Abstract
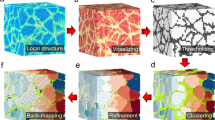
We took the classic ‘guess the number of beans in a jar game’ and amplified the research question. Rather than estimate the quantity of particles in the jar, we sought to characterize the spaces between them. Here we present an approach for delineating the pockets of empty space (three-dimensional pores) between packed particles, which are hotspots for activity in applications and natural phenomena that deal with particulate materials. We utilize techniques from graph theory to exploit information about particle configuration that allows us to locate important spatial landmarks within the void space. These landmarks are the basis for our pore segmentation, where we consider both interior pores as well as entrance and exit pores into and out of the structure. Our method is robust for particles of varying size, form, stiffness and configuration, which allows us to study and compare three-dimensional pores across a range of packed particle types. We report striking relationships between particles and pores that are described mathematically, and we offer a visual library of pore types. With a meaningful discretization of void space, we demonstrate that packed particles can be understood not by their solid space, but by their empty space.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source datasets for Figs. 3–6 are available with this paper. Raw datasets are available in a Code Ocean compute capsule (https://doi.org/10.24433/CO.4876664.v1)81 and at https://seguralab.duke.edu/what/highlights/lovamap/.
Code availability
LOVAMAP segmentation and descriptor code are available through a Code Ocean compute capsule (https://doi.org/10.24433/CO.4876664.v1)81. The copyrights of the LOVAMAP code are owned by Duke University. The code was developed by L.R., P.C. and T.S. at the Segura Lab, Duke University Pratt School of Engineering. To request a license for the code, please contact the Digital Innovations department at the Duke Office for Translation and Commercialization (OTC) (https://olv.duke.edu/software/) at otcquestions@duke.edu with reference to ‘OTC File No. 7784’. CC BY-NC-SA licenses for the code are free for academics. To learn more about LOVAMAP, visit https://seguralab.duke.edu/what/highlights/lovamap/.
References
Cohn, H. A conceptual breakthrough in sphere packing. Not. Am. Math. Soc. 64, 102–115 (2017).
Torquato, S. Random Heterogeneous Materials: Microstructure and Macroscopic Properties 2nd edn (Springer, 2013).
Corwin, E. I., Clusel, M., Siemens, A. O. N. & Brujić, J. Model for random packing of polydisperse frictionless spheres. Soft Matter 6, 2949–2959 (2010).
Desmond, K. W. & Weeks, E. R. Influence of particle size distribution on random close packing of spheres. Phys. Rev. E 90, 022204 (2014).
Torquato, S. Basic understanding of condensed phases of matter via packing models. J. Chem. Phys. 149, 020901 (2018).
Seckendorff, J. & Hinrichsen, O. Review on the structure of random packed-beds. Can. J. Chem. Eng. 99, S703–S733 (2021).
Culligan, K. A., Wildenschild, D., Christensen, B. S. B., Gray, W. G. & Rivers, M. L. Pore-scale characteristics of multiphase flow in porous media: a comparison of air–water and oil–water experiments. Adv. Water Res. 29, 227–238 (2006).
Landry, C. J., Karpyn, Z. T. & Piri, M. Pore-scale analysis of trapped immiscible fluid structures and fluid interfacial areas in oil-wet and water-wet bead packs. Geofluids 11, 209–227 (2011).
Roozbahani, M. M., Huat, B. B. K. & Asadi, A. The effect of different random number distributions on the porosity of spherical particles. Adv. Powder Technol. 24, 26–35 (2013).
Rauter, M., Viroulet, S., Gylfadottir, S. S., Fellin, W. & Lovholt, F. Granular porous landslide tsunami modelling—the 2014 Lake Askja flank collapse. Nat. Commun. 13, 678 (2022).
Doerr, F. J. S. & Florence, A. J. A micro-XRT image analysis and machine learning methodology for the characterisation of multi-particulate capsule formulations. Int. J. Pharm. X 2, 100041 (2020).
