Abstract
Fungi comprise relevant human pathogens, causing over a billion infections each year. Plastic pollution alters niches of fungi by providing trillions of artificial microhabitats, mostly in the form of microplastics, where pathogens might accumulate, thrive, and evolve. However, interactions between fungi and microplastics in nature are largely unexplored. To address this knowledge gap, we investigated the assembly, architecture, and ecology of mycobiomes in soil (micro)plastispheres near human dwellings in a model- and network-based metagenome study combined with a global-scale trait data annotation. Our results reveal a strong selection of important human pathogens, in an idiosyncratic, otherwise predominantly neutrally assembled plastisphere, which is strongly linked to generic fungal virulence traits. These findings substantiate our niche expansion postulate, demonstrate the emergence of plastiphily among fungal pathogens and imply the existence of a plastisphere virulence school, underpinning the need to declare microplastics as a factor of global health.
Similar content being viewed by others
Introduction
One of humanity’s irrevocable legacies for centuries to come is that all life on Earth must interact with plastics. Plastic is a pervasive pollutant in the biosphere, where it affects all levels of biological organisation1. Elusive microplastic (MP; <5 mm)2 and nanoplastic particles (<1 µm), primarily derived from fragmentation of larger plastic objects, can invade tissues and organs, and even infiltrate cells, where they can cause inflammatory responses and subcellular dysfunctions3. Organismic interactions with plastic debris, such as ingestion and entanglement, are known to sometimes have severe consequences in the fitness of individuals, including obstruction4, asphyxiation5 and impairment of life history traits6. Plastic debris also impacts populations and trophic networks by introducing allochthonous chemicals7, disrupting intraspecific communication8 or altering environmental conditions9. While direct and indirect impacts have been extensively studied primarily in relation to larger organisms and at local scales, microorganisms are increasingly being studied for cascading impacts of plastic pollution at community level that could affect entire ecosystems10,11.
For microbes, MP pollution is essentially a terraforming event, with trillions of manmade, long-lived, physico-chemically diverse substrates permeating all of Earth’s biomes12,13 and providing a variety of ecological niches for microbial life14. It is in these interaction landscapes that MP pollution probably unfolds its greatest ecological disruption potential. Therefore, in recent years, the focus of MP research has shifted to the microbial colonisation of plastic debris in different ecosystems, with the common goal of elucidating the nature of the plastisphere10,11,15. The scientific consensus is that plastispheres harbour multipartite, dynamic microbial communities that generally differ in structure and composition from biological assemblages in the surrounding compartment or other natural substrates11,14. These altered microbiomes have potentially serious ecological and functional implications, such as the concentration of antibiotic resistance genes16, the facilitation of horizontal gene transfer17, the accumulation as well as transmission of pathogenic microbes10,18. Therefore, exploring the mechanisms that lead to the formation of epiplastic communities is crucial for assessing the ecological risks posed by the plastisphere microbiota.
Fungal pathogens have a huge impact on plant and animal life, driving species extinctions, causing ecosystem failure, and threatening food security19,20,21. Fungal infections in humans represent a devastating and worsening yet overlooked global health crisis, with over one billion cases and more than 1.6 million deaths annually22,23. A critical factor in the emergence of fungal diseases is anthropogenic alteration of the natural environment, which entails new opportunities for fungal pathogens to thrive, diversify and evolve20,24. Yet, plastisphere mycobiomes, especially in terrestrial environments, have been almost entirely omitted, although plastic pollution is one of the most profound alterations to the Earth’s surface25 and fungal pathogens are ecologically predisposed and biologically fine-tuned to benefit from it. Pathogenic fungi are excellent colonisers of hydrophobic substrates such as phyllospheres26 and animal skin27 that are also prevalent in all types of extreme and oligotrophic habitats in the natural and built environment28,29, such as medical devices30 and even dishwashers31, where they form highly resistant, polymycobial biofilms32. This niche width is enabled by a spectrum of ecological traits, such as extremotolerance28 and melanisation33,34. However, while in nature those traits enable general adaptability and survival28, in humans they promote opportunistic pathogenicity and are therefore considered virulence factors35,36,37. Preliminary evidence that MPs act as a reservoir for extremotolerant and melanised opportunists in terrestrial ecosystems10 suggests that the plastisphere is an ideal ‘home’ for fungal pathogens and implies a link to these complex virulence traits, raising fundamental questions about their relevance in the assembly of plastisphere mycobiomes.
Microbial community assembly is generally driven by two complementary processes: stochastic processes (neutral theory)38, which emphasise the role of probabilistic dispersal and ecological drift, and deterministic processes (niche theory), which emphasise environmental filtering and biological interactions as drivers of community structure39. Adopting niche theory to epiplastic communities would thus imply that certain microbes are positively selected by MPs and can thrive in the plastisphere due to differences in their relative ecological fitness, a phenomenon we call ‘plastiphily’. Recent model integrations of these theories, however, have shown that neutral processes dominate the assembly of epiplastic bacteriomes in both aquatic40 and soil environments16,41, indicating that the bulk of these communities arise by chance. On the other hand, network-based studies have highlighted the importance of biotic interactions such as competition and facilitation, as well as so-called keystone taxa in the formation of microbial biofilms on environmental plastics42,43, and polymer type was considered as a deterministic driver in plastisphere bacteriome development under experimental field conditions11. Selective factors play a crucial role in structuring mycobiomes during colonisation of extreme habitats44 and the succession of dysbiotic mycobiota45, which are potentially linked to pathogenicity outcomes. Considering their systematic co-occurrence on soil MPs10 and assuming fitness advantages due to the above-mentioned complex virulence traits, it is likely that certain fungal pathogens are selected by MPs and thus ‘plastiphilic’. To date, reports on the ecological mechanisms underlying the assembly of soil plastisphere mycobiomes are lacking, so at this stage virtually nothing is known about the origin of epiplastic fungal pathogens in terrestrial systems.
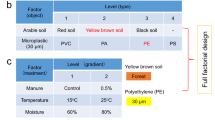
Here, we seek to decipher the ecological processes and biotic interactions that determine the assembly of soil (micro)plastisphere mycobiomes in an explorative field study in tropical soils of Kenya, where extreme plastic pollution meets some of the largest fungal pathogen reservoirs in the world24. Therefore, we applied a neutral community model and co-occurrence network analysis to ITS sequence data obtained by barcoding fungal metagenomes from different interface systems between the natural and built environment, which are designated hotspots for fungal-plastic interaction landscapes where mature communities have formed10. By using a randomised sampling design and analysing indeterminate plastispheres regardless of polymer type, the focus of our work was on intrinsic representativeness and the description of emergence effects. We contextualise model and network outputs with phylogenetic and community diversity analyses and combine them with a global-scale trait data annotation to answer the following questions: (i) does the plastisphere metacommunity (MC) exhibit an idiosyncratic architecture across abundance fractions? (ii) are epiplastic mycobiomes structured by an interplay of stochastic and deterministic processes? (iii) are human pathogens particularly selected by MP (fungal pathogen plastiphily hypothesis)? and (iv) do these pathogens prefer the in situ plastisphere over the ambient soil (niche expansion postulate) (Fig. 1a)? Virulence is a context-dependent and multifactorial, relative extended phenotype of microorganisms37. Unlike bacteriologists, mycologists have not been able to define classical virulence factors, even for notorious human pathogens, due to a teleological-holistic view of such traits28 and the largely opportunistic nature of fungal pathogenicity35,36,37. As a result, to date there is no effective and reliable (meta-)omics method to broadly assess virulence potentials in complex fungal communities. To address the final question of our work still adequately, namely (v) whether there is a relationship between virulence traits and plastiphily (plastiphilic virulence traits hypothesis) (Fig. 1a), we introduce here for the first time the concept of ‘generic virulence traits’ (GVTs) (Fig. 1b). We assign ecological multi-use traits that are phylogenetically widespread and known to support virulence in a variety of fungi, namely melanisation, extremotolerance, dimorphism and oligotrophy28,35,36,37, to all phylotypes identified to genus level to approximate generic virulence potential (GVP). Based on these findings, we shed light on the mechanisms underlying the emergence of plastisphere mycobiomes and pathogenic fungi in soil environments, describe how conserved, ecological, non-specific virulence factors correlate with plastiphily in human pathogens, and discuss the global health relevance of these phenomena in an increasingly plasticised world.
a The central hypotheses of the study. Plastisphere and soil metacommunities show architectural idiosyncrasies in the form of diversity variations and even compartment-specific taxa when all abundance fractions are considered (i). General plastisphere MC assembly is determined by a relatively balanced interplay between deterministic (selection, biotic interactions, etc.) and stochastic processes (e.g., dispersal and drift) (ii), while particularly pathogenic fungi are subject to selection effects and keystone taxa (fungal pathogen plastiphily hypothesis) (iii). Positive selection of pathogens will be stronger in the in situ plastisphere compared to the surrounding soil, indicating a niche preference (niche expansion postulate) (iv). Finally, positive selection by MP, i.e., plastiphily, will correlate with the (co-)incidence of generic virulence traits (GVTs), that is a higher generic virulence potential (GVP) (plastiphilic virulence traits hypothesis) (v). b The GVT concept. Fungal pathogenicity in humans is largely opportunistic, so their virulence is strongly linked to their environmental (out-of-host) ecology28. Consequently, classical virulence traits for human pathogens have not been identified35,36,37. However, ancient, (1) phylogenetically widespread, (2) ecologically multifunctional (and thus non-specific) traits exist that (3) support virulence in a variety of fungi and thus generically (ecological conditions). Among these GVTs, we selected those that are also (4) well-described and frequently observed on a global scale (e.g., at different sites, independent records), (5) without a specific link to plastisphere colonisation according to the current state of research (unbiased), and (6) for which annotation by means of taxonomic profiling at the genus level can be performed comprehensively (operational conditions). From the four selected GVTs, namely melanisation, extremotolerance, oligotrophy and dimorphism, we deduce for all identified fungal genera a reductionistic approximation metric for the GVP (values between 0 and 4; integers only) and thus an ordinal scale for eco-evolutionary projections.
