Abstract
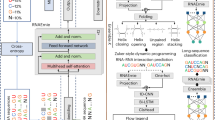
Here we represent human lives in a way that shares structural similarity to language, and we exploit this similarity to adapt natural language processing techniques to examine the evolution and predictability of human lives based on detailed event sequences. We do this by drawing on a comprehensive registry dataset, which is available for Denmark across several years, and that includes information about life-events related to health, education, occupation, income, address and working hours, recorded with day-to-day resolution. We create embeddings of life-events in a single vector space, showing that this embedding space is robust and highly structured. Our models allow us to predict diverse outcomes ranging from early mortality to personality nuances, outperforming state-of-the-art models by a wide margin. Using methods for interpreting deep learning models, we probe the algorithm to understand the factors that enable our predictions. Our framework allows researchers to discover potential mechanisms that impact life outcomes as well as the associated possibilities for personalized interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data used in this study are not publicly available due to Danish Data Protection regulations. Access to the data can be obtained via Statistics Denmark for Researchers in accordance with the rules of Statistics Denmark’s Research Scheme: https://www.dst.dk/en/TilSalg/Forskningsservice/Dataadgang. Source data are provided with this paper.
Code availability
The source code for the data processing, life2vec training, statistical analysis and visualization is available on GitHub at https://github.com/SocialComplexityLab/life2vec (ref. 82). The model weights, experiment logs and associated model outputs can be obtained in accordance with the rules of Statistics Denmark’s Research Scheme: https://www.dst.dk/en/TilSalg/Forskningsservice/Dataadgang.
References
Mansfield, L. A. et al. Predicting global patterns of long-term climate change from short-term simulations using machine learning. NPJ Clim. Atmos. Sci. 3, 44 (2020).
Alali, Y., Harrou, F. & Sun, Y. A proficient approach to forecast COVID-19 spread via optimized dynamic machine learning models. Sci. Rep. 12, 2467 (2022).
Zuboff, S. The Age of Surveillance Capitalism: The Fight for a Human Future at the New Frontier of Power (PublicAffairs, 2019).
Weber, M. The Theory of Social and Economic Organization (Simon & Schuster, 2009).
Salganik, M. J. et al. Measuring the predictability of life outcomes with a scientific mass collaboration. Proc. Natl Acad. Sci. USA 117, 8398–8403 (2020).
Lynge, E., Sandegaard, J. L. & Rebolj, M. The Danish National Patient Register. Scand. J. Public Health 39, 30–33 (2011).
Pedersen, C. B. The Danish civil registration system. Scand. J. Public Health 39, 22–25 (2011).
Salganik, M. J. Bit by Bit: Social Research in the Digital Age (Princeton Univ. Press, 2019).
Grimmer, J., Roberts, M. E. & Stewart, B. M. Text as Data: A New Framework for Machine Learning and the Social Sciences (Princeton Univ. Press, 2022).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), (ed. O’Conner L.) 770–778 (IEEE, 2016).
Silver, D. et al. Mastering the game of go with deep neural networks and tree search. Nature 529, 484–489 (2016).
Rumelhart, D. E., Hinton, G. E. & Williams, R. J. Learning representations by back-propagating errors. Nature 323, 533–536 (1986).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30, 5999–6009 (2017).
Brown, T. et al. Language models are few-shot learners. Proc. NeurIPS 33, 1877–1901 (2020).
Grechishnikova, D. Transformer neural network for protein-specific de novo drug generation as a machine translation problem. Sci. Rep. 11, 321 (2021).
Li, Y. et al. BEHRT: transformer for electronic health records. Sci. Rep. 10, 7155 (2020).
Bojesomo, A., Al-Marzouqi, H. & Liatsis, P. Spatiotemporal vision transformer for short time weather forecasting. In Proc. 2021 IEEE International Conference on Big Data (Big Data) (eds. Chen Y. et al.) 5741–5746 (IEEE, 2021).
Huang, C.-Z. A. et al. Music transformer: generating music with long-term structure. Preprint at https://openreview.net/forum?id=rJe4ShAcF7 (2023).
Vafa, K. et al. CAREER: Economic prediction of labor sequence data under distribution shift. In NeurIPS 2022 Workshop DistShift Spotlight (2022).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: pre-training of deep bidirectional transformers for language understanding. Proc. NAANCL Hum. Lang. Tech. 1, 4171–4186 (2019).
Choromanski, K. M. et al. Rethinking attention with performers. Preprint at https://openreview.net/forum?id=Ua6zuk0WRH (2023).
Kozlowski, A. C., Taddy, M. & Evans, J. A. The geometry of culture: analyzing the meanings of class through word embeddings. Am. Sociol. Rev. 84, 905–949 (2019).
