival than those with low ERCC1 expression. The DCS treatment regimen was inferior to CS in patients with low ERCC1 expression and a favorable prognosis. Therefore, the triplet therapy is not required in this patient subset. Recurrent gastric cancer patients with ERCC1 rs3212964, ERCC1 rs2298881, or GSTP1 rs1695 SNPs had higher ERCC1 expression and had a worse OS. Moreover, DCS was inferior to CS in terms of PFS if patients had the ERCC1 rs11615 AA or ERCC1 rs3212948 GG SNPs. DCS was also inferior to CS in terms of OS in patients with the XRCC1 rs25487 TT or GSTP1 rs1695 GG SNPs. Our data confirmed that ERCC1 expression, as well as specific SNPs in ERCC1, XRCC1, and GSTP1, are significant prognostic indicators that could guide the choice between DCS or CS treatment regimens. There were no significant differences in the impact of ERCC1 mRNA expression on prognosis between TP53 mutant and wild-type (P = 0.5, Pearson’s Chi-squared test).
Our previous ancillary investigation of another randomized controlled trial, JCOG9912, showed that low ERCC1 expression was a significant independent favorable prognostic factor in patients with advanced gastric cancer who were also receiving first-line chemotherapy. The baseline patient characteristics were different between JCOG9912 and the current JCOG1013 trial. About 50% of analyzed patients in JCOG1013 had peritoneal metastasis compared with 27% in JCOG9912. The frequency of liver metastasis was 30% in this study but 48% in JCOG9912. Although the treatment regimens were different (CS and DCS were used in JCOG1013 whereas 5-FU monotherapy, S-1 monotherapy, or cisplatin plus irinotecan combination therapy were used in JCOG9912) the prognostic effect of ERCC1 expression was still evident in both cases. The ERCC1 rs3212964 and ERCC1 rs2298881 SNPs were found in patients with higher ERCC1 expression, which explains why they were associated with a poorer prognosis. Because genotyping from FFPE breast cancer specimens was significantly concordant with genotyping from germline DNA, the effects of cytotoxic chemotherapy and its impact on survival can be predicted by DNA analysis of blood or buccal mucosa [33].
Since high ERCC1 expression is an indicator of poor prognosis, it has been a challenge to show the superiority of alternative combination therapies without platinum with regard to survival [14, 34]. From our results, some patients with low ERCC1 expression have a good prognosis, and this is compromised if they are given the more toxic triplet therapy. Hence, the administration of DCS to patients with an otherwise favorable prognosis, particularly those who are eligible for curative resection. should be avoided. Commercially available methods to evaluate ERCC1 mRNA expression status are warranted, as they will guide the choice of triplet DCS or doublet CS, which in turn will reduce the incidence of toxicity-related death and will improve patients’ quality of life. S-1 was effective in ERCC1-high patients with resectable stage II or III gastric cancer after surgery in the adjuvant setting. However, S-1 monotherapy did not impart a statistically significant survival benefit in ERCC1-low patients in the ACTS-GC trial [35]. Therefore, ERCC1 mRNA expression could be predictive marker in the adjuvant setting. Thus, ERCC1 is not only related to the resistance of cisplatin but other chemotherapeutic agents. The results of our present study, which show that DCS is more effective than CS in ERCC1-high patients, are consistent with these previously published data.
The ERCC1 gene generates four isoforms designated 201, 202, 203, and 204 by alternative splicing. Currently, available antibodies such as 8F1 cannot discriminate between these isoforms and thus cannot guide therapeutic decision-making regarding cisplatin combined therapy in patients with non-small-cell lung cancer; this requires specific detection of the unique functional ERCC1-202 isoform [26]. ERCC1 expression was analyzed by western blot in seventeen human gastric cancer cell lines, and all were found to express either 201, 202, and/or 203, but not 204 [31]. Although domain-specific functions that are clinically relevant to the 202 isoform of ERCC1 have been identified, there is a lack of structure-function data for the other isoforms with regard to cisplatin resistance. Considering this, we suggest that antibodies capable of detecting not only 202 but also other major ERCC1 isoforms may be useful for evaluation of cisplatin sensitivity. Therefore, we used 9D11, a novel antibody that recognizes ERCC1 isoforms 201, 202, and 203 [31]. We did not, however, identify a significant prognostic impact of ERCC1 protein expression when using this 9D11 antibody. Evaluating ERCC1 protein expression levels using anti-ERCC1 antibodies is not useful for predicting the prognosis of patients receiving cisplatin combination therapy. In conclusion, we believe that this is the first study to evaluate the prognostic and predictive value of ERCC1 gene alteration, ERCC1 mRNA expression, and GSTP1 polymorphism in patients with unresectable or recurrent gastric cancer. We demonstrate that genomic and transcriptomic analyses can guide the selection of cytotoxic chemotherapy and recommend that gene sequencing is performed before selecting patients for specific treatment regimens.