Results
Plastisphere and soil metacommunity
After consolidation of the data and exclusion of singletons, we obtained a total dataset consisting of 95 samples and 4,772,721 high-quality ITS sequences, which were clustered into 2441 fungal OTUs (2208 after rarefaction) (Supplementary Data 1). About 37% of all phylotypes (838 OTUs) were identified at the genus or species level and subjected to meta-analysis (species-level synopsis in Supplementary Data 2). The soil MC was much more species-rich (1991 OTUs) than the plastic MC (927 OTUs) (Fig. 2a), indicating that the plastisphere is the more selective compartment. The five most species-rich classes on MP, Dothideomycetes, Sordariomycetes, Agaricomycetes, Eurotiomycetes and Tremellomycetes, accounted for a similar proportion of plastisphere (89%) and soil (86%) reads, while they were systematically and significantly more diverse in soil than in the plastisphere (Fig. 2b, Supplementary Tables 1–3). Assignment to guilds resulted in four main guilds (Fig. 2c, g, Supplementary Data 2), of which saprotrophs were the most diverse in the soil (p < 0.05), while MP showed similar diversity levels for saprotrophs and pathogens (Fig. 2c, Supplementary Tables 4–6, Supplementary Data 3). Among abundance fractions, conditionally rare and abundant taxa (CRAT) were similarly diverse between compartments, while most other fractions were significantly more diverse (p < 0.05) in the soil (Supplementary Tables 7–9, Supplementary Data 3).
a Schematic illustration of soil and plastisphere metacommunities. LC = local community, corresponding to a soil or MP sample, respectively. Soil MC comprised 49 samples and 1991 OTUs, while plastisphere MC comprised 46 samples and 927 OTUs. b Shannon diversity among the most speciose classes found on MP and in the soil, where Agar. = Agaricomycetes, Trem. = Tremellomycetes, Euro. = Eurotiomycetes, Sord. = Sordariomycetes and Doth. = Dothideomycetes. ‘other’ codes for all remaining identified and unassigned classes of the respective compartments. c Shannon diversity among ecological guilds found on MP and in the soil. d Shannon diversity among abundance fractions of either compartment. ART, always rare taxa; CRT, conditionally rare taxa; CRAT, conditionally rare and abundant taxa; MT, moderate taxa; CAT, conditionally abundant taxa; AAT, always abundant taxa. The lower-case letters indicate significant differences between groups (p < 0.05) via one-way ANOVA. Box plots indicate median (middle line), 25th, 75th percentile (box) and 5th and 95th percentile (whiskers). e Venn diagram showing the number of shared and specific OTUs of the metacommunities. Compartment-specific and shared relative abundances of f speciose classes, including Mala. = Malasseziomycetes, g ecological guilds and h abundance fractions separated by sites (S-1 to S-5). Colour-coding is consistent for all panels.
Compartment-specific and shared diversity
Around 64% of the soil-inhabiting and 23% of the plastic-colonising phylotypes were found exclusively in local communities of the respective compartments (specific MC), with 710 OTUs shared between both metacommunities (Fig. 2e). Across all sites, the soil-specific MC was dominated by Sordariomycetes and Agaricomycetes (Fig. 2f left), had large proportions of unidentified guilds, and small and low numbers of pathogens (Fig. 2g left), with most reads assigned to low-abundance fractions (Fig. 2h left). The shared MC comprised predominantly Dothideomycetes and Sordariomycetes (Fig. 2f middle) and between 36 to over 50% pathogens (Fig. 2g middle), with reads assigned to all six abundance fractions similarly across all sites (Fig. 2h middle), resembling an ‘intermediate state’ between soil- and plastic-specific MC. In terms of abundance, the plastic-specific MC was solely dominated by Dothideomycetes and pathogens, which reached proportions of over 90%, respectively (Fig. 2f, g right), with always abundant taxa (AAT) comprising between around 50–90% of reads (Fig. 2h right). These observations demonstrate both the nestedness and strong idiosyncrasy of the metacommunities, as well as the presence and dominance of plastic-specific fungal pathogens.
PCoA ordinations and ANOSIM tests of each fraction revealed a complete compositional separation of soil and plastisphere fungal communities (p < 0.01), with the highest differences in beta diversity between CRAT (R = 0.603) and AAT fractions (R = 0.494, Fig. 3a, Supplementary Table 10). Mean abundance correlated significantly (p < 0.01) and strongly (r > 0.6) with niche width in both MCs (Fig. 3b), with no difference between average niche width between compartments.
a Beta diversity between abundance fractions of soil and plastisphere metacommunities based on Bray-Curtis dissimilarity and visualised by PCoA ordination. ART, always rare taxa; CRT, conditionally rare taxa; CRAT, conditionally rare and abundant taxa; MT, moderate taxa; CAT, conditionally abundant taxa; AAT, always abundant taxa. Ellipsoids represent 95% confidence intervals. Similarities (sample statistic ‘R’) and significance (included p values) were assessed using ANOSIM. b Spearman correlation between fungal niche width and mean relative abundance of OTUs in the plastisphere and soil MC. Bubble colours code for the six distinct abundance fractions mentioned above. Correlation coefficient (r) and significance (p) value included.
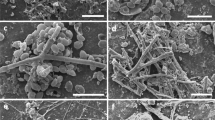
To further characterise the plastic-specific MC, we assessed the phylogenetic distributions of the 84 plastic-specific phylotypes identified to species or genus level (Fig. 4). These fungi made up 9% of OTUs and ca. 13% of reads of the entire plastisphere MC and belonged predominantly (79 taxa) to the subkingdom Dikarya, indicating an overall low evolutionary distance. Ecologically, plastic-specific fungi could be roughly categorised into saprotrophic or mutualistic non-opportunists with low to no virulence score (37 taxa, e.g., Agaricomycetes, Leotiomycetes, Mortierellomycetes) and predominantly pathogenic opportunists with high virulence score (34 taxa, e.g., diverse Dothideomycetes, most Tremellomycetes and all Eurotiomycetes). Among fungi explicitly relevant to human health, we found dimorphic, polyextremotolerant black yeasts, such as Exophiala and Knufia; ubiquitous, hyaline multi-host pathogens like Fusarium and Aspergillus; several highly clinically relevant, cryptococcal yeasts, including Papiliotrema, Filobasidium and Cryptococcus; and multiple allergenic, toxigenic and pathogenic black fungi, such as Alternaria, Curvularia, Chaetomium and Stachybotrys, which were by far the most abundant plastic-specific fungi (ca. 80% of reads).
Displayed are those 84 OTUs that have been found exclusively on MP and identified at genus or species level. Clade colours code for fungal classes. Phylotypes were classified into ecological guilds (green, red and brown bubbles) and annotated with trait data including opportunism (dark red triangle) and the four generic virulence traits, namely melanisation (black bubble), extremotolerance (dark grey), oligotrophy (light grey) and dimorphism (white). Traits were only annotated when present. Pez. = Pezizomycetes. (*) indicates ‘unclassified’.
Mycobiome assembly
The neutral community model (NCM) of Sloan et al.46 was used to assess the contribution of stochastic and deterministic processes to mycobiome assembly in microplastispheres and the bulk soil (Fig. 5). As indicated by the overall lower Akaike’s information criterion (AIC) scores, the NCM outperformed the binomial distribution model for both compartments (Supplementary Tables 11 and 12), indicating an influence of passive dispersal and ecological drift beyond purely random sampling from the respective metacommunities47. The NCM estimated much of the correlation between occurrence frequency and relative abundance variation of plastisphere and soil OTUs, explaining 62% (plastic) and 67.5% (soil) of the community variance (Fig. 5), indicating the stronger effect of stochastic processes on soil community assembly. Migration rates (m) were estimated to be higher for soil than for plastisphere fungal communities (Supplementary Table 11), suggesting greater dispersal of fungi from the soil MC. In both compartments, there were several phylotypes that occurred more or less frequently than predicted by the NCM given their overall abundance in the MC (Fig. 5 and Supplementary Data 3). Points that are above the prediction (cyan) represent fungi that are more frequent than expected, suggesting that they are actively selected and maintained by the compartment, while points below the prediction frame (dark red) represent fungi that occur less frequently than expected, suggesting that the environmental compartment is either selecting against them or limiting their ability to disperse (Fig. 5). Plastisphere taxa above prediction are therefore considered plastiphilic fungi.
The predicted occurrence frequencies for plastic, soil and all representing fungal communities from MP, soil and both compartments, respectively. Each point represents a fungal OTU and different colours indicate OTUs that occur more (cyan) or less frequently (dark red) than predicted by the neutral model. Solid blue lines indicate the optimum fit to the NCM and dashed blue lines represent 95% confidence intervals around the model prediction. R2 indicates the fit of the neutral model and Nm equals the MC size times immigration.
Ecological distinction between neutral and non-neutral partitions
To disentangle the effects of selective processes on community structure and ecology, we separated each MC into three partitions comprising the OTUs found above, below, and within the neutral prediction, respectively (Supplementary Data 2). Taxa from the neutral and non-neutral partitions showed different community composition, diversity, and abundance in soil and plastic (Fig. 6, Supplementary Data 3). Beta diversity among partitions and compartments were assessed using Hellinger distance based on square-root transformed read counts and visualised by non-metric multidimensional scaling (NMDS). Community structure between partitions differed significantly in plastic (ANOSIM R = 0.765), soil (R = 0.872) and between compartments (R = 0.845) at p < 0.01 (Fig. 6a, Supplementary Tables 13 and 14). In both compartments, the non-neutral partitions of MC exhibited higher within-group and between-group heterogeneity than the neutral partitions (Supplementary Table 14), as shown by the density of each cluster and the ordination distances, respectively (Fig. 6a). Overall, neutral partitions accounted for the majority of fungal OTUs for plastic (85%) and soil (88%) (Fig. 6b). However, the fitting proportions of abundance differed markedly between compartments, with neutral partitions accounting for 93% of soil reads but only 60% of MP reads (Fig. 6b), indicating the prevalence of dominant taxa within the non-neutral plastisphere. Moreover, the plastisphere above-partition harboured the most even (ANOVA p < 0.05) and second most diverse (p < 0.05) community among all six partitions (Fig. 6c, Supplementary Tables 17 and 18), suggesting the existence of a diverse, relatively equitable plastiphilic mycobiome.
Communities were pooled and OTUs were then divided into separate sample partitions based on their consistence with (black), deviation above (cyan) or below (dark red) the neutral prediction. Colour coding is consistent for all panels. a NMDS ordination based on Hellinger distance resemblance matrices of square-root transformed OTU abundances. Significant variation between partitions was tested by PERMANOVA (p-values) and ANOSIM (global R). NMDS ordination stress values included. b Fitting proportions of fungal OTU richness and abundance by the Sloan NCM. c Inter-partitional alpha diversity metrics according to compartment, including species richness, Shannon diversity and Pielou’s species evenness estimated for plastic and soil. Box plots indicate median (middle line), 25th, 75th percentile (box) and 5th and 95th percentile (whiskers). d Distinct fitting proportions of richness (OTU number) and e abundance (sequence number) of plastisphere and soil fungal classes, including only those identified classes comprising at least 0.1% of sequences.
Network properties and fungal keystone taxa
To explore the fungal relationships in the soil and in the plastisphere, co-occurrence network analysis based on the most prevalent OTUs (present in >10% of respective samples) was performed using only significant and strong Spearman correlations (r > 0.6 or <−0.6, p < 0.01) (Supplementary Data 4). The two networks showed considerable differences in their structure and topological properties (Fig. 7a and Supplementary Table 19). The soil network consisted of 259 nodes and 498 edges (1 negative), with a modularity of 0.77, an average path length of 5.82 and an average degree of 3.85. The plastisphere network was considerably smaller, with 97 nodes and 166 exclusively positive edges, and had lower values for modularity (0.63), path length (3.73) and degree (3.42). However, the plastisphere network had the higher clustering coefficient (0.41) and harboured around twice the proportion of pathogens compared to the soil network with an overall markedly higher average virulence score (Fig. 7a, Supplementary Table 19). To investigate the keystone taxa behind the assembly of plastisphere and soil MC, the fungi with the highest degree and betweenness centrality (BC) were identified for each network (Supplementary Table 20)48,49. Among the two networks, the OTUs functioning as central hubs (highest degree), gatekeepers (highest BC), and thus also keystones were completely different. The keystone taxa of the plastisphere were mostly pathogenic, mainly plastiphilic Sordariomycetes and Dothideomycetes, with varying frequency and relatively high relative abundance, some of which are also known plastic biodegraders, such as Fusarium and Phoma (Fig. 7b–d, Supplementary Table 20).
