Abstract
Availability of the essential amino acid methionine affects cellular metabolism and growth, and dietary methionine restriction has been implicated as a cancer therapeutic strategy. Nevertheless, how liver cancer cells respond to methionine deprivation and underlying mechanisms remain unclear. Here we find that human liver cancer cells undergo irreversible cell cycle arrest upon methionine deprivation in vitro. Blocking methionine adenosyl transferase 2A (MAT2A)-dependent methionine catabolism induces cell cycle arrest and DNA damage in liver cancer cells, resulting in cellular senescence. A pharmacological screen further identified GSK3 inhibitors as senolytics that selectively kill MAT2A-inhibited senescent liver cancer cells. Importantly, combined treatment with MAT2A and GSK3 inhibitors therapeutically blunts liver tumor growth in vitro and in vivo across multiple models. Together, methionine catabolism is essential for liver tumor growth, and its inhibition can be exploited as an improved pro-senescence strategy for combination with senolytic agents to treat liver cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Human liver cancer data were derived from the TCGA Research Network (http://cancergenome.nih.gov/), and were downloaded and analyzed at the Molecular Profiling Facility at the University of Pennsylvania as previously12. For single-cell RNA-seq data analysis, MAT2A transcript levels were compared between 1,497 non-malignant and 7,285 tumorigenic hepatocytes using published dataset under accession code GSE149614 (ref. 65). Expression and the percentage of cells in which gene expression was analyzed using the non-parametric Wilcoxon rank-sum test. Each point represents expression value of each single cell and the yellow dot represents the mean expression for each group. We further compared the mRNA levels of MAT2A in tumorigenic hepatocytes by integrating uninjured hepatocytes (4,800 cells) from a separate dataset under accession code GSE174748 (ref. 66). RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE184065, the analyzed data are deposited in figshare (https://doi.org/10.6084/m9.figshare.22215361), and raw image data files are deposited in figshare (10.6084/m9.figshare.24225508). The software and algorithms for data analyses used in this study are all well established from previous work and are referenced throughout the manuscript. Source data for Figs. 1a,c,i,j, 2a,g,j,l,n, 3c,g, 4c,d and 6b,d,f–h,j and Extended Data Figs. 1a,b,e,f, 2a–c, 4a–d,f,h, 5a,d–f,i,j, 6b, 7a, 8b,f,g and 9b–d,f,g have been provided as Source Data files. All other data supporting the findings of this study are available from the corresponding author upon reasonable requests. Source data are provided with this paper.
Code availability
No custom code was used in this study.
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 71, 7–33 (2021).
The Cancer Genome Atlas Research Network. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341 (2017).
Finn, R. S. et al. Atezolizumab plus bevacizumab in unresectable hepatocellular carcinoma. N. Engl. J. Med. 382, 1894–1905 (2020).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
DeBerardinis, R. J. & Chandel, N. S. We need to talk about the Warburg effect. Nat. Metab. 2, 127–129 (2020).
Faubert, B., Solmonson, A. & DeBerardinis, R. J. Metabolic reprogramming and cancer progression. Science 368, eaaw5473 (2020).
Li, F. & Simon, M. C. Cancer cells don’t live alone: metabolic communication within tumor microenvironments. Dev. Cell 54, 183–195 (2020).
Anderson, N. M. & Simon, M. C. The tumor microenvironment. Curr. Biol. 30, R921–R925 (2020).
Lyssiotis, C. A. & Kimmelman, A. C. Metabolic interactions in the tumor microenvironment. Trends Cell Biol. 27, 863–875 (2017).
Satriano, L., Lewinska, M., Rodrigues, P. M., Banales, J. M. & Andersen, J. B. Metabolic rearrangements in primary liver cancers: cause and consequences. Nat. Rev. Gastroenterol. Hepatol. 16, 748–766 (2019).
Missiaen, R. et al. GCN2 inhibition sensitizes arginine-deprived hepatocellular carcinoma cells to senolytic treatment. Cell Metab. 34, 1151–1167 e1157 (2022).
