Abstract
The lifetime of chemical bonds shortens exponentially with force. Oddly, some protein-ligand complexes called catch bonds exhibit a sharp increase in lifetime when pulled with greater force. Inventing catch bond interfaces in synthetic materials would enable force-enhanced kinetics or self-strengthening under mechanical stress. Here, we present a molecular design that recapitulates catch bond behavior between nanoparticles tethered with macromolecules, consisting of one looped and one straight tether linking particles with weak adhesion. We calibrate the loop stiffness such that it opens around a target force to enable load-sharing among tethers, which facilitates a sequential to coordinated failure transition that reproduces experimental catch bond force-lifetime curve characteristics. We derive an analytical relation validated by molecular simulations to prove that loop and adhesion interactions can be tailored to achieve a spectrum of catch bond lifetime curves with this simple design. Our predictions break new ground towards designing tunable, catch-bond inspired self-strengthening materials.
Similar content being viewed by others
Introduction
The force-enhanced lifetime behavior of catch bonds1, which occurs only at small scales where thermal forces are relevant, starkly contrasts with ordinary chemical interactions called ‘slip bonds’, which exhibit an exponential lifetime decay with force, as predicted by Kramers2 and Bell3, and Zhurkov’s earlier work on fracture4. Experiments and simulations on various catch bond proteins (P-selectin/L-selectin5, pili adhesin FimH6, integrin7, actin8, von Willebrand factor9, and kinetochore10) have revealed that catch-bonds are common and play critical roles in many physiological processes. Different phenomenological models11, such as the two-pathway model12 and structural models, i.e., allosteric13 and the sliding re-binding models14, have been developed to fit the nonmonotonic lifetime curve of catch bonds with empirical parameters obtained from experiments. Phenomenological and structural models on proteins excel at fitting biophysical lifetimes curves but do not illustrate the dynamic molecular mechanism of a catch-bond in action11. The complex 3D structure and dynamics of proteins make it difficult to understand which features are necessary for catch bond formation, and existing molecular simulation approaches offer limited resolution into probing the conformational dynamics of a catch bond protein in action.
The self-strengthening and the reversible nature of the catch bonds make them uniquely important for materials science because a discontinuous transition to high resistance to failure is often desired in extreme loading scenarios that arise in applications spanning sports equipment, garments, adhesives, protective gear, and structural components. Recent modeling efforts by the authors have established that it is possible to design shape-changing nanoparticles that form catch bonds with each other, specifically by creating soft switches that facilitate force-induced affinity changes around a target force15,16. Unlike shear jamming seen in colloids and granular matter, these systems don’t require volume-preserving deformations to exhibit strengthening. Instead, they utilize instabilities to trigger an affinity switch that works even in pure tension. However, designing such instabilities with nanoparticles is challenging synthetically. Alternatively, it has been hypothesized that nanoparticle networks held together by polymer tethers17 that exhibit catch bonds18,19,20 could improve mechanical properties such as toughness beyond a threshold strain-rate or force level. Yet, macromolecular designs that reproduce a catch bond mechanism in such polymer-grafted nanoparticle networks are yet to be theoretically proposed and synthesized.
With this premise in mind, we choose to take a minimalistic approach to designing interfaces that replicate catch bond features. Instead of complex topologies, we utilize linear polymer tethers covalently grafted onto a nanoparticle and interacting with another particle with weak adhesive interactions (slip bonds) that we call an adhesin interaction (Fig. 1a). While both tethers have the same contour length (LI = LII), one of the tethers can form a loop stabilized by an ordinary slip bond that will ‘store’ some of its length, whereas the other lacks the interaction that facilitates a loop. These design elements, consisting of weak adhesin and loop interactions, are quite universal. An adhesin interaction can be formed simply from any non-covalent secondary interaction, such as hydrogen bonds or electrostatics21. Loops are ubiquitous in macromolecules, with various examples illustrated in Fig. 1b. Functionally, reversible loops facilitate sacrificial bonds and hidden length in proteins like titin, which is vital for muscle elasticity22. Looped or hairpin conformations are also attainable by DNA23 and RNA molecules24. In foldamers25 and DNA containing gels26, it has been possible to make synthetic looped polymers as well27. To our knowledge, the use of loops to create interfaces with catch bond kinetics has not been previously reported.
a Polymer-grafted nanoparticle network. Nanoparticles (gray spheres) are interconnected by two (red and blue) tethers. Blown-up image shows details of how two nanoparticle surfaces are interlinked by two tethers, with one having a loop (blue) and the other being non-looped (red). Adhesive parts and loop ends are represented by green and blue circles respectively. The adhesin and the loop interactions are represented with pink dashed lines. b Loop motifs in biological and biomimetic systems (i) protein-mediated DNA looping, (ii) mRNA hairpin loop, (iii) biomimetic design of a modular polymer that is inspired by titin that can reversibly unfold. Red dashed lines represent hydrogen bonds. c, d The two tether interface has two modes of failure under constant tensile force f: sequential and coordinated. In sequential failure, \({S}_{1}\) and \({S}_{2}\) mark adhesin dissociation events. In coordinated failure, \({C}_{1}\) marks the unfolding of the loop, \({C}_{2}\) and \({C}_{3}\) mark adhesin dissociations. The graphs show force exposure of the individual tethers in sequential and coordinated failure scenarios. The solid line represents the looped and the dashed line represents the tether without a loop.