Averardi, A., Cola, C., Zeltmann, S. E. & Gupta, N. Effect of particle size distribution on the packing of powder beds: a critical discussion relevant to additive manufacturing. Mater. Today Commun. 24, 100964 (2020).
Walker, D. M. et al. Self-assembly in a near-frictionless granular material: conformational structures and transitions in uniaxial cyclic compression of hydrogel spheres. Soft Matter 11, 2157–2173 (2015).
Zhao, S., Evans, T. M. & Zhou, X. Three-dimensional Voronoi analysis of monodisperse ellipsoids during triaxial shear. Powder Technol. 323, 323–336 (2018).
Yi, L. Y., Zou, R. P., Pinson, D., Dong, K. J. & Yu, A. B. An assessment of the mathematical models for estimating the coordination number of the packing of multisized particles. Powder Technol. 379, 58–68 (2021).
Zhang, C., Zhao, S., Zhao, J. & Zhou, X. Three-dimensional Voronoi analysis of realistic grain packing: an XCT assisted set Voronoi tessellation framework. Powder Technol. 379, 251–264 (2021).
Zhao, S., Zhao, J. & Guo, N. Universality of internal structure characteristics in granular media under shear. Phys. Rev. E 101, 012906 (2020).
Wilson-Whitford, S. R., Gao, J., Chiara Roffin, M., Buckley, W. E. & Gilchrist, J. F. Microrollers flow uphill as granular media. Nat. Commun. 14, 5829 (2023).
Ketcham, R. A., Meth, C., Hirsch, D. M. & Carlson, W. D. Improved methods for quantitative analysis of three-dimensional porphyroblastic textures. Geosphere 1, 42–59 (2005).
Videla, A., Lin, C.-L. & Miller, J. D. Watershed functions applied to a 3D image segmentation problem for the analysis of packed particle beds. Part. Part. Syst. Charact. 23, 237–245 (2006).
Manoharan, V. N. Colloidal matter: packing, geometry, and entropy. Science 349, 1253751 (2015).
Lu, F. et al. Unusual packing of soft-shelled nanocubes. Sci. Adv. 5, eaaw2399 (2019).
Al-Raoush, R. Microstructure characterization of granular materials. Physica A 377, 545–558 (2007).
Batys, P. & Weroński, P. Porosity and tortuosity of layer-by-layer assemblies of spherical particles. Model. Simul. Mater. Sci. Eng. 22, 065017 (2014).
Miyabe, K. New moment equations for chromatography using various stationary phases of different structural characteristics. Anal. Chem. 79, 7457–7472 (2007).
Larachi, F. et al. X-ray micro-tomography and pore network modeling of single-phase fixed-bed reactors. Chem. Eng. J. 240, 290–306 (2014).
Saadatfar, M., Takeuchi, H., Robins, V., Francois, N. & Hiraoka, Y. Pore configuration landscape of granular crystallization. Nat. Commun. 8, 15082 (2017).
Steinhaus, H. Mathematical Snapshots 3rd edn (Dover, 2011).
Houdoux, D., Amon, A., Marsan, D., Weiss, J. & Crassous, J. Micro-slips in an experimental granular shear band replicate the spatiotemporal characteristics of natural earthquakes. Commun. Earth Environ. 2, 90 (2021).
Darling, N. J. et al. Click by click microporous annealed particle (MAP) scaffolds. Adv. Healthc. Mater. 9, e1901391 (2020).
Fang, J. et al. Injectable drug-releasing microporous annealed particle scaffolds for treating myocardial infarction. Adv. Funct. Mater. 30, 2004307 (2020).
Griffin, D. R. et al. Activating an adaptive immune response from a hydrogel scaffold imparts regenerative wound healing. Nat. Mater. 20, 560–569 (2021).
Dumont, C. M. et al. Aligned hydrogel tubes guide regeneration following spinal cord injury. Acta Biomater. 86, 312–322 (2019).