References
Yamada Y. Present status and perspective of chemotherapy for patients with unresectable advanced or metastatic gastric cancer in Japan. Glob Health Med. 2020;2:156–63.
Van Cutsem E, Moiseyenko VM, Tjulandin S, Majlis A, Constenla M, Boni C, et al. Phase III study of docetaxel and cisplatin plus fluorouracil compared with cisplatin and fluorouracil as first-line therapy for advanced gastric cancer. J Clin Oncol. 2006;24:4991–7.
Yamada Y, Boku N, Mizusawa J, Iwasa S, Kadowaki S, Nakayama N, et al. Docetaxel plus cisplatin and S-1 versus cisplatin and S-1 in patients with advanced gastric cancer (JCOG1013): an open-label, phase 3, randomized controlled trial. Lancet Gastroenterol Hepatol. 2019;4:501–10.
Grenader T, Waddell T, Peckitt C, Oates J, Starling N, Cunningham D, et al. Prognostic value of neutrophil-to-lymphocyte ratio in advanced oesophago-gastric cancer: exploratory analysis of the REAL-2 trial. Ann Oncol. 2016;27:687–92.
Yamada Y, Higuchi K, Nishikawa K, Gotoh M, Fuse N, Sugimoto N, et al. Phase III study comparing oxaliplatin plus S-1 with cisplatin plus S-1 in chemotherapy-naïve patients with advanced gastric cancer. Ann Oncol. 2015;26:141–8.
Al-Batran SE, Homann N, Pauligk C, Goetze TO, Meiler J, Kasper S, et al. Perioperative chemotherapy with fluorouracil plus leucovorin, oxaliplatin, and docetaxel versus fluorouracil or capecitabine plus cisplatin and epirubicin for locally advanced, resectable gastric or gastro-oesophageal junction adenocarcinoma (FLOT4): a randomized, phase 2/3 trial. Lancet. 2019;393:1948–57.
Kang YK, Yook JH, Park YK, Lee JS, Kim YW, Kim JY, et al. PRODIGY: a phase III study of neoadjuvant docetaxel, oxaliplatin, and S-1 plus surgery and adjuvant S-1 versus surgery and adjuvant S-1 for resectable advanced gastric cancer. J Clin Oncol. 2021;39:2903–13.
Reed E, Kohn KW. Platinum analogues. In Chabner BA, Collins JM, editors. Cancer chemotherapy: principles and practice. Philadelphia, PA: J.B. Lippincott Company; 1990. p. 465–90.
Yamada Y, Boku N, Nishina T, Yamaguchi K, Denda T, Tsuji A, et al. Impact of excision repair cross-complementing gene 1 (ERCC1) on the outcomes of patients with advanced gastric cancer: correlative study in Japan Clinical Oncology Group Trial JCOG9912. Ann Oncol. 2013;24:2560–5.
Metzger R, Leichman CG, Danenberg KD, Danenberg PV, Lenz HY, Hayashi K, et al. ERCC1 mRNA levels complement thymidylate synthase mRNA levels in predicting response and survival for gastric cancer patients receiving combination cisplatin and fluorouracil chemotherapy. J Clin Oncol. 1998;16:309–16.
Lenz HJ, Leichman CG, Danenberg KD, Danenberg PV, Groshen S, Cohen H, et al. Thymidylate synthase mRNA level in adenocarcinoma of the stomach: a predictor for primary tumor response and overall survival. J Clin Oncol. 1996;14:176–82.
Wei J, Zou Z, Qian X, Ding Y, Xie L, Sanchez JJ, et al. ERCC1 mRNA levels and survival of advanced gastric cancer patients treated with a modified FOLFOX regimen. Br J Cancer. 2008;98:1398–402.