Networks only include OTUs with >10% occurrence frequency across the soil or plastisphere MC, respectively. Nodes represent OTUs and edges represent significant (p < 0.01) and strong (p > 0.6 or <−0.6) Spearman correlations between nodes, where grey = positive and orange = negative correlations. a Comparison between soil (left) and plastisphere network (right). Node colour indicates guild affiliation and node size is proportional to the number of GVTs of each fungal OTU at the genus level (i.e., virulence score). b Correlations between plastiphilic and non-plastiphilic OTUs in the plastisphere. Node colour encodes the respective partition of each OTU, with above-partition OTUs (cyan) representing plastiphilic fungi. Node size is proportional to the number of connections of a node (i.e., degree). c Co-occurrences of putative plastic biodegraders in the plastisphere. Node colour indicates plastic biodegradability, where PBD = plastic biodegraders, non = non-degraders and n. a. = unassigned fungi without corresponding annotation. Node size is proportional to the number of shortest paths between any pair of nodes passing through an individual node (i.e., betweenness centrality). d Relationships between plastisphere fungi of distinct classes. Node colour indicates class affiliation, where Trem. = Tremellomycetes, Euro. = Eurotiomycetes, Sord. = Sordariomycetes, Doth. = Dothideomycetes, other = all other assigned classes and unid. = unidentified classes. Node size is proportional to the relative occurrence frequency of each OTU across the entire plastisphere MC.
Plastiphily among ubiquitous human pathogens
To test for differential selection of fungal pathogens by soil and MP, different taxonomic and ecological groups of pathogenic fungi were selected (Supplementary Table 21, Supplementary Data 2). These included general ‘pathogens’ and ‘opportunists’ and their subgroups of Dothideomycetes and Tremellomycetes, as these classes were not only relatively the most selected by MP (Figs. 6d, 8a) but also characteristic of the plastisphere10. Furthermore, we classified genera shared between compartments that have a ‘cosmopolitan’ distribution, ‘high’ or ‘extremely high’ adaptability and are ‘opportunistic’ as the ‘CAO’ (cosmopolitan adaptable opportunists) group (21 genera) and the genera Aspergillus, Candida, and Cryptococcus additionally as “hidden killers”19, to individually assess selection effects associated with globally relevant and the most important fungal pathogens for humans, respectively. Selection was quantified using within-group selection (WGS), i.e., the proportion of above OTUs in a group to the total OTUs in that group, across-metacommunity selection (AMS), i.e., the proportion of above OTUs in a group to the above OTUs in the entire MC, and a selection index (SI) calculated from the product of the two. In addition, we determined the times average selection (TAS) of groups from the ratio of the groups’ WGS and the average selection in the respective MC, i.e., the proportion of the above OTUs in the MC (Supplementary Tables 21 and 22).
a Ratios of relative abundance, relative richness, and selection of different taxonomic and ecological groups of pathogenic fungi between MP and soil. The first 21 groups listed represent CAO (cosmopolitan adaptable opportunist) genera separated according to growth form into ‘filamentous fungi’ and ‘yeasts’. The ‘ecological groups’ include pathogens, opportunists, taxonomic subgroups of them, the total CAO group and ‘hidden killers’ comprising the genera Aspergillus, Candida, and Cryptococcus. Here, doth. = dothideomycetous, trem. = tremellomycetous, path. = pathogens and opp. = opportunists. Blue colour indicates MP:soil ratio >1, red colour indicates ratio <1 and white colour indicates ratio = 1. Darker colours indicate more extreme ratio values. Striped boxes indicate exclusive selection of the group on MP (blue) or soil (red). Selection was assessed using the selection index (SI) introduced in this study, which is calculated from the product of within-group selection (WGS) and across-metacommunity selection (AMS). b Correlation between WGS and AMS for pathogenic groups, where P = pathogens, O = opportunists, DP = dothideomycetous pathogens, DO = dothideomycetous opportunists, TP = tremellomycetous pathogens, TO = tremellomycetous opportunists and HK = hidden killers. Bubble size corresponds to SI value. c Correlation between WGS and AMS for CAO genera. Genera were abbreviated with the first two letters. Bubble size corresponds to SI value. d Correlation heatmap showing Spearman correlations between different combinations of traits. PLS = pathogenic lifestyle, OP = opportunism, ET = extremotolerance, OT = oligotrophy, ML = melanisation, PM = polymorphism and VIR = virulence score. Darker colours indicate stronger positive correlations. e Correlation between WGS and mean virulence score for pathogenic groups. Correlation coefficient (r) and significance (p) value included. Shaded areas represent the 95% confidence intervals.
Over 25% of epiplastic pathogens were in the above-partition (TAS of 2.4), accounting for just under half of all plastiphilic fungi, and showed selection 3 to 22 times stronger than any other guild according to the SI (Supplementary Tables 21–23). In contrast, only 9% of all soil pathogens were in the above-partition (AMS of 13%), which showed slightly below-average selection (TAS < 1), and a 2.5-fold lower SI than saprotrophs. WGS and TAS of all eight pathogenic groups investigated were significantly higher on MP than in the soil (p < 0.001) (Fig. 8b, Supplementary Table 22). The correlation between AMS and WGS was highly significant and almost perfectly inverse for the pathogenic groups on MP (r = −0.95, p < 0.01) but positive in soil (r = 0.76, p < 0.05). According to the SI values, fungal species belonging to opportunists, CAO, hidden killers, dothideomycetous and tremellomycetous opportunists were selected 6, 9, 18, 37 and 275 times more strongly on MP than in soil, respectively (Fig. 8a, Supplementary Table 22). Overall, 19 out of 21 CAO genera were relatively more speciose on MP (compared to 2 in soil), while species of 13 out of 21 CAO genera were more strongly or exclusively selected by MP (compared to 5 in soil) (Fig. 8a). The positive correlation between WGS and AMS was equally strong and significant (r = 0.86, p < 0.01) for CAO genera on plastic and in the soil, averaging 27% and 1.6% on MP, and 9% and 0.4% in soil, respectively (Fig. 8c, Supplementary Tables 22 and 23). Among these genera, Curvularia (about 100 times higher than in the soil), Hannaella and Cryptococcus (both selected by MP only) had the highest SI in the plastisphere (Fig. 8a, c, Supplementary Table 22).
Link between plastiphily and GVTs
Mean GVP was significantly (p < 0.01) positively correlated with opportunism (r = 0.65 and 0.59) and pathogenic lifestyle (r = 0.4 and 0.39) in both MCs (Fig. 8d). Opportunism and pathogenic lifestyle were significantly positively correlated with each individual GVT in both compartments, with oligotrophy and extremotolerance each having the strongest correlations (Fig. 8d, Supplementary Table 24). Overall, extremotolerance and oligotrophy were the two GVTs correlated most strongly with each other (r = 0.73 and 0.65, p < 0.01). Pathogenic groups had systematically and significantly higher GVPs than their non-pathogenic or non-opportunistic counterparts for both compartments (Supplementary Fig. 1, Supplementary Table 26). The GVP of the pathogenic groups were thereby higher on MP than in soil, although not significantly (Supplementary Tables 22, 25, 26). We found a strong and significant positive correlation between WGS and GVP for pathogenic groups on MP (r = 0.75, p < 0.05), which was not the case in the soil (r = −0.40, p > 0.05) (Fig. 8e).
Discussion
The methodological adequacy of the molecular analyses and the strict compliance of the research data management with the FAIR + + guiding principles50 were approved in the previous study10. The variation between soil and MP might be more pronounced than found here, as a complete separation of the compartments is technically not possible at this point51, leading to cross-contamination that artificially increases the degree of mycobiome nestedness. Describing ecological phenomena from community barcoding data requires derivation of functional information from taxonomic profiles52. Fungal metabarcoding studies, however, suffer from a general scarcity of appropriate databases, insufficient taxon coverage, and a limited number of investigated traits52,53. For a comprehensive and accurate assignment of trait data in our study, we collected and harmonised records and information from five relevant databases and hundreds of scientific publications, annotating nine traits for all identified fungal genera at an appropriate granularity (Supplementary Data 1). Fungal pathogenicity in humans is largely opportunistic, so fungal virulence is strongly linked to their environmental (out-of-host) ecology28. Consequently, classical virulence traits for human pathogens have not been identified35,36,37. Here, we used a set of well-described, commonly observed, and multi-use symplesiomorphic autecological28,37 and physiological traits33,54,55 that contribute to the virulence of a wide variety of fungi (Supplementary Data 1), and thus qualify as ‘generic virulence traits’ (GVTs) to approach the generic virulence potential (GVP) of fungi (Fig. 1b). Overall, we therefore argue that our meticulous implementation, fair documentation, and comprehensive contextualisation allow a clear interpretation of the obtained results in a synecological context and thus inferences and predictions of real-world phenomena.
Our analyses revealed significant compositional, ecological, and phylogenetic differences between soil and plastisphere MCs (Figs. 2–4), indicating the emergence of idiosyncratic mycobiomes on MP. Previous studies have also reported structural differences between fungal communities on MP and in the surrounding medium56,57, but primarily focused on high-abundance taxa10. Comparing different abundance fractions, we found clear differences in alpha (Fig. 2d) and beta diversity (Fig. 3a) between epiplastic and soil fungal communities in each fraction, including the so-called “rare biosphere”58 (ART and CRT). Plastic appears to favour the presence of Dothideomycetes and Tremellomycetes, as well as fungal pathogens in general (Fig. 2b, c, f, g). Saprotrophic fungi dominate fungal biodiversity outside symbioses or on non-living substrates59,60. The observed convergence of saprotroph and pathogen diversity on plastic (Fig. 2c) makes it possibly the only fungal habitat recognised to date with such a pattern of pathogen (co-)dominance, reminiscent of dysbiotic mycobiomes during disease61,62. Among the plastic colonisers, we detected many plastic-specific fungi that were not detected in the surrounding soil (Fig. 4). Unlike the rare biosphere inhabiting plastic-specific bacteria from marine MP described by Scales et al.63, the plastic-specific fungi here were among the conditionally and always abundant taxa (Fig. 2h), suggesting a high ecological relevance for the plastisphere biome. We hypothesise that these specific fungi were alien species introduced from other habitats by windblown MP or singular disposal events. The discovery of specificity and cryptic diversity in the plastisphere (Fig. 2f, g) demonstrates the potential of MPs as a reservoir of exclusive and unexplored fungi, and a vector for alien species, exposing how little we know about the origin, scope, and complexity of the fungal-plastic reciprocity.