Results
Analytical model
In our model, we consider two adhesin interactions to be identical and have different bond characteristics from the loop interaction. As shown in Fig. 1a, in equilibrium, the minimum energy configuration of this system is such that the folded tether keeps the two surfaces together under thermal undulations, while the tether without the loop is slack. When a constant tensile force is applied to pull the two surfaces apart, the tether with the loop (tether II) becomes taut and carries most of the load. Thus, its adhesin will dissociate before the adhesin of the tether without the loop (tether I). Conceptually, there are two primary kinetic pathways through which the interface can fail. If the tether II adhesin dissociates before its loop opens, the entire load will be transferred to tether I. Since adhesins are sequentially exposed to the entire load and fail one after the other, this will result in sequential failure. Alternatively, if the loop opens before tether II adhesin dissociates, the load is shared between the adhesins as both tethers become stretched. The overall lifetime will be greatly extended as each adhesin is exposed to half of the load, and the system exhibits coordinated failure. These failure mechanisms are schematically illustrated in Fig. 1c, which also shows the load distribution between the tethers. It should be noted that the existence of these two failure mechanisms is analogous to low and high affinity states of proteins, or failure pathways of different resistance often described in two-state or two-pathway phenomenological models of catch bonds28.
The central question that must be addressed is how the energy landscapes of the loop and the tether interactions should be designed to facilitate a transition from one failure mode to the other as the magnitude of the force increases. To answer this question, we derived an analytical expression for the lifetime of the interface, which denotes the time required for the dissociation of both adhesin interactions. To describe the kinetics of the loop and adhesin interactions in a simple way, we use Bell’s theorem3, which can describe rate of bond dissociation under external force f as
where ωo is the natural vibration frequency of the bond and Eb is the energy barrier, Δx is the transition state distance, and \({k}_{B}T\) is the thermal energy respectively. It is convenient to combine the natural frequency and exponential of the energy barrier by defining a thermal off rate \({r}^{0}\), which is the rate of bond dissociation in the absence of force. Hence, adhesin dissociation rate \({r}_{A}\) and loop opening rate \({r}_{L}\) have the following forms
Note that ∆x controls the force dependence of r, and r with the larger ∆x will be more force-sensitive and result in a sharper decrease in bond lifetime with force.
The timeline of the adhesin dissociation versus loop opening events on the looped tether governs which failure pathway a single pulling trial will follow. Thus, the probability of adhesin breaking before loop opening, or vice versa, dictates the system lifetime. If we call average sequential trial lifetime as \({{{\langle }}{{{{{\rm{\tau }}}}}}{{\rangle }}}_{S}\) and average coordinated lifetime as \({{{\langle }}{{{{{\rm{\tau }}}}}}{{\rangle }}}_{C},\) the average lifetime of the system, \(\left\langle {{{{{\rm{\tau }}}}}}\right\rangle ,\) will have the form of
where \({P}_{A < L}\) is the probability of adhesin dissociation happening prior to loop opening. This probability can be expressed as
With these definitions, it is possible to derive the analytical lifetime equation for the two-tether system in terms of Eqs. 2, 3 as
Note that r, \({P}_{A < L}\) and \(\left\langle {{{{{\rm{\tau }}}}}}\right\rangle\) are all functions of the applied force. Derivations of Eqs. 4–6 and the underlying assumptions are provided in the Supplementary Note 1–6.
