Abstract
Cancer immunotherapy with chimeric antigen receptor (CAR) T cells can cause immune effector cell-associated neurotoxicity syndrome (ICANS). However, the molecular mechanisms leading to ICANS are not well understood. Here we examined the role of microglia using mouse models and cohorts of individuals with ICANS. CD19-directed CAR (CAR19) T cell transfer in B cell lymphoma-bearing mice caused microglia activation and neurocognitive deficits. The TGFβ-activated kinase-1 (TAK1)–NF-κB–p38 MAPK pathway was activated in microglia after CAR19 T cell transfer. Pharmacological TAK1 inhibition or genetic Tak1 deletion in microglia using Cx3cr1CreER:Tak1fl/fl mice resulted in reduced microglia activation and improved neurocognitive activity. TAK1 inhibition allowed for potent CAR19-induced antilymphoma effects. Individuals with ICANS exhibited microglia activation in vivo when studied by translocator protein positron emission tomography, and imaging mass cytometry revealed a shift from resting to activated microglia. In summary, we prove a role for microglia in ICANS pathophysiology, identify the TAK1–NF-κB–p38 MAPK axis as a pathogenic signaling pathway and provide a rationale to test TAK1 inhibition in a clinical trial for ICANS prevention after CAR19 T cell-based cancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The microarray data in this study were deposited in the Gene Expression Omnibus (GEO) repository under accession code GSE193696. The single-nucleus transcriptomic data were deposited in the GEO repository under accession number GSE223647. Kinase assay data were deposited in the GEO repository under accession code GSE225811. IMC data were deposited in Zenodo at https://doi.org/10.5281/zenodo.10695442 (ref. 78). Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
No custom code was generated.
References
Santomasso, B. D. et al. Clinical and biological correlates of neurotoxicity associated with CAR T-cell therapy in patients with B-cell acute lymphoblastic leukemia. Cancer Discov. 8, 958–971 (2018).
Maude, S. et al. Tisagenlecleucel in children and young adults with B-cell lymphoblastic leukemia. N. Engl. J. Med. 378, 439–448 (2018).
Neelapu, S. S. et al. Axicabtagene ciloleucel CAR T-cell therapy in refractory large B-cell lymphoma. N. Engl. J. Med. 377, 2531–2544 (2017).
Belin, C. et al. Description of neurotoxicity in a series of patients treated with CAR T-cell therapy. Sci. Rep. 10, 18997 (2020).
Sterner, R. C. & Sterner, R. M. Immune effector cell associated neurotoxicity syndrome in chimeric antigen receptor-T cell therapy. Front. Immunol. 13, 879608 (2022).
Gust, J. et al. Endothelial activation and blood–brain barrier disruption in neurotoxicity after adoptive immunotherapy with CD19 CAR-T cells. Cancer Discov. 7, 1404–1419 (2017).
Parker, K. R. et al. Single-cell analyses identify brain mural cells expressing CD19 as potential off-tumor targets for CAR-T immunotherapies. Cell 183, 126–142 (2020).
Deng, Q. et al. Characteristics of anti-CD19 CAR T cell infusion products associated with efficacy and toxicity in patients with large B cell lymphomas. Nat. Med. 12, 1878–1887 (2020).
Norelli, M. et al. Monocyte-derived IL-1 and IL-6 are differentially required for cytokine-release syndrome and neurotoxicity due to CAR T cells. Nat. Med. 24, 739–748 (2018).
Giavridis, T. et al. CAR T cell-induced cytokine release syndrome is mediated by macrophages and abated by IL-1 blockade. Nat. Med. 24, 731–738 (2018).
Sterner, R. M. et al. GM-CSF inhibition reduces cytokine release syndrome and neuroinflammation but enhances CAR-T cell function in xenografts. Blood 133, 697–709 (2019).
Lee, D. W. et al. ASTCT consensus grading for cytokine release syndrome and neurologic toxicity associated with immune effector cells. Biol. Blood Marrow Transplant. 4, 625–638 (2019).
Maus, M. V. et al. Society for Immunotherapy of Cancer (SITC) clinical practice guideline on immune effector cell-related adverse events. J. Immunother. Cancer 2, e001511 (2020).
Strati, P. et al. Prognostic impact of corticosteroids on efficacy of chimeric antigen receptor T-cell therapy in large B-cell lymphoma. Blood 137, 3272–3276 (2021).
Zeiser, R. & Lee, S. J. Three US Food and Drug Administration-approved therapies for chronic GVHD. Blood 139, 1642–1645 (2022).
Zeiser, R. et al. Ruxolitinib for glucocorticoid-refractory chronic graft-versus-host disease. N. Engl. J. Med. 385, 228–238 (2021).
Zeiser, R. et al. Ruxolitinib for glucocorticoid-refractory acute graft-versus-host disease. N. Engl. J. Med. 382, 1800–1810 (2020).
Prinz, M., Jung, S. & Priller, J. Microglia biology: one century of evolving concepts. Cell 179, 292–311 (2019).
Prinz, M., Masuda, T., Wheeler, M. A. & Quintana, F. J. Microglia and central nervous system-associated macrophages-from origin to disease modulation. Annu. Rev. Immunol. 39, 251–277 (2021).
Kochenderfer, J. N., Yu, Z., Frasheri, D., Restifo, N. P. & Rosenberg, S. A. Adoptive transfer of syngeneic T cells transduced with a chimeric antigen receptor that recognizes murine CD19 can eradicate lymphoma and normal B cells. Blood 116, 3875–3886 (2010).
d’Errico, P. et al. Microglia contribute to the propagation of Aβ into unaffected brain tissue. Nat. Neurosci. 25, 20–25 (2022).
Goldmann, T. et al. Origin, fate and dynamics of macrophages at central nervous system interfaces. Nat. Immunol. 17, 797–805 (2016).
Mathew, N. R. et al. Graft-versus-host disease of the CNS is mediated by TNF upregulation in microglia. J. Clin. Invest. 130, 1315–1329 (2020).
Manouchehri, N. et al. CD11c+CD88+CD317+ myeloid cells are critical mediators of persistent CNS autoimmunity. Proc. Natl Acad. Sci. USA 118, e2014492118 (2021).