Truong, N. F. et al. Microporous annealed particle hydrogel stiffness, void space size, and adhesion properties impact cell proliferation, cell spreading, and gene transfer. Acta Biomater. 94, 160–172 (2019).
Matsiko, A., Gleeson, J. P. & O’Brien, F. J. Scaffold mean pore size influences mesenchymal stem cell chondrogenic differentiation and matrix deposition. Tissue Eng. Part A 21, 486–497 (2015).
McWhorter, F. Y., Wang, T., Nguyen, P., Chung, T. & Liu, W. F. Modulation of macrophage phenotype by cell shape. Proc. Natl Acad. Sci. USA 110, 17253–17258 (2013).
Zadpoor, A. A. Bone tissue regeneration: the role of scaffold geometry. Biomater. Sci. 3, 231–245 (2015).
Denais, C. M. et al. Nuclear envelope rupture and repair during cancer cell migration. Science 352, 353–358 (2016).
Werner, M. et al. Surface curvature differentially regulates stem cell migration and differentiation via altered attachment morphology and nuclear deformation. Adv. Sci. 4, 1600347 (2017).
Natsui, S., Sawada, A., Nogami, H., Kikuchi, T. & Suzuki, R. O. Topological consideration of 3-D local void structure for static holdup site in packed bed. ISIJ Int. 60, 1453–1460 (2020).
Shelepova, E. A., Paschek, D., Ludwig, R. & Medvedev, N. N. Comparing the void space and long-range structure of an ionic liquid with a neutral mixture of similar sized molecules. J. Mol. Liq. 299, 112121 (2020).
Li, Z., Wang, Y. H., Chow, J. K., Su, Z. & Li, X. 3D pore network extraction in granular media by unifying the Delaunay tessellation and maximal ball methods. J. Pet. Sci. Eng. 167, 692–701 (2018).
van der Linden, J. H., Sufian, A., Narsilio, G. A., Russell, A. R. & Tordesillas, A. A computational geometry approach to pore network construction for granular packings. Comput. Geosci. 112, 133–143 (2018).
Schaller, F. M. et al. Non-universal Voronoi cell shapes in amorphous ellipsoid packs. Europhys. Lett. 111, 24002 (2015).
Weis, S., Schönhöfer, P. W. A., Schaller, F. M., Schröter, M. & Schröder-Turk, G. E. Pomelo, a tool for computing generic set Voronoi diagrams of aspherical particles of arbitrary shape. EPJ Web Conf. 140, 5–8 (2017).
Willems, T. F., Rycroft, C. H., Kazi, M., Meza, J. C. & Haranczyk, M. Algorithms and tools for high-throughput geometry-based analysis of crystalline porous materials. Microporous Mesoporous Mater. 149, 134–141 (2012).
Li, X. & Li, X. S. Micro–macro quantification of the internal structure of granular materials. J. Eng. Mech. 135, 641–656 (2009).
Fu, P. & Dafalias, Y. F. Relationship between void- and contact normal-based fabric tensors for 2D idealized granular materials. Int. J. Solids Struct. 63, 68–81 (2015).
Roozbahani, M. M., Borela, R. & Frost, J. D. Pore size distribution in granular material microstructure. Materials 10, 1237 (2017).
Caldwell, A. S., Campbell, G. T., Shekiro, K. M. T. & Anseth, K. S. Clickable microgel scaffolds as platforms for 3D cell encapsulation. Adv. Healthc. Mater. 6, 1700254 (2017).
Sideris, E. et al. Particle hydrogels based on hyaluronic acid building blocks. ACS Biomater. Sci. Eng. 2, 2034–2041 (2016).
Sheikhi, A. et al. Microfluidic-enabled bottom-up hydrogels from annealable naturally-derived protein microbeads. Biomaterials 192, 560–568 (2019).
Khorasani, H. et al. A quantitative approach to scar analysis. Am. J. Pathol. 178, 621–628 (2011).
