Abstract
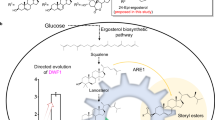
Jasmonates are a class of plant hormones with many agricultural applications and potential medicinal properties. However, the low content of jasmonates in plants and environmental issues with their production make their supply challenging. In the present study, we report the de novo microbial biosynthesis of jasmonic acid and its derivatives, methyl jasmonate and jasmonoyl isoleucine, from glucose using an engineered baker’s yeast. The study uses enzymes located in the endoplasmic reticulum and cytosol to generate the intermediates α-linolenic acid and cis-12-oxophytodienoic acid. Our final engineered stain, which integrates 15 heterologous genes from diverse plants and fungi and had 3 of its native genes deleted, produces jasmonic acid at titres of 19.0 mg l−1 in flask cultures through in vitro supplementation of α-linolenic acid. In addition to the well-known natural structures (−)-jasmonic acid and (+)-epi-jasmonic acid, the engineered yeast also synthesized the previously unobserved unnatural structures (+)-jasmonic acid and (−)-epi-jasmonic acid. These results demonstrate that yeast is a scalable and sustainable platform to produce both naturally occurring jasmonates and those structures not found naturally in plants.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated in the present study are provided within the paper and its Supplementary Information files. Source data are provided with this paper.
References
Wang, J., Song, L., Gong, X., Xu, J. & Li, M. Functions of jasmonic acid in plant regulation and response to abiotic stress. Int. J. Mol. Sci. 21, 1446 (2020).
Yukimune, Y., Tabata, H., Higashi, Y. & Hara, Y. Methyl jasmonate-induced overproduction of paclitaxel and baccatin III in Taxus cell suspension cultures. Nat. Biotechnol. 14, 1129–1132 (1996).
Deepthi, S. & Satheeshkumar, K. Cell line selection combined with jasmonic acid elicitation enhance camptothecin production in cell suspension cultures of Ophiorrhiza mungos L. Appl. Microbiol. Biotechnol. 101, 545–558 (2017).
Shen, Z. et al. High production of jasmonic acid by Lasiodiplodia iranensis using solid-state fermentation: optimization and understanding. Biotechnol. J. 17, 2100550 (2022).
Scognamiglio, J., Jones, L., Letizia, C. S. & Api, A. M. Fragrance material review on methyl jasmonate. Food Chem. Toxicol. 50, S572–S576 (2012).
Sá-Nakanishi, A. B. et al. Anti-inflammatory and antioxidant actions of methyl jasmonate are associated with metabolic modifications in the liver of arthritic rats. Oxid. Med. Cell. Longev. 2018, 2056250 (2018).
Fingrut, O. & Flescher, E. Plant stress hormones suppress the proliferation and induce apoptosis in human cancer cells. Leukemia 16, 608–616 (2002).
Bömer, M. et al. Jasmonates induce Arabidopsis bioactivities selectively inhibiting the growth of breast cancer cells through CDC6 and mTOR. New Phytol. 229, 2120–2134 (2021).
Creelman, R. A. & Mullet, J. E. Jasmonic acid distribution and action in plants: regulation during development and response to biotic and abiotic stress. Proc. Natl Acad. Sci. USA 92, 4114–4119 (1995).
Ghasemi Pirbalouti, A., Sajjadi, S. E. & Parang, K. A review (research and patents) on jasmonic acid and its derivatives. Arch. Pharm. 347, 229–239 (2014).
Chapuis, C. The jubilee of methyl jasmonate and Hedione®. Helv. Chim. Acta 95, 1479–1511 (2012).
Carroll, A. L., Desai, S. H. & Atsumi, S. Microbial production of scent and flavor compounds. Curr. Opin. Biotechnol. 37, 8–15 (2016).
Eng, F. et al. Jasmonic acid biosynthesis by fungi: derivatives, first evidence on biochemical pathways and culture conditions for production. PeerJ 9, e10873 (2021).
