Abstract
The border zone (BZ) of the infarcted heart is a geographically complex and biologically enigmatic interface separating poorly perfused infarct zones (IZs) from remote zones (RZs). The cellular and molecular mechanisms of myocardial BZs are not well understood because microdissection inevitably combines them with uncontrolled amounts of RZs and IZs. Here, we use single-cell/nucleus RNA sequencing, spatial transcriptomics and multiplexed RNA fluorescence in situ hybridization to redefine the BZ based on cardiomyocyte transcriptomes. BZ1 (Nppa+Xirp2−) forms a hundreds-of-micrometer-thick layer of morphologically intact cells adjacent to RZs that are detectable within an hour of injury. Meanwhile, BZ2 (Nppa+Xirp2+) forms a near-single-cell-thick layer of morphologically distorted cardiomyocytes at the IZ edge that colocalize with matricellular protein-expressing myofibroblasts and express predominantly mechanotransduction genes. Surprisingly, mechanical injury alone is sufficient to induce BZ genes. We propose a ‘loss of neighbor’ hypothesis to explain how ischemic cell death mechanically destabilizes the BZ to induce its transcriptional response.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
sc/snRNA-seq data and spatial transcriptomic sequencing data have been deposited to the Gene Expression Omnibus under accession no. GSE214611. All other data supporting the findings in this study are included in the main article and associated files.
Code availability
The code used to process this data is publicly available at Zenodo (https://zenodo.org/record/7055957#.Y00deezMIqu)
References
Factor, S. M., Sonnenblick, E. H. & Kirk, E. S. The histologic border zone of acute myocardial infarction—islands or peninsulas? Am. J. Pathol. 92, 111–124 (1978).
Janse, M. J. et al. The “border zone” in myocardial ischemia. An electrophysiological, metabolic, and histochemical correlation in the pig heart. Circ. Res. 44, 576–588 (1979).
van Duijvenboden, K. et al. Conserved NPPB+ border zone switches from MEF2- to AP-1-driven gene program. Circulation 140, 864–879 (2019).
Vivien, C. J., Hudson, J. E. & Porrello, E. R. Evolution, comparative biology and ontogeny of vertebrate heart regeneration. NPJ Regen. Med. 1, 16012 (2016).
Smith, R. M., Black, A. J., Velamakanni, S. S., Akkin, T. & Tolkacheva, E. G. Visualizing the complex 3D geometry of the perfusion border zone in isolated rabbit heart. Appl. Opt. 51, 2713–2721 (2012).
Pirolo, J. S., Hutchins, G. M. & Moore, G. W. Infarct expansion: pathologic analysis of 204 patients with a single myocardial infarct. J. Am. Coll. Cardiol. 7, 349–354 (1986).
Erlebacher, J. A., Richter, R. C., Alonso, D. R., Devereux, R. B. & Gay, W. A. Jr. Early infarct expansion: structural or functional? J. Am. Coll. Cardiol. 6, 839–844 (1985).
Gao, X. M., Xu, Q., Kiriazis, H., Dart, A. M. & Du, X. J. Mouse model of post-infarct ventricular rupture: time course, strain- and gender-dependency, tensile strength, and histopathology. Cardiovasc Res. 65, 469–477 (2005).
Mounier-Vehier, F. et al. Borderzone infarct subtypes: preliminary study of the presumed mechanism. Eur. Neurol. 34, 11–15 (1994).
Wetstein, L., Michelson, E. L., Simson, M. B., Moore, E. N. & Harken, A. H. Increased normoxic-to-ischemic tissue borderzone as the cause for reentrant ventricular tachyarrhythmias. J. Surg. Res. 32, 526–534 (1982).
Hearse, D. J. & Yellon, D. M. The “Border zone” and myocardial protection: a time for reassessment? Acta Med. Scand. Suppl. 651, 37–46 (1981).
Duncanson, E. R. & Mackey-Bojack, S. M. Histologic examination of the heart in the forensic autopsy. Acad. Forensic Pathol. 8, 565–615 (2018).
Jackson, B. M. et al. Extension of borderzone myocardium in postinfarction dilated cardiomyopathy. J. Am. Coll. Cardiol. 40, 1160–1167 (2002).