Morris, E. C., Neelapu, S. S., Giavridis, T. & Sadelain, M. Cytokine release syndrome and associated neurotoxicity in cancer immunotherapy. Nat. Rev. Immunol. 2, 85–96 (2022).
Hay, K. A. et al. Kinetics and biomarkers of severe cytokine release syndrome after CD19 chimeric antigen receptor-modified T-cell therapy. Blood 130, 2295–2306 (2017).
Spangenberg, E. et al. Sustained microglial depletion with CSF1R inhibitor impairs parenchymal plaque development in an Alzheimer’s disease model. Nat. Commun. 10, 3758 (2019).
Mildner, A. et al. CCR2+Ly-6Chi monocytes are crucial for the effector phase of autoimmunity in the central nervous system. Brain 132, 2487–2500 (2009).
Jordão, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, 6425 (2019).
Absinta, M. et al. A lymphocyte–microglia–astrocyte axis in chronic active multiple sclerosis. Nature 597, 709–714 (2021).
Schafer, D. P. et al. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 74, 691–705 (2012).
Teachey, D. T. et al. Identification of predictive biomarkers for cytokine release syndrome after chimeric antigen receptor T cell therapy for acute lymphoblastic leukemia. Cancer Discov. 6, 664–679 (2017).
Nagai, A. et al. Involvement of cystatin C in pathophysiology of CNS diseases. Front. Biosci. 13, 3470–3479 (2008).
Maldonado-Ruiz, R. et al. MCP-1 signaling disrupts social behavior by modulating brain volumetric changes and microglia morphology. Mol. Neurobiol. 59, 932–949 (2022).
Faulhaber, L. D. et al. Brain capillary obstruction during neurotoxicity in a mouse model of anti-CD19 chimeric antigen receptor T-cell therapy. Brain Commun. 4, fcab309 (2021).
Haroon, E., Miller, A. H. & Sanacora, G. Inflammation, glutamate, and glia: a trio of trouble in mood disorders. Neuropsychopharmacology 42, 193–215 (2016).
Raud, S. et al. Relation between increased anxiety and reduced expression of α1 and α2 subunits of GABAA receptors in WFS1-deficient mice. Neurosci. Lett. 460, 138–142 (2009).
Pribiag, H. & Stellwagen, D. TNF-α downregulates inhibitory neurotransmission through protein phosphatase 1-dependent trafficking of GABAA receptors. J. Neurosci. 33, 15879–15893 (2013).
Masuda, T. et al. Novel Hexb-based tools for studying microglia in the CNS. Nat. Immunol. 21, 802–815 (2020).
Ouyang, C. et al. Transforming growth factor (TGF)-β-activated kinase 1 (TAK1) activation requires phosphorylation of serine 412 by protein kinase A catalytic subunit α (PKACα) and X-linked protein kinase (PRKX). J. Biol. Chem. 289, 24226–24237 (2014).
Liu, X. et al. CaMKII promotes TLR-triggered proinflammatory cytokine and type I interferon production by directly binding and activating TAK1 and IRF3 in macrophages. Blood 112, 4961–4970 (2008).
Matsumoto-Ida, M. et al. Activation of TGF-β1–TAK1–p38 MAPK pathway in spared cardiomyocytes is involved in left ventricular remodeling after myocardial infarction in rats. Am. J. Physiol. Heart. Circ. Physiol. 290, H709–H715 (2006).
Goldmann, T. et al. A new type of microglia gene targeting shows TAK1 to be pivotal in CNS autoimmune inflammation. Nat. Neurosci. 16, 1618–1626 (2013).
McKenzie, K. J. et al. Is β-APP a marker of axonal damage in short-surviving head injury? Acta Neuropathol. 92, 608–613 (1996).
Schwabenland, M. et al. Deep spatial profiling of human COVID-19 brains reveals neuroinflammation with distinct microanatomical microglia–T-cell interactions. Immunity 54, 1594–1610 (2021).
Sharma, M. et al. Pathological spectrum and β-APP immunoreactivity as a diagnostic tool of diffuse axonal injury following traumatic brain injury: a novel classification. J. Lab. Physicians 15, 399–408 (2023).
Casellas, P., Galiegue, S. & Basile, A. S. Peripheral benzodiazepine receptors and mitochondrial function. Neurochem. Int. 40, 475–486 (2002).
Kuhlmann, A. C. & Guilarte, T. R. Cellular and subcellular localization of peripheral benzodiazepine receptors after trimethyltin neurotoxicity. J. Neurochem. 74, 1694–1704 (2002).
Ikawa, M. et al. 11C-ER176, a radioligand for 18-kDa translocator protein, has adequate sensitivity to robustly image all three affinity genotypes in human brain. J. Nucl. Med. 58, 320–325 (2017).
Unterrainer, M. et al. TSPO PET with [18F]GE-180 sensitively detects focal neuroinflammation in patients with relapsing-remitting multiple sclerosis. Eur. J. Nucl. Med. Mol. Imaging 45, 1423–1431 (2018).
Volk, L., Chiu, S. L., Sharma, K. & Huganir, R. L. Glutamate synapses in human cognitive disorders. Annu. Rev. Neurosci. 38, 127–149 (2015).
Xiu, M. H. et al. Tumor necrosis factor-α-1031T/C polymorphism is associated with cognitive deficits in chronic schizophrenia patients versus healthy controls. Am. J. Med. Genet. B Neuropsychiatr. Genet. 177, 379–387 (2018).
Medeiros, R. et al. Connecting TNF-α signaling pathways to iNOS expression in a mouse model of Alzheimer’s disease: relevance for the behavioral and synaptic deficits induced by amyloid β protein. J. Neurosci. 27, 5394–5404 (2007).
Nakajima, M. et al. L-17F induces IL-6 via TAK1–NFκB pathway in airway smooth muscle cells. Immun. Inflamm. Dis. 5, 124–131 (2017).
Klatt, A. R. et al. TAK1 downregulation reduces IL-1β induced expression of MMP13, MMP1 and TNF-α. Biomed. Pharmacother. 60, 55–61 (2006).
Xu, Y. R. & Lei, C. Q. TAK1–TABs complex: a central signalosome in inflammatory responses. Front. Immunol. 11, 608976 (2021).