The soil plastisphere in its entirety represents a fragmental environmental compartment embedded in the three-dimensional continuum of the soil matrix. This habitat fragmentation implies community nestedness64, while the spatial proximity suggests community coalescence between the two compartments65. We found more than 75% of epiplastic OTUs also in the soil, where they accounted for slightly more than one-third of the MC (Fig. 2e), indicating that the plastisphere is largely nested in the soil mycobiome. This finding is consistent with the ‘nested subset theory’ according to which communities with lower species richness in habitat patches are nested in species-rich communities of larger habitats66. In terms of dominance structure of taxonomic and ecological groups, the shared MC clearly represented an ‘intermediate state’ between specific MCs, as shown by the transition in relative abundance of Dothideomycetes and pathogens (Fig. 2f, g). We interpret these patterns as indicative of intercompartmental coalescence. While the observed mycobiome nestedness indicates strong community coalescence in form of migration of soil fungi into the plastisphere, the distinct dominance patterns of pathogens imply a dynamic exchange of pathogens emanating from the plastisphere (Fig. 2c, g). Falling leaves feed global fungal coalescence processes by introducing plant-associated species into the soil in a cyclic equilibrium, thereby influencing planetary nutrient cycling65,67. Assuming a global plastisphere mycobiome, deposited MP could drive the coalescence of fungal communities on a similar scale but with unpredictable dynamics, due to its ubiquity, distinctiveness, persistence, and sheer numbers, especially in open areas such as savannas, deserts, oceans, and parts of the anthroposphere. The coalescence of pathogens may be a consequence of plastic pollution that has the potential to alter the proportion, identity, and diversity of fungal pathogens in virtually any environmental compartment.
The NCM46 successfully predicted most fungal distributions across local communities in both compartments (Fig. 5), following the basic trend of abundant taxa being widespread in the MC68, consistent with neutral theory38,39. Passive dispersal and ecological drift alone explained a considerable proportion of mycobiome structure on plastic and in soil. This finding is consistent with the results of Sun et al.69 and Zhu et al.16, who, using the same model, found that epiplastic bacteriomes in aquatic and terrestrial systems, respectively, were mainly formed by neutral processes. However, we found several taxa that deviated from neutral predictions (Fig. 6) and are thus likely to be under environmental selection pressure (positive/negative)47, which was also the case for certain soil bacteria being impacted by the plastic type11. Therefore, we partially reject our third hypothesis of balanced complementary interplay by remarking that plastisphere mycobiomes are strongly influenced by stochasticity. In general, taxa that occurred less frequently than predicted (below-partition) are likely to be selected against by the respective compartment47,70. In contrast, taxa that were more widespread than expected (above-partition) are likely to be selected by the compartment47,70. Specifically, epiplastic taxa in the above-partition are likely to show plastiphily, i.e., adaptation to the plastic habitat with correspondingly increased ecological fitness. Plastiphilic fungi can thus systematically colonise plastics, form colonies, and successfully reproduce in the plastisphere10. Importantly, the non-neutral partitions of plastic and soil differed significantly in composition and ecology (Fig. 6), proving compartment-specific selection. Given the intrinsically similar environmental conditions, the differential selection must be due to substrate particularities as previously proposed11,16, with MP artificially widening the fungal niche. Furthermore, plastiphilic fungi formed the most equitable and second most diverse partition (Fig. 6c), suggesting that plastiphily plays an important role in the emergence of fungal community diversity in terrestrial systems.
Our co-occurrence network analysis revealed numerous distinct correlations among fungi in soil and on MP (Fig. 7a, Supplementary Table 19), demonstrating the importance of selective forces in the assembly of mycobiomes and implying the emergence of fundamentally different interaction landscapes on MP. The lower overall size and average path length of the plastisphere network indicates a compact network with ‘small world’ properties, implying rapid material and information transport among co-occurring species71,72, while the higher clustering coefficient suggests a more complex network with stronger interactions than in soil48,49. Overall, however, the degree to which these correlations represent real-world organismic interactions remains to be investigated. Interestingly, plastiphilic fungi were overrepresented in an exclusively positive network (Fig. 7b), suggesting that well-adapted, presumably active fungi10 interact synergistically in assembling the plastisphere mycobiome. Conversely, the higher geodesic distance and number of neutrally assembled taxa in the soil network could mean that the soil fungi are mainly dormant spores from a historic contingent. However, both networks showed relatively high modularity and average degree, indicating distinct functional groups, and realised niches73 in both compartments. As expected, both networks contained predominantly positive edges (Fig. 7a), which is consistent with the stress gradient hypothesis44,74, according to which facilitation and niche sharing prevail over competition in harsher environments such as the investigated low-water, oligotrophic, and UV-irradiated ferralsols75. Positive networks, however, are unstable due to possible fitness interdependences, resulting in rapid regime shifts and low resilience of both MCs when perturbed74. While the keystone taxa in soil were neutrally assembled saprotrophic or unidentified fungi, plastisphere keystones were abundant, primarily plastiphilic fungal pathogens with the capability to biodegrade plastics (Fig. 7b, c, Supplementary Table 20). Apparently, metabolically versatile pathogens such as Phoma and Fusarium76,77,78 drive plastisphere assembly based on trophic interactions as ‘keystone plastic degraders’. By co-opting their repertoire of degradative traits76,78 to modify or degrade polymer components (i.e., ‘exaptative plastibiome’), these pathogens may facilitate co-colonisation, co-degradation and co-metabolisation of MP by other members of the community, as previously described for other substrates79,80, ultimately enabling plasticlastic commensalism and syntrophy. It has been shown that some microbes can completely degrade, assimilate, and mineralise at least certain plastics under experimental conditions81,82,83, a feat that is yet to be observed in nature. The co-selection of fungi in the plastisphere based on the plastic biodegradation capability may indicate that fungal metabolism has already become better adapted to plastic polymers. Given MP in the deep sea, atmosphere, and Arctic ice84, the presumed evolving trophic plastiphily of fungi offers a vague but real prospect for future bioremediation of plastic if we ever stop the emission of plastic waste into the environment. As hypothesised, the indices used revealed that MP selects pathogens more strongly than members of any other guild (Fig. 8). Moreover, selection of pathogens was much stronger in the plastisphere than in the soil, where it did not even exceed MC average (Fig. 8b, c). Remarkably, this relation was observed for all ecological and phylogenetic groups of pathogens considered (Fig. 8a–c, Supplementary Table 22). Plastic is therefore arguably the more suitable habitat for fungal pathogens in the investigated systems compared to soil. Among the plastiphilic fungi, we found numerous opportunistic human pathogens with cosmopolitan distribution and high ecological adaptability (the CAO group), including allergenic, toxigenic, and pathogenic black fungi like Alternaria, Bipolaris, Curvularia and Epicoccum (Supplementary Data 1). Melanised opportunists, primarily Dothideomycetes, also dominated the plastic-specific mycobiome (Fig. 4). These versatile pathogens infect crops24, livestock and domestic animals85, cause building damage86 and a spectrum of opportunistic phaeohyphomycoses in humans85. Several polymorphic cryptococcal yeasts, such as Hannaella and Papiliotrema, which are known to cause a range of potentially fatal cryptococcoses87, were in fact only selected by plastic. In addition, polyextremotolerant black yeasts of the genera Exophiala and Knufia88 were exclusive to plastic here. These pleomorphic fungi are common in indoor plastic niches like dishwashers32 and water storage tanks31 and able to cause systemic infections in humans31,88. Alarmingly, we found some of the most important human pathogens on plastic waste, namely Candida, Cryptococcus and Aspergillus fumigatus, which are responsible for millions of life-threatening respiratory and cerebral infections each year19. The observed exclusive selection of some of these health-relevant human pathogens19,53 by MP (Fig. 8), however, represents an ecological paradox. According to competitive exclusion theory89, selection of these fungi (e.g., Cryptococcus species) outside their primary niches, e.g., bird faeces or the human body19,35,53, is highly unlikely as they will be outcompeted and eliminated by other better adapted species in the environment89. Therefore, it could be argued that the plastisphere resembles their ancestral habitat so closely that it represents a surrogate primary niche for these pathogens outside the host where they can thrive and evolve, presumably convergently. Considering that most plastiphilic fungal pathogens are not only cosmopolitan but also some of the most abundant soil fungi worldwide68, we argue that selection of fungal pathogens may become a global emergent consequence of plastic pollution.
According to the ‘exaptation theory’90,91, virulence traits of opportunistic fungi are also present in non-opportunistic species, as they have originally evolved for different purposes, i.e., environmental stress tolerance, and thus inherently exhibit ‘dual-use’. Our meta-analysis supports this hypothesis, as we found that all GVTs are present in both opportunistic and non-opportunistic fungi (Supplementary Data 1). Pathogenic and opportunistic groups, however, showed systematically and significantly higher GVP than their non-pathogenic or non-opportunistic counterparts in both compartments, with GVPs of epiplastic pathogens being tentatively higher than those of soil equivalents (Supplementary Fig. 1, Supplementary Table 22). Although it is axiomatic that pathogens are more virulent than non-pathogens, no inferences about in vivo pathogenicity or virulence can be made from the presented data. GVTs and the GVP are linked to both pathogenic lifestyle and opportunism, with oligotrophy and extremotolerance showing the strongest association (Fig. 8d). This is consistent with previous genomic studies that found a strong link between extremotolerance and opportunistic fungal pathogenicity28. Our results show that plastiphily correlates with GVP in fungi (Fig. 8e). Therefore, adaptation to the plastisphere appears to be linked to the number or coincidence of fungal GVTs as proposed in our plastiphilic virulence traits hypothesis. The observation that GVTs are strongly linked to both opportunism and to plastiphily supports the hypothesis that phenotypes relevant to fungal pathogenicity are shaped by plastisphere selection pressures. This corresponds to the ‘accidental virulence’ hypothesis91 and the concept of the ‘environmental virulence school’36 according to which exposure to selection pressures in the environment outside the host most likely result in (pre)adaptations with benefits for pathogenesis. Although caution is required when inferring phenomena from patterns in ecology52,53, it is evident that all GVTs considered are associated with both opportunistic pathogenicity and affinity for plastics, which has even been shown using several pathogens found in this study to be selected by or exclusive to plastic28,32,34. Therefore, the plastisphere must be considered a ‘school for virulence’, where plastiphilic fungi can ‘exercise’ their exaptations toward higher virulence potentials (‘plastiphilic virulence’) (Fig. 9).
a Extract of the terrestrial microplastic cycle according to the global odyssey of plastic pollution137 extended by the fungal-plastic reciprocity as observed in this study. Along the cycle, MP is permanently habitat, interaction landscape and vector of associated fungi and therefore important driver of fungal ecology, biogeography, and epidemiology. b Origin and ecology of epiplastic mycobiomes in the soil environment according to this study and Gkoutselis et al.10. Each symbol type represents a distinct fungal species. Plastisphere mycobiomes on deposited microplastics in soils, consisting of non-pathogenic (black symbols), and several pathogenic species (red symbols), assemble predominantly via dispersal and drift (neutral processes) from the MC (symbols without frame = neutrally assembled fungi). Several, mainly pathogenic fungi, however, are selected by MP and are therefore plastiphilic (symbols with cyan frame) or appear to co-select each other into the plastisphere through biotic interactions (systematically co-occurring species, especially keystone taxa) (deterministic processes). The resultant epiplastic communities show a high diversity (pathogen convergence) and selective accumulation of certain pathogens. Despite compositional and ecological idiosyncrasies of the different mycobiomes, with diverse plastic- and soil-specific fungal taxa, a large proportion of the plastic-associated species overlap with the planktonic ones in the soil (nestedness), indicating intercompartmental coalescence. c ‘The plastisphere virulence school’. Fungal plastiphily is directly linked to the incidence of symplesiomorphic traits in fungi that, inter alia, (co)determine virulence in humans, such as melanisation, polymorphism, extremotolerance and oligotrophy (exaptations). Adapting to the plastisphere will therefore result in the ‘fine-tuning’ of these and other relevant traits through natural selection, eventually increasing the generic virulence potentials of epiplastic pathogens in humans (plastiphilic virulence). According to the ‘host invasion trait congruency’ theorem, exposure of fungi to the plastisphere micro-niches will convergently favour evolutionary (pre)adaptations to all four criteria relevant for the emergence of opportunistic pathogenicity in humans (1–4)35. Ecological phenomena found in this study and in Gkoutselis et al.10 are highlighted in bold. Many other empirically grounded mechanisms have been adapted and proposed. (↑) = increase, (+) = acquisition/emergence and CH = hydrocarbons. Created with biorender.com.