Li, F. et al. FBP1 loss disrupts liver metabolism and promotes tumorigenesis through a hepatic stellate cell senescence secretome. Nat. Cell Biol. 22, 728–739 (2020).
Wang, C. et al. Inducing and exploiting vulnerabilities for the treatment of liver cancer. Nature 574, 268–272 (2019).
Sanderson, S. M., Gao, X., Dai, Z. & Locasale, J. W. Methionine metabolism in health and cancer: a nexus of diet and precision medicine. Nat. Rev. Cancer 19, 625–637 (2019).
Ducker, G. S. & Rabinowitz, J. D. One-carbon metabolism in health and disease. Cell Metab. 25, 27–42 (2017).
Wang, Z. et al. Methionine is a metabolic dependency of tumor-initiating cells. Nat. Med. 25, 825–837 (2019).
Fang, L. et al. Methionine restriction promotes cGAS activation and chromatin untethering through demethylation to enhance antitumor immunity. Cancer Cell 41, 1118–1133 e1112 (2023).
Gao, X. et al. Dietary methionine influences therapy in mouse cancer models and alters human metabolism. Nature 572, 397–401 (2019).
Machado, M. V. et al. Mouse models of diet-induced nonalcoholic steatohepatitis reproduce the heterogeneity of the human disease. PLoS ONE 10, e0127991 (2015).
Itagaki, H., Shimizu, K., Morikawa, S., Ogawa, K. & Ezaki, T. Morphological and functional characterization of non-alcoholic fatty liver disease induced by a methionine-choline-deficient diet in C57BL/6 mice. Int. J. Clin. Exp. Pathol. 6, 2683–2696 (2013).
Ma, C. et al. NAFLD causes selective CD4+ T lymphocyte loss and promotes hepatocarcinogenesis. Nature 531, 253–257 (2016).
Campisi, J. Aging, cellular senescence, and cancer. Annu. Rev. Physiol. 75, 685–705 (2013).
Rodier, F. et al. Persistent DNA damage signalling triggers senescence-associated inflammatory cytokine secretion. Nat. Cell Biol. 11, 973–979 (2009).
Campisi, J. & d’Adda di Fagagna, F. Cellular senescence: when bad things happen to good cells. Nat. Rev. Mol. Cell Biol. 8, 729–740 (2007).
Li, Z. et al. Methionine metabolism in chronic liver diseases: an update on molecular mechanism and therapeutic implication. Signal Transduct. Target Ther. 5, 280 (2020).
Frau, M., Feo, F. & Pascale, R. M. Pleiotropic effects of methionine adenosyltransferases deregulation as determinants of liver cancer progression and prognosis. J. Hepatol. 59, 830–841 (2013).
Cai, J., Mao, Z., Hwang, J. J. & Lu, S. C. Differential expression of methionine adenosyltransferase genes influences the rate of growth of human hepatocellular carcinoma cells. Cancer Res. 58, 1444–1450 (1998).
Martinez-Chantar, M. L. et al. l-methionine availability regulates expression of the methionine adenosyltransferase 2A gene in human hepatocarcinoma cells: role of S-adenosylmethionine. J. Biol. Chem. 278, 19885–19890 (2003).
Jeggo, P. A., Pearl, L. H. & Carr, A. M. DNA repair, genome stability and cancer: a historical perspective. Nat. Rev. Cancer 16, 35–42 (2016).
Lee, S. & Schmitt, C. A. The dynamic nature of senescence in cancer. Nat. Cell Biol. 21, 94–101 (2019).
Zhang, W. et al. Fluorinated N,N-dialkylaminostilbenes repress colon cancer by targeting methionine S-adenosyltransferase 2A. ACS Chem. Biol. 8, 796–803 (2013).
Li, J. T. et al. Dietary folate drives methionine metabolism to promote cancer development by stabilizing MAT IIA. Signal Transduct. Target Ther. 7, 192 (2022).