Park, J. H. et al. CD19 CAR T-cell therapy and prophylactic anakinra in relapsed or refractory lymphoma: phase 2 trial interim results. Nat. Med. 7, 1710–1717 (2023).
Takaesu, G. et al. Interleukin-1 (IL-1) receptor-associated kinase leads to activation of TAK1 by inducing TAB2 translocation in the IL-1 signaling pathway. Mol. Cell. Biol. 7, 2475–2484 (2001).
Blonska, M. et al. TAK1 is recruited to the tumor necrosis factor-α (TNF-α) receptor 1 complex in a receptor-interacting protein (RIP)-dependent manner and cooperates with MEKK3 leading to NF-κB activation. J. Biol. Chem. 280, 43056–43063 (2005).
Rodrigues, M., Petrova, T., Tibbs, B., Arthur, J. S. C. & Cohen, P. TAK1 protein kinase activity is required for TLR signalling and cytokine production in myeloid cells. Biochem. J 479, 1891–1907 (2022).
Palleis, C. et al. In vivo assessment of neuroinflammation in 4-repeat tauopathies. Mov. Disord. 36, 883–894 (2021).
Hammers, A. et al. Three-dimensional maximum probability atlas of the human brain, with particular reference to the temporal lobe. Hum. Brain Mapp. 4, 224–247 (2003).
Rauchmann, B. S. et al. Microglial activation and connectivity in Alzheimer disease and aging. Ann. Neurol. 92, 768–781 (2022).
Kreisl, W. C. et al. A genetic polymorphism for translocator protein 18 kDa affects both in vitro and in vivo radioligand binding in human brain to this putative biomarker of neuroinflammation. J. Cereb. Blood Flow Metab. 33, 53–58 (2013).
Rojo, R. et al. Deletion of a Csf1r enhancer selectively impacts CSF1R expression and development of tissue macrophage populations. Nat. Commun. 10, 3215 (2019).
Crowell, H., Zanotelli, V., Chevrier, S. & Robinson, M. CATALYST: Cytometry dATa anALYSis Tools. R package version 1.22.0. https://github.com/HelenaLC/CATALYST (2022).
Bernot, K. M. et al. Eradicating acute myeloid leukemia in a MllPTD/wt:Flt3ITD/wt murine model: a path to novel therapeutic approaches for human disease. Blood 122, 3778–3783 (2013).
Warner, N. L., Moore, M. A. & Metcalf, D. A transplantable myelomonocytic leukemia in BALB-c mice: cytology, karyotype, and muramidase content. J. Natl Cancer Inst. 43, 963–982 (1969).
Schwab, L. et al. Neutrophil granulocytes recruited upon translocation of intestinal bacteria enhance GvHD via tissue damage. Nat. Med. 20, 648–654 (2014).
Leger, M. et al. Object recognition test in mice. Nat. Protoc. 12, 2531–2537 (2013).
Mazziotti, R. et al. Mir-132/212 is required for maturation of binocular matching of orientation preference and depth perception. Nat. Commun. 8, 15488 (2017).
R Core Team. R: a language and environment for statistical computing (R Foundation for Statistical Computing, 2022).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Jeppson, H., Hofmann, H., Cook, D. & Wickham H. ggmosaic: mosaic plots in the ‘ggplot2’ framework. R package version 0.3.3. https://CRAN.R-project.org/package=ggmosaic (2021).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Royal Stat. Soc. B 57, 289–300 (1995).
Bengsch, B. et al. Imaging mass cytometry analysis of CNS tissues from patients with CAR T immune effector cell-associated neurotoxicity syndrome (ICANS). Zenodo https://doi.org/10.5281/zenodo.10695442 (2024).
Acknowledgements
We thank M. Roessler for help with the graphical abstract. We thank the Lighthouse Core Facility for cell sorting and support with flow cytometry, J. Bodenek-Wersing and U. Jagadeshwaran for cell sorting, E. Barleon and the Neuropathology technical team in Freiburg for their help with immunohistochemistry, B. Sauer for tissue processing and staining and B. Fritsch for providing the grip strength meter. This study was supported by the Deutsche Forschungsgemeinschaft (DFG), Germany, project ID 259373024–TRR 167 (to R.Z., D.E., T.B., M. Boerries. and M.P.), SFB-1479 project ID 441891347 to N.K., M. Boerries. and R.Z., SFB1160 TP B09 to N.K., M. Boerries. and R.Z., CRC1453 project ID 431984000-S1 to M. Boerries., project ID 256073931-Z02 to M. Boerries., SFB 850 (Z1 M.B.), DFG individual grant 872/4-1 to R.Z., the European Union EU proposal number ERC-2022-ADG project 101094168 AlloCure (ERC Advanced grant to R.Z.), the Deutsche Krebshilfe (grant number 70114655) and the Jose-Carreras Leukemia foundation grant number DJCLS 09R/2022 (R.Z.), the Leukemia and Lymphoma Society (award number 7030-23 to R.Z.) and Germany’s Excellence Strategy (CIBSS EXC-2189 project ID 390939984 to R.Z. and N.K.). N.K. and D.E. were supported by the Else Kröner-Fresenius-Stiftung (EKFS 2015_A147,2022 EKFS). D.E., M. Schwabenland. and J.B. are supported by the Berta-Ottenstein-Programme for Clinician Scientists, Faculty of Medicine, University of Freiburg. M. Schwabenland. is supported by the IMM-PACT-Programme for Clinician Scientists, Department of Medicine II, Medical Center, University of Freiburg and Faculty of Medicine, University of Freiburg, funded by the DFG (German Research Foundation) 413517907. M. Brendel. was supported by grants from the DFG under Germany’s Excellence Strategy within the framework of the Munich Cluster for Systems Neurology (EXC 2145 SyNergy ID 390857198). We also acknowledge 2858 project number 421887978 to L.v.B., N.L.A. and M. Brendel., SFB TRR 338 452881907 to L.v.B. and M. Subklewe. and Advanced Munich Clinical Scientist Program to L.v.B. B.R.B. is supported by P01 CA 065493. M. Boerries. is supported by the German Federal Ministry of Education and Research (BMBF) within the framework of the e:Med research and funding concept (coNfirm, FKZ 01ZX1708F) and within the Medical Informatics Funding Scheme MIRACUM-FKZ 01ZZ1801B (M.B.) and EkoEstMed–FKZ 01ZZ2015 (G.A.). M.R. was supported by the Berman Foundation. The project was supported by the Lighthouse Core Facility (Medical Faculty, University of Freiburg, 2021/A2-Fol and 2021/B3-Fol; DFG project ID 450392965), DKTK (German Cancer Consortium) project NoviCARAZA and BMBF funding ERA-NET TRANSCAN-3, EC cofunded call 2021, SmartCAR-T and PIXEL (JTC2021) to R.Z., EU project 101119855, exTra.