Borodina, I. & Nielsen, J. Advances in metabolic engineering of yeast Saccharomyces cerevisiae for production of chemicals. Biotechnol. J. 9, 609–620 (2014).
Sheng, J. & Feng, X. Metabolic engineering of yeast to produce fatty acid-derived biofuels: bottlenecks and solutions. Front. Microbiol. 6, 554 (2015).
Reider Apel, A. et al. A Cas9-based toolkit to program gene expression in Saccharomyces cerevisiae. Nucleic Acids Res. 45, 496–508 (2017).
Mans, R. et al. CRISPR/Cas9: a molecular Swiss army knife for simultaneous introduction of multiple genetic modifications in Saccharomyces cerevisiae. FEMS Yeast Res. 15, fov004 (2015).
Jakočiūnas, T. et al. Multiplex metabolic pathway engineering using CRISPR/Cas9 in Saccharomyces cerevisiae. Metab. Eng. 28, 213–222 (2015).
Yu, T. et al. Reprogramming yeast metabolism from alcoholic fermentation to lipogenesis. Cell 174, 1549–1558.e1514 (2018).
Pollmann, S. et al. Substrate channeling in oxylipin biosynthesis through a protein complex in the plastid envelope of Arabidopsis thaliana. J. Exp. Bot. 70, 1483–1495 (2019).
Wasternack, C. & Hause, B. The missing link in jasmonic acid biosynthesis. Nat. Plants 5, 776–777 (2019).
Theodoulou, F. L. et al. Jasmonic acid levels are reduced in COMATOSE ATP-binding cassette transporter mutants. Implications for transport of jasmonate precursors into peroxisomes. Plant Physiol. 137, 835–840 (2005).
Li, C. et al. Role of beta-oxidation in jasmonate biosynthesis and systemic wound signaling in tomato. Plant Cell 17, 971–986 (2005).
Kainou, K., Kamisaka, Y., Kimura, K. & Uemura, H. Isolation of Δ12 and ω3-fatty acid desaturase genes from the yeast Kluyveromyces lactis and their heterologous expression to produce linoleic and α-linolenic acids in Saccharomyces cerevisiae. Yeast 23, 605–612 (2006).
Zhang, H. T., Yang, J. S., Shan, L. & Bi, Y. P. Functional expression of a ω-3 fatty acid desaturase gene from Glycine max in Saccharomyces cerevisiae. Chinese J. Biotechnol. 22, 33–38 (2006).
Xue, Y. et al. Omega-3 fatty acid desaturase gene family from two ω-3 sources, Salvia hispanica and Perilla frutescens: cloning, characterization and expression. PLoS ONE 13, e0191432 (2018).
Rodríguez-Rodríguez, M. F., Salas, J. J., Venegas-Calerón, M., Garcés, R. & Martínez-Force, E. Molecular cloning and characterization of the genes encoding a microsomal oleate Δ12 desaturase (CsFAD2) and linoleate Δ15 desaturase (CsFAD3) from Camelina sativa. Ind. Crops Prod. 89, 405–415 (2016).
Oura, T. & Kajiwara, S. Saccharomyces kluyveri FAD3 encodes an omega3 fatty acid desaturase. Microbiology 150, 1983–1990 (2004).
Sibirny, A. A. Yeast peroxisomes: structure, functions and biotechnological opportunities. FEMS Yeast Res. 16, fow038 (2016).
Yazawa, H., Iwahashi, H., Kamisaka, Y., Kimura, K. & Uemura, H. Production of polyunsaturated fatty acids in yeast Saccharomyces cerevisiae and its relation to alkaline pH tolerance. Yeast 26, 167–184 (2009).
Yazawa, H., Iwahashi, H., Kamisaka, Y., Kimura, K. & Uemura, H. Improvement of polyunsaturated fatty acids synthesis by the coexpression of CYB5 with desaturase genes in Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 87, 2185–2193 (2010).
Yazawa, H. et al. Heterologous production of dihomo-gamma-linolenic acid in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 73, 6965–6971 (2007).