Pilehvar, M. T. & Camacho-Collados, J. Embeddings in natural language processing: theory and advances in vector representations of meaning. Synth. Lect. Hum. Lang. Technol. 13, 1–175 (2020).
Arbejdsmarkedsregnskab (Danmarks Statistik, 2022); https://www.dst.dk/da/Statistik/emner/arbejde-og-indkomst/befolkningens-arbejdsmarkedsstatus/arbejdsmarkedsregnskab
International Standard Classification of Occupations: ISCO-08 (International Labour Office, 2012).
Dansk Branchekode 2007: DB07 (Danish Industrial Classification of All Economic Activities 2007) v3 edn (Danmarks Statistik, 2015).
International Classification of Diseases, 10th Revision (ICD-10) (World Health Organization, 1994).
Yadav, P., Steinbach, M., Kumar, V. & Simon, G. Mining electronic health records (EHRS) a survey. ACM Comput. Surv. 50, 1–40 (2018).
Han, Z., Zhao, J., Leung, H., Ma, K. F. & Wang, W. A review of deep learning models for time series prediction. IEEE Sens. J. 21, 7833–7848 (2019).
Moncada-Torres, A., van Maaren, M. C., Hendriks, M. P., Siesling, S. & Geleijnse, G. Explainable machine learning can outperform Cox regression predictions and provide insights in breast cancer survival. Sci. Rep. 11, 6968 (2021).
Rogers, A., Kovaleva, O. & Rumshisky, A. A primer in BERTology: what we know about how BERT works. Trans. Assoc. Comput. Ling. 8, 842–866 (2021).
Kazemi, S. M. et al. Time2Vec: learning a vector representation of time. Preprint at https://openreview.net/forum?id=rklklCVYvB (2023).
Bachlechner, T., Majumder, B. P., Mao, H., Cottrell, G. & McAuley, J. ReZero is all you need: fast convergence at large depth. Proc. Conf. Uncertainty Artif. Intell. 161, 1352–1361 (2021).
Ramachandran, P., Zoph, B. & Le, Q. V. Searching for activation functions. Preprint at https://openreview.net/forum?id=SkBYYyZRZ (2023).
Nguyen, T. Q. & Salazar, J. Transformers without tears: improving the normalization of self-attention. Proc. 16th International Conference on Spoken Language Translation (eds Niehues, J. et al.) 2019.iwslt-1.17 (ACL, 2019).
Pappas, N., Miculicich, L. & Henderson, J. Beyond weight tying: learning joint input-output embeddings for neural machine translation. Proc. Third Conference on Machine Translation (eds Borar, O. et al.) W18-6308 (ACL, 2018).
Kanai, S., Fujiwara, Y., Yamanaka, Y. & Adachi, S. Sigsoftmax: reanalysis of the softmax bottleneck. Proc. NeurIPS (eds Bengio S. et al.) 31, 286–296 (2018).
Wang, Y., Huang, H., Rudin, C. & Shaposhnik, Y. Understanding how dimension reduction tools work: an empirical approach to deciphering t-SNE, UMAP, TriMAP and PaCMAP for data visualization. JMLR 22, 9129–9201 (2021).
Naemi, A. et al. Machine learning techniques for mortality prediction in emergency departments: a systematic review. BMJ Open 11, e052663 (2021).
Jiang, L., Li, D., Wang, Q., Wang, S. & Wang, S. Improving positive unlabeled learning: practical AUL estimation and new training method for extremely imbalanced data sets. Preprint at https://arxiv.org/abs/2004.09820 (2020).
Wang, C., Pu, J., Xu, Z. & Zhang, J. Asymmetric loss for positive-unlabeled learning. In Proc. 2021 IEEE International Conference on Multimedia and Expo (ICME) 1–6 (IEEE, 2021).
Hansen, A. V., Mortensen, L. H., Ekstrøm, C. T., Trompet, S. & Westendorp, R. Predicting mortality and visualizing health care spending by predicted mortality in Danes over age 65. Sci. Rep. 13, 1203 (2023).
Ramola, R., Jain, S. & Radivojac, P. Estimating classification accuracy in positive-unlabeled learning: characterization and correction strategies. Pac. Symp. Biocomput. 24, 124–135 (2019).
Geifman, Y. & El-Yaniv, R. Selective classification for deep neural networks. In Proc Advances in Neural Information Processing Systems (eds Guyon, I et al.) 30 (Curran Associates, 2017).
Kim, B. et al. Interpretability beyond feature attribution: quantitative testing with concept activation vectors (TCAV). Proc. ICML 30, 2668–2677 (2018).