The essential design principle to ensure catch bond behavior in our system can be stated as follows. At lower forces, the tethers should predominantly fail sequentially. As the force increases, the system should become more likely to exhibit coordinated failure, which will prolong the lifetime. Analytically, this translates into \({P}_{A < L}\) transitioning sigmoidally from 1 (\({r}_{A}\gg {r}_{L}\)) to 0 (\({r}_{L}\gg {r}_{A}\)) with increasing f (Fig. 2a). Hence, \({r}_{A}^{0}\) should be chosen greater than \({r}_{L}^{0}\) and \(\Delta {x}_{A}\) should be chosen as less than \(\Delta {x}_{L}\). Based on this design principle, we heuristically propose an original set of parameters for rA and rL, which are listed in Table 1. The units are chosen to match the typical force and lifetime ranges of existing catch bond curves. For these parameters, Fig. 2b shows that the lifetime of the interface is non-monotonic with force, showing a positive slope and peak at an intermediate range of forces. This proves our original conjecture that the dual tether interfaces with loops can exhibit catch bond behavior for a calibrated set of parameters. For interpreting and generalizing the lifetime curves, we specify four quantities of interest (Fig. 2c), which are the maximum value of the lifetime curve after the initial slip regime, \({\left\langle \tau \right\rangle }_{\max }\), the critical force at which it occurs, fc and gain in lifetime, \({\tau }_{G}\), which is the ratio of \({\left\langle \tau \right\rangle }_{\max }\) and \({\left\langle \tau \right\rangle }_{\min }\) and normalized force range of the catch bond peak, ∆f, which is defined as the difference between the force values of at \({\tau }_{\max }\) and \({\tau }_{\min }\) divided by the critical force fc. \({\left\langle \tau \right\rangle }_{\min }\) and \({\left\langle \tau \right\rangle }_{\max }\) are determined by numerically computing the zeros of the derivative of Eq. 6. These two points mark the slip-to-catch and catch-to-slip transitions, respectively. Systems that have τG > 1 show catch bond behavior. Figure 2d shows how varying \({r}_{A}^{0}/{r}_{L}^{0}\) while keeping Δx values constant governs catch bond performance (black line). As \({r}_{A}^{0}/{r}_{L}^{0}\) grows, increase in \({\tau }_{G}\) reaches a plateau. The same observation is true for ∆f, i.e., the width of the lifetime behavior approaches to limit 0.255 with increasing \({r}_{A}^{0}/{r}_{L}^{0}\) (red line). On the other hand, according to Fig. 2e, \({\tau }_{G}\) is maximum when the \(\Delta {x}_{L}/\Delta {x}_{A}\) is 1.93, going above or below this value decreases the gain till catch bond behavior is no longer possible, i.e., there is a narrow range of ratios of parameters that catch bond behavior is possible. Similarly, ∆f has a maximum, it reaches 0.255 when \(\Delta {x}_{L}/\Delta {x}_{A}\) is 1.71. The relations depicted in Fig. 2d, e, are true for all values of r0 and Δx that gives the same \({r}_{A}^{0}/{r}_{L}^{0}\) and \(\Delta {x}_{L}/\Delta {x}_{A}\); thus, we conclude that the relative values of the parameters chiefly govern the existence of catch bond behavior, and their magnitudes are less important. In addition, it should be noted that our lifetime curve is triphasic (slip-catch-slip behavior29), which is also seen in various biological catch bonds7,30. \({\tau }_{G}\) and ∆f values we find for near-optimal systems we report here are comparable to what’s seen in some of these biological systems.
a In order to observe the catch bond behavior, the probability of adhesin dissociating prior to loop opening \({P}_{A < L}\) should transition in a sigmoidal fashion from 1 (\({r}_{A}\gg {r}_{L}\)) to 0 (\({r}_{L}\gg {r}_{A}\)) with increasing pulling force. b The analytical expression (black line) and Markov Chain Monte Carlo simulations (diamonds) show that the two-tether system with tuned kinetic parameters can exhibit catch bond behavior. The sequential trial lifetime \({{{\langle }}{{{{{\rm{\tau }}}}}}{{\rangle }}}_{{{{{{\rm{S}}}}}}}\) (blue line), average coordinated lifetime \({{{\langle }}{{{{{\rm{\tau }}}}}}{{\rangle }}}_{C}\) (orange line) and the probability of adhesin dissociation happening prior to loop opening (black line) are shown. c Four quantities of interest are defined to characterize catch bond curves: (1) value of the peak lifetime, \({\left\langle \tau \right\rangle }_{\max }\), (2) the force at which it \({\left\langle \tau \right\rangle }_{\max }\) occurs, \({f}_{c}\), (3) gain in lifetime, \({\tau }_{G}\), defined as the ratio of \({\left\langle \tau \right\rangle }_{\max }\) and \({\left\langle \tau \right\rangle }_{\min }\), and (4) normalized force range of the catch bond peak, \(\Delta f\), which is defined as the difference between the force values of at \({\tau }_{\max }\) and \({\tau }_{\min }\) divided by the critical force \({f}_{c}.\) \({\left\langle \tau \right\rangle }_{\min }\) is defined as the value of the lifetime curve, at which the lifetime starts to increase. In this plot, the lifetime and force are normalized by \({\left\langle \tau \right\rangle }_{\max }\) and \({f}_{c}\), respectively. d, e The effects of varying the kinetic parameters on \({\tau }_{G}\) (black) and \(\Delta f\) (red). For d, the ranges of \({r}_{L}^{0}\) and \({r}_{A}^{0}\) are \(1.0\times {10}^{-5}\) to \(1.0\times {10}^{-3}\) and 200 to 800 respectively. \(\varDelta {x}_{A}\) and \(\varDelta {x}_{L}\) values are 0.5 and 1. For e the ranges of \(\varDelta {x}_{A}\) and \(\varDelta {x}_{L}\) are \(0.1\) to \(1.0\) and 0.5 to 1.2 respectively. \({r}_{L}^{0}\) and \({r}_{A}^{0}\) values are \(1.0\times {10}^{-3}\) and 400.
Markov chain Monte Carlo simulation
To validate these analytical predictions, we carry out Markov chain Monte Carlo (MCMC) simulations to calculate lifetime from rate parameters. This method is commonly used to describe bond rupture and formation events31,32, where rate constants are used to simulate probabilistic events with acceptance checks that account for both thermal effects and force contributions. As illustrated in Fig. 2b, the simulation results (black diamonds) are in excellent agreement with the analytical expression.