Oura, T. & Kajiwara, S. Substrate specificity and regioselectivity of delta12 and omega3 fatty acid desaturases from Saccharomyces kluyveri. Biosci. Biotechnol. Biochem. 72, 3174–3179 (2008).
Ferreira, R., Teixeira, P. G., Siewers, V. & Nielsen, J. Redirection of lipid flux toward phospholipids in yeast increases fatty acid turnover and secretion. Proc. Natl Acad. Sci. USA 115, 1262–1267 (2018).
Bonner, W. M. & Bloch, K. Purification and properties of fatty acyl thioesterase I from Escherichia coli. J. Biol. Chem. 247, 3123–3133 (1972).
Paton, C. M. & Ntambi, J. M. Biochemical and physiological function of stearoyl-CoA desaturase. Am. J. Physiol. Endocrinol. Metab. 297, E28–E37 (2009).
Yazawa, H., Kamisaka, Y., Kimura, K., Yamaoka, M. & Uemura, H. Efficient accumulation of oleic acid in Saccharomyces cerevisiae caused by expression of rat elongase 2 gene (rELO2) and its contribution to tolerance to alcohols. Appl. Microbiol. Biotechnol. 91, 1593–1600 (2011).
Zhai, Q., Yan, C., Li, L., Xie, D. & Li, C. in Hormone Metabolism and Signaling in Plants (eds Li, J., Li, C. & Smith, S. M.) 243–272 (Academic Press, 2017).
Chauvin, A., Caldelari, D., Wolfender, J. L. & Farmer, E. E. Four 13-lipoxygenases contribute to rapid jasmonate synthesis in wounded Arabidopsis thaliana leaves: a role for lipoxygenase 6 in responses to long-distance wound signals. New Phytol. 197, 566–575 (2013).
Bannenberg, G., Martínez, M., Hamberg, M. & Castresana, C. Diversity of the enzymatic activity in the lipoxygenase gene family of Arabidopsis thaliana. Lipids 44, 85–95 (2009).
Mandal, S., Dahuja, A. & Santha, I. M. Lipoxygenase activity in soybean is modulated by enzyme-substrate ratio. J. Plant Biochem. Biotechnol. 23, 217–220 (2014).
Browse, J., Warwick, N., Somerville, C. R. & Slack, C. R. Fluxes through the prokaryotic and eukaryotic pathways of lipid synthesis in the ‘16:3’ plant Arabidopsis thaliana. Biochem. J. 235, 25–31 (1986).
Ishiguro, S., Kawai-Oda, A., Ueda, J., Nishida, I. & Okada, K. The DEFECTIVE IN ANTHER DEHISCIENCE gene encodes a novel phospholipase A1 catalyzing the initial step of jasmonic acid biosynthesis, which synchronizes pollen maturation, anther dehiscence, and flower opening in Arabidopsis. Plant Cell 13, 2191–2209 (2001).
Löwe, J., Dietz, K. J. & Gröger, H. From a biosynthetic pathway toward a biocatalytic process and chemocatalytic modifications: three-step enzymatic cascade to the plant metabolite cis-(+)-12-OPDA and metathesis-derived products. Adv. Sci. 7, 1902973 (2020).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Koo, A. J. K., Chung, H. S., Kobayashi, Y. & Howe, G. A. Identification of a peroxisomal acyl-activating enzyme involved in the biosynthesis of jasmonic acid in Arabidopsis. J. Biol. Chem. 281, 33511–33520 (2006).
Wasternack, C. & Kombrink, E. Jasmonates: structural requirements for lipid-derived signals active in plant stress responses and development. ACS Chem. Biol. 5, 63–77 (2010).
Kragler, F., Lametschwandtner, G., Christmann, J., Hartig, A. & Harada, J. J. Identification and analysis of the plant peroxisomal targeting signal 1 receptor NtPEX5. Proc. Natl Acad. Sci. USA 95, 13336–13341 (1998).
Koch, T., Bandemer, K. & Boland, W. Biosynthesis of cis-jasmone: a pathway for the inactivation and the disposal of the plant stress hormone jasmonic acid to the gas phase? Helv. Chim. Acta 80, 838–850 (1997).