Yang, Y. et al. Age-related differences in postinfarct left ventricular rupture and remodeling. Am. J. Physiol. Heart Circ. Physiol. 294, H1815–H1822 (2008).
Cavasin, M. A., Tao, Z., Menon, S. & Yang, X.-P. Gender differences in cardiac function during early remodeling after acute myocardial infarction in mice. Life Sci. 75, 2181–2192 (2004).
Tsao, C. W. et al. Heart Disease and Stroke Statistics-2022 Update: A Report From the American Heart Association. Circulation 145, e153–e639 (2022).
von Lueder, T. G., Kotecha, D., Atar, D. & Hopper, I. Neurohormonal blockade in heart failure. Card. Fail. Rev. 3, 19–24 (2017).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
Calcagno, D. M. et al. The myeloid type I interferon response to myocardial infarction begins in bone marrow and is regulated by Nrf2-activated macrophages. Sci. Immunol. 5, eaaz1974 (2020).
Calcagno, D. M. et al. SiglecF(HI) marks late-stage neutrophils of the infarcted heart: a single-cell transcriptomic analysis of neutrophil diversification. J. Am. Heart Assoc. 10, e019019 (2021).
Dick, S. A. et al. Self-renewing resident cardiac macrophages limit adverse remodeling following myocardial infarction. Nat. Immunol. 20, 29–39 (2019).
King, K. R. et al. IRF3 and type I interferons fuel a fatal response to myocardial infarction. Nat. Med. 23, 1481–1487 (2017).
Forte, E. et al. Dynamic interstitial cell response during myocardial infarction predicts resilience to rupture in genetically diverse mice. Cell Rep. 30, 3149–3163.e6 (2020).
Hu, P. et al. Single-nucleus transcriptomic survey of cell diversity and functional maturation in postnatal mammalian hearts. Genes Dev. 32, 1344–1357 (2018).
Martini, E. et al. Single-cell sequencing of mouse heart immune infiltrate in pressure overload-driven heart failure reveals extent of immune activation. Circulation 140, 2089–2107 (2019).
Ren, Z. et al. Single-cell reconstruction of progression trajectory reveals intervention principles in pathological cardiac hypertrophy. Circulation 141, 1704–1719 (2020).
Tucker, N. R. et al. Transcriptional and cellular diversity of the human heart. Circulation 142, 466–482 (2020).
Wang, Z. et al. Cell-type-specific gene regulatory networks underlying murine neonatal heart regeneration at single-cell resolution. Cell Rep. 33, 108472 (2020).
Cui, M. et al. Dynamic transcriptional responses to injury of regenerative and non-regenerative cardiomyocytes revealed by single-nucleus RNA sequencing. Dev. Cell 53, 102–116.e8 (2020).
Kuppe, C. et al. Spatial multi-omic map of human myocardial infarction. Nature 608, 766–777 (2022).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Sinn, H. W., Balsamo, J., Lilien, J. & Lin, J. J. Localization of the novel Xin protein to the adherens junction complex in cardiac and skeletal muscle during development. Dev. Dyn. 225, 1–13 (2002).
Gustafson-Wagner, E. A. et al. Loss of mXinalpha, an intercalated disk protein, results in cardiac hypertrophy and cardiomyopathy with conduction defects. Am. J. Physiol. Heart Circ. Physiol. 293, H2680–H2692 (2007).
Long, P. A., Larsen, B. T., Evans, J. M. & Olson, T. M. Exome sequencing identifies pathogenic and modifier mutations in a child with sporadic dilated cardiomyopathy. J. Am. Heart Assoc. 4, e002443 (2015).
Huang, Y.-S., Xing, Y.-L. & Li, H.-W. Heterozygous desmin gene (DES) mutation contributes to familial dilated cardiomyopathy. J. Int. Med. Res. 49, 3000605211006598 (2021).
McCalmon, S. A. et al. Modulation of angiotensin II-mediated cardiac remodeling by the MEF2A target gene Xirp2. Circ. Res. 106, 952–960 (2010).