After establishing the catch bond behavior in our two-tether system, we systematically vary kinetic parameters to examine the factors that affect \({f}_{c}\) and \({\tau }_{\max }\). Figure 3 shows that \({f}_{c}\) can be decreased by increasing \({r}_{L}^{0}\) and \(\Delta {x}_{L}\) and increased by increasing \({r}_{A}^{0}\). \(\Delta {x}_{A}\) shows a more complex relationship, where the value of \({f}_{c}\) has a peak. On the other hand, \({\tau }_{\max }\) can be increased by increasing \({r}_{L}^{0}\) and \(\Delta {x}_{L}\) and decreased by increasing \({r}_{A}^{0}\) and \(\Delta {x}_{A}\). The important takeaway is that we can target certain \({f}_{c}\) and \({\tau }_{\max }\) values by varying \(\Delta {x}_{A}\) and \(\Delta {x}_{L}\). However, there is a tradeoff between maximizing \({\tau }_{\max }\) and tuning \({f}_{c}\) when varying kinetic parameters. We emphasize that the plots we obtain are specific to the system at hand, but nevertheless, the conceptual design framework we have established here can be applied to diverse systems with similar soft switch mechanisms that regulate low to high affinity transitions.
a \({r}_{L}^{0}\) b \({r}_{A}^{0}\) c \(\Delta {x}_{L}\) d \(\Delta {x}_{A}\). The \({f}_{c}\) and \({\tau }_{\max }\) can be tuned by changing these kinetic parameters. Decreasing \({r}_{L}^{0}\) and \({r}_{A}^{0}\), respectively increase and decrease \({f}_{c}.\) However, \(\varDelta {x}_{A}\) shows a more complex relationship, where the value of \({f}_{c}\) has a peak. On the other hand, \({\tau }_{\max }\) can be increased by increasing \({r}_{L}^{0}\) and \(\varDelta {x}_{L}\) and decreased by increasing \({r}_{A}^{0}\) and \(\varDelta {x}_{A}\).
Molecular dynamics simulation
Next, we test our findings in a computational experiment using molecular dynamics (MD) simulations, to illustrate that the theoretical concepts do hold for realistic molecular systems. A coarse-grained representation of the system is shown in Fig. 4a. To generate landscapes that qualitatively have similar kinetics as the analytical study, we represent \({E}_{A}\) and \({E}_{L}\) with Morse potentials, which have the following general form:
Here, D0 is the depth of the energy well, \({x}_{o}\) is the equilibrium bond distance and α is the parameter that controls the width of the well (the smaller α is, the broader the well), and x is the bond length. The base interaction parameters that we used in the MD simulations are listed in Table 2. In the methods section, we discuss our justification of D0 and α parameters.
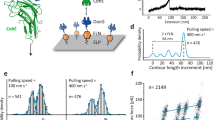
a Molecular setup of two tethers. The system uses the same color code in the Fig. 1a. The weak interactions are visualized by transparent purple lines. b Proposed energy landscapes of adhesin and loop interactions. Loop interaction has a deeper and broader energy well compared to the adhesin interaction. c–f Mean lifetime 〈τ〉 versus force f curve for tether systems with varying interaction parameters (\({D}_{o,L},{\alpha }_{L}{,D}_{o,A},{\alpha }_{A}\)). The error bars represent 95% confidence interval, derived by the bootstrapping method.
As in the case of the theoretical model, we explore how the lifetime curve characteristics depend on the energy landscape parameters. Figure 4c demonstrates the effect of \({D}_{0,L}\), where increasing its value lowers \({\tau }_{\max }\) and increases \({f}_{c}\). This means that overly stable and stiff loops will require a larger force to be opened and that naturally causes a reduction in the lifetime at both low and high force levels. Conversely, Fig. 4d shows that increasing \({D}_{0,A}\) brings the whole lifetime curve up considerably, given the exponential dependence of the lifetimes on the adhesin strength. As shown in Fig. 4e, f, increasing \({\alpha }_{L}\) lowers \({\tau }_{\max }\) and increases \({f}_{c}\). Increasing \({\alpha }_{A}\) has an opposite effect, since stiffer adhesin interactions concentrate the work more greatly on the loops, triggering a transition to the coordinated failure around a lower force threshold.