Wershof, E. et al. A FIJI macro for quantifying pattern in extracellular matrix. Life Sci. Alliance 4, e202000880 (2021).
Jiang, Z. et al. Machine-learning-revealed statistics of the particle-carbon/binder detachment in lithium-ion battery cathodes. Nat. Commun. 11, 2310 (2020).
Blum, H. A. in Models for the Perception of Speech and Visual Form (ed. Wathen-Dunn, W.) 362–380 (MIT Press, 1967).
Sherbrooke, E. C., Patrikalakis, N. M. & Brisson, E. An algorithm for the medial axis transform of 3D polyhedral solids. IEEE Trans. Vis. Comput. Graph. 2, 44–61 (1996).
Pizer, S. M., Siddiqi, K., Székeley, G., Damon, J. N. & Zucker, S. W. Multiscale medial loci and their properties. Int. J. Comput. Vis. 55, 155–179 (2003).
Hesselink, W. H. & Roerdink, J. B. T. M. Euclidean skeletons of digital image and volume data in linear time by the integer medial axis transform. IEEE Trans. Pattern Anal. Mach. Intell. 30, 2204–2217 (2008).
Xiong, Q., Baychev, T. G. & Jivkov, A. P. Review of pore network modelling of porous media: experimental characterisations, network constructions and applications to reactive transport. J. Contam. Hydrol. 192, 101–117 (2016).
Shaked, D. & Bruckstein, A. M. Pruning medial axes. Comput. Vis. Image Underst. 69, 156–169 (1998).
Lindquist, W. B. & Venkatarangan, A. Investigating 3D geometry of porous media from high resolution images. Phys. Chem. Earth A 25, 593–599 (1999).
Silin, D. & Patzek, T. Pore space morphology analysis using maximal inscribed spheres. Physica A 371, 336–360 (2006).
Jones, A. C. et al. The correlation of pore morphology, interconnectivity and physical properties of 3D ceramic scaffolds with bone ingrowth. Biomaterials 30, 1440–1451 (2009).
Chiang, S.-C. The Euclidean Distance Transform. PhD thesis, Purdue Univ. (1992).
Liang, Z., Ioannidis, A. & Chatzis, I. Geometric and topological analysis of three-dimensional porous media: pore space partitioning based on morphological skeletonization. J. Colloid Interface Sci. 221, 13–24 (2000).
Youssef, S. et al. High resolution CT and pore-network models to assess petrophysical properties of homogeneous and heterogeneous carbonates. In SPE/EAGE Reservoir Characterization and Simulation Conference SPE-111427-MS (SPE, 2007).
Thomson, P.-R., Hazel, A. & Hier-Majumder, S. The influence of microporous cements on the pore network geometry of natural sedimentary rocks. Front. Earth Sci. https://doi.org/10.3389/feart.2019.00048 (2019).
Hormann, K., Baranau, V., Hlushkou, D., Höltzel, A. & Tallarek, U. Topological analysis of non-granular, disordered porous media: determination of pore connectivity, pore coordination, and geometric tortuosity in physically reconstructed silica monoliths. New J. Chem. 40, 4187–4199 (2016).
Jiang, Z. et al. Efficient extraction of networks from three-dimensional porous media. Water Resour. Res. 43, W12S03 (2007).
Rabbani, A., Ayatollahi, S., Kharrat, R. & Dashti, N. Estimation of 3-D pore network coordination number of rocks from watershed segmentation of a single 2-D image. Adv. Water Res. 94, 264–277 (2016).
Huaimin, D., Jianmeng, S., Likai, C., Naser, G. & Weichao, Y. Characteristics of the pore structure of natural gas hydrate reservoir in the Qilian Mountain Permafrost, Northwest China. J. Appl. Geophys. 164, 153–159 (2019).
Dong, H. & Blunt, M. J. Pore-network extraction from micro-computerized-tomography images. Phys. Rev. E 80, 036307 (2009).