Schmitt, C. A., Wang, B. & Demaria, M. Senescence and cancer—role and therapeutic opportunities. Nat. Rev. Clin. Oncol. 19, 619–636 (2022).
Narita, M. et al. Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 113, 703–716 (2003).
Chen, R. Z., Pettersson, U., Beard, C., Jackson-Grusby, L. & Jaenisch, R. DNA hypomethylation leads to elevated mutation rates. Nature 395, 89–93 (1998).
Sato, T., Issa, J. J. & Kropf, P. DNA hypomethylating drugs in cancer therapy. Cold Spring Harb. Perspect. Med. 7, a026948 (2017).
Christman, J. K. 5-Azacytidine and 5-aza-2′-deoxycytidine as inhibitors of DNA methylation: mechanistic studies and their implications for cancer therapy. Oncogene 21, 5483–5495 (2002).
Leite de Oliveira, R. & Bernards, R. Anti-cancer therapy: senescence is the new black. EMBO J. 37, e99386 (2018).
Kirkland, J. L. & Tchkonia, T. Cellular senescence: a translational perspective. EBioMedicine 21, 21–28 (2017).
Toso, A. et al. Enhancing chemotherapy efficacy in Pten-deficient prostate tumors by activating the senescence-associated antitumor immunity. Cell Rep. 9, 75–89 (2014).
Dorr, J. R. et al. Synthetic lethal metabolic targeting of cellular senescence in cancer therapy. Nature 501, 421–425 (2013).
Rizzieri, D. A. et al. An open-label phase 2 study of glycogen synthase kinase-3 inhibitor LY2090314 in patients with acute leukemia. Leuk. Lymphoma 57, 1800–1806 (2016).
Beurel, E., Grieco, S. F. & Jope, R. S. Glycogen synthase kinase-3 (GSK3): regulation, actions, and diseases. Pharmacol. Ther. 148, 114–131 (2015).
Milanovic, M. et al. Senescence-associated reprogramming promotes cancer stemness. Nature 553, 96–100 (2018).
Grassilli, E. et al. Inhibition of GSK3B bypass drug resistance of p53-null colon carcinomas by enabling necroptosis in response to chemotherapy. Clin. Cancer Res. 19, 3820–3831 (2013).
Yang, J. et al. GSK-3β promotes cell survival by modulating Bif-1-dependent autophagy and cell death. J. Cell Sci. 123, 861–870 (2010).
Galluzzi, L. et al. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 25, 486–541 (2018).
Cabon, L. et al. BID regulates AIF-mediated caspase-independent necroptosis by promoting BAX activation. Cell Death Differ. 19, 245–256 (2012).
DeBerardinis, R. J. & Chandel, N. S. Fundamentals of cancer metabolism. Sci. Adv. 2, e1600200 (2016).
DeBerardinis, R. J., Lum, J. J., Hatzivassiliou, G. & Thompson, C. B. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 7, 11–20 (2008).
Marjon, K. et al. MTAP deletions in cancer create vulnerability to targeting of the MAT2A/PRMT5/RIOK1 axis. Cell Rep. 15, 574–587 (2016).
Kryukov, G. V. et al. MTAP deletion confers enhanced dependency on the PRMT5 arginine methyltransferase in cancer cells. Science 351, 1214–1218 (2016).
Kalev, P. et al. MAT2A inhibition blocks the growth of MTAP-deleted cancer cells by reducing PRMT5-dependent mRNA splicing and inducing DNA damage. Cancer Cell 39, 209–224 e211 (2021).
Mavrakis, K. J. et al. Disordered methionine metabolism in MTAP/CDKN2A-deleted cancers leads to dependence on PRMT5. Science 351, 1208–1213 (2016).
Golbourn, B. J. et al. Loss of MAT2A compromises methionine metabolism and represents a vulnerability in H3K27M mutant glioma by modulating the epigenome. Nat. Cancer 3, 629–648 (2022).
Chen, H. A. et al. Senescence rewires microenvironment sensing to facilitate anti-tumor immunity. Cancer Discov. 13, 432–453 (2023).