Discussion
To compare the analytical and MD results, we need to establish relations between the kinetic and interaction parameters. According to Bell’s theorem, \({r}^{0}\) is proportional to \({e}^{-{E}_{b}}\). In the Morse potential, \({D}_{0}\) is equivalent to the energy barrier \({E}_{b}\), thus we plot Fig. 5a, b with respect to \({e}^{-{D}_{0}}\). Since the Morse potential does not have a transition state, we do not have a definite \(\Delta x\) to define the width of the energy landscape. However, we can use the inflection point of the Morse potential \({x}^{* }={x}_{o}+{{{{{\rm{ln}}}}}}\left(2\right){\alpha }^{-1}\) and infer that the width is inversely proportional to \({\alpha }^{-1}\). Thus, we plot Fig. 5c, d with respect to \({\alpha }^{-1}.\) Overall, Fig. 5 follows the same trends as Fig. 3, i.e., the shift in \({f}_{c}\) and value of \({\tau }_{\max }\) depend similarly on both parameter spaces. Although there are differences in Figs. 3b and 5b, where the right shift in \({f}_{c}\) is sharper in MD simulations and Figs. 3c and 5c, where the increase in \({\tau }_{\max }\) is concave in analytical equations and convex in MD simulations. These minor differences, which don’t change any of our main conclusions, can be attributed to the nonlinearity of the energy landscape or additional rebinding events that we have not considered in this iteration of the analytical model. Bell’s theorem is the simplest description of force-dependence in dissociation kinetics and may not accurately describe trends in nonlinear systems. Additionally, our analytical theory does not account for adhesin rebinding to occur, and in the MD simulations, some of the cases do exhibit rebinding. MD analysis of rebinding effect on lifetime curves is provided in Supplementary Note 7. We note that there are models to study the force-dependent kinetics of parallel bonds33 including rebinding effects, however, in our case utilizing parallel tethers alone doesn’t result in catch bond behavior.
a \({e}^{-{D}_{0,L}}\) b \({e}^{-{D}_{0,A}}\) c \({\alpha }_{L}^{-1}\) d \({\alpha }_{A}^{-1}\). It shows the comparison between analytical and MD results and establishes a relation between the kinetic and interaction parameters. The change in \({f}_{c}\) and \({\tau }_{\max }\) with \({e}^{-{D}_{0}}\) are very similar to \({r}_{0}\).
Lastly, we investigate the effects of the tether length on the catch bond behavior using MD simulations. For these simulations, we vary the number of segments in our two-tether system. As an example, the tethers in Fig. 4a have 5 segments. Figure 6 shows that as the number of segments increases, the peak lifetime of the system starts to decrease, ultimately transitioning to slip behavior in 15-unit tether. This behavior is expected because longer tethers exhibit lower stiffness, and the stiffness of a polymer can play an important role in transmitting the force to the bond. It has been shown in the literature34 that for polymer tethers with adhesive tips, softer tethers will result in a higher off rate, i.e., shorter lifetime.
However, we rely on our established insights to re-calibrate the adhesin and loop kinetics. According to Fig. 5, we modified the MD interaction parameter to see if a catch bond behavior can be obtained with long tether systems. This data is shown as 15* in Fig. 6, showing the re-emergence of the catch bond behavior. The \({\alpha }_{A}\) is increased to 12 from 10 and \({\alpha }_{L}\) is reduced to 1.54 from 2. The \({D}_{0,L}\) and \({D}_{0,A}\) is kept as 2.8 kcal/mol and 1.25 kcal/mol, respectively, for all systems. This improves the gain and thus facilitates a slip-catch transition, without considering tether elasticity. This demonstrates that our theoretical and computational simulations still provide valuable insight into systems with longer chains, but also that the loop and adhesin kinetics would need to be tailored for specific tether characteristics.
Although our two tether systems do not replicate any existing structure, we estimated two of our key parameters, Δf and \({\tau }_{G}\) for different catch bond proteins (Table 3) using the same methodology discussed above. We also calculate ∆f and \({\tau }_{G}\) for our MD and MCMC simulations to compare with experimental results (Table 4). In experimental data on triphasic catch bonds, we find \(\Delta f\) ranges from 0.38 to 0.68, whereas in MCMC simulations the range is 0.03 to 0.255 and in MD it is 0.18 to 0.3. This indicates simulations are not that different in terms of the breadth of the force range relative to experiments. In the case of \({\tau }_{G}\), our results are also in a range of experimental values as shown in Table 4. Our models, despite having much simpler energy landscapes than actual protein-ligand complexes, recapitulate similar gain values.
Conclusion
This work demonstrates a catch bond mechanism that could be employed in polymer or macromolecule grafted nanoparticles. The essential feature of the tethers is to control the lifetime of the interface by tuning the opening kinetics of a loop that facilitates load-sharing by revealing hidden length in one of two tethers interconnecting the particles. This work establishes an analytical theory validated by both Monte Carlo and MD simulations, which provides insights into how in such a model system one could precisely dictate the shape of the force vs. lifetime curve by tuning loop and adhesin kinetics. By tailoring the stiffness and strength of a loop in one of the tethers, it is possible to control sequential vs. coordinated failure of the adhesive tethers, which can be used to engineer force dependence of the kinetic pathways of interfacial dissociation. Tuning the energy landscapes or rate parameters clearly revealed that a target peak lifetime, gain, and the normalized force range over which catch bond behavior occurs can be programmed. While we have not exhaustively explored all possible designs, this versatile strategy opens the possibility of creating catch bonds that exhibit lifetime characteristics reaching beyond the feasible range seen in biological proteins (typically pN forces and seconds in peak lifetimes). This has major implications for mechanosensitive materials, nanocomposites, and drug delivery systems that could utilize mechanical cues for tailoring interfacial strength. Moreover, the fact that polymer-like tethers, loops, and weak bonds can be generated even in macroscopic thermalized magneto-mechanical systems35, we envision that it is possible to create catch bonds in a scale-invariant fashion. Lastly, using MD simulations, we generated reproducible lifetime curves reminiscent of highly complex catch bond proteins. While our design is, to the best of our knowledge, unprecedented and distinct from biological catch bonds, our study also raises the questions as to whether catch bonds can occur through load-sharing and programmed unfolding of multiple adhesion proteins, where soft fold and stiff ligand interactions might hold the key to achieving force enhanced lifetimes in diverse biomolecular systems. Future studies may build on these findings to study the mechanical properties of catch-bond capable networks of nanoparticles or investigate modifications to the design to further optimize performance metrics such as the lifetime gain and normalized force range of synthetic catch bonds. In summary, this work presents a versatile strategy for making man-made catch bond mechanisms, suggests that catch bond behavior can occur readily in interfaces consisting of multiple folded macromolecules, and provides insight into how self-strengthening interfaces can be engineered in nanoparticle networks with man-made catch-bonds.