Author information
Authors and Affiliations
Contributions
J.M.V. performed and analyzed the majority of the mouse and in vitro experiments and helped to develop the overall concept and writing of the paper. F.B., R.C.A., D.E., S.D., D.A., N.E.K., D.S., V.F., A.Z. and L.M.B. helped with in vitro and in vivo experiments. K.K., P.H. and J.P. helped with genetic microglia depletion experiments. M. Schwabenland., F.B., H.S. and B.B. performed IMC. D.P., G.A. and M. Boerries. performed and analyzed the microarray data. C. Chhatbar. and G.M. performed the snRNA-seq. F. Simonetta., C. Capitini, T.J.F. and C.L.M. provided the stably transduced CAR19 T cell lines and B-ALL cell lines. N.R.J., J.N.H.G.H., R.M., R.W., S.V., A.M., R.C., M.R., S.G., E.W., M.T., C.S., M. Stelljes, A.J., J.S., A.R., M.K., L.R., H.E., C.-M.M. and C.R. helped with human data collection. P.T.M., J.B., M.v.B.-B., N.L.A., L. M. Bartos, M. Brendel, A.H., F. Strübing, M. Subklewe and L.v.B. provided TSPO-PET imaging and participant data. M.M. helped with monocyte depletion experiments. T.B. provided help with behavioral studies in mice. N.K., D.E., M. Subklewe., J.D., M.R., B.R.B., S.G. and M.P. contributed to critical analysis of the data, edited the paper and provided reagents. R.Z. developed the overall concept, supervised the experiments, analyzed data and wrote the paper.
Corresponding author
Ethics declarations
Ethics and inclusion statement
All authors that made significant contributions are listed according to the Nature Portfolio guidelines.
Competing interests
M.R. holds multiple patents related to CAR T cell immunotherapy that are managed by the University of Pennsylvania. M.R. has served as a consultant for nanoString, BMS, GSK, GLG, Bayer and AbClon. M.R. receives research funding from AbClon, Beckman Coulter and Oxoford Nano Imaging. M.R. is the scientific founder of viTToria biotherapeutics. R.Z. has served as a consultant for Novartis, Incyte, MNK and Sanofi. S.G. has patents related to CAR therapy with royalties paid from Novartis to the University of Pennsylvania. S.G. is a scientific cofounder and holds equity in Interius Biotherapeutics and Carisma Therapeutics. S.G. is a scientific advisor to Carisma, Cartography, Currus, Interius, Kite, NKILT, Mission Bio and Vor Bio. All other authors declare no conflicts of interest.
Peer review
Peer review information
Nature Cancer thanks Wenjie Luo, Marcela Maus, Michel Sadelain and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 In vivo CD19-CAR-T-cell efficacy.
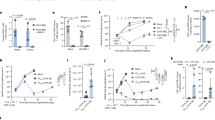
(a, b) The plot represents the percentage of specific lysis of B-NHL (a) B-ALL (b) cells upon co-culture with NT (mice, n = 2 or 3) or CD19-CAR-T-cells (mice, n = 3) at escalating effector to target (E:T) ratios. (c) Representative images showing the lymphoma-derived bioluminescence-signal (BLI) of mice receiving B-NHL + NT-T-cells, or B-NHL + CD19-CAR-T-cells. (d) Scatter plot representing the quantification of tumor burden via BLI in B-NHL + NT (mice, n = 5), B-NHL + CD19-CAR (mice, n = 5) T-cells treated at indicated time points. (e, f) The scatter plots show the percentage (e) or absolute counts (f) of CD19+B220+ cells within the CD45+ cell population in the spleens of B-NHL-bearing mice treated with NT (mice, n = 5) or CD19-CAR (mice, n = 6) T-cells. (g, h) The scatter plots show the percentage (g) or absolute counts (h) of CD19+B220+ cells within the CD45+ cell population in the peripheral blood of B-NHL-bearing mice treated with NT (mice, n = 5) or CD19-CAR (mice, n = 5) T-cells. (i, j) The scatter plot shows the percentage (i), absolute counts (j) of CD19+B220+ cells within the CD45+ cell population in the spleens of B-ALL-bearing mice treated with NT (mice, n = 5) or CD19-CAR (mice, n = 6) T-cells. (k, l) The scatter plots show the percentage (k) or absolute counts (l) of CD19+B220+ cells within the CD45+ cell population in the peripheral blood of B-ALL-bearing mice treated with NT (mice, n = 5) or CD19-CAR (mice, n = 5) T-cells. (m) The scatter plot shows the quantification (percentage) of GFP+ A20 lymphoma cells of all CD45+ population in the CNS (mice, n = 9), BM (mice, n = 9), and spleen (mice, n = 8). (n) The scatter plot shows the quantification (cells per 20x HPF) of GFP+ A20 lymphoma cells that received NT (mice, n = 2) or CD19-CAR (mice, n = 3) T-cells in the meninges. (o) Representative images of the meninges of mice bearing GFP+ A20 lymphoma cells, staining for DAPI. Scale bar: 100 µm. Experiments and staining were performed once (c-l, n), three independent times (a-b, m), results (mean ± s.e.m) were plotted. P-values were calculated using the two-tailed Mann-Whitney- test (d (d3, 22), e-I, k-l) one-way-Kruskal-Wallis-test (m, n) (mixed-effect-analysis)(a, b) two-tailed students-unpaired t-test (d (d0, 15), j).
Extended Data Fig. 2 Cytokines on the protein level in the CNS of mice without tumor or with tumor (B-NHL or B-ALL) after CD19-CART cell transfer.