Matsubara J, Nishina T, Yamada Y, Moriwaki T, Shimoda T, Kajiwara T, et al. Impacts of excision repair cross-complementing gene 1 (ERCC1), dihydropyrimidine dehydrogenase, and epidermal growth factor receptor on the outcomes of patients with advanced gastric cancer. Br J Cancer. 2008;98:832–9.
Iqbal S, McDonough S, Lenz HJ, Ilson D, Burtness B, Nangia CS, et al. Randomized, phase II study prospectively evaluating treatment of metastatic esophageal, gastric, or gastroesophageal cancer by gene expression of ERCC1: SWOG S1201. J Clin Oncol. 2019;38:472–9.
Thompson LH, Brookman KW, Jones NJ, Allen SA, Carrano AV. Molecular cloning of the human XRCC1 gene, which corrects defective DNA strand break repair and sister chromatid exchange. Mol Cell Biol. 1990;10:6160–71.
Liu JY, Liu QM, Li LR. Association of GSTP1 and XRCC1 gene polymorphisms with clinical outcomes of patients with advanced non-small cell lung cancer. Genet Mol Res. 2015;14:10331–7.
Murata T, Hatayama I, Kakizaki I, Satoh K, Sato K, Tsuchida S. Lentinan enhances sensitivity of mouse colon 26 tumor to cis-diamminedichloroplatinum (II) and decreases glutathione transferase expression. Jpn J Cancer Res. 1996;87:1171–8.
Zou M, Hu X, Xu B, Tong T, Jing Y, Xi L, et al. Glutathione S‑transferase isozyme alpha 1 is predominantly involved in the cisplatin resistance of common types of solid cancer. Oncol Rep. 2019;41:989–98.
Seo BG, Kwon HC, Oh SY, Lee S, Kim SG, Kim SH, et al. Comprehensive analysis of excision repair complementation group 1, glutathione S-transferase, thymidylate synthase and uridine diphosphate glucuronosyl transferase 1A1 polymorphisms predictive for treatment outcome in patients with advanced gastric cancer treated with FOLFOX or FOLFIRI. Oncol Rep. 2009;22:127–36.
Yin M, Yan J, Martinez-Belibrea E, Graziano F, Lenz HJ, Kim HJ, et al. ERCC1 and ERCC2 polymorphisms predict clinical outcomes of oxaliplatin-based chemotherapies in gastric and colorectal cancer: a systemic review and meta-analysis. Clin Cancer Res. 2011;17:1632–40.
Li Y, Liu Z, Liu H, Wang LE, Tang D, Ajani JA, et al. ERCC1 and ERCC2 variants predict survival in gastric cancer patients. PLoS One. 2013;8:e71994.
Lv Y, Xu M, Sun Y, Liu Y, Zhao L, Liu X, et al. Prognostic significance of excision repair cross complementation group 1 rs2298881 in patients with gastric cancer receiving platinum-based chemotherapy: a PRISMA-compliant meta-analysis. Medicine (Baltimore). 2021;100:e26850.
Smyth A, Zhang S, Cunningham D, Wotherspoon A, Soong R, Peckitt C, et al. Pharmacogenetic analysis of the UK MRC (Medical Research Council) MAGIC Trial: association of polymorphisms with toxicity and survival in patients treated with perioperative epirubicin, cisplatin, and 5-fluorouracil (ECF) Chemotherapy. Clin Cancer Res. 2017;23:7543–9.
Kim JS, Kim MA, Kim TM, Lee SH, Kim DW, Im SA, et al. Biomarker analysis in stage III-IV (M0) gastric cancer patients who received curative surgery followed by adjuvant 5-fluorouracil and cisplatin chemotherapy: epidermal growth factor receptor (EGFR) associated with favourable survival. Br J Cancer. 2009;100:732–8.
Wang J, Zhou XQ, Li JY, Cheng JF, Zeng XN, Li X, et al. Prognostic significance of ERCC1 expression in postoperative patients with gastric cancer. Chin J Cancer Res. 2014;26:323–30.
Friboulet L, Olaussen KA, Pignon JP, Shepherd FA, Tsao MS, Graziano S, et al. ERCC1 isoform expression and DNA repair in non-small-cell lung cancer. N Engl J Cancer. 2013;368:1101–10.
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM, et al. Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst. 2005;97:1180–4.
Horikoshi T, Danenberg KD, Stadlbauer THW, Volkenandt M, Shea LC, Aigner K, et al. Quantitation of thymidylate synthase, dihydrofolate reductase, and DT diaphorase gene expression in human tumors using the polymerase chain reaction. Cancer Res. 1992;52:108–16.