Methods
Markov Chain Monte Carlo
The transition events of both the loop and the adhesins are modeled as a two-state first-order (Markov) process, where the dissociation and unfolding probabilities, under a short time interval \(\Delta t,\) are \({P}_{A}={r}_{A}\Delta t\) and \({P}_{L}={r}_{L}\Delta t\) respectively. Here, \({r}_{A}\) and \({r}_{L}\) are the force dependent rate equations that are defined in Eqs. 4–5 and Δt is the time step (polling interval) of the simulation. The time step of 10−8 time units is found to be sufficient to make large changes to the system at each time step improbable and is employed in the simulations. To determine what would happen in each time step, a random number (uniformly distributed between 0 and 1) is compared with \({P}_{A}\) and \({P}_{L}\). Depending on which event happens first, the simulation branches into two paths. If adhesin dissociates first, the simulation will calculate the lifetime for sequential failure, where event times S1 and S2 are recorded (Fig. 1c). If loop opens first, the simulation will calculate lifetime for coordinated failure, where event times C1, C2, and C3 are recorded (Fig. 1c). The accumulated time steps are the lifetime of the two-tether system in that simulation run, which is repeated 100,000 times for a given force to obtain an ensemble of exponentially distributed lifetimes and their average at that force. The four model parameters (\({r}_{A}^{0}\), \({r}_{L}^{0}\), \(\Delta {x}_{A}\) and \(\Delta {x}_{L}\)) are chosen to be the same as the analytical theory for comparison.
Molecular dynamics
The MD simulations were performed using the LAMMPS software package36. The simulations were run in the NVT ensemble at 50 K using a Langevin thermostat with a damping factor of 100 time steps. A lower temperature is used to reduce noise while ensuring thermal fluctuations give rise to stochastic bond breaking events. The adopted time step of 1 fs was found to be sufficiently small to ensure the accuracy of the lifetime measurements. The initial configuration of the system is shown in Fig. 4. Each simulation starts with 50,000 time step equilibration process, which is adequate given the simple nature and small size of our system. After equilibration, an instantaneous constant force is applied perpendicular to the top surface, while the bottom surface is held stationary. We measure lifetimes under forces ranging from 160 to 380 pN. Both surfaces are treated as rigid bodies that are constrained to move only in the direction normal to their planes. The flexible polymer tethers consist of harmonic bond interactions with no angle terms. The mass of the tether beads and the surface beads are set as 1000 g mol−1. Each tether has 6 beads that are connected with 2 Å long harmonic bonds with stiffness of 1000 kcal mol−1 Å−2.
In line with the two-state model representation of catch bonds, the mean lifetime of the system \(\left\langle \tau \right\rangle\) must increase when a tensile force is applied, which occurs if the force facilitates a propensity for the system to transition into the high-affinity state. At small forces, most trials should result in failure in the low-affinity state, which in this study corresponds to the sequential failure path. At large forces, most trials should result in transition to a high-affinity state, which in this study corresponds to the coordinated failure path. Interaction energies between the adhesins and the bottom plate are measured. To obtain meaningful statistics for each force value, 10,000 trials were performed. The interface lifetime was recorded when the interaction energies for both adhesins drop below −0.1 kcal mol−1, where we considered adhesins to be dissociated. Animations of sample simulation run that end with sequential and coordinated failure are provided in as Supplementary Movie 1 and Supplementary Movie 2, respectively. It should be noted that the temperature and the force ranges used in the simulations are arbitrary and do not aim to reproduce any particular experimental setup. We note that goal here is to prove the existence of catch bond behavior in these systems in a generic way and quantify its dependence on parameters in a theory-driven fashion, rather than matching a particular experimental observation.