(a-b) Representative images show Imaris-(Bitplane)-based 3D-reconstruction of Iba-1+ microglia from the cortex of tumor-free mice treated with either NT (mice, n = 3) or CD19-CAR (mice, n = 3) T-cells, as indicated. Scale bar: 5 µm. (b) Scatter plots depict Imaris-based automated quantification of microglial morphology. (c-j) The scatter plots show the quantification (fold change of MFI) of TREM2 (c, g), MHC-II (d, h), TNF (e, i) and GM-CSF (f, j), expression on CD45loCD11b+cells isolated from the CNS of tumor free C57BL/6 (c-f) or BALB/c (g-j) mice, treated with either NT (mice, n = 5) or CD19 CAR (mice, n = 6) T-cells. (k) The scatter plots show the quantification (pg/ml) of TNF, IL-6, CCL2 in the homogenized CNS of BALB/c mice, treated with CD19-CAR-T-cells, 72 h after CD19-CAR-T-cell transfer (mice, n = 6). When indicated mice were B-NHL-(A20)-bearing (mice, n = 6). (l) The scatter plots show the quantification of TNF, IL-6, and CCL2 in the homogenized CNS of B-NHL-(A20)-bearing BALB/c mice treated with CD19-CAR-T-cells. The CNS was isolated at the indicated time points (12 h (mice, n = 5), 24 h (mice, n = 5) and 72 h (mice, n = 6) after CD19-CAR-T- cell transfer). (m) The scatter plot shows the quantification of TNF, IL-6, CCL2 in the homogenized CNS of C57/BL6 mice, treated with CD19-CAR-T-cells, 72 h after CD19-CAR-T-cell transfer (mice n = 6). When indicated mice were B-ALL-(E2A-PBX1)-bearing (mice, n = 6). (n) The scatter plots show the quantification (pg/ml) of TNF, IL-6, CCL2 in the serum of B-NHL- bearing mice. When indicated mice were without T-cell transfer (mice, n = 5), treated with NT (mice, n = 4) or CD19-CAR (mice, n = 5) T cells. Analysis was done 72 h after T-cell transfer. (o-p) The scatter plots show the quantification (pg/ml) of TNF, IL-6, CCL2 in the serum of B-ALL -bearing mice. When indicated mice were treated with NT (mice, n = 4) or CD19-CAR (mice, n = 5) T-cells. Analysis was done 12 h (o) or 24 h (p) after T-cell transfer. Experiments and staining were performed once (a-h), two times (i-x), results (mean ± s.e.m) were plotted. P-values were calculated using two-tailed Mann-Whitney test (b,g,i, k(middle), m (Left, right)), two-tailed students unpaired t test (c-f,h, j, k(left, right), m(right), o, p) or one-way ANOVA (i (Middle, right), n (left, right)) or Kruskal-Wallis-test (i (left), n(middle)).
Extended Data Fig. 3 Expression of VCAM1 and ICAM1 in the CNS of B-NHL-bearing mice after NT T cell or CD19-CART cell transfer.
(a-b) Immunofluorescence imaging representing VCAM1 (red) and DAPI (blue) positive cells in the cortex of B-NHL-bearing mice receiving either B-NHL alone (mice, n = 3), B-NHL + NT-T-cells (mice, n = 5) or B-NHL + CD19-CAR-T-cells(mice, n = 5) (Scale bar: 50 µm). Staining was performed once. (b) Quantification of the VCAM1 positive area occupied in % of the total area. (c-e) The scatter plot shows the VCAM1 (c, d) or ICAM1 (e) expression on CD31+ cells (Fold change of MFI). Analysis of the CNS of BALB/c mice receiving either B-NHL + NT (mice, n = 7-9) or B-NHL + CD19-CAR (mice, n = 4-10) T-cells 24 h or 72 h after T-cell transfer. (f) The scatter plot shows the VCAM1 expression on CD31+ cells (Fold change of MFI). Analysis of the CNS of B-NHL-bearing BALB/c mice receiving CD19-CAR-T-cells. (72 h after T cell transfer). When indicated the mice received control (mice, n = 5) or PLX5622diet (mice, n = 5). (g, h) The scatter plots show ICAM2 (g) and VCAM1 (h) expression on CD31+ cells (Fold change of MFI) in the CNS of tumor free BALB/c mice receiving NT (mice, n = 5), or CD19-CAR (mice, n = 5) T-cells. The analysis was done 72 h after T-cell transfer. (i-t) The scatter plot shows the quantification (fold change of MFI) of TREM2 (i, o), CD80 (j, p) CD11c (k, q), p38MAPK (l, r), TNF (m, s), GM-CSF (n, t) expression on CD45loCD11b+ cells isolated from the CNS of B-NHL bearing (i-n) or B-ALL bearing mice (o-t) at 24 h after treatment with either NT (mice, n = 4-5) or CD19 CAR (mice, n = 4-5) T-cells as indicated. The experiments and staining were performed once (a,f-g, i-t), two times (b-e) and results (mean ± s.e.m.) were plotted. P-values were calculated using two-tailed student’s unpaired t-test (c,e-j, m, n, p, q) or Mann-Whitney test (d, k, l, o, r), Kruskal-wallis test (b).
Extended Data Fig. 4 Marker profiles of CD19-CAR-T-cell (CD45.1+) in B-ALL-bearing or tumor free recipient mice (CD45.2+) after CD19-CAR-T-cell transfer.