Narayan, A., Berger, B. & Cho, H. Assessing single-cell transcriptomic variability through density-preserving data visualization. Nat. Biotechnol. 39, 765–774 (2021).
Atanasova, P., Simonsen, J. G., Lioma, C. & Augenstein, I. A diagnostic study of explainability techniques for text classification. In Proc. 2020 Conference on Empirical Methods in Natural Language Processing (EMNLP) (eds Webber, B. et al.) 3256–3274 (ACL, 2020).
Bastings, J. & Filippova, K. The elephant in the interpretability room: why use attention as explanation when we have saliency methods? In Proc. Third Blackbox NLP Workshop on Analyzing and Interpreting Neural Networks for NLP (eds. Alishashi A. et al.) 149–155 (ACL, 2020).
Bommasani, R. et al. On the opportunities and risks of foundation models. Preprint at https://arxiv.org/abs/2108.07258 (2021).
Roberts, B. W., Kuncel, N. R., Shiner, R., Caspi, A. & Goldberg, L. R. The power of personality: the comparative validity of personality traits, socioeconomic status, and cognitive ability for predicting important life outcomes. Perspect. Psychol. Sci. 2, 313–345 (2007).
Stewart, R. D., Mõttus, R., Seeboth, A., Soto, C. J. & Johnson, W. The finer details? The predictability of life outcomes from Big Five domains, facets and nuances. J. Pers. 90, 167–182 (2022).
McCrae, R. R. & Costa, P. T. Jr. in Handbook of Personality: Theory and Research (eds John, O. P. & Robins, R. W.) 159–181 (Guilford Press, 2008).
Zettler, I., Thielmann, I., Hilbig, B. E. & Moshagen, M. The nomological net of the HEXACO model of personality: a large-scale meta-analytic investigation. Perspect. Psychol. Sci. 15, 723–760 (2020).
Det Danske Personligheds Og Sociale Adfærdspanel https://copsy.dk/posap/ (accessed 21 March 2021).
Gangl, M. Changing labour markets and early career outcomes: labour market entry in Europe over the past decade. Work Employ. Soc. 16, 67–90 (2002).
Halleröd, B., Ekbrand, H. & Bengtsson, M. In-work poverty and labour market trajectories: poverty risks among the working population in 22 European countries. J. Eur. Public Policy 25, 473–488 (2015).
Mackenbach, J. P. et al. Socioeconomic inequalities in health in 22 European countries. N. Engl. J. Med. 358, 2468–2481 (2008).
Adler, N. E. & Ostrove, J. M. Socioeconomic status and health: what we know and what we don’t. Ann. N. Y. Acad. Sci. 896, 3–15 (1999).
Liao, T. F. et al. Sequence analysis: its past, present and future. Soc. Sci. Res. 107, 102772 (2022).
Regulation (EU) 2016/679 of the European Parliament and of the Council of 27 April 2016 on the protection of natural persons with regard to the processing of personal data and on the free movement of such data, and repealing Directive 95/46/EC (General Data Protection Regulation) (European Parliament & Council of the European Union); https://data.europa.eu/eli/reg/2016/679/oj
Mehrabi, N., Morstatter, F., Saxena, N., Lerman, K. & Galstyan, A. A survey on bias and fairness in machine learning. ACM Comput. Surv. 54, 115 (2021).
Burkart, N. & Huber, M. F. A survey on the explainability of supervised machine learning. J. Artif. Intell. Res. 70, 245–317 (2021).
Madiega, T. Artificial Intelligence Act (European Parliament, 2023); https://www.europarl.europa.eu/thinktank/en/document/EPRS_BRI(2021)698792
Eurostat. European system of accounts. ESA 2010 Publications Office of the European Union, 2013. Off. J. Eur. Un. 174, 56 (2013).
Biś, D., Podkorytov, M. & Liu, X. Too much in common: shifting of embeddings in transformer language models and its implications. In Proc. 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (eds Toutanova, K. et al.) 5117–5130 (ACL, 2021).
Beltagy, I., Peters, M. E. & Cohan, A. Longformer: the long-document transformer. Preprint at https://arxiv.org/abs/2004.05150 (2020).
Wettig, A., Gao, T., Zhong, Z. & Chen, D. Should you mask 15% in masked language modeling? In Proc. 17th Conference of the European Chapter of the Association for Computational Linguistics (eds Vlachos, A. & Augenstein, I.) 2985–3000 (ACL, 2023).
Jawahar, G., Sagot, B. & Seddah, D. What does BERT learn about the structure of language? In Proc. 57th Annual Meeting of the Association for Computational Linguistics (eds Korhonen, A. et al.) 3651–3657 (ACL, 2019).
