Abstract
Obtaining the free energy of large molecules from quantum mechanical energy functions is a long-standing challenge. We describe a method that allows us to estimate, at the quantum mechanical level, the harmonic contributions to the thermodynamics of molecular systems of large size, with modest cost. Using this approach, we compute the vibrational thermodynamics of a series of diamond nanocrystals, and show that the error per atom decreases with system size in the limit of large systems. We further show that we can obtain the vibrational contributions to the binding free energies of prototypical protein–ligand complexes where exact computation is too expensive to be practical. Our work raises the possibility of routine quantum mechanical estimates of thermodynamic quantities in complex systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
Data for all systems can be regenerated by running our stochastic Lanczos code on the geometries available at https://doi.org/10.6084/m9.figshare.22258447.v1 (ref. 35). Source Data are provided with this paper.
Code availability
The stochastic Lanczos code is available in Supplementary Software 1. The Gaussian and plane waves implementation is available through PySCF at www.pyscf.org.
References
Grimme, S. & Schreiner, P. R. Computational chemistry: the fate of current methods and future challenges. Angew. Chem. Int. Ed. 57, 4170–4176 (2018).
Li, H. & Jensen, J. H. Partial Hessian vibrational analysis: the localization of the molecular vibrational energy and entropy. Theor. Chem. Acc. 107, 211–219 (2002).
Woodcock, H. L., Zheng, W., Ghysels, A., Shao, Y., Kong, J. & Brooks, B. R. Vibrational subsystem analysis: a method for probing free energies and correlations in the harmonic limit. J. Chem. Phys. 129, 214109 (2008).
Filippone, F. & Parrinello, M. Vibrational analysis from linear response theory. Chem. Phys. Lett. 345, 179–182 (2001).
Karplus, M. & Kushick, J. N. Method for estimating the configurational entropy of macromolecules. Macromolecules 14, 325–332 (1981).
Brooks, B. R., Janezic, D. & Karplus, M. Harmonic analysis of large systems. I. Methodology. J. Comput. Chem. 16, 1522–1542 (1995).
Ubaru, S., Chen, J. & Saad, Y. Fast estimation of tr(f(A)) via stochastic Lanczos quadrature. SIAM J. Matrix Anal. Appl. 38, 1075–1099 (2017).
Hellman, O., Steneteg, P., Abrikosov, I. A. & Simak, S. I. Temperature dependent effective potential method for accurate free energy calculations of solids. Phys. Rev. B 87, 1–8 (2013).
Errea, I., Calandra, M. & Mauri, F. Anharmonic free energies and phonon dispersions from the stochastic self-consistent harmonic approximation: application to platinum and palladium hydrides. Phys. Rev. B 89, 1–16 (2014).
Baer, R., Neuhauser, D. & Rabani, E. Self-averaging stochastic Kohn–Sham density-functional theory. Phys. Rev. Lett. 111, 1–5 (2013).
Han, I., Malioutov, D. & Shin, J. Large-scale log-determinant computation through stochastic chebyshev expansions. In International Conference on Machine Learning 908–917 (PMLR, 2015).
Kaledin, A. L. Gradient-based direct normal-mode analysis. J. Chem. Phys. 122, 184106 (2005).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865 (1996).
Grimme, S., Bannwarth, C. & Shushkov, P. A robust and accurate tight-binding quantum chemical method for structures, vibrational frequencies, and noncovalent interactions of large molecular systems parametrized for all spd-block elements (z = 1–86). J. Chem. Theory Comput. 13, 1989–2009 (2017).
Ehrlich, S., Göller, A. H. & Grimme, S. Towards full quantum-mechanics-based protein–ligand binding affinities. ChemPhysChem 18, 898–905 (2017).
Spicher, S. & Grimme, S. Efficient computation of free energy contributions for association reactions of large molecules. J. Phys. Chem. Lett. 11, 6606–6611 (2020).
Grimme, S. Supramolecular binding thermodynamics by dispersion-corrected density functional theory. Chemistry 18, 9955–9964 (2012).
Reiling, K. K., Endres, N. F., Dauber, D. S., Craik, C. S. & Stroud, R. M. Anisotropic dynamics of the JE-2147-HIV protease complex: drug resistance and thermodynamic binding mode examined in a 1.09 Å structure. Biochemistry 41, 4582–4594 (2002).
Mardirossian, N., Wang, Y., Pearlman, D.A., Chan, G.K. & Shiozaki, T. Novel algorithms and high-performance cloud computing enable efficient fully quantum mechanical protein–ligand scoring. Preprint at https://arxiv.org/abs/2004.08725 (2020).
Chen, W., Gilson, M. K., Webb, S. P. & Potter, M. J. Modeling protein–ligand binding by mining minima. J. Chem. Theory Comput. 6, 3540–3557 (2010).
Meyer, R. A., Musco, C., Musco, C. & Woodruff, D. P. Hutch++: optimal stochastic trace estimation. In Symposium on Simplicity in Algorithms (SOSA) 142–155 (Society for Industrial and Applied Mathematics, 2021).