Medvedev, N. N., Voloshin, V. P., Luchnikov, V. A. & Gavrilova, M. L. An algorithm for three-dimensional Voronoi S-network. J. Comput. Chem. 27, 1676–1692 (2006).
Joon Lee, C., Kang, Y.-M., Cho, K.-H. & No, K. T. A robust method for searching the smallest set of smallest rings with a path-included distance matrix. Proc. Natl Acad. Sci. USA 106, 17355–17358 (2009).
Jones, J. R., Atwood, R. C., Poologasundarampillai, G., Yue, S. & Lee, P. D. Quantifying the 3D macrostructure of tissue scaffolds. J. Mater. Sci. Mater. Med. 20, 463–471 (2009).
Bashoor-Zadeh, M., Baroud, G. & Bohner, M. Geometric analysis of porous bone substitutes using micro-computed tomography and fuzzy distance transform. Acta Biomater. 6, 864–875 (2010).
Gostick, J. T. Versatile and efficient pore network extraction method using marker-based watershed segmentation. Phys. Rev. E 96, 023307 (2017).
Rong, L. W., Dong, K. J. & Yu, A. B. Lattice-Boltzmann computation of hydraulic pore-to-pore conductance in packed beds of uniform spheres. Chem. Eng. Sci. 224, 115798 (2020).
Sweijen, T., Hassanizadeh, S. M., Chareyre, B. & Zhuang, L. Dynamic pore-scale model of drainage in granular porous media: the pore-unit assembly method. Water Resour. Res. 54, 4193–4213 (2018).
Riley, L., Cheng, P. & Segura, T. Identification and analysis of 3D-pores in packed particulate materials. Code Ocean https://doi.org/10.24433/CO.4876664.v1 (2023).
Acknowledgements
We extend a special thanks to M. Roozbahani, who was a senior graduate student under D. Frost at Georgia Tech, when he kindly shared his experience and expertise during the early stages of the project. We thank the National Institutes of Health, the National Institutes of Neurological Disorders and Stroke (1R01NS112940 (T.S.), 1R01NS079691 (T.S.), R01NS094599 (T.S.)) and the National Institute of Allergy and Infectious Disease (1R01AI152568 (T.S.)).
Author information
Authors and Affiliations
Contributions
L.R. headed the development of the work by designing the computational framework, writing and implementing code, generating data, analyzing data, writing the paper and creating figures. P.C. provided computational expertise through guidance and discussion, contributed to the design of the computational framework, wrote code, played a key role in improving software runtime speeds, designed and simulated input domains, contributed to writing methods, and organized the flow of the final paper. T.S. was pivotal in the conception and direction of the project, including applications to real data, provided guidance and discussion throughout, and made substantial contribution to data analysis, figure preparation, and paper organization and editing.
Corresponding author
Ethics declarations
Competing interests
P.C. is the founder of Ninjabyte Computing, a company that offers computational consulting and simulation expertise. L.R., P.C. and T.S. have filed for patent protection pertaining to LOVAMAP as an analytical tool to facilitate material optimization (reference number DU7784PCT).
Peer review
Peer review information
Nature Computational Science thanks Florian Bouvill, T. Matthew Evans and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Jie Pan, in collaboration with the Nature Computational Science team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–20, Table 1 and Notes 1–3.
Source data
Source Data Fig. 3
Source data for Fig. 3 plots.
Source Data Fig. 4
Source data for Fig. 4 plots.
Source Data Fig. 5
Source data for Fig. 5 plots.
Source Data Fig. 6
Source data for Fig. 6 plots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Riley, L., Cheng, P. & Segura, T. Identification and analysis of 3D pores in packed particulate materials. Nat Comput Sci 3, 975–992 (2023). https://doi.org/10.1038/s43588-023-00551-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-023-00551-x
This article is cited by
-
Investigating the elegance of empty space
Nature Computational Science (2023)