Seo, H. S. et al. Jasmonic acid carboxyl methyltransferase: a key enzyme for jasmonate-regulated plant responses. Proc. Natl Acad. Sci. USA. 98, 4788–4793 (2001).
Fu, W. et al. The jasmonic acid-amino acid conjugates JA-Val and JA-Leu are involved in rice resistance to herbivores. Plant Cell Environ. 45, 262–272 (2022).
Suza, W. P. & Staswick, P. E. The role of JAR1 in jasmonoyl-l-isoleucine production during Arabidopsis wound response. Planta 227, 1221–1232 (2008).
Yu, T. et al. Metabolic reconfiguration enables synthetic reductive metabolism in yeast. Nat. Metab. 4, 1551–1559 (2022).
Dar, A. A., Choudhury, A. R., Kancharla, P. K. & Arumugam, N. The FAD2 gene in plants: occurrence, regulation and role. Front. Plant Sci. 8, 1789 (2017).
Wang, Y. et al. Directed evolution: methodologies and applications. Chem. Rev. 121, 12384–12444 (2021).
Romero, P. A. & Arnold, F. H. Exploring protein fitness landscapes by directed evolution. Nat. Rev. Mol. Cell Biol. 10, 866–876 (2009).
Hammer, S. K. & Avalos, J. L. Harnessing yeast organelles for metabolic engineering. Nat. Chem. Biol. 13, 823–832 (2017).
Kim, J. E. et al. Tailoring the Saccharomyces cerevisiae endoplasmic reticulum for functional assembly of terpene synthesis pathway. Metab. Eng. 56, 50–59 (2019).
Arendt, P. et al. An endoplasmic reticulum-engineered yeast platform for overproduction of triterpenoids. Metab. Eng. 40, 165–175 (2017).
Zhou, Y. J. et al. Harnessing yeast peroxisomes for biosynthesis of fatty-acid-derived biofuels and chemicals with relieved side-pathway competition. J. Am. Chem. Soc. 138, 15368–15377 (2016).
DeLoache, W. C., Russ, Z. N. & Dueber, J. E. Towards repurposing the yeast peroxisome for compartmentalizing heterologous metabolic pathways. Nat. Commun. 7, 11152 (2016).
Liu, G. S. et al. The yeast peroxisome: a dynamic storage depot and subcellular factory for squalene overproduction. Metab. Eng. 57, 151–161 (2020).
Jarocka-Karpowicz, I. & Markowska, A. Therapeutic potential of jasmonic acid and its derivatives. Int. J. Mol. Sci. 22, 8437 (2021).
Zheng, T. et al. Upcycling CO2 into energy-rich long-chain compounds via electrochemical and metabolic engineering. Nat. Catal. 5, 388–396 (2022).
Kim, S. & Gonzalez, R. Selective production of decanoic acid from iterative reversal of β-oxidation pathway. Biotechnol. Bioeng. 115, 1311–1320 (2018).
Smith, S. M. Strategies for the purification of membrane proteins. Methods Mol. Biol. 1485, 389–400 (2017).
Pan, X., Welti, R. & Wang, X. Quantitative analysis of major plant hormones in crude plant extracts by high-performance liquid chromatography-mass spectrometry. Nat. Protoc. 5, 986–992 (2010).
Luzarowski, M. et al. Global mapping of protein–metabolite interactions in Saccharomyces cerevisiae reveals that Ser-Leu dipeptide regulates phosphoglycerate kinase activity. Commun. Biol. 4, 181 (2021).
Chen, N. et al. Systematic genetic modifications of cell wall biosynthesis enhanced the secretion and surface-display of polysaccharide degrading enzymes in Saccharomyces cerevisiae. Metab. Eng. 77, 273–282 (2023).