Jaklič, J. & Prelovšek, P. Lanczos method for the calculation of finite-temperature quantities in correlated systems. Phys. Rev. B 49, 5065 (1994).
Neese, F. Software update: the ORCA program system, version 4.0. WIREs Comput. Mol. Sci. 8, 4–9 (2018).
Bannwarth, C., Ehlert, S. & Grimme, S. GFN2-xTB—an accurate and broadly parametrized self-consistent tight-binding quantum chemical method with multipole electrostatics and density-dependent dispersion contributions. J. Chem. Theory Comput. 15, 1652–1671 (2019).
Bannwarth, C. et al. Extended tight-binding quantum chemistry methods. WIREs Comput. Mol. Sci. 11, 1–49 (2021).
McClain, J., Sun, Q., Chan, G. K.-L. & Berkelbach, T. C. Gaussian-based coupled-cluster theory for the ground-state and band structure of solids. J. Chem. Theory Comput. 13, 1209–1218 (2017).
Schindler, C. E. M. et al. Large-scale assessment of binding free energy calculations in active drug discovery projects. J. Chem. Inf. Model. 60, 5457–5474 (2020).
Maier, J. A. et al. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general Amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Case, D.A. et al. Amber 2021 (Univeristy of California, San Francisco, 2021); https://ambermd.org/doc12/Amber21.pdf
Schrödinger, L. The PyMOL Molecular Graphics System v.1.8 (PyMOL, 2015).
Waaler, J. et al. Preclinical lead optimization of a 1,2,4-triazole based tankyrase inhibitor. J. Med. Chem. 63, 6834–6846 (2020).
Berman, H. M. et al. The protein data bank. Acta Crystallog. Section D 58, 899–907 (2002).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
White, A.F., Li, C., Zhang, X. & Chan, G. K.-L. Stochastic Harmonic Free Energy (Figshare, 2023); https://doi.org/10.6084/m9.figshare.22258447.v1
Buchstaller, H.-P. et al. Discovery and optimization of 2-arylquinazolin-4-ones into a potent and selective tankyrase inhibitor modulating wnt pathway activity. J. Medicinal Chem. 62, 7897–7909 (2019).
Acknowledgements
We thank S. Murlidaran for help with protein preparation. A.F.W. (theoretical analysis) was supported by the US Department of Energy (grant no. DE-SC0018140). C.L. (computational studies) was supported by the US National Science Foundation (grant no. 1931328). X.Z. (improved DFT implementation for proteins) was supported by the US Department of Energy (grant no. DE-SC0019330). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.F.W. and G.K.C. conceived the project. A.F.W., C.L. and X.Z. performed the work. All authors contributed to the writing of the paper.
Corresponding author
Ethics declarations
Competing interests
Garnet K. Chan is a co-owner of QSimulate, Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Jan Jensen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Jie Pan, in collaboration with the Nature Computational Science team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
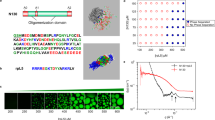
Extended Data Fig. 1 Harmonic binding free energies of TNKS.
Harmonic thermal contributions for the TNKS2 complex computed at the xTB level. a) An image of the TNKS2 complex. The truncated part of the protein is shown as transparent, while the remaining two protein chains and the ligand are colored orange, yellow and grey, respectively. Image rendered by VMD. b) The thermal enthalpy and entropy, and free energy of binding for the TNKS2 system for varying Lanczos order (m = 8, 16, 32 from above to below). The exact value is given by the grey vertical line. Box edges represent mean ± one standard error of 100 samples, center line represents the median, whiskers represent two standard errors, and fliers are data beyond two standard errors. c) Statistical errors in binding free energy quantities as a function of % of cost of the exact calculation (error bars denote error of error). The detailed error estimation protocol is in Supplementary Section 1.
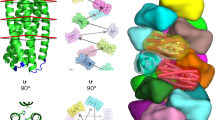
Extended Data Fig. 2 Harmonic binding free energies of HIV protease.
Harmonic thermal contributions for the JE-2147-HIV protease complex computed at the xTB level. a) An image of the JE-2147-HIV protease complex. The two protein chains and the ligand are shown in yellow, green, and grey respectively. Image rendered by VMD. b) Statistical errors in binding free energy quantities as a function of % of cost of the exact calculation (error bars denote error of error). The detailed error estimation protocol is in Supplementary Section 1.
Supplementary information
Supplementary Information
Supplementary Sections 1–4, Figs. 1–4, Tables 1 and 2, and Discussion.
Supplementary Data 1
Source Data for Supplementary Fig. 1. Random samples of harmonic free energies.
Supplementary Software 1
Computer code for stochastic sampling of harmonic free energies.
Source data
Source Data Fig. 1
Random samples of harmonic free energies of diamonds.
Source Data Extended Data Table/Fig. 1
Random samples of harmonic free energies of TNKS.
Source Data Extended Data Table/Fig. 2
Random samples of harmonic free energies of HIV protease.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
White, A.F., Li, C., Zhang, X. et al. Quantum harmonic free energies for biomolecules and nanomaterials. Nat Comput Sci 3, 328–333 (2023). https://doi.org/10.1038/s43588-023-00432-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-023-00432-3