Marin, I. et al. Cellular senescence is immunogenic and promotes anti-tumor immunity. Cancer Discov. (2022).
Wang, L. et al. High-throughput functional genetic and compound screens identify targets for senescence induction in cancer. Cell Rep. 21, 773–783 (2017).
Wang, L. et al. cFLIP suppression and DR5 activation sensitize senescent cancer cells to senolysis. Nat. Cancer 3, 1284–1299 (2022).
Thorne, C. A. et al. GSK-3 modulates cellular responses to a broad spectrum of kinase inhibitors. Nat. Chem. Biol. 11, 58–63 (2015).
Agrawal, S. et al. El-MAVEN: a fast, robust, and user-friendly mass spectrometry data processing engine for metabolomics. Methods Mol. Biol. 1978, 301–321 (2019).
Reiberger, T. et al. An orthotopic mouse model of hepatocellular carcinoma with underlying liver cirrhosis. Nat. Protoc. 10, 1264–1274 (2015).
GSEA Home. Broad Institute http://software.broadinstitute.org/gsea/index.jsp
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Lu, Y. et al. A single-cell atlas of the multicellular ecosystem of primary and metastatic hepatocellular carcinoma. Nat. Commun. 13, 4594 (2022).
Filliol, A. et al. Opposing roles of hepatic stellate cell subpopulations in hepatocarcinogenesis. Nature 610, 356–365 (2022).
Acknowledgements
We thank the members of the Simon and Li laboratories for their helpful discussions and insights on the manuscript. We are grateful to J. Tobias for help with processing the human TCGA data, and to H. Ji and L. Hu for the critical reading of the manuscript. This work was supported by the National Key R&D Program of China (2022YFA1103900 to F.L.), the National Natural Science Foundation of China (82273223 to F.L., 32270798 to P.L.), and Shanghai Pujiang Program (21PJ1400800 to F.L.), and the National Institutes of Health grants P01CA104838 and R35CA197602 to M.C.S.
Author information
Authors and Affiliations
Contributions
F.L. and M.C.S. conceived the project and designed the experiments, B.K. and M.C.S. supervised the overall study. F.L., P.L., W.M., L. L., N.M.A., N.P.L., M.B., J.P. and A.M. performed the experiments. M.B. provided help with animal husbandry and technical assistance in the mouse experiments. G.W., J.L. and L.Z. performed single cell and regular RNA-seq data analysis. F.L. and P.L. analyzed the data and wrote the manuscript. F.L. and M.C.S. revised the manuscript. All authors revised and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no potential conflicts of interest.
Peer review
Peer review information
Nature Cancer thanks Jason Locasale and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 HCC cells undergo senescence upon methionine starvation.
a, Cell numbers of HepG2, PLC/PRF/5 and SNU449 cells with or without 72-hour methionine (Met) deprivation. b, Cell death quantification (Annexin V+%) of indicated HCC cells with or without 72-hour methionine (Met) deprivation. c, Western blot analysis of Huh7 cells with or without 72-hour methionine (Met) deprivation. Actin was used as a loading control. d, Representative crystal violet staining of control and methionine-deprived Huh7 and Hep3B cells replated in methionine-containing medium for 3-week clonogenicity assays. e, Quantification of cells with >2 53BP1 foci in control and methionine-deprived Huh7 and Hep3B cells. To quantify, 5 random fields containing at least 30 cells were counted and averaged. n = 5 fields for each group. f, Quantification of HepG2, PLC/PRF5 and SNU449 cell numbers when cultured in medium with or without homocysteine (Hcy, 100 µM). Data presented as mean ± s.e.m. of three independent experiments; statistical significance was determined by a two-tailed Student’s t−test (a,b,e,f). Experiments were repeated three times independently, with similar results (c,d).
Extended Data Fig. 2 MAT2A as a potential oncogene in HCC.