Bootstrapping
The events we simulated all followed exponential distributions, where the mean and the standard deviation of the lifetime data were equal. Since our data were far from the normal distribution, we decided to use the bootstrapping method instead of using standard deviations, to calculate the variability of \(\left\langle \tau \right\rangle\) of the data. For each force value, a histogram of means was created from 10,000 bootstrap samples with the size of the original data set both for MCMC and MD simulations. A confidence interval of \(\left\langle \tau \right\rangle\) at 95% confidence level is calculated and plotted as error bars.
Interaction energy landscapes
Based on our previous works on catch bond systems15,16, we hypothesized that the loop interaction, which serves as a force-sensitive switch, should have a deeper and a broader energy well compared to adhesin energy landscape to enable catch bond formation, as shown in Fig. 4b. The work done on the bonds by the force, tilts the loop and adhesin energy landscapes \(\left({E}_{L},{E}_{A}\right)\), and the resulting reduction in the energy barrier \(({E}_{b})\) is sensitive to the distance to the transition state \(\left(\Delta x\right)\). Therefore, in Eq. 7, ensuring that \({E}_{L}\) has a lower α (broader landscape) compared to \({E}_{A}\) results in the \({E}_{b}\) of the folded interaction to be reduced greatly when the applied force is adequately large. According to Kramers’ theory2, \(\left\langle \tau \right\rangle\) is directly proportional to \(\exp [{E}_{b}]\), thus the loop lifetime \({\left\langle \tau \right\rangle }_{L}\), compared to the adhesin lifetime \({\left\langle \tau \right\rangle }_{A}\), declines more sharply when subjected to force. In addition, since \({E}_{L}\) has a deeper barrier compared to \({E}_{A}\), loop lifetime \({\left\langle \tau \right\rangle }_{L}\) is greater than the adhesin \({\left\langle \tau \right\rangle }_{A}\) in the absence of force. However, since \({E}_{L}\) has larger reduction of \({E}_{b}\) under force, \({\left\langle \tau \right\rangle }_{L}\) crosses over \({\left\langle \tau \right\rangle }_{A}\) and becomes shorter beyond a threshold force value. This facilitates the transition to coordinated failure through loop opening at large forces.
Data availability
All data used for analysis is made available upon request to the corresponding author.
Code availability
All simulation input scripts and codes can be found at the following repository: https://github.com/keten-group/2tethermodel
References
Dembo, M., Torney, D. C., Saxman, K. & Hammer, D. The reaction-limited kinetics of membrane-to-surface adhesion and detachment. Proc. R. Soc. London. Ser. B. Biol. Sci. 234, 55–83 (1988).
Kramers, H. A. Brownian motion in a field of force and the diffusion model of chemical reactions. Physica. 7, 284–304 (1940).
Bell, G. I. Models for the specific adhesion of cells to cells. Science (1979) 200, 618–627 (1978).
Zhurkov, S. N. Kinetic concept of the strength of solids. Int. J. Fract. 26, 295–307 (1984).
Sarangapani, K. K. et al. Low force decelerates L-selectin dissociation from P-selectin glycoprotein ligand-1 and endoglycan. J. Biol. Chem. 279, 2291–2298 (2004).
Thomas, W. E., Trintchina, E., Forero, M., Vogel, V. & Sokurenko, E. V. Bacterial adhesion to target cells enhanced by shear force. Cell 109, 913–923 (2002).
Kong, F., García, A. J., Mould, A. P., Humphries, M. J. & Zhu, C. Demonstration of catch bonds between an integrin and its ligand. J. Cell Biol. 185, 1275–1284 (2009).
Buckley, C. D. et al. The minimal cadherin-catenin complex binds to actin filaments under force. Science 346, 1254211 (2014).
Yago, T. et al. Platelet glycoprotein Iba forms catch bonds with human WT vWF but not with type 2B von Willebrand disease vWF. J. Clin. Invest. 118, 20 (2008).
Sarangapani, K. K. & Asbury, C. L. Catch and release: how do kinetochores hook the right microtubules during mitosis? Trends Genet. 30, 150–159 (2014).
Thomas, W. E., Vogel, V. & Sokurenko, E. Biophysics of catch bonds. Ann. Rev. Biophys. 37, 399–416 (2008).
Thomas, W. et al. Catch-bond model derived from allostery explains force-activated bacterial adhesion. Biophys. J. 90, 753–764 (2006).
Pereverzev, Y. V., Prezhdo, O. V. & Sokurenko, E. V. Allosteric role of the large-scale domain opening in biological catch-binding. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 79, 1–7 (2009).
Lou, J. & Zhu, C. A structure-based sliding-rebinding mechanism for catch bonds. Biophys. J. 92, 1471–1485 (2007).
Dansuk, K. C., Keten, S., Dansuk, K. C. & Keten, S. A simple mechanical model for synthetic catch bonds a simple mechanical model for synthetic catch bonds. Matter 1–15 https://doi.org/10.1016/j.matt.2019.06.005.
Dansuk, K. C. & Keten, S. Self-strengthening biphasic nanoparticle assemblies with intrinsic catch bonds. Nature Communications 12, 85 (2021).
Iyer, B. V. S. et al. Ductility, toughness and strain recovery in self-healing dual cross-linked nanoparticle networks studied by computer simulations. Progress in Polymer Science 40, 121–137 (2014).