Acknowledgements
This work was financially supported by National Key Research and Development Program of China (grant no. 2018YFA0903200 to X.L. and H.T.), the National Natural Science Foundation of China (grant no. 32071421 to X.L.), Guangdong Basic and Applied Basic Research Foundation (grant no. 2021A1515010842 to H.T.) and the Shenzhen Science and Technology Program (grant nos. ZDSYS20210623091810032 and RCYX20200714114736026 to X.L.). We thank T. Yu for critical discussion and Z. Wei for helping to organize the meeting about this project.
Author information
Authors and Affiliations
Contributions
H.T., J.D.K. and X.L. conceived the project. H.T., J.D., S.L. and X.L. designed the experiments. H.T. and S.L. performed experiments. H.T., J.D., S.L. and X.L. analysed the results and wrote the manuscript. H.T., J.D.K. and X.L. revised the manuscript. All authors revised and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.D.K. has a financial interest in Amyris, Lygos, Demetrix, Napigen, Maple Bio, Apertor Labs, Zero Acre Farms, Berkeley Yeast and Ansa Biotechnology. X.L. has a financial interest in Demetrix and Synceres. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Synthesis thanks Mattheos Koffas, Jens Nielsen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Thomas West, in collaboration with the Nature Synthesis team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Production of linoleic acid and α-LeA in SJ01.
The control strain (WT) and the α-LeA-producing strain SJ01 were cultured in nitrogen-limited minimal medium containing 20 g l−1 glucose. WT, wild-type strain; Kl, Kluyveromyces lactis. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 2 Production of oleic acid.
The control strain (WT) and the α-LeA-producing strain SJ01 were cultured in nitrogen-limited minimal medium containing 20 g l−1 glucose. α-LeA, α-linolenic acid; WT, wild-type; Kl, Kluyveromyces lactis. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 3 Cultivation condition optimization for α-LeA production.
The effect of various temperatures, medium and fermentation times on production of α-LeA (A), linoleic acid (B) and oleic acid (C). α-LeA, α-linolenic acid; NSD, nitrogen-limited minimal; YPD, yeast extract peptone dextrose; d, days. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 4 Production of linoleic acid (A) and oleic acid (B) by engineering the fatty acid biosynthesis pathway and introduction of additional copies of FAD2/FAD3.
Expression of Ec’tesA enhanced the conversion of fatty acyl-ACP into free fatty acids, deletion of native POX1 and FAA1/4 reduced the consumption of free fatty acids, and the copy number of genes encoding the rate-limiting enzymes were increased. Cs, Camelina sativa. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 5 The effect of RnELO2 expression on the production of fatty acids.
(a) Pathway engineering by overexpressing RnELO2. Rn, R. norvegicus. (b) Production of fatty acids in strains SJ05 and SJ06. SJ05 is the control strain without expression of RnELO2. SJ06 is the RnELO2-expressing strain. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 6 Presence of linoleic acid and α-LeA in the membranes of α-LeA-producing strain SJ05.
WT, wild-type strain. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 7 Cell phenotype characterization of the wild-type strain Lab001 and α-LeA-producing strain SJ05.
a. Cell growth curves of S. cerevisiae strains. b. Description of maximum specific growth rate. μmax, maximum specific growth rate. Spotting growth assay of strains under different cultivation temperatures (c) and ethanol concentrations (d). WT, wild-type strain. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 8 Proteomic analysis of enzymes in the OPDA synthetic pathway.
a. OPDA synthetic pathway. b. The expression of AtLOX2, AtAOS and AtAOC2 in SJ07. At, A. thaliana. All data are presented as mean ± s.d. of biological triplicates.
Extended Data Fig. 9 Production of JA from exogenously added α-linolenic acid.
JA, jasmonic acid. All data are presented as mean ± s.d. of biological triplicates.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 1–7.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tang, H., Lin, S., Deng, J. et al. Engineering yeast for the de novo synthesis of jasmonates. Nat. Synth 3, 224–235 (2024). https://doi.org/10.1038/s44160-023-00429-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44160-023-00429-w
This article is cited by
-
Engineered yeast brews plant hormones
Nature Synthesis (2024)
-
Exploring the plant lipidome: techniques, challenges, and prospects
Advanced Biotechnology (2024)