Koizumi W, Tanabe S, Azuma M, Ishido K, Nishimura K, Sasaki T, et al. Impacts of fluorouracil-metabolizing enzymes on the outcomes of patients treated with S-1 alone or S-1 plus cisplatin for first-line treatment of advanced gastric cancer. Int J Cancer. 2010;126:162–70.
Sunami K, Ichikawa H, Kubo T, Kato M, Fujiwara Y, Shimomura A, et al. Feasibility and utility of a panel testing for 114 cancer-associated genes in a clinical setting: a hospital-based study. Cancer Sci. 2019;110:1480–90.
Oishi T, Sasaki Y, Tong Y, Chen L, Onodera T, Iwasa S, et al. A newly established monoclonal antibody against ERCC1 detects major isoforms of ERCC1 in gastric cancer. Glob Health Med. 2021;3:226–35.
Japanese Gastric Cancer Association. Japanese classification of gastric carcinoma:3rd English edition. Gastric Cancer. 2011;14:101–12.
Hertz DL, Kidwell KM, Thibert JN, Gersch C, Regan MM, Skaar TC, et al. Genotyping concordance in DNA extracted from formalin-fixed paraffin embedded (FFPE) breast tumor and whole blood for pharmacogenetic analyses. Mol Oncol. 2015;9:1868–76.
Cobo M, Isla D, Montes A, Sanchez JM, Provencio M, Viñolas N, et al. Customizing cisplatin based on quantitative excision repair cross-complementing 1 mRNA expression: a phase III trial in non–small-cell lung cancer. J Clin Oncol. 2007;25:2747–54.
Kitada K, Ochiai A, Ichikawa W, Terashima M, Kurahashi I, Sakuramoto S, et al. Interaction between intratumoral ERCC1 expression and adjuvant treatment with S-1 on the survival of patients enrolled in the ACTS-GC study. J Clin Oncol. 2012;30:53.
Acknowledgements
We thank Ms. Hideko Morita, National Center for Global Health and Medicine, and Ms. Kazue Horio, Kitasato University, for their technical support and fruitful discussion.
Funding
Financial support for this research was provided by the NCGM Intramural Research Fund (20A1014) and National Cancer Research and Development Fund (2020-J-3, 2023-J-03). The funding sources had no role in the study design, data collection, analysis, interpretation, or writing of the manuscript. The corresponding author had full access to all of the data and the final responsibility to submit for publication.
Author information
Authors and Affiliations
Contributions
Y. Yamada: conceptualization, investigation, writing-original draft, data curation, formal analysis, editing, funding acquisition, and project administration. Investigation, writing-original draft, and editing. K. Nagashima: data curation, formal analysis, and editing. M. Azuma: investigation, data curation, and editing. M. Masutani: investigation, data curation, and editing. H. Ichikawa: investigation, data curation, formal analysis, and editing. S. Iwasa: investigation and editing. N. Takahashi: investigation and editing. H. Hirano: investigation and editing. K. Kanato: investigation and editing. N. Machida: investigation and editing. T. Kinoshita: investigation and editing. H. Hata: investigation and editing. H. Kawakami: investigation and editing. D. Takahari: investigation and editing. N. Boku: investigation and editing. Y. Kurokawa: investigation and editing. M. Terashima: investigation and editing. T. Yoshikawa: Investigation and editing. S. Sekine: investigation and editing. N. Hiraoka: investigation, data curation, and editing.