In summary, our results support the notion that plastic pollution is linked to the occurrence of fungal infections by providing reservoirs, vectors and a selective environment for pathogenic fungi in terrestrial systems, which can favour all four criteria relevant for the emergence of fungal pathogenicity35 (Fig. 9c). The assembly of a diversity of locally concentrated fungal pathogens (pathogen convergence) on MP, together with selective accumulation10 and coalescence effects will certainly increase the number and density of infectious agents (propagule size), while particle dispersibility, invasiveness and persistence could increase the range and number of transmission events (propagule number), altogether enhancing pathogen pressure92. This in turn would, according to the principles of biological invasions93, increase both accessibility and colonisation success of adherent pathogens (criterium 1). In oligotrophic soil environments such as savannas, deserts, and Arctic dry valleys, where plastic debris might represent ‘islands of carbon’, plastiphilic fungi may evolve to overcome longer periods of nutrient deprivation by utilising trace additives on the particle surface or even modify metabolic pathways to exploit the polymer itself. Potent hydrocarbonoclastic species could evolve into potent degrader fungi capable of decomposing epithelia of the human host94,95 (criterium 2). In the very same environments, MP in topsoil, especially in equatorial regions, are exposed to high UV radiation and are becoming more attractive to fungi as carbon residues become more accessible through polymer weathering96. Melanised, plastic degraders would thrive under those conditions and quickly evolve to tolerate high temperatures. Such thermotolerant and plastiphilic fungi may overcome the thermal repulsion imposed by mammal endothermy and homeothermy35,97, and thus enhance the ability to thrive inside the human body (criterium 3). Finally, exposure to immense oxidative stress during polymer weathering and biodegradation98 may select for plastiphiles to resist reactive oxygen species deployed by immune cells99, arming them to resist the human immune system (criterium 4). Plastisphere life may further promote a pathogenic phenotype by fine-tuning thigmo responses36,100, disrupting fungistasis101, favouring biofilm formation10,102, and increasing rates of mutation, hybridisation, and gene transfer53,103. Thus, plastiphilic fungal pathogens would become increasingly virulent to humans, while even previously non-pathogenic species could emerge as ‘accidental’ opportunists91 once they acquired a critical repertoire of GVTs through their plastisphere passage (Fig. 9c). As global plastic pollution coincides with global warming24,104, habitat fragmentation68, desertification104 and an unprecedented number of immunosuppressed human individuals19,21, plastiphilic fungal virulence is likely to play an increasing role in global human health.
Given the strong link between virulence potential, opportunistic pathogenicity, and the selection of ubiquitous, extremely abundant, and significant human pathogens by MP, plastic pollution may contribute substantially to the occurrence of fungal infections worldwide and might have been doing so for a long time. Therefore, there is an urgent need to classify microplastics as a global health factor. Taken together, our findings provide first insights into the origins of epiplastic mycobiomes and the mechanisms behind the emergence of fungal plastiphily and demonstrate the important role of plastic debris in the ecology and evolution of pathogenic fungi. This new knowledge is not only crucial for developing strategies to combat both global burdens—fungal infections and plastic pollution—but also forms the basis for a deeper understanding of the biodiversity, ecology, functionality, and adaptability of plastic-colonising soil microbes in an increasingly plasticised world.
Materials and methods
Sampling and selective subsampling
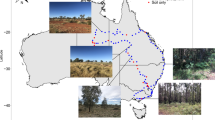
The environmental samples were identical to those used in the previous study by Gkoutselis et al.10. Briefly, the study design involved the selection of five different sites with high levels of plastic contamination and human activities within the town limits of Siaya (Siaya County, Western Kenya) followed by the random collection of five environmental samples from the contaminated topsoil of each plot (total n = 25). The sites included two open landfills, a roadside, a marketplace, and a courtyard. Further details on the sampling procedure, the contextual data collection and the site descriptions are provided in Gkoutselis et al.10. All relevant data of the sampling campaign have been deposited in DiversityCollection105 at the Staatliche Naturwissenschaftliche Sammlungen Bayerns (SNSB), Germany, in accordance with the FAIR + + guiding principles for scientific data management50. Here, the subsamples from the previous study10 were used and further subsamples were generated and included by multistage subsampling with intermittent homogenisation. Briefly, about 5 g of plastic-contaminated soil was collected from the remaining environmental samples (about 10 g) under sterile conditions and homogenised (no global homogenisation to preserve the heterogeneous compartmentalisation of the initial sampling campaign). In accordance with the methodology used previously10, 100-mg soil and plastic subsamples were generated here to ensure comparability. In this way, the subsample pool was expanded from n = 50 (25 + 25) to n = 100 (50 + 50). The size range of the plastic debris was determined by examining 100 randomly selected particles per site and sample for each study and ranged from 1 to 30 mm (combined), with the majority (81%) of particles <5 mm. Due to this high share of microplastics in the entire mix, all plastic subsamples were considered as microplastic-dominated2. As before, MPs were combined into subsamples regardless of shape and polymer type. All subsamples were frozen at −20 °C until further processing.
DNA extraction, PCR and Illumina sequencing
All steps were conducted as previously described10. Briefly, plastic fragments were visually identified and attaching soil particles were meticulously separated from plastic surfaces forming the respective soil and plastic compartment. Total metagenomic DNA from soil and plastic subsamples was extracted using the NucleoSpin® Soil kit (MACHEREY NAGEL, Düren, Germany) according to the manufacturer’s instructions after initial adjustments. NanoDrop® ND-1000 UV-Vis spectrophotometer (Thermo Fisher Scientific, Waltham, USA) and 0.8% agarose gel electrophoresis were used to determine the quality and concentration of the isolated DNA. ITS1 and ITS2 regions were amplified by nested PCR based on ITS1F and ITS4 primers, each containing a unique combination of TAG and INDEX sequences, part of the Illumina sequencing primer, and P5 and P7 adapters, respectively, to generate dual-indexed amplicons. An enzymatic purification step (ExoSAP digestion) was interposed to remove or deactivate excess reactants prior to PCR2. Cycling and purification were performed in a thermocycler (BioRad, Hercules, USA). The combination of TAG (PCR1) and INDEX (PCR2) sequences allowed the generation of unique sequence patterns for each sample for multiplexing of amplicons. Quality control and pooling were performed as previously described10. Gel bands were visualised using Gel Doc™ EZ gel imaging system (BioRad, Hercules, USA). Amplicons were paired-end sequenced on an Illumina MiSeq 3000 sequencer (Illumina, Inc., San Diego, USA) at the Department of Genetics, LMU Munich, Germany.
Bioinformatics
Barcodes (TAG sequences) were removed from the obtained raw reads and the reads were demultiplexed using extract_barcodes.py and split_libraries_fastq.py as implemented in QIIME106. The demultiplexed reads were then imported to the QIIME2 pipeline107 and remaining adapter and primer sequences were trimmed, according to the applied sequences motives, using the cutadapt tool108. Quality filtering, including removal of chimeric sequences and dereplication of exact sequence variants was performed using the DADA2 plugin under QIIME2107 based on a maximum expected error of 2 and a minimum fold abundance of parent sequences of 1 over potential chimeras. OTUs were clustered from the amplicon sequence variants (ASVs) from the DADA2 output using VSEARCH and a threshold of 97% sequence similarity. Taxonomy assignment was performed using the feature classifier classify-sklearn command implemented in QIIME2 and a Naïve Bayesian classifier previously trained on the UNITE (v.8.0) dynamic database109 with a chunk size of 20,000. OTU representative sequences were then taxonomically classified using a threshold of 97% sequence similarity and a minimum confidence score of 70%. Samples containing less than 1000 sequences110 and singletons70 were removed from the dataset. The processed sequencing results of the two separate subsampling events of the original environmental samples were merged at the level of the de-noised and dereplicated ASV tables, resulting from the DADA2 analysis, using the ‘QIIME feature-tables merge’ command for the ensuing OTU clustering.
Neutral community model
To determine the processes driving mycobiome assembly, we implemented the neutral community model developed by Sloan et al.46, which is an adaptation of neutral theory adjusted to large microbial populations. The model predicts the relationship between frequency with which taxa occur in a set of local communities (fungal communities of individual plastic or soil subsamples) and their abundance across the wider metacommunity (fungal communities of all plastic or soil subsamples). In principle, the NCM predicts that taxa that are abundant in the MC will be widespread as they are more likely to be randomly dispersed among different sampling sites, whereas rare taxa are more likely to be lost at different sites due to ecological drift (i.e., stochastic loss and replacement of individuals). Unlike many other contemporary neutral models, such as the unified neutral theory of biodiversity38, the NCM used here does not incorporate the process of speciation. Here, dispersal among communities is estimated by Nm, which determines the correlation between observed occurrence frequency of fungi (proportion of local communities comprising each OTU) and their abundance in the MC (mean relative abundance across all local communities), with N describing the MC size and m being the migration rate. The migration rate represents the probability that a random loss of an individual in a local community will be replaced by dispersal from the MC, as opposed to reproduction within the local community. The parameter R2 indicates the overall fit to the neutral model46, assessed by comparing the sum of squares of residuals with the total sum of squares47. To test whether incorporating ecological drift and dispersal limitation improve the fit of the model beyond just random sampling from the MC46, we compared the NCM fit with the fit of a binomial distribution model. To compare the fit of the neutral and binomial model, the Akaike information criterion (AIC) was calculated and compared for each model16,47. Fitting of the free parameter m was performed in R (R 4.2.0)111 using non-linear least-squares fitting from the minpack.lm package112. Wilson score interval from the HMisc package in R113 was used to calculate the binomial 95% confidence intervals. Computation of the AIC was also conducted in R (R 4.2.0)111. Calculation of 95% confidence intervals around all fitting statistics was done by bootstrapping based on 1000 replications (Supplementary Data 2). We used the R code of Burns et al.47 for all analyses. The NCM was applied separately to soil and plastic as well as to the combined dataset on rarefied OTU matrices. To analyse deviations from the neutral predictions, OTUs from each dataset were separated into three partitions depending on whether they occurred more frequently than (above-partition), less frequently than (below-partition) or as frequently as predicted by the NCM (neutral-partition) based on 95% confidence intervals. Ecologically, fungi above or below prediction are therefore considered as being actively selected for (selection) or against (exclusion) by the target compartment, respectively (deterministic processes), while taxa within prediction are considered randomly assembled (neutral processes). Fungi in the above-partition of the plastisphere were defined as ‘plastiphilic’ or ‘plastiphiles’. Mean relative abundance, as well as observed and predicted occurrence frequency of each OTU of each compartment can be found in Supplementary Data 3.