Verdonschot, J. A. J. et al. A mutation update for the FLNC gene in myopathies and cardiomyopathies. Hum. Mutat. 41, 1091–1111 (2020).
Ortiz-Genga, M. F. et al. Truncating FLNC mutations are associated with high-risk dilated and arrhythmogenic cardiomyopathies. J. Am. Coll. Cardiol. 68, 2440–2451 (2016).
Hall, C. L. et al. Filamin C variants are associated with a distinctive clinical and immunohistochemical arrhythmogenic cardiomyopathy phenotype. Int. J. Cardiol. 307, 101–108 (2020).
Oz, S. et al. Reduction in filamin C transcript is associated with arrhythmogenic cardiomyopathy in Ashkenazi Jews. Int. J. Cardiol. 317, 133–138 (2020).
Begay, R. L. et al. Filamin C truncation mutations are associated with arrhythmogenic dilated cardiomyopathy and changes in the cell–cell adhesion structures. JACC Clin. Electrophysiol. 4, 504–514 (2018).
Valdés-Mas, R. et al. Mutations in filamin C cause a new form of familial hypertrophic cardiomyopathy. Nat. Commun. 5, 5326 (2014).
Han, P. et al. A long noncoding RNA protects the heart from pathological hypertrophy. Nature 514, 102–106 (2014).
Günthel, M. Epigenetic state changes underlie metabolic switch in mouse post-infarction border zone cardiomyocytes. J. Cardiovasc. Dev. Dis. 8, 134 (2021).
Guo, H. et al. Intercalated disc protein Xinβ is required for Hippo-YAP signaling in the heart. Nat. Commun. 11, 4666 (2020).
Tao, G. et al. Pitx2 promotes heart repair by activating the antioxidant response after cardiac injury. Nature 534, 119–123 (2016).
Nadadur, R. D. et al. Pitx2 modulates a Tbx5-dependent gene regulatory network to maintain atrial rhythm. Sci. Transl. Med. 8, 354ra115 (2016).
Man, J. C. K. et al. Genetic dissection of a super enhancer controlling the Nppa-Nppb cluster in the heart. Circ. Res. 128, 115–129 (2021).
Cao, D. J. et al. Cytosolic DNA sensing promotes macrophage transformation and governs myocardial ischemic injury. Circulation 137, 2613–2634 (2018).
Farbehi, N. et al. Single-cell expression profiling reveals dynamic flux of cardiac stromal, vascular and immune cells in health and injury. eLife 8, e43882 (2019).
Muhl, L. et al. Single-cell analysis uncovers fibroblast heterogeneity and criteria for fibroblast and mural cell identification and discrimination. Nat. Commun. 11, 3953 (2020).
Dela Paz, N. G. & D’Amore, P. A. Arterial versus venous endothelial cells. Cell Tissue Res. 335, 5–16 (2009).
Vanlandewijck, M. et al. A molecular atlas of cell types and zonation in the brain vasculature. Nature 554, 475–480 (2018).
Kalucka, J. et al. Single-cell transcriptome atlas of murine endothelial cells. Cell 180, 764–779.e20 (2020).
Fujimoto, N. et al. Single-cell mapping reveals new markers and functions of lymphatic endothelial cells in lymph nodes. PLoS Biol. 18, e3000704 (2020).
Gerarduzzi, C., Hartmann, U., Leask, A. & Drobetsky, E. The matrix revolution: matricellular proteins and restructuring of the cancer microenvironment. Cancer Res. 80, 2705–2717 (2020).
Nishioka, T. et al. Tenascin C may aggravate left ventricular remodeling and function after myocardial infarction in mice. Am. J. Physiol. Heart Circ. Physiol. 298, H1072–H1078 (2010).
McCurdy, S. M. et al. SPARC mediates early extracellular matrix remodeling following myocardial infarction. Am. J. Physiol. Heart Circ. Physiol. 301, H497–H505 (2011).
Murry, C. E., Giachelli, C. M., Schwartz, S. M. & Vracko, R. Macrophages express osteopontin during repair of myocardial necrosis. Am. J. Pathol. 145, 1450–1462 (1994).