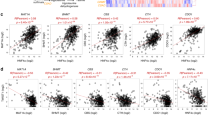
a, Quantification of DAB intensity of MAT2A IHC staining of liver sections from MYCOE;Trp53KO (n = 7 sections from 4 mice for AL, n = 12 sections from 5 mice for tumour) orthotopic model and DEN (n = 7 sections from 3 mice for AL, n = 16 sections from 4 mice for tumour) model. b, The frequency and normalized UMI of MAT2A transcripts in 7,285 pro-tumorigenic hepatocytes as compared to 1497 non-malignant hepatocytes (left) or 4800 uninjured hepatocytes from healthy livers (Right) using public single cell RNA-seq data. c, GSEA plots of indicated gene sets based on MAT2A transcript levels in 374 HCC patient samples from TCGA dataset. MAT2A-high: top 50%, n = 187; MAT2A-low: bottom 50%, n = 187. Data presented as mean ± s.e.m.; statistical significance was determined by a two-tailed Student’s t-test (a), one-side non-parametric Wilcoxon rank-sum test (b) or the Kolmogorov Smirnov (K-S) test (c).
Extended Data Fig. 3 Genetic MAT2A inhibition induces DNA damage and promotes SASP gene expression.
a, Representative immunofluorescence images of Huh7-sgR26, Huh7-sgMAT2A-1 and Huh7-sgMAT2A-2 cells stained with 53BP1 (red), γH2AX (red) and DAPI (blue). Scale bar: 50 µm. b, Quantification of cells with >2 53BP1 foci in Huh7-sgR26 (n = 5 200x fields), Huh7-sgMAT2A-1 (n = 5 200x fields) and Huh7-sgMAT2A-2 (n = 5 200x fields) cells. To quantify, 5 random 200x fields containing at least 30 cells were counted and averaged. c, qRT-PCR analysis of indicated gene expression in Huh7-sgR26, Huh7-sgMAT2A-1 and Huh7-sgMAT2A-2 cells. Data are presented as the mean ± s.e.m of three independent experiments; statistical significance was determined by a two-tailed Student’s t-test (b,c). Experiments were repeated three times independently, with similar results (a).
Extended Data Fig. 4 MAT2A inhibition by FIDAS-5 induces senescence in liver cancer cells.
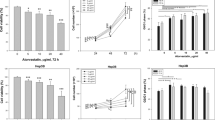
a, Cell viabilities of Huh7 and Hep3B cells treated with indicated doses of FIDAS-5 for 72 hours. b, Huh7 and Hep3B cell growth curves in the presence of 1 µM FIDAS-5. c, Relative SAM and SAH abundance determined by LC-MS analysis of control (n = 4) and FIDAS-5 (n = 4) treated Huh7 cells for 72 hours. d, Cell death quantification (Annexin V+%) of Huh7 and Hep3B cells with or without 72-hour FIDAS-5 exposure (Huh7: 5 µM; Hep3B:1 µM). e, Representative immunofluorescence images of vehicle control and FIDAS-5-induced senescent (FIS) Huh7 cells stained with 53BP1 (red), γH2AX (red) and DAPI (blue). Scale bar: 50 µm. f, Quantification of cells with >2 53BP1 foci in control (n = 5 200x fields) and FIS (n = 5 200x fields) Huh7 cells. To quantify, 5 random fields containing at least 30 cells were counted and averaged. g, Representative crystal violet staining of control and FIS Huh7 cells replated in growth medium for 3-week clonogenicity assays. h, qRT-PCR analysis of indicated gene expression in control and FIS Huh7 cells. i, j, Representative SA-β-gal staining (i) and quantification (j) of vehicle control (Veh) and FIDAS-5 treated TOV21G cells. To quantify in (j), 5 random 100x fields were counted and averaged. n = 5 200x fields for each group. Scale bar: 100 µm. k, Mouse body weight curves over time in vehicle control (Veh, n = 7 female BAB/c mice) and FIDAS-5 (n = 5 female BAB/c mice) treatment groups. Data presented as mean ± s.e.m. of three independent experiments (a,b,d,f,h,j,k) or mean ± s.e.m.(c); statistical significance was determined by a two-tailed Student’s t-test (a-d,f,h,j,k). Experiments were repeated three times independently, with similar results (e,g,i).