(a-d) The scatter plot shows the quantification (fold change of MFI) of CD25 (a) CD69 (b) on CD8+CD45.1+, (c) GM-CSF (d) TNF (d) in CD4+CD45.1+ cells isolated from the spleen of C57BL/6 mice (CD45.2+) treated with CD45.1+ CD19-CAR-T-cells. When indicated mice were B-ALL (E2A-PBX1)-bearing (mice, n = 6 in each group). (e) The scatter plot shows the quantification (absolute counts) of CD3+-T-cells from all CD45.1+ cells isolated from the CNS of B-ALL-bearing(CD45.2+) mice treated with PBS (mice, n = 5) or CD45.1+CD19-CAR-T-cells (mice, n = 14). (f-h)The scatter plot shows the quantification (absolute counts) of CD163+CD11bhi (f) CD206+CD11bhi (g) Lyve1+CD11bhi (h) cells isolated from the CNS of B-NHL or B-ALL-bearing mice treated with NT (mice, n = 5-6) or CD19-CAR (mice, n = 5-6) T-cells. (i-p) The scatter plot shows the quantification (fold change of MFI) of TNF (i, m) and CCL2 (j, n, l, p) phospho-p38-MAPK (k, o) expression in CD11b+ cells isolated from the spleen (i-l) or BM (m-p) of B-NHL or B-ALL-bearing mice treated with PBS (mice, n = 3-6), NT (mice, n = 3-7), or CD19-CAR (mice, n = 4-9) T-cells. (q, r) The scatter plot shows the percentage of CD19+ of CD45+ (q) CD19+ of CD45− (non-hematopoietic) (r) cells in the CNS of B-NHL-bearing mice that received NT (mice, n = 5-11), or CD19-CAR (mice, n = 6-10) T-cells. (s) Heat map of mRNA microarray data showing significantly regulated genes (*) with the annotation IL-1 related genes isolated from sorted microglia from the CNS of B-NHL-bearing mice treated with NT (mice, n = 4) or CD19-CAR (mice, n = 4) T-cells. Color codes represent the Z-score log2 intensity (left panel) and log2 fold change (right panel). (t-x) The scatter plots show the quantification (absolute counts) of CD11c+(t), Ly6C+(u), Ly6G+(v) F4/80+(w) of CD45hiCD11b+cells or CD3+CD45+ cells (x) isolated from the CNS of B-NHL-bearing mice treated with PBS (mice, n = 3-6), NT (mice, n = 5-8), or CD19-CAR (mice, n = 5-9) T-cells. The experiment was performed once (a-d, i, j, l, n), two times (k, m, o, p, s, t-x), three times (e, r), results (mean ± s.e.m) were plotted. P-values were calculated using two-tailed-students unpaired t-test (a-c, e, f-h(middle), r), two-tailed Mann-Whitney-test (d, f-h(left), q) or Kuskal-Wallis-test (i,j,l-p,v,x) or one-way ANOVA (k, t, u, w).
Extended Data Fig. 5 Impact of anti-CCR2 antibody treatment on immune cells in the brain of B-ALL-bearing mice after NT T cell or CD19 CAR T cell transfer.
(a) The scatter plot shows the quantification (absolute counts) of Ly6C+CD45hiCD11b+ cells isolated from the CNS of B-ALL-bearing mice that received CD19-CAR T cells and when indicated treated with vehicle (mice, n = 7) or anti-CCR2 antibody (MC21) (mice, n = 7). (b) The scatter plot shows the percentage of time spent by B-ALL-bearing mice that received CD19-CAR-T-cells and when indicated treated with vehicle (mice, n = 7) or anti-CCR2 antibody (MC21) (mice, n = 7) in exploring a novel object. (c) The scatter plot shows the percentage of open arm entries made by B-ALL-bearing mice that received CD19-CAR-T-cells and when indicated treated with vehicle (mice, n = 7) or anti-CCR2 antibody (MC21) (mice, n = 7) in an elevated plus maze. (d) The scatter plot shows the grip strength of B-ALL-bearing mice that received CD19-CAR-T-cells and when indicated treated with vehicle (mice, n = 7) or anti-CCR2 antibody (MC21) (mice, n = 7). (e-l) Flow cytometry analysis was performed on CD45loCD11b+ cells isolated from the CNS of B-ALL-bearing mice that received CD19-CAR-T-cells and when indicated treated with vehicle (mice, n = 7) or anti-CCR2 antibody (MC21) (mice, n = 7). The scatter plot shows the quantification (Fold change of MFI) of CD11c (e), TREM2 (f), CD80 (g), P2RY12 (h), p38- MAPK (phospho) (i), expression and quantification (absolute counts) of Ly6G+ (j), F4/80+ (k), CD45hi CD11b+ (l) cells from the indicated groups. (m-n) The scatter plot shows the quantification (in %) of Ly6Chigh in CD11b+ Ly6G−cells isolated from the bone marrow (m) or Spleen (n) of B-ALL-bearing mice treated with CD19-CAR-T-cells and when indicated with vehicle or anti-CCR2 antibody (MC21)(mice, n = 7 per group). (o-p) The scatter plot shows the quantification (Fold change of MFI of vehicle) of CCR2 on CD11b+Ly6G−Ly6Chi cells isolated from the BM (o) or Spleen (p) of B-ALL-bearing mice treated with CD19-CAR-T-cells and when indicated with vehicle or anti-CCR2 antibody (MC21)(mice, n = 7 per group). (a-p) The experiments were performed once (mean ± s.e.m.). (a-p) P-values were calculated using two-tailed student’s unpaired t-test (a-n, p) or Mann-Whitney-test (o).
Extended Data Fig. 6 snRNA sequence based analysis of the brain of CAR-T-cell treated mice.
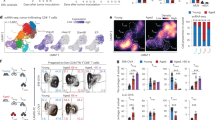
(a) Heat map depicting top genes (rows) enriched per cell type from the CNS of indicated groups. Colors in the heat map correspond to scaled expression. (b) The Dot plot shows marker gene expression of selected genes in cells from the CNS of B-NHL (A20) bearing mice treated with NT compared to CD19-CAR T-cells. (c) UMAP visualization of microglia from the CNS of B-NHL-bearing mice treated with NT-T-cells compared to CD19-CAR-T-cells. Color-coding is based on cluster assignment into 7 clusters. A total of 5,110 cells were included in the UMAP. (d) Marimekko plot showing frequency of microglia derived from the indicated groups across 7 clusters. (e) Dot plot for gene ontology (GO) terms associated with each cluster of microglia. Size and color of the dots correspond to gene ratio and adjusted p-value respectively. (f) Volcano plot showing the differentially expressed genes in microglia from the indicated groups. The –log10-transformed adjusted P value (P-adj, y-axis) is plotted against the log2-transformed fold change in expression between the two treatment groups (x-axis). (g) The violin plot shows the number of genes detected per nucleus of cells from the CNS of the indicated groups. (h-j) The scatter plot shows neurocognitive assessment of non-tumor-bearing C57BL/6 mice treated with NT (mice, n = 5), or CD19-CAR (mice, n = 5) T-cells. Time spent exploring the novel object (h), Time spent in safe zone (i). Neurophenotypic scores from day1-5 post T-cell transfer (j). (k-p) Immunofluorescence imaging representing the NeuN+, DAPI+ cells from the Cortex (k, l, m and o) or hippocampus (n and p) from the CNS of B-NHL or B-ALL-bearing mice treated with either PBS (mice, n = 4-9), NT (mice, n = 4-10) or CD19 CAR (mice, n = 4-8). (k, m, n) Representative images (k) (Scale bar: 100 µm), (m, n) (Scale bar: 400 µm), (l, o, p) quantification. The experiment or staining was performed once (a-j, m-p), twice (k, l) and the results (mean ± s.e.m) were plotted. P-values were calculated using two-tailed students unpaired students t-test (h, i). P-values were calculated using one-way ANOVA (l) with mixed effect analysis (j) or Kruskal-Wallis test (o, p).