Schellings, M. W. M. et al. Absence of SPARC results in increased cardiac rupture and dysfunction after acute myocardial infarction. J. Exp. Med. 206, 113–123 (2009).
Oka, T. et al. Genetic manipulation of periostin expression reveals a role in cardiac hypertrophy and ventricular remodeling. Circ. Res. 101, 313–321 (2007).
Trueblood, N. A. et al. Exaggerated left ventricular dilation and reduced collagen deposition after myocardial infarction in mice lacking osteopontin. Circ. Res. 88, 1080–1087 (2001).
Richardson, W. J. & Holmes, J. W. Why is infarct expansion such an elusive therapeutic target? J. Cardiovasc. Transl. Res. 8, 421–430 (2015).
Luther, D. J., Thodeti, C. K. & Meszaros, J. G. Injury models to study cardiac remodeling in the mouse: myocardial infarction and ischemia–reperfusion. Methods Mol. Biol. 1037, 325–342 (2013).
Monroe, T. O. et al. YAP partially reprograms chromatin accessibility to directly induce adult cardiogenesis in vivo. Dev. Cell 48, 765–779.e7 (2019).
Xiao, Y., Leach, J., Wang, J. & Martin, J. F. Hippo/Yap signaling in cardiac development and regeneration. Curr. Treat. Options Cardiovasc. Med. 18, 38 (2016).
Acknowledgements
This publication includes data generated at the UC San Diego IGM Genomics Center utilizing an Illumina NovaSeq 6000 that was purchased with funding from a National Institutes of Health SIG grant (#S10 OD026929). We thank IGM and the Nikon Imaging Center at UCSD for technical assistance. The work was funded by National Institutes of Health (NIH) grants NIH UL1TR001442 (UCSD), AHA17IRG33410543 (K.R.K.), NIH R00HL129168 (K.R.K.), NIH DP2AR075321 (K.R.K.), NIH HL142251 (F.S.) and NIH T32HL105373 (D.M.C.), National Heart Lung and Blood Institute (NHLBI) grants NHLBI T32HL007444 (V.K.N.) and NHLBI HL162369 (F.S.) and Department of Defense grant no. W81XWH1810380 (F.S.).
Author information
Authors and Affiliations
Contributions
D.M.C., N.T., V.K.N. and K.R.K. designed and performed the experiments, analyzed the data and wrote the manuscript. J.M.M., A.T., R.S., J.L. and Y.L. performed the experiments. J.M.D. and E.A. provided the human tissue samples. K.L.C., K.Z. and F.S. provided guidance on experimental design. Z.F. performed all mouse surgeries and developed new methods including the NP models. K.R.K. conceived the project and provided funding. All authors reviewed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
F.S. is a cofounder and has an equity interest in Papillon Therapeutics; he is a consultant and has equity interest and a research grant from LEXEO Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Rafael Kramann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Integration of single cell and nuclei RNA-seq heart datasets during acute responses to MI.
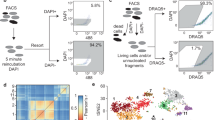
(a) Overall experimental design and integration of sc/snRNA-seq data with whole transcriptome spatial data. Hearts were harvested at several time points following experimental MI and collected for snRNA-seq. The resulting data matrices were integrated with available scRNA-seq data. (b) Experimental timepoints post-MI that were examined in our study with a summary table of total numbers of cells, nuclei and spatial pixels analyzed to support the robustness of our claims across biological replicates. (c) Gating strategy to isolate nuclei using DAPI and FACS. (d) Mitochondrial QC metrics of samples and replicates for both single nuclei, single cell and integrated sn/sc data. (e) UMAP plots annotated by major cell types (left) and subsets (right) after removing nuclei and cells that have more than 5% mitochondrial counts. (f) UMAP plots displaying composition of single nuclei (left) and single cell (right) derived samples. (g) Subcluster composition as derived from UMAPs shown in (f). (h) UMAP plots split by timepoint and across biological replicates. (i) Average subcluster composition displayed in (h). (l) QC metric of samples and replicates for both single nuclei and single cell represented in counts per sample (nCounts) and features per sample (nGenes).