Iyer, B. V. S., Yashin, V. V. & Balazs, A. C. Harnessing biomimetic catch bonds to create mechanically robust nanoparticle networks. Polymer (United Kingdom) 69, 310–320 (2015).
Mbanga, B. L., Yashin, V. V. & Balazs, A. C. Tailoring the Mechanical Properties of Nanoparticle Networks That Encompass Biomimetic Catch Bonds. (2017) https://doi.org/10.1002/polb.24542.
Dansuk, K. C. & Keten, S. Tunable seat belt behavior in nanocomposite interfaces inspired from bacterial adhesion pili. Soft Matter 14, 1530–1539 (2018).
Busch, A. & Waksman, G. Chaperone-usher pathways: Diversity and pilus assembly mechanism. Philos. Trans. R. Soc. B: Biol. Sci. 367, 1112–1122 (2012).
Kellermayer, M. S. Z., Smith, S. B., Granzier, H. L. & Bustamante, C. Folding-unfolding transitions in single titin molecules characterized with laser tweezers. Science (1979) 276, 1112–1116 (1997).
Chen, Y. J., Johnson, S., Mulligan, P., Spakowitz, A. J. & Phillips, R. Modulation of DNA loop lifetimes by the free energy of loop formation. Proc. Natl Acad. Sci. USA 111, 17396–17401 (2014).
Hyeon, C. & Thirumalai, D. Mechanical unfolding of RNA hairpins. Proc. Natl Acad. Sci. USA 102, 6789–6794 (2005).
Adhikari, R. & Makarov, D. E. Mechanochemical kinetics in elastomeric polymer networks: heterogeneity of local forces results in nonexponential kinetics. J. Phys. Chem. B https://doi.org/10.1021/acs.jpcb.6b12758 (2017).
Cangialosi, A. et al. DNA sequence–directed shape change of photopatterned hydrogels via high-degree swelling. Science (1979) 357, 1126–1130 (2017).
Chung, J., Kushner, A. M., Weisman, A. C. & Guan, Z. Direct correlation of single-molecule properties with bulk mechanical performance for the biomimetic design of polymers. Nat. Mater. 13, 1055–1062 (2014).
Bullerjahn, J. T. & Kroy, K. Analytical catch-slip bond model for arbitrary forces and loading rates. Phys. Rev. E 93, 1–8 (2016).
Chakrabarti, S., Hinczewski, M. & Thirumalai, D. Phenomenological and microscopic theories for catch bonds. J. Struct. Biol. 197, 50–56 (2017).
Harder, A. et al. Catch bond interaction between cell-surface sulfatase Sulf1 and Glycosaminoglycans. Biophys. J. 108, 1709–1717 (2015).
Björnham, O., Axner, O. & Andersson, M. Modeling of the elongation and retraction of Escherichia coli P pili under strain by Monte Carlo simulations. Eur. Biophys. J. 37, 381–391 (2008).
Zakrisson, J., Wiklund, K., Axner, O. & Andersson, M. The shaft of the type 1 Fimbriae regulates an external force to match the FimH catch bond. Biophys. J. 104, 2137–2148 (2013).
Ackbarow, T., Chen, X., Keten, S. & Buehler, M. J. Hierarchies, multiple energy barriers, and robustness govern the fracture mechanics of α-helical and β-sheet protein domains. Proc. Natl Acad. Sci. USA 104, 16410–16415 (2007).
Evans, E. Probing the relation between force—lifetime—and chemistry in single molecular bonds. Ann. Rev. Biophys. Biomol. Struct. 30, 105–128 (2001).
Niu, R. et al. Magnetic handshake materials as a scale-invariant platform for programmed self-assembly. Proc. Natl Acad. Sci. USA 116, 24402–24407 (2019).
Plimpton, S., National, L. S. Fast parallel algorithms for short–range molecular dynamics. J. Comput. Phys. 117, 1–42 (1995).
Wayman, A. M., Chen, W., McEver, R. P. & Zhu, C. Triphasic force dependence of E-selectin/ligand dissociation governs cell rolling under flow. Biophys. J. 99, 1166–1174 (2010).
Acknowledgements
This research was sponsored by the Office of Naval Research Early Career Award (PECASE, grant # N00014-16-1-3175), the Departments of Civil & Environmental and Mechanical Engineering at Northwestern University, and a supercomputing grant from Northwestern University High-Performance Computing Center.
Author information
Authors and Affiliations
Contributions
K.D. and S.K. ideated and supervised the research. K.D. and S.P. performed the MCMC and MD simulations and data analysis. All authors contributed to the writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Materials thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: John Plummer.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dansuk, K.C., Pal, S. & Keten, S. A catch bond mechanism with looped adhesive tethers for self-strengthening materials. Commun Mater 4, 60 (2023). https://doi.org/10.1038/s43246-023-00387-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43246-023-00387-y
This article is cited by
-
Emergence of slip-ideal-slip behavior in tip-links serve as force filters of sound in hearing
Nature Communications (2024)