Extended Data Fig. 7 Genetic microglia depletion.
(a-f) Neurocognitive tests and flow cytometry analysis were performed on B-ALL-bearing littermate controls (WT) (mice, n = 5-7) or Cx3cr1CreER:Csf1rfl/fl mice (mice, n = 6-8) mice treated with CD19-CAR-T-cells. (g-l) Neurocognitive tests and flow cytometry analysis were performed on B-ALL-bearing littermate controls (WT) (mice, n = 6) or Csf1rΔFIRE/ΔFIRE mice (mice, n = 6) treated with CD19-CAR-T- cells. (m-r) Neurocognitive tests and flow cytometry were performed on B-ALL-bearing littermate controls (WT) (mice, n = 5-8) or HexBCreER: Csf1rfl/fl mice (mice, n = 5-8) treated with CD19-CAR- T-cells. In each model, the following were assessed: Neurocognitive function, Depletion efficacy of CD45loCD11b+cells, CNS-associated-macrophages (Perivascular, border associated and meningeal macrophages) quantification and tumor burden quantification by CD19+B220+CD45+ cells. (a, g, m) The scatter plot shows the percentage of open arm entries by WT littermate or Cx3cr1CreER:Csf1rfl/fl or Csf1rΔFIRE/ΔFIRE or HexBCreER: Csf1rfl/fl mice in an elevated plus maze test. (b, h, n) The scatter plot shows the time spent by WT littermate or Cx3cr1CreER:Csf1rfl/fl or Csf1rΔFIRE/ΔFIRE or HexBCreER: Csf1rfl/fl mice in exploring a novel object (exploration time/total time in %) in a novel object recognition test. (c, i, o) The scatter plot shows the grip strength of WT littermate or Cx3cr1CreER:Csf1rfl/fl or Csf1rΔFIRE/ΔFIRE or HexBCreER: Csf1rfl/fl mice normalized to body weight (N) in a grip strength test. (d, j, p) The scatter plot shows the quantification (absolute counts) of CD45loCD11b+ cells isolated from the CNS of the respective models as indicated. (e, k, q) The scatter plot shows the quantification (absolute counts) of CAMs cells of all CD11b+ cells isolated from the CNS. (f, l, r) The scatter plot shows the quantification (absolute counts) of CD19+B220+ of all CD45+ cells isolated from the spleen. Experiments were performed once (g-l, p-q), two independent times (a-f, m-o, r), and the results (mean ± s.e.m) were pooled. P-values were calculated using two-tailed unpaired student’s t-test (a,b,e (left),g, j, k(middle and right) (m-p, q (middle and right)) or two-tailed Mann-Whitney test (c, d, e(middle and right), f, h, j, k(left), l, q(left), r).
Extended Data Fig. 8 Takinib reduces microglia activation and improves cognitive function.
(a) Heat-map-based on microarray depicting the significantly regulated genes (*) related to the TAK1 pathway from sorted microglia from B-NHL-bearing mice treated with NT (mice, n = 4) or CD19- CAR (mice, n = 4) T-cells. Color codes represents the Z-score log2 intensity. (b-e) The scatter plot shows the quantification (fold change of MFI) of (b) phospho-p38-MAPK (pT180/pY182) (d) p70s6k in microglia from the CNS of B-NHL-bearing mice treated with CD19-CAR-T-cells receiving vehicle (mice, n = 10-11) or takinib (0.5 mg/kg/mouse/day) (mice, n = 11). (c, e) Representative flow cytometry plots. (f) The scatter plot shows the percentage of open arm entries made by B-ALL-bearing mice treated with CD19-CAR-T-cells receiving vehicle (mice, n = 10) or takinib (0.5 mg/kg/mouse/day) (mice, n = 10) with respect to the total time in an elevated plus maze test. (g) The scatter plot shows the percentage of time spent by B-ALL-bearing mice treated with CD19-CAR-T-cells receiving vehicle (mice, n = 10) or takinib (0.5 mg/kg/mouse/day) (mice, n = 10) in exploring a novel object in a novel object recognition test. (h) The scatter plot shows the grip strength normalized to body weight of B-ALL-bearing mice treated with CD19-CAR-T-cells receiving vehicle (mice, n = 10) or takinib (0.5 mg/kg/mouse/day) (mice, n = 10) in a grip strength test. (i-p) The scatter plot shows the quantification of CD45+ (absolute counts) (i, j), CD11b+ (frequency) (k, l), CD3+ (absolute counts) (m, n), NK1.1+ (absolute counts)) (o, p), cells from the spleen and BM respectively of B-NHL (A20)-bearing mice that received CD19-CAR-T-cells and treated with vehicle (mice, n = 5) or takinib (mice, n = 5). (q, r) Imaging of intravascular, extravascular fluorescence via extravasation of FITC-Dextran (indicative of BBB damage) from the CNS of B-ALL-bearing mice that received CD19-CAR-T-cells and treated with vehicle (mice, n = 10) or takinib (mice, n = 10). (q) Representative images. Scale bar- 50 µm, (r) quantification. Experiments or Staining’s were performed once (a, i-p), two (f-h, q-r), three (b, d) independent times and results (mean ± s.e.m.) were pooled. P-values were calculated using two-tailed student’s unpaired t-test (b, d, f-n, p, q) or Mann-Whitney-test (o).