Extended Data Fig. 2 Integration and quantification of spatial transcriptomic datasets.
(a) UMAP plots split by sample. (b) Quantification of cluster classification as a percentage of total pixels captured by sample (data presented as mean values ± SEM; n = 2, sham; n = 1, 1 hr and 4hrs; n = 3, 72 hrs and 168hrs post-MI). (c) Quality control metrics, split by sample. (d) Spatial plot showing results of clustering across different time points after MI and replicates.
Extended Data Fig. 3 High resolution clustering of CMs at 24 hrs and 72 hrs post-MI.
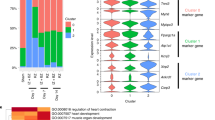
(a) UMAP plots of subset and reclustered CMs as shown in Fig. 1b with higher resolution (top) and annotated based on spatial mapping (bottom). (b) Heatmap of resulting DEGs based on resolution. (f) Heatmap of DEGs based on spatial mapping. (c) Select gene ontology terms enriched in BZ2 CMs related to Ras/Rho signaling. (d) Average scaled expression of guanine nucleotide exchange factors (GEFs) and GTPase activating proteins (GAPs) in CM subsets. (e) Spatial feature plots of representative Ras/Rho related genes showing distribution in BZ.
Extended Data Fig. 4 Mapping of snRNA-seq derived CM subsets to space.
(a) Strategy for mapping labels (IZ, RZ, BZ1 and BZ2) from CM nuclei to spatial clusters (see methods). (b) UMAP plot of integrated dataset composed of 34,116 pixels in 16 samples (summarized in Extended Data Fig. 1b, see Extended Data fig. 8). (c) DEGs based on high-resolution clustering (top) and post-classification regions (bottom). (d) Histogram of counts from representative samples (green, sham; red, 72 hrs post-MI). (e) Histogram of BZ1 scores. (f) Dot plot of BZ1 and BZ2 scores. (g,h) Feature plots (g) and violin plots (h) of CM, BZ1 and BZ2 scores. (i) Results of ROC analysis (AUC, area under the curve) and stepwise label-mapping (reference cluster indicated above). (j) Classification results shown in UMAP space. (k) AUC as a function of gene-set length.
Extended Data Fig. 5 Spatial transcriptomics of human STEMI tissue.
(a) H&E of cross section of a single human heart sample from a patient presenting with anterior wall STEMI. (b) Spatial transcriptomic clustering results based on DEG analysis and assessment of BZ marker genes shown in space in (c) and by violin plots in (d). (e) Heatmap of cluster defining DEGs. LQ, low quality.
Extended Data Fig. 6 Segmentation, classification and quantification of mFISH data.
(a) After mFISH imaging, slides were stained for WGA to label the CM membrane. (b) Areas with well-defined perimeters were converted to regions-of-interest (shown in white). (c) Expression of BZ1 markers (Nppa, Clu) and BZ2 markers (Xirp2, Flnc) across ROIs. Scatter plot was visually inspected to establish thresholds and classify ROIs as RZ, BZ1, or BZ2 (blue, purple, red). (d) Quantification of neighbor composition in 25-pixel radius (n = 625 cells, and 1008 cells examined over 1 sample for BZ1 and BZ2, respectively). (e) Quantification of minimum distance to the IZ border defined by Tnnt2 staining. (f) To quantify direct contact of CMs, the percentage of Tnnt2 + pixels were calculated in ROI perimeters with various degrees of thickness (n = 625 cells, and 1008 cells examined over 1 sample for BZ1 and BZ2, respectively). (g) Contiguous regions of respective CM subsets were measured. Images shown are representative of 2 separate experiments. Scale bars indicate 500 μm. Boxplots presented with mean; box: 25th-75th%; whiskers: 2.5th-97.5th %. **** P-value < .0001; Mann-Whitney Test, two-sided.
Extended Data Fig. 7 Colocalization of immune and fibroblast subsets across time.