Meta-analysis
Only identified genera in the combined rarefied dataset were subjected to meta-analysis. For all other phylotypes, ‘unidentified’ was set as default for any annotation or assignment. For inferences about physiology, ecology, biogeography, and relevance to humans, fungi were annotated with four relevant traits, namely growth form, distribution, adaptability, and opportunism (Supplementary Data 1). Growth form was annotated using the comprehensive FungalTraits database114, with the categories ‘filamentous’, ‘yeast-like’ (= yeasts and dimorphic yeasts) and ‘zoosporic’ (= any zoosporic group). Distribution was determined using the GlobalFungi database115, with genera classified according to the number of continents on which corresponding specimens have been recorded into ‘endemic’ (1), ‘widespread’ (2–5) and ‘cosmopolitan’ (>5). Adaptability was categorised according to the number of possible lifestyles (FungalTraits)114 or guilds (FUNGuild)116 into ‘low’ (1 or 2), ‘high’ (3 or 4) and ‘extremely high’ (>4). Finally, opportunism (= opportunistic human pathogenicity10,28) was determined by querying FungalTraits114, FUNGuild116, Encyclopedia of Life (EOL)117, Index Fungorum (http://www.indexfungorum.org) and all results (up to a maximum of 100) of a PubMed search using the respective genus name. Any relevant record of a specimen as an aetiological agent for an infection or disease in humans qualified the corresponding genus as ‘opportunistic’, otherwise it was considered ‘non-opportunistic’.Generally, in the absence of entries for any trait in the relevant database(s), the appropriate literature was consulted instead. To further elaborate on the functionalities, epidemiology, and relevance to human health, recovered fungi were assigned to three distinct ecological groups, namely guilds, cosmopolitan adaptable opportunists (CAO) and so-called “hidden killers”19 (Supplementary Data 1). Guilds were assigned by consolidating the entries for ‘primary_lifestyle’ in FungalTraits114. As we focused primarily on human pathogenicity in this study, we chose to simplify the assignment by using more generic versions of the classical fungal guilds116, where ‘pathogen’ = any pathogen or parasite, ‘saprotroph’ = any saprotroph, sooty mold, endophyte, or epiphyte, and ‘mutualist’ = any mycorrhizal and lichenised group. Missing records in the database(s) were compensated by relevant literature. CAO were those fungi that showed ‘cosmopolitan’ distribution, ‘high’ or ‘extremely high’ adaptability and ‘opportunistic’ potential according to our trait data annotation. Hidden killers were named and defined according to the study of Brown et al.19 and consisted of only the most significant human pathogens causing opportunistic invasive mycoses, in our study Aspergillus, Candida and Cryptococcus.
To approach fungal virulence potential, we introduced the concept of generic virulence traits (GVTs, Fig. 1b). Data for each GVT, namely melanisation33, polymorphism55, extremotolerance28 and oligotrophy118, were acquired from FungalTraits113, GlobalFungi114, EOL116, and all results (up to a maximum of 100) of a Google Scholar search using the respective genus and trait name (e.g., ‘Curvularia, melanisation’). It is impossible to make a general statement at the genus level regarding the presence of any of these GVTs, as even plesiomorphic and ancestral traits of a genus could have been lost in individual species119. At the same time, it is impossible to annotate such traits for each species identified in a metagenome study due to the lack of appropriate databases, publications, and a generally limited number of observed traits51,52. In our study, each GVT was only assigned to a particular genus when at least three different species of that genus were found to have the trait. The four GVTs were approximated as follows: (1) Melanisation: based on the pigmentation of vegetative structures (e.g., hyphae, yeast cells), fungi were categorised into ‘hyaline’, ‘melanised’ or ‘pigmented’ (unspecified or non-melanin pigmentation). (2) Polymorphism: fungi with the ability to shift from a multicellular hyphal to a unicellular yeast growth form55 were classified as ‘dimorphic’, all other ones were considered ‘monomorphic’. (3) Extremotolerance: fungi were classified as ‘extremotolerant’, if they met any of the several criteria proposed by Gostinčar et al.28, including superficial growth in exposed habitats (e.g., rock-inhabiting fungi) subjected to varying temperatures, dryness, and solar irradiation; lichenised lifestyle (mycobiont in an algal-fungus combination); osmotolerant or osmophilic from dry indoor habitats, desert soils, and similar habitats and halotolerant or halophilic fungi from hypersaline and highly contaminated environments; and psychrotolerant or psychrophilic fungi from deep ocean waters and polar regions and similar habitats. All other fungi were classified as ‘non-extremotolerant’. (4) Oligotrophy: fungi were classified as ‘oligotrophic’, if they met any of the criteria described by Gostinčar et al.118, including growth in extremely nutrient-poor natural habitats and environments, such as rock surface or subsurface, Arctic dry valleys, deserts, and high mountain areas; growth in (oligotrophic) anthropogenic habitats such as monuments, concrete walls, biofilters and other indoor habitats; growth on silicon, metals, glass and on a variety of more or less durable organic surfaces, including plastic materials and similar polymers; or biodegradation of complex, non-natural compounds, such as phenolic hydrocarbons, TNT, radioactive materials, plastics, etc.76,77,78,94. All other fungi were classified as ‘non-oligotrophic’. To achieve a simple parametrisation, we introduced a ‘virulence score’ based on the number of GVTs found in each phylotype (0–4). In a real virulence context, the GVP boasts only a vague informative value, due to the intrinsic methodological imprecision and the complex context-dependency of fungal infections28. Instead, we argue that it constitutes a suitable reductionistic approximation metric for fungal virulence potentials and an ordinal scale for eco-evolutionary projections, as similar presence, identity, and coincidence of the described GVTs translate into similar options regarding ecological niches and positions in the evolutionary landscape120,121.
Abundance fractions
To accurately analyse the architecture of the target mycobiomes, we defined thresholds for relative abundance and frequency of occurrence to classify all OTUs of each MC into six different abundance fractions as previously described70. Therefore, for each compartment, samples were grouped according to sites to form site-specific (bulk) samples. (1) always abundant taxa (AAT) were defined as OTUs with an abundance of ≥1% in all samples; (2) conditionally abundant taxa (CAT) were defined as taxa with an abundance of ≥0.01% in all samples and ≥1% at least one sample; (3) moderate taxa (MT) were defined as OTUs with an abundance between 0.01 and 1% in all samples; (4) conditionally rare and abundant taxa (CRAT) were defined as OTUs ranging from rare (<0.01%) to abundant (≥1%); (5) conditionally rare taxa (CRT) were defined as OTUs with an abundance of >1% in all samples and <0.01% in at least one sample; and (6) always rare taxa (ART) were defined as OTUs with an abundance <0.01% in all samples.
Statistical analysis
Beta diversity analysis using Hellinger distance, ANOSIM and PERMANOVA (Fig. 6a) were performed with the Primer 7 software package and the add-on package PERMANOVA+ (PRIMER-e Ltd, Plymouth, United Kingdom)122. All other statistical analyses and data visualisations were performed in R (R 4.2.0)111, using the vegan123, phyloseq124, ggplot125, microbiome126, reshape2127, ggpubr128, spaa129 and dplyr130 packages. Correlation analyses used in this study were Spearman tests conducted in R (R 4.2.0)111.
Phylogenetic analysis
To demonstrate the evolutionary distance of plastic-specific taxa, a phylogenetic tree was constructed, including only phylotypes from the rarefied dataset identified at least to the genus level (n = 84). Nucleotide sequences were aligned using MUSCLE131 as implemented in MEGA11132. Phylogenetic relationships were inferred using the maximum likelihood method, and Tamura-Nei distance133 as a substitution model and a bootstrap method based on 1000 replications as a phylogeny test. The phylogram was calculated in MEGA11132 and visualised using iTOL version 6134.
Selection indices
Positive selection of fungi was quantified separately for epiplastic and soil fungi based on the results of the NCM using four distinct indices:
-
1.
Within-group selection (WGS), i.e., the proportion of above OTUs in a group to the total OTUs in that group:
$${{WGS}}_{i}=\frac{{n}_{{iap}}}{{n}_{i}}$$ -
2.
Across-metacommunity selection (AMS), i.e., the proportion of above OTUs in a group to the above OTUs in the total MC:
$${{AMS}}_{i}=\frac{{n}_{{iap}}}{{n}_{{jap}}}$$ -
3.
Selection index (SI) calculated from the product of the WGS and AMS of each group:
$${{SI}}_{i}={{WGS}}_{i}\times {{AMS}}_{i}$$ -
4.
Times average selection (TAS), i.e., the ratio of the groups’ WGS and the proportion of above OTUs in the MC:
where n is the number of OTUs, i is the group under examination, j is the corresponding MC and ap refers to the respective above-partition. All calculated values can be found in Supplementary Tables 22 and 23.
Network analysis
For co-occurrence network construction, the most frequent 10% of OTUs of each MC were chosen. The Spearman correlation matrix was calculated using the psych package135 in R. To highlight the most important relationships, only strong positive (ρ > 0.6) or strong negative (ρ < −0.6) and highly significant (p < 0.01) correlations were used for network construction49,72,80. Network analysis and visualisation was conducted in Gephi 0.9.7136. The network properties, clustering coefficient, modularity, average path length, average normalised degree, and positive ratio were included in the analyses (Supplementary Table 19). OTUs with maximum degree and betweenness centrality were considered keystone taxa49,72. Phylotypes in the plastisphere network were annotated with the trait of plastic biodegradability (Supplementary Data 3). Relevant data were obtained from the sources mentioned in Ekanayaka et al.77 and with the phrases used there extended by the respective genus name.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequences from community barcoding are linked under BioProject accession number PRJNA705067. Environmental samples are stored in the collection at the Mycology Department, University of Bayreuth. All other data are directly accessible in the digital repository accessible via https://zenodo.org/records/10012828.
Code availability
R codes used in this study are accessible via https://zenodo.org/records/10012828.
References
Villarrubia-Gómez, P., Cornell, S. E. & Fabres, J. Marine plastic pollution as a planetary boundary threat—the drifting piece in the sustainability puzzle. Mar. Policy 96, 213–220 (2018).
Hartmann, N. B. et al. Are we speaking the same language? Recommendations for a definition and categorization framework for plastic debris. Environ. Sci. Technol. 53, 1039–1047 (2019).
Lu, Y. et al. Uptake and accumulation of polystyrene microplastics in zebrafish (Danio rerio) and toxic effects in liver. Environ. Sci. Technol. 50, 4054–4060 (2016).