Sun, C., Qiu, X., Xu, Y. & Huang, X. How to fine-tune BERT for text classification? Proc. CCl 11856, 194–206 (2019).
Huang, S., Wang, S., Li, D. & Jiang, L. AUL is a better optimization metric in PU learning. Preprint at https://openreview.net/forum?id=2NU7a9AHo-6 (2023).
Wilmoth, J. R. et al. in Methods Protocol for the Human Mortality Database 10–11 (Univ. California Berkeley and Max Planck Institute for Demographic Research, 2007).
Lee, K. & Ashton, M. C. Psychometric properties of the HEXACO personality inventory. Multivariate Behav. Res. 39, 329–358 (2004).
Yu, S. et al. A re-balancing strategy for class-imbalanced classification based on instance difficulty. In Proc. 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) (ed. O'Conner L.) 70–79 (IEEE, 2022).
Müller, R., Kornblith, S. & Hinton, G. E. When does label smoothing help? In Adv. Neural Information Processing Systems 32 (NeurIPS 2019) (eds H. Wallach. et al.). 32, 4694–4703 (Curran Associates, 2019).
Polat, G. et al. Class distance weighted cross-entropy loss for ulcerative colitis severity estimation. Proc. MIUA 13413, 157–171 (2022).
Lin, T.-Y., Goyal, P., Girshick, R., He, K. & Dollár, P. Focal loss for dense object detection. Proc. IEEE PAMI 2, 318–327 (2018).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the inception architecture for computer vision. In Proc. 2016 IEEE Conference on Computer Vision and Pattern Recognition (ed. O'Conner L.) (CVPR) 2818–2826 (IEEE, 2016).
Groenendijk, R., Karaoglu, S., Gevers, T. & Mensink, T. Multi-loss weighting with coefficient of variations. In Proc. 2021 IEEE Winter Conference on Applications of Computer Vision (WACV) 1468–1477 (IEEE, 2021).
Kriegeskorte, N., Mur, M. & Bandettini, P. A. Representational similarity analysis-connecting the branches of systems neuroscience. Front. Syst. Neurosci. 2, 4 (2008).
Liang, Y., Cao, R., Zheng, J., Ren, J. & Gao, L. Learning to remove: towards isotropic pre-trained BERT embedding. In Proc. Artificial Neural Networks and Machine Learning – ICANN 2021: 30th International Conference on Artificial Neural Networks (eds. Farkaš I. et al.) 448–459 (ACM, 2021).
Mu, J., Bhat, S. & Viswanath, P. All-but-the-top: simple and effective postprocessing for word representations. Preprint at https://openreview.net/forum?id=HkuGJ3kCb (2023).
Savcisens, G. Socialcomplexitylab/life2vec. Zenodo https://doi.org/10.5281/zenodo.10118621 (2023).
Acknowledgements
We thank S. M. Hartmann for help with structuring and refactoring the code and M. F. Odgaard as well as the entire Social Complexity Lab for helpful feedback and discussions. The work was funded by the Villum Foundation Grant Nation-Scale Social Networks (to S.L.).
Author information
Authors and Affiliations
Contributions
S.L. and G.S. conceived and designed the analysis. G.S. implemented the computational framework and performed the analysis, and T.E.-R., L.K.H., L.H.M., L.L., A.R., I.Z. and S.L. supported the analysis. T.E.-R., A.R. and L.K.H. contributed to the algorithmic auditing. A.R. contributed to the methodology of the transformer architecture. S.L., L.K.H. and L.H.M. refined the statistical evaluations and methodology. L.L. and I.Z. contributed data and refined the methodology for the personality nuances predictions. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Inclusion and ethics statement
This study relies on secondary analysis of administrative data and does not require approval from the Danish committee system established under the Danish Act on Research Ethics Review of Health Research Projects. The data analysis was conducted in accordance with the rules set by the Danish Data Protection Agency and the information security and data confidentiality policies of Statistics Denmark. See Methods section Ethics and broader impacts, for further information.
Peer review
Peer review information
Nature Computational Science thanks Michal Kosinski, Denis Helic and Dashun Wang for their contribution to the peer review of this work. Primary Handling Editor: Fernando Chirigati, in collaboration with the Nature Computational Science team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary sections 1–5, Figs. 1–10 and Tables 1–12.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Savcisens, G., Eliassi-Rad, T., Hansen, L.K. et al. Using sequences of life-events to predict human lives. Nat Comput Sci 4, 43–56 (2024). https://doi.org/10.1038/s43588-023-00573-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-023-00573-5
This article is cited by
-
Combining the strengths of Dutch survey and register data in a data challenge to predict fertility (PreFer)
Journal of Computational Social Science (2024)