(a) Spatial plots showing subclustering results (shown in 3c) across 72 hrs post-MI and 168 hrs post-MI replicates. (b) Colocalized immune subsets in 72 hrs post-MI replicate. (c) Spatially correlated and clustered gene-set scores. (d) Spatially correlated and clustered genes based on spearman rank correlation analysis for all regions (left) and infarct zone (right) of 72 hrs post-MI sample. (e) All gene-set scores projected to clusters. (f) Volcano plot comparing gene-set scores in BZ1 and BZ2 (Wilcoxon Ranked Sum Test, two-sided).
Extended Data Fig. 8 Spatial transcriptomics on mechanical injury models.
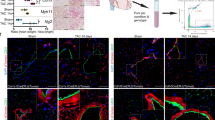
(a) H&E sections at low magnification (lower panel) and high magnification (top panel). Representative of at least 2 separate experiments. (b) Clustering results of spatial transcriptomic data based on integrated dataset as shown in Extended Data Fig. 3. (c) Hif1 scores applied to representative 72 hrs post-MI sample (left) and NP (right) with quantifications plotted per pixel in (d). In comparison to post-MI hearts, needle pass injuries have a significantly lower Hif1 score compared to the positive control (72 hr MI) samples and is unchanged from negative controls in sham samples. (e) Experimental design of fluorescent dye assay to assess regional perfusion. Wheat germ agglutinin (WGA) conjugated to AlexaFluor 488 was injected directly into LV 10 min or 72 hrs post NP injury and hearts were harvested 20 minutes after injection for imaging and quantification (data presented as mean values ± SEM n = 3 biologically independent samples). The representative image in the left panel designates the visualization of the needle pass area designated as the negative control and the reference site designated as neighbors, and the right panel represents a positive control area remote from the site of injury. Quantification of samples harvested 30 minutes and 72 hrs post-NP injury reveal that in both time courses, Neighbors and Positive controls are significantly increased as compared to the Negative Control (Needle pass) sites and are not different from each other (f) Representative H&E-stained sections of NP injuries highlighting area of injury in dash lines and directions of adjacent myocyte bundles in arrows used to quantify gene expression. Representative of 2 separate experiments, each of which had tissue sections aligned with the long axis of the needle insertion. (g) Anisotropy ratios in each sample with non-injured area (right) showing no significant anisotropy in the borderzones of needle pass injuries (n = 2 biologically independent samples). Data presented as mean values ± SEM. *P-value < 0.05, **P-value < 0.01, ***P-value < 0.001, ****P-value < 0.0001; One-way ANOVA with Tukey’s post-hoc analysis.
Supplementary information
Supplementary Table 1
Differentially expressed genes of major cell types: (1.1) cardiomyocytes; (1.2) immune cells; (1.3) and stromal cells. P values were derived from Seurat and FDR-adjusted for multiple hypothesis testing
Supplementary Table 2
Differentially expressed genes of cell type subsets: (2.1–2.7) monocyte and macrophage subsets; (2.8) dendritic cell subsets; (2.9 and 2.10) neutrophil subsets; (2.11–2.18) endothelial cell subsets; (2.19–2.25) fibroblast subsets; (2.26–2.30) cardiomyocyte subsets. P values were derived from Seurat and FDR-adjusted for multiple hypothesis testing
Supplementary Table 3
Differentially expressed genes derived from spatial transcriptomics after label mapping. P values were derived from Seurat and FDR-adjusted for multiple hypothesis testing
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Calcagno, D.M., Taghdiri, N., Ninh, V.K. et al. Single-cell and spatial transcriptomics of the infarcted heart define the dynamic onset of the border zone in response to mechanical destabilization. Nat Cardiovasc Res 1, 1039–1055 (2022). https://doi.org/10.1038/s44161-022-00160-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-022-00160-3
This article is cited by
-
The role of cardiac pericytes in health and disease: therapeutic targets for myocardial infarction
Nature Reviews Cardiology (2024)
-
YAP induces a neonatal-like pro-renewal niche in the adult heart
Nature Cardiovascular Research (2024)
-
Don’t go breakin’ my heart: cardioprotective alterations to the mechanical and structural properties of reperfused myocardium during post-infarction inflammation
Biophysical Reviews (2023)
-
Mapping the border zone in myocardial infarction
Nature Cardiovascular Research (2022)