Gregory, M. R. Environmental implications of plastic debris in marine settings—entanglement, ingestion, smothering, hangers-on, hitch-hiking and alien invasions. Philos. Trans. R. Soc. B 364, 2013–2025 (2009).
Gall, S. C. & Thompson, R. C. The impact of debris on marine life. Mar. Pollut. Bull. 92, 170–179 (2015).
Trotter, B. et al. Long-term exposure of Daphnia magna to polystyrene microplastic (PS-MP) leads to alterations of the proteome, morphology and life-history. Sci. Total Environ. 795, 148822 (2021).
Sheridan, E. A. et al. Plastic pollution fosters more microbial growth in lakes than natural organic matter. Nat. Commun. 13, 1–9 (2022).
Trotter, B., Ramsperger, A. F. R. M., Raab, P., Haberstroh, J. & Laforsch, C. Plastic waste interferes with chemical communication in aquatic ecosystems. Sci. Rep. 9, 5889 (2019).
Lwanga, E. H. et al. Field evidence for transfer of plastic debris along a terrestrial food chain. Sci. Rep. 7, 1–7 (2017).
Gkoutselis, G. et al. Microplastics accumulate fungal pathogens in terrestrial ecosystems. Sci. Rep. 11, 1–13 (2021).
Rohrbach, S. et al. Microplastic polymer properties as deterministic factors driving terrestrial plastisphere microbiome assembly and succession in the field. Environ. Microbiol. 25, 2681–2697 (2022).
Van Sebille, E. et al. A global inventory of small floating plastic debris. Environ. Res. Lett. 10, 124006 (2015).
Weithmann, N. et al. Organic fertilizer as a vehicle for the entry of microplastic into the environment. Sci. Adv. 4, eaap8060 (2018).
Amaral-Zettler, L. A., Zettler, E. R. & Mincer, T. J. Ecology of the plastisphere. Nat. Rev. Microbiol. 18, 139–151 (2020).
Zettler, E. R., Mincer, T. J. & Amaral-Zettler, L. A. Life in the “plastisphere”: microbial communities on plastic marine debris. Environ. Sci. Technol. 47, 7137–7146 (2013).
Zhu, D., Ma, J., Li, G., Rillig, M. C. & Zhu, Y. G. Soil plastispheres as hotpots of antibiotic resistance genes and potential pathogens. ISME J. 25, 2681–2697 (2021).
Imran, M., Das, K. R. & Naik, M. M. Co-selection of multi-antibiotic resistance in bacterial pathogens in metal and microplastic contaminated environments: an emerging health threat. Chemosphere 215, 846–857 (2019).
Lamb, J. B. et al. Plastic waste associated with disease on coral reefs. Science 359, 460–462 (2018).
Brown, G. D. et al. Hidden killers: human fungal infections. Sci. Transl. Med. 4, 165rv13 (2012).
Fisher, M. C. et al. Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 186–194 (2012).
Fisher, M. C. et al. Threats posed by the fungal kingdom to humans, wildlife, and agriculture. MBio 11, e00449–20 (2020).
Bongomin, F., Gago, S., Oladele, R. O. & Denning, D. W. Global and multi-national prevalence of fungal diseases—estimate precision. J. Fungi 3, 57 (2017).
Kainz, K., Bauer, M. A., Madeo, F. & Carmona-Gutierrez, D. Fungal infections in humans: the silent crisis. Microb. Cell 7, 143 (2020).
Delgado-Baquerizo, M. et al. The proportion of soil-borne pathogens increases with warming at the global scale. Nat. Clim. Change 10, 550–554 (2020).
Barnes, D. K., Galgani, F., Thompson, R. C. & Barlaz, M. Accumulation and fragmentation of plastic debris in global environments. Philos. Trans. R. Soc. B: Biol. Sci 364, 1985–1998 (2009).
Perreault, R. & Laforest-Lapointe, I. Plant-microbe interactions in the phyllosphere: facing challenges of the anthropocene. ISME J. 16, 339–345 (2022).
Seyedmousavi, S. et al. Fungal infections in animals: a patchwork of different situations. Med. Mycol. 56, S165–S187 (2018).
Gostinčar, C. Fungi between extremotolerance and opportunistic pathogenicity on humans. Fungal Div. 93, 195–213 (2018).
Gueidan, C. et al. A rock-inhabiting ancestor for mutualistic and pathogen-rich fungal lineages. Stud. Mycol. 61, 111–119 (2008).
Pappas, P. et al. Invasive candidiasis. Nat. Rev. Dis. Primers 4, 18026 (2018).
Gümral, R. et al. Dishwashers provide a selective extreme environment for human-opportunistic yeast-like fungi. Fungal Div. 76, 1–9 (2016).
Novak Babič, M., Gostinčar, C. & Gunde-Cimerman, N. Microorganisms populating the water-related indoor biome. Appl. Microbiol. Biotechnol. 104, 6443–6462 (2020).
Cordero, R. J. & Casadevall, A. Functions of fungal melanin beyond virulence. Fungal Biol. Rev. 31, 99–112 (2017).
Gostinčar, C., Grube, M. & Gunde-Cimerman, N. Evolution of fungal pathogens in domestic environments? Fungal Biol. 115, 1008–1018 (2011).
Köhler, J. R., Casadevall, A. & Perfect, J. The spectrum of fungi that infects humans. Cold Spring Harb. Perspect. Med. 5, a019273 (2015).
Brunke, S., Mogavero, S., Kasper, L. & Hube, B. Virulence factors in fungal pathogens of man. Curr. Opin. Microbiol. 32, 89–95 (2016).
Rokas, A. Evolution of the human pathogenic lifestyle in fungi. Nat. Microbiol. 7, 607–619 (2022).
Hubbell, S. P. A Unified Neutral Theory of Biodiversity and Biogeography (PUP, 2001).
Zhou, J. & Ning, D. Stochastic community assembly: Does it matter in microbial ecology? Microbiol. Mol. Biol. Rev. 81, e00002–e00017 (2017).
Zhang, S. J. et al. The structure and assembly mechanisms of plastisphere microbial community in natural marine environment. J. Hazard. Mater. 421, 126780 (2022).
Sun, Y., Shi, J., Wang, X., Ding, C. & Wang, J. Deciphering the mechanisms shaping the plastisphere microbiota in soil. Msystems 7, e00352–22 (2022).
Li, C. et al. The ecology of the plastisphere: microbial composition, function, assembly, and network in the freshwater and seawater ecosystems. Water Res. 202, 117428 (2021).
Ji, L., Tanunchai, B., Wahdan, S. F. M., Schädler, M. & Purahong, W. Future climate change enhances the complexity of plastisphere microbial co-occurrence networks, but does not significantly affect the community assembly. Sci. Total Environ. 844, 157016 (2022).
Gao, C. et al. Co-occurrence networks reveal more complexity than community composition in resistance and resilience of microbial communities. Nat. Commun. 13, 1–12 (2022).
Gutierrez, M. W., van Tilburg Bernardes, E., Changirwa, D., McDonald, B. & Arrieta, M. C. “Molding” immunity-modulation of mucosal and systemic immunity by the intestinal mycobiome in health and disease. Mucosal Immunol. 15, 573–583 (2022).
Sloan, W. T. et al. Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ Microbiol. 8, 732–740 (2006).
Burns, A. R. et al. Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J. 10, 655–664 (2016).
de Vries, F. T. et al. Soil bacterial networks are less stable under drought than fungal networks. Nat. Commun. 9, 1–12 (2018).
Guo, B. et al. Microbial co-occurrence network topological properties link with reactor parameters and reveal importance of low-abundance genera. NPJ Biofilms Microbiomes 8, 1–13 (2022).
Harjes, J., Link, A., Weibulat, T., Triebel, D. & Rambold, G. FAIR digital objects in environmental and life sciences should comprise workflow operation design data and method information for repeatability of study setups and reproducibility of results. Database 2020, baaa059 (2020).
Möller, J. N., Löder, M. G. & Laforsch, C. Finding microplastics in soils: a review of analytical methods. Environ. Sci. Technol 54, 2078–2090 (2020).
Djemiel, C. et al. Inferring microbiota functions from taxonomic genes: a review. Gigascience 11, giab090 (2022).
Nilsson, R. H. et al. Mycobiome diversity: high-throughput sequencing and identification of fungi. Nat. Rev. Microbiol. 17, 95–109 (2019).
Treseder, K. K. & Lennon, J. T. Fungal traits that drive ecosystem dynamics on land. Microbiol. Mol. Biol. Rev. 79, 243–262 (2015).
Boyce, K. J. & Andrianopoulos, A. Fungal dimorphism: the switch from hyphae to yeast is a specialized morphogenetic adaptation allowing colonization of a host. FEMS Microbiol. Rev. 39, 797–811 (2015).
Kettner, M. T., Rojas-Jimenez, K., Oberbeckmann, S., Labrenz, M. & Grossart, H. P. Microplastics alter composition of fungal communities in aquatic ecosystems. Environ. Microbiol. 19, 4447–4459 (2017).
Lacerda, A. L. D. F., Proietti, M. C., Secchi, E. R. & Taylor, J. D. Diverse groups of fungi are associated with plastics in the surface waters of the Western South Atlantic and the Antarctic Peninsula. Mol. Ecol. 29, 1903–1918 (2020).
Lynch, M. D. & Neufeld, J. D. Ecology and exploration of the rare biosphere. Nat. Rev. Microbiol 13, 217–229 (2015).
Tedersoo, L. et al. Global diversity and geography of soil fungi. Science 346, 1256688 (2014).
Watkinson, S. C., Boddy, L. & Money, N. The Fungi (Academic, 2015).
Luan, C. et al. Dysbiosis of fungal microbiota in the intestinal mucosa of patients with colorectal adenomas. Sci. Rep. 5, 1–9 (2015).
Wheeler, M. L. et al. Immunological consequences of intestinal fungal dysbiosis. Cell Host Microbe 19, 865–873 (2016).
Scales, B. S. et al. Cross-hemisphere study reveals geographically ubiquitous, plastic-specific bacteria emerging from the rare and unexplored biosphere. mSphere 6, e00851–20 (2021).
Ladau, J. & Eloe-Fadrosh, E. A. Spatial, temporal, and phylogenetic scales of microbial ecology. Trends Microbiol. 27, 662–669 (2019).
Rillig, M. C. et al. Interchange of entire communities: microbial community coalescence. Trends Ecol. Evol. 30, 470–476 (2015).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Voříšková, J. & Baldrian, P. Fungal community on decomposing leaf litter undergoes rapid successional changes. ISME J. 7, 477–486 (2013).
Egidi, E. et al. A few Ascomycota taxa dominate soil fungal communities worldwide. Nat. Commun. 10, 1–9 (2019).
Sun, Y. et al. Contribution of stochastic processes to the microbial community assembly on field‐collected microplastics. Environ. Microbiol. 23, 6707–6720 (2021).
Chen, W. et al. Stochastic processes shape microeukaryotic community assembly in a subtropical river across wet and dry seasons. Microbiome 7, 1–16 (2019).
Deng, Y. et al. Molecular ecological network analyses. BMC Bioinform. 13, 1–20 (2012).
Varsadiya, M., Urich, T., Hugelius, G. & Bárta, J. Fungi in permafrost-affected soils of the Canadian Arctic: Horizon- and site-specific keystone taxa revealed by co-occurrence network. Microorganisms 9, 1943 (2021).