Extended Data Fig. 5 Characterization of early FIDAS-5 treated HCC cells.
a, Relative numbers of Huh7 cells cultured in growth medium with indicated ROS scavengers for 72 hours. b, Western blot analysis of indicated histone methylation marks in Huh7 cells treated with indicated concentration of FIDAS-5 for indicated time. H3 was used as loading control. c, d, Representative immunofluorescence images (c) and statistical analysis (d) of Veh and treatment-induced senescent Huh7 cells stained with H3K9me3 antibody (green). To quantify in d, 10 random fields containing at least 80 cells were counted and averaged for the percentage of H3K9me3+ nuclei. n = 10 200x fields for each group. Scale bar: 50 µm. e, Relative gene expression by qRT-PCR analysis of FIDAS-5 and vehicle treated Huh7 and Hep3B cells for 48 hours. f, Quantification of cells with >2 53BP1 foci in FIDAS5-treated Huh7, Hep3B cells for indicated time. To quantify, 5 random fields containing at least 30 cells were counted and averaged. n = 5 200x fields for each group. g, Western blot analysis of p21 and GAPDH from Huh7-sgR26, Huh7-sgCDKN1A-1, Huh7-sgCDKN1A-2 and Huh7-sgCDKN1A-3 cells. h, i, Representative SA-β-gal staining (h) and quantification (i) of Huh7 cells and FIDAS-5 treated Huh7-sgR26, Huh7-sgCDKN1A-1 and Huh7-sgCDKN1A-2 cells. Scale bar: 100 µm. j, Relative cell numbers of FIDAS-5 treated Huh7-sgR26, Huh7-sgP21-1 and Huh7-sgP21-2 cells. Data presented as mean ± s.e.m. of three independent experiments; statistical significance was determined by a two-tailed Student’s t-test (a,e,f,i,j). Experiments were repeated three times independently, with similar results (b,c,g,h).
Extended Data Fig. 6 DNA damage in early decitabine-treated HCC cells.
a, Representative immunofluorescence images of vehicle control and 48-hour decitabine-treated Huh7 and Hep3B cells stained with 53BP1 (red), γH2AX (red) and DAPI (blue). The arrows indicate cytoplasmic DNA. Scale bar: 10 µm. b, Quantification of cells with >2 53BP1 foci or >2 γH2AX foci in decitabine-treated Huh7 and Hep3B cells for 48 h. To quantify, 5 random fields containing at least 30 cells were counted and averaged. n = 5 200x fields for each group. c, Representative crystal violet staining of control and decitabine-induced senescent Huh7 cells replated in growth medium for 3-week clonogenicity assays. Data presented as mean ± s.e.m. of three independent experiments; statistical significance was determined by a two-tailed Student’s t-test (b). Experiments were repeated three times independently, with similar results (a,c).
Extended Data Fig. 7 GSK3 inhibition induces cell death in TIS liver cancer cells.
a, Relative cell numbers of Huh7 cells treated with different doses of BIO and CHIR99021 for 72 hours. b, Representative crystal violet staining of control and FIS Huh7 and Hep3B cells treated with indicated doses of CHIR98014 for 6 days. c, Representative crystal violet staining of vehicle control and TIS Huh7 cells treated with indicated doses of LY2090314 for 6 days. d, Representative crystal violet staining of Huh7-sgR26, Huh7-sgMAT2A-1 and Huh7-sgMAT2A-7 cells treated with 0.5 µM LY2090314 for 6 days. e, Representative crystal violet staining of Huh6 cells treated with indicated drug combinations for 6 days. Data presented as mean ± s.e.m. of three independent experiments; statistical significance was determined by a two-tailed Student’s t-test (a). Experiments were repeated three times independently, with similar results (b-e).