Corresponding author
Ethics declarations
Competing interests
Y. Yamada has received lecture fee from Taiho Pharmaceutical. K. Nagashima has received consulting fee from Senju Pharmaceutical, Toray Industries, and Kowa. S. Iwasa is the employee of Chugai Pharma, and has received honoraria from Taiho Pharmaceutical, Lilly Japan, Chugai Pharma, Ono Pharmaceutical, Daiichi-Sankyo, Bristol-Myers Squibb Japan, and Agilent, institutional research funding from Daiichi-Sankyo, Bristol-Myers Squibb, Eisai, Merck Serono, Bayer, Ono Pharmaceutical, Astellas Pharma, Pfizer, Seagen, Zymeworks, Taiho Pharmaceutical, and AstraZeneca. N. Takahashi has received honoraria from Ono Pharmaceutical, Bristol-Myers Squibb Japan, and Taiho Pharmaceutical. H. Hirano has received honoraria Bristol-Myers Squibb Japan, Chugai Pharma, Novartis, Taiho Pharmaceutical, Fujifilm, and Teijin Pharma, and institutional research funding BeiGene. N. Machida has received honoraria from Taiho Pharmaceutical, Bristol-Myers Squibb Japan, Ono Pharmaceutical, Daiichi-Sankyo, Lilly Japan, Yakult Honsha, Takeda, MSD K.K, Chugai Pharma, Merck KGaA, Novartis, and Astellas Pharma, and institutional research funding from MSD, AstraZeneca, Amgen, Ono Pharmaceutical, Taiho Pharmaceutical, ALX Oncology, and Bristol-Myers Squibb Japan. T. Kinoshita has received lecture fees from Johnson & Johnson, Intuitive Surgical, Medtronic, Olympus Medical Systems, Daiichi Sankyo, Lilly Japan, Bristol-Myers Squibb Japan, Kaken Pharmaceutical, and Taiho Pharmaceutical. H. Kawakami has received cousulting fee from Taiho Pharmaceutical, Bristol-Myers Squibb Japan, Ono Pharmaceutical, Lilly Japan, and Daiichi-Sankyo, honoraria from Chugai Pharma/ Roche, Taiho Pharmaceutical, Ono Pharmaceutical, Bristol-Myers Squibb Japan, Yakult Honsha, Takeda, MSD K.K, Merck Serono, Lilly Japan, Daiichi-Sankyo, Bayer, and institutional research funding from Chugai Pharma, Daiichi-Sankyo, Eisai, Kobayashi Pharmaceutical. D. Takahari has received honoraria, Taiho Pharmaceutical, Lilly Japan, Ono Pharmaceutical, Yakult Honsha, Bristol-Myers Squibb Japan, and Daiichi-Sankyo/UCB Japan, and institutional research funding from Ono Pharmaceutical and Taiho Pharmaceutical. N. Boku has received honoraria from Taiho Pharmaceutical, Ono Pharmaceutical, Bristol-Myers Squibb Japan, Daiichi-Sankyo, Lilly, and institutional research funding from Ono Pharmaceutical and Takeda. Y. Kurokawa has received lecture fees from Taiho Pharmaceutical, Ono Pharmaceutical, Daiichi Sankyo, Lilly, Nippon Kayaku, and Kaken Pharmaceutical, and research fundings from Taiho Pharmaceutical, Yakult Honsha, AstraZeneca, Ono Pharmaceutical, and MSD Oncology outside of the submitted work. M. Terashima has received honoraria from Chugai Pharma, Taiho Pharmaceutical, Lilly, Ono Pharmaceutical, Bristol-Myers Squibb, Daiichi-Sankyo, Yakult Honsha, Johnson & Johnson, Olympus, and Intuitive Surgical. T. Yoshikawa has received consulting fee from MSD Oncology and TERUMO, honoraria from Chugai Pharma, Taiho Pharmaceutical, Daiichi-Sankyo, Ono Pharmaceutical, Medtronic, Bristol-Myers Squibb, EA Pharma, Johnson & Johnson/ Janssen, AstraZeneca, Intuitive Surgical, Japan Blood Products Organization, Olympus, Lilly, Astellas Pharma, and research funding from Lilly. S. Sekine has received honoraria from MSD and Roche, and institutional research funding from AstraZeneca. M. Azuma, M. Masutani, H. Ichikawa, K. Kanato, and H. Hata, and N. Hiraoka have no relationships to disclose.
Ethics approval and consent to participate
Written informed consent to be enrolled in JCOG1013 was obtained before registration and the opportunity to refuse to provide tumor samples was provided through web sites of the National Cancer Center and the Japan Clinical Oncology Group (JCOG) according to the Japanese Ethical Guidelines for Medical and Biological Research Involving Human Subjects. The protocol of this translational study was approved by the institutional review board of the National Center for Global Health and Medicine and each participating hospital and complied according to the criteria of REMARK (reporting recommendations for tumor marker prognostic studies.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamada, Y., Nagashima, K., Azuma, M. et al. Predictive and prognostic value of excision repair cross-complementing group 1 in patients with advanced gastric cancer. BJC Rep 2, 18 (2024). https://doi.org/10.1038/s44276-024-00046-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s44276-024-00046-w