Tong, X., Leung, M. H., Wilkins, D., Cheung, H. H. & Lee, P. K. Neutral processes drive seasonal assembly of the skin mycobiome. Msystems 4, e00004–e00019 (2019).
Hernandez, D. J., David, A. S., Menges, E. S., Searcy, C. A. & Afkhami, M. E. Environmental stress destabilizes microbial networks. ISME J. 15, 1722–1734 (2021).
Owino, C. O., Owuor, P. O. & Sigunga, D. O. Elucidating the causes of low phosphorus levels in ferralsols of Siaya County, Western Kenya. J. Soil Sci. Environ. 6, 260–267 (2015).
Harms, H., Schlosser, D. & Wick, L. Y. Untapped potential: exploiting fungi in bioremediation of hazardous chemicals. Nat. Rev. Microbiol. 9, 177–192 (2011).
Ekanayaka, A. H. et al. A review of the fungi that degrade plastic. J. Fungi 8, 772 (2022).
Navarro, D. et al. Large-scale phenotyping of 1,000 fungal strains for the degradation of non-natural, industrial compounds. Commun. Biol. 4, 1–10 (2021).
Tang, Q. et al. Keystone microbes affect the evolution and ecological coexistence of the community via species/strain specificity. J. Appl. Microbiol. 132, 1227–1238 (2022).
Yang, S. et al. Hydrocarbon degraders establish at the costs of microbial richness, abundance and keystone taxa after crude oil contamination in permafrost environments. Sci. Rep. 6, 1–13 (2016).
Yoshida, S. et al. A bacterium that degrades and assimilates poly (ethylene terephthalate). Science 351, 1196–1199 (2016).
Meyer-Cifuentes, I. E. et al. Synergistic biodegradation of aromatic-aliphatic copolyester plastic by a marine microbial consortium. Nat Commun. 11, 5790 (2020).
Nelson, T. F. et al. Biodegradation of poly(butylene succinate) in soil laboratory incubations assessed by stable carbon isotope labelling. Nat Commun. 13, 5691 (2022).
Allen, D. et al. Microplastics and nanoplastics in the marine-atmosphere environment. Nat. Rev. Earth Environ. 3, 393–405 (2022).
Revankar, S. G. & Sutton, D. A. Melanized fungi in human disease. Clin. Microbiol. Rev. 23, 884–928 (2010).
Kuhn, D. M. & Ghannoum, M. Indoor mold, toxigenic fungi, and Stachybotrys chartarum: infectious disease perspective. Clin. Microbiol. Rev. 16, 144–172 (2003).
Morales-López, S. E. & Garcia-Effron, G. Infections due to rare Cryptococcus species. A literature review. J. Fungi 7, 279 (2021).
Teixeira, M. D. M. et al. Exploring the genomic diversity of black yeasts and relatives (Chaetothyriales, Ascomycota). Stud. Mycol. 86, 1–28 (2017).
Segre, H. et al. Competitive exclusion, beta diversity, and deterministic vs. stochastic drivers of community assembly. Ecol. Lett. 17, 1400–1408 (2014).
Casadevall, A., Steenbergen, J. N. & Nosanchuk, J. D. ‘Ready made’ virulence and ‘dual use’ virulence factors in pathogenic environmental fungi—the Cryptococcus neoformans paradigm. Curr. Opin. Microbiol. 6, 332–337 (2003).
Casadevall, A. & Pirofski, L. A. Accidental virulence, cryptic pathogenesis, martians, lost hosts, and the pathogenicity of environmental microbes. Eukaryot. Cell 6, 2169–2174 (2007).
Plowright, R. K. et al. Pathways to zoonotic spillover. Nat. Rev. Microbiol. 15, 502–510 (2017).
Vilà, M. et al. Viewing emerging human infectious epidemics through the lens of invasion biology. Bioscience 71, 722–740 (2021).
Prenafeta-Boldu, F. X., Summerbell, R. & Sybren de Hoog, G. Fungi growing on aromatic hydrocarbons: Biotechnology’s unexpected encounter with biohazard? FEMS Microbiol. Rev. 30, 109–130 (2006).
Blasi, B. et al. Pathogenic yet environmentally friendly? Black fungal candidates for bioremediation of pollutants. Geomicrobiol. J. 33, 308–317 (2016).
Meides, N. et al. Reconstructing the environmental degradation of polystyrene by accelerated weathering. Environ. Sci. Technol. 55, 7930–7938 (2021).
Robert, V. A. & Casadevall, A. Vertebrate endothermy restricts most fungi as potential pathogens. J. Infect. Dis. 200, 1623–1626 (2009).
Shi, X., Chen, Z., Liu, X., Wei, W. & Ni, B. J. The photochemical behaviors of microplastics through the lens of reactive oxygen species: photolysis mechanisms and enhancing photo-transformation of pollutants. Sci. Total Environ. 846, 157498 (2022).
Romani, L. Immunity to fungal infections. Nat. Rev. Immunol. 11, 275–288 (2011).
Almeida, M. C. & Brand, A. C. Thigmo responses: the fungal sense of touch. Microbiol. Spectr. 5, 5–2 (2017).
Garbeva, P., Hol, W. G., Termorshuizen, A. J., Kowalchuk, G. A. & De Boer, W. Fungistasis and general soil biostasis—a new synthesis. Soil Biol. Biochem. 43, 469–477 (2011).
Gravelat, F. N. et al. Aspergillus galactosaminogalactan mediates adherence to host constituents and conceals hyphal β-glucan from the immune system. PLoS Pathog. 9, e1003575 (2013).
Arias-Andres, M., Klümper, U., Rojas-Jimenez, K. & Grossart, H. P. Microplastic pollution increases gene exchange in aquatic ecosystems. Environ. Pollut. 237, 253–261 (2018).
Jansson, J. K. & Hofmockel, K. S. Soil microbiomes and climate change. Nat. Rev. Microbiol. 18, 35–46 (2020).
Triebel, D., Hagedorn, G. & Rambold, G. (eds). Diversity Workbench—A Virtual Research Environment for Building and Accessing Biodiversity and Environmental Data. https://www.diversityworkbench.net (1999).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Nilsson, R. H. et al. The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 47, 259–264 (2019).
Weiss, S. et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5, 27 (2017).
R_Core_Team. A language and environment for statistical computing (R Foundation for Statistical Computing, 2017).
Elzhov, T. V., Mullen, K. M., Spiess, A. N. & Bolker, B. minpack.lm: R interface to the Levenberg-Marquardt nonlinear least-squares algorithm found in MINPACK, plus support for bounds. R package version 1.1-8. http://CRAN.R-project.org/package=minpack.lm (2013).
Harrell, F. E. Hmisc: Harrell Miscellaneous. R package version 3.14-5. http://CRAN.R-project.org/package=Hmisc (2014).
Põlme, S. et al. FungalTraits: a user-friendly traits database of fungi and fungus-like stramenopiles. Fungal Div. 105, 1–16 (2020).
Větrovský, T. et al. GlobalFungi, a global database of fungal occurrences from high-throughput-sequencing metabarcoding studies. Sci. Data 7, 1–14 (2020).
Nguyen, N. H. et al. FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 20, 241–248 (2016).
Parr, C. S. et al. Te encyclopedia of life v2: providing global access to knowledge about life on earth. Biodivers. Data J. 2, 1–28 (2014).
Gostinčar, C., Muggia, L. & Grube, M. Polyextremotolerant black fungi: oligotrophism, adaptive potential, and a link to lichen symbioses. Front. Microbiol. 3, 390 (2012).
James, T. Y. et al. Reconstructing the early evolution of fungi using a six-gene phylogeny. Nature 443, 818–822 (2006).
Anderson, J. B. Evolution of antifungal-drug resistance: mechanisms and pathogen fitness. Nat. Rev. Microbiol. 3, 547–556 (2005).
Cowen, L. E. The evolution of fungal drug resistance: modulating the trajectory from genotype to phenotype. Nat. Rev. Microbiol. 6, 187–198 (2008).
Clarke, K. R. & Gorley, R. N. Getting started with PRIMER v7. PRIMER-E: Plymouth, Plymouth Marine Laboratory Vol. 20 (2015).
Oksanen, F. J. et al. Vegan: community ecology package. R package version 2.4-3. https://CRAN.R-project.org/package=vegan (2017).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Villanueva, R. A. M. & Chen, Z. J. ggplot2: Elegant graphics for data analysis. 160–167 (2019).
Lahti, L. & Shetty, S. Tools for microbiome analysis in R 2.1. 26 (2017).
Wickham, H. Reshaping data with the reshape package. J. Stat. Softw. 21, 1–20 (2007).
Kassambara, A. & Kassambara, M. A. Package ‘ggpubr’. R package version 0.1, 6 (2020).
Zhang, J., Ding, Q. & Huang, J. spaa: Species association analysis. R package version 0.2, 2, 33 (2016).
Wickham, H., Francois R., Henry, L. & Müller, K. dplyr: A grammar of data manipulation. R package version 0.7, 4 (2017).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Tamura, K. & Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10, 512–526 (1993).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Revelle, W. & Revelle, M. W. Package ‘psych’. The Comprehensive R Archive Network Vol. 337 (2015).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: an open source software for exploring and manipulating networks. ICWSM 3, 361–362 (2009).
Rochman, C. M. & Hoellein, T. The global odyssey of plastic pollution. Science 368, 1184–1185 (2020).
Acknowledgements
This study was funded by the German Research Foundation (DFG) - SFB 1357-39197756; Subproject C04, and the Open Access Publishing Fund of the University of Bayreuth. We wish to thank Derek Peršoh (Faculty of Biology, Chemistry and Earth Sciences, University of Bayreuth) for his help in the optimisation of sequence data processing, Stella Franca Rösch (Chair of Marketing and Innovation, University of Bayreuth) for her support in graphic formatting and Luca Söhnel (Dept. of Mycology, University of Bayreuth) for his assistance in the subsampling process. Gratitude is also dedicated to the Ministry of Agriculture and Rural Development Kenya Plant Health Inspectorate Service (KEPHIS) for providing the export permit (Phytosanitary certificate no. 207/2019), the Bayerische Landesanstalt für Landwirtschaft, Institut für Pflanzenschutz for issue of the import authorisation (IPS 4a-7322.451) and the ITCER e.V. (https://itcer.org) for on-site support.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
G.G. and S.R. carried out the laboratory work and wrote the original draft. A.B. carried out Illumina sequencing. Sequence data processing was done by J.H. G.G., S.R. and J.H. performed the statistical analyses. G.G. and G.R. were responsible for the meta-analysis. The creation of the final version of the manuscript was supported by all authors. M.A.H. and G.R. acquired funding, designed the study, and supervised the project. G.R. collected and provided the environmental samples from Kenya.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth & Environment thanks Jie Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Clare Davis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gkoutselis, G., Rohrbach, S., Harjes, J. et al. Plastiphily is linked to generic virulence traits of important human pathogenic fungi. Commun Earth Environ 5, 51 (2024). https://doi.org/10.1038/s43247-023-01127-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43247-023-01127-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.