Extended Data Fig. 8 GSK3 maintains viability of TIS liver cancer cells.
a, Western blot analysis of GSK3A, GSK3B and GAPDH in lysates from control and FIS Huh7 cells with or without 48-hour LY2090314 treatment at indicated doses. b, Normalized TOP Flash and FOP Flash luciferase activity in control and FIS Huh7 cells. c, Subcellular fractionation and western blot analysis of indicated proteins in vehicle and FIS Huh7 cells. d, Kaplan–Meier overall survival plots stratified by GSK3A or GSK3B mRNA levels from HCC TCGA database. e, Western blot analysis of GSK3A, GSK3B and GAPDH in lysates from Huh7 cells expressing indicated sgRNAs. f, Cell growth curves of Huh7 cells expressing indicated sgRNAs. g, Cell death quantification (Annexin V+%) of Huh7 cells expressing indicated sgRNAs. h, Western blot analysis of GSK3A, GSK3B and GAPDH from Huh7 cells expressing indicated shRNAs. i, Representative crystal violet staining of Huh7-shCtrl, Huh7-shGSK3A/B-1 and Huh7-shGSK3A/B-2 cells with or without 0.5 µM LY2090314 treatment for 6 days. j, Flow cytometry plots of Annexin-V/PI apoptosis assay in Huh7 cells treated with indicated doses of LY2090314 for 72 hours. k, Western blot analysis of GSK3A, GSK3B and GAPDH from FIS Huh7 cells expressing indicated siRNAs. Data presented as mean ± s.e.m. of three independent experiments; statistical significance was determined by a two-tailed Student’s t-test (b,f,g) or log-rank Mantel–Cox test (d). Experiments were repeated three times independently, with similar results (a,c,e,h-k).
Extended Data Fig. 9 Combined MAT2A and GSK3 inhibition limits liver tumor growth.
a, Representative crystal violet staining of Huh7 cells treated with indicated doses of drug combination for 12 days. b-d, Quantification of end point Huh7 and Hep3B xenograft tumor weight (b), volume (c) and mouse body weight (d) from vehicle control (Veh), FIDAS-5, LY2090314 and combined FIDAS-5/LY2090314 treatment groups. Huh7: n = 6 female BAB/c mice for each group. Hep3B: n = 5 female NSG mice for Veh, n = 6 female NSG mice for FIDAS-5, n = 6 female NSG mice for LY2090314, n = 7 female NSG mice for combination group. e, f, Representative IHC images (e) and quantification (f) of PCNA and γH2AX staining in Hep3B xenograft tumor sections from indicated treatment groups. To quantify, 3–5 representative images with a ×200 field of view (FOV) were used for quantification using Image J and averaged for each animal. n = 6 mice for each group. Scale bar: 100 µm. g, Body weight change of vehicle control (n = 3 male mice) and combined FIDAS-5/LY2090314 (n = 3 male mice)-treated C57BL/6 J mice. h, Representative HE staining of liver sections from vehicle and combined FIDAS-5/LY2090314-treated wild type mice. Scale bar: 100 µm. Data presented as mean ± s.e.m. (b-g); statistical significance was determined by a two-tailed Student’s t-test (b-f) Experiments were repeated three times independently, with similar results (a).
Supplementary information
Supplementary Information
Supplementary Fig. 1. Representative SAM and SAH chromatogram peaks from liquid chromatography–mass spectrometry.
Supplementary Tables 1–3
Supplementary Tables 1–3.
Supplementary Table 4
Supplementary Table 4.
Source data
Source Data
Unprocessed western blots.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
IF images.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
IF images.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
IF images.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
IF images.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
IF images.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
IF images.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
IF images.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
IF images.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
IF images.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
IF images.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
IF images.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
IF images.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
IF images.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
IF images.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, F., Liu, P., Mi, W. et al. Blocking methionine catabolism induces senescence and confers vulnerability to GSK3 inhibition in liver cancer. Nat Cancer 5, 131–146 (2024). https://doi.org/10.1038/s43018-023-00671-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-023-00671-3