Extended Data Fig. 9 Takinib does not interfere with cytotoxicity of CAR19 T cells.
(a) Representative images showing the bioluminescence signal derived from mice that received B-NHL and vehicle, B-NHL + CD19-CAR-T-cells and vehicle, or B-NHL + CD19-CAR-T-cells and takinib at 48 h post CD19-CAR-T-cell infusion. (b) The scatter plot shows the absolute cell counts of CD19+ cells isolated from the BM of CD19+B-NHL-bearing mice treated with CD19-CAR-T-cells and either vehicle (mice, n = 10) or takinib (mice, n = 10). (c) Scatter plot depicting percentage cell viability of B-NHL cells upon co-culture with NT (mice, n = 3) or CD19-CAR (mice, n = 3) T-cells at an effector-(CD19 CAR/NT T-cells) to target-(A20) (E: T) ratio of 5:1 with 10 µM takinib or vehicle. (d-f) Bar diagram representing the viability of B-NHL (d) and B-ALL (e) NT/CD19-CAR-T-cells cells upon treatment with the indicated concentrations of takinib for 24 h. (Mice, n = 3 in all groups). (g-j) The scatter plots show the quantification (fold change of MFI) of Granzyme B (g), Perforin (h),TNF (i), CD107 (j) expression in CD3+-T-cells derived from a B-NHL and NT/CD19-CAR T-cell co-culture experiment. (Mice, n = 3 in all groups). (k) Survival rates of B-NHL-bearing mice that received PBS (no T cells) or CD19-CAR-T-cells and were treated with vehicle (mice, n = 10 (grey), n = 12 (blue)) or takinib (mice, n = 10 (green), n = 12 (red)) from day 3 to day 10 post B-NHL transfer as indicated. (l) Survival rates of B-NHL-bearing mice that received PBS (no T cells) (mice, n = 10) or CD19-CAR-T-cells (mice, n = 10) and were treated with prednisolone. (m) Survival rates of B-ALL-bearing mice that received PBS or CD19-CAR-T-cells and were treated with vehicle (mice, n = 10 (grey), n = 10 (blue)) or takinib (mice, n = 10 (green) n = 10 (red)) from day 3 to day 10 post B-ALL transfer as indicated. (n) Survival rates of B-ALL-bearing mice that received PBS (no T cells) (mice, n = 10) or CD19-CAR-T-cells (mice, n = 10) and were treated with prednisolone. (o) The heatmap shows unsupervised clustering of 23 myeloid markers obtained from post-mortem FFPE ICANS (patients, n = 7) or age-matched-control (patients, n = 6) samples. Experiments were performed two (b, l, n) three (c-k, m) times and the results (mean ± s.e.m.) were pooled. P-values were calculated using two-tailed Mann-whitney-test (b), one- sided-Kruskal-Wallis-test (c-j) (pairwise-comparisons) (c), two-sided Mantel-Cox-test (k-n).
Extended Data Fig. 10 TSPO-PET imaging to characterize ICANS.
(a) The immunohistochemistry depicts staining for APP (anti-APP staining in brown, H&E) of a CNS biopsy isolated from a patient developing ICANS. The black arrows point towards APP deposits. (b) The immunohistochemistry (staining for APP and Iba-1) depicts analysis of the CNS derived from a patient without ICANS (control patient). The staining was performed once (a-b). (c, d) Visual comparison of TSPO-PET z-scores and white matter hyperintensities in MRI FLAIR sequences. Patient #1 showed hyperintensities of the subcortical white matter in regions matching TSPO-PET increases, whereas patients #2 and #3 displayed no alterations of the MRI FLAIR signal. (d) Mean standardized uptake value in whole-brain parenchyma using TSPO-PET with ([18F]GE-180 control patients (patients, n = 22) and 3 patients that had developed ICANS. P-values were calculated using Mann-whitney test. (e, f) TSPO-PET imaging of patients with HGBCL (axicabtagene ciloleucel) and MCL (brexucabtagene autoleucel) that received CAR-T-cells against CD19 and developed ICANS. Images (e) depict standardized uptake values on a representative transaxial slice of [11C]ER176 PET before (first column) and after CAR-T-cell therapy (second column). Mean standardized uptake value in whole-brain parenchyma (f). Shown are n = 2 independent patients. (g, h) TSPO-PET imaging of patients with multiple myeloma that received CAR-T-cells against B cell maturation antigen (BCMA) (patients, idecabtagene vicleucel, n = 4; ciltacabtagene autoleucel, n = 1) and developed no clinical signs of ICANS. (g) Representative transaxial slices of [11C]ER176 PET before (first column) and after BCMA CAR-T-cell therapy (second column). Mean standardized uptake value in whole-brain parenchyma (h). Filled triangles depict patients were high-affinity binding, filled squares show patients with mixed-affinity binding. Shown are n = 5 independent patients. (d, f, h) P-values were calculated using Mann-Whitney test. (i) Proposed mechanism: CAR T cells that eliminate B-cell malignancies induce endothelial cell activation and BBB leakage, which is followed by microglia activation, leading to increased production of GM-CSF, CCL-2 (MCP-1) and TNF, as well as neurocognitive deficits in B-cell lymphoma-bearing mice. The TGF-β-activated kinase-1 (TAK1)-NF-κB p38-MAPK-pathway is activated in microglia cells upon CD19-CAR-T-cell transfer. Pharmacological TAK1-inhibition or selective genetic Tak1 deletion in microglia results in reduced microglia activation, lower TNF and GM-CSF production, and improved neurocognitive activity.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and legends.
Supplementary Table 1
Supplementary Tables 1–8.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Figs. 1, 2, 4, 7 and 8
Imaging source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Figs. 1–3, 6 and 8
Imaging source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vinnakota, J.M., Biavasco, F., Schwabenland, M. et al. Targeting TGFβ-activated kinase-1 activation in microglia reduces CAR T immune effector cell-associated neurotoxicity syndrome. Nat Cancer (2024). https://doi.org/10.1038/s43018-024-00764-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43018-024-00764-7