Abstract
To improve our understanding of longstanding disparities in incidence and mortality in lung cancer across ancestry, we performed a systematic comparative analysis of molecular features in tumors from African Americans (AAs) and European Americans (EAs). We find that lung squamous cell carcinoma tumors from AAs exhibit higher genomic instability—the proportion of non-diploid genome—aggressive molecular features such as chromothripsis and higher homologous recombination deficiency (HRD). In The Cancer Genome Atlas, we demonstrate that high genomic instability, HRD and chromothripsis among tumors from AAs is found across many cancer types. The prevalence of germline HRD (that is, the total number of pathogenic variants in homologous recombination genes) is higher in tumors from AAs, suggesting that the somatic differences observed have genetic ancestry origins. We also identify AA-specific copy-number-based arm-, focal- and gene-level recurrent features in lung cancer, including higher frequencies of PTEN deletion and KRAS amplification. These results highlight the importance of including under-represented populations in genomics research.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Human TCGA cohort mutation data were derived from the publicly available mSignatureDB database (http://tardis.cgu.edu.tw/msignaturedb/). For the corresponding samples and copy-number profiles, level 3 segmented files were retrieved from the firehose pipeline (https://gdac.broadinstitute.org/) where a consistent version of reference hg19 was used. The NCI-MD data were derived from patients enrolled in the ongoing NCI-MD Case-Control Study. All relevant data in this work are available upon reasonable request, except for the TCGA pathogenic variant calls that required dbGaP controlled access and any sequence information that would make it possible to identify study participants. Anonymized level 3 segmented files for each sample, in addition to the raw files for copy-number profiles of the NCI-MD patients and their corresponding expression profiles, are deposited in dbGAP with the accession number phs001895.
Code availability
We used open-source R version 3.6 to generate the figures. Wherever required, commercially available Adobe Illustrator 23.0.3 (2019) was used to create the figure grids. All of the scripts for analysis and figure production were built in-house and are provided on GitHub at https://github.com/sanjusinha7/Scripts_MolCharAAvsEA.
References
DeSantis, C. E., Miller, K. D., Goding Sauer, A., Jemal, A. & Siegel, R. L. Cancer statistics for African Americans, 2019. CA Cancer J. Clin. 69, 211–233 (2019).
Yuan, J. et al. Integrated analysis of genetic ancestry and genomic alterations across cancers. Cancer Cell 34, 549–560.e9 (2018).
Sherman, R. M. et al. Assembly of a pan-genome from deep sequencing of 910 humans of African descent. Nat. Genet. 51, 30–35 (2018).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2019. CA Cancer J. Clin. 69, 7–34 (2019).
Ryan, B. M. Lung cancer health disparities. Carcinogenesis 39, 741–751 (2018).
Mitchell, K. A., Zingone, A., Toulabi, L., Boeckelman, J. & Ryan, B. M. Comparative transcriptome profiling reveals coding and noncoding RNA differences in NSCLC from African Americans and European Americans. Clin. Cancer Res. 23, 7412–7425 (2017).
Wallace, T. A. et al. Tumor immunobiological differences in prostate cancer between African-American and European-American men. Cancer Res. 68, 927–936 (2008).
Guda, K. et al. Novel recurrently mutated genes in African American colon cancers. Proc. Natl Acad. Sci. USA 112, 1149–1154 (2015).
Chaisaingmongkol, J. et al. Common molecular subtypes among Asian hepatocellular carcinoma and cholangiocarcinoma. Cancer Cell 32, 57–70.e3 (2017).
Foster, J. M. et al. Cross-laboratory validation of the OncoScan® FFPE Assay, a multiplex tool for whole genome tumour profiling. BMC Med. Genom. 8, 5 (2015).
Knijnenburg, T. A. et al. Genomic and molecular landscape of DNA damage repair deficiency across the cancer genome Atlas. Cell Rep. 23, 239–254.e6 (2018).
Telli, M. L. et al. Homologous recombination deficiency (HRD) score predicts response to platinum-containing neoadjuvant chemotherapy in patients with triple-negative breast cancer. Clin. Cancer Res. 22, 3764–3773 (2016).
Swisher, E. M. et al. Rucaparib in relapsed, platinum-sensitive high-grade ovarian carcinoma (ARIEL2 Part 1): an international, multicentre, open-label, phase 2 trial. Lancet Oncol. 18, 75–87 (2017).
Jin, Y., Schaffer, A. A., Feolo, M., Holmes, J. B. & Kattman, B. L. GRAF-pop: a fast distance-based method to infer subject ancestry from multiple genotype datasets without principal components analysis. G3 (Bethesda) 9, 2447–2461 (2019).
Ratnaparkhe, M. et al. Defective DNA damage repair leads to frequent catastrophic genomic events in murine and human tumors. Nat. Commun. 9, 4760 (2018).
Zhang, C. Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
Zhang, C. Z., Leibowitz, M. L. & Pellman, D. Chromothripsis and beyond: rapid genome evolution from complex chromosomal rearrangements. Gene Dev. 27, 2513–2530 (2013).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Taylor, A. M. et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell 33, 676–689.e3 (2018).
Polimanti, R. et al. Haplotype differences for copy number variants in the 22q11.23 region among human populations: a pigmentation-based model for selective pressure. Eur. J. Hum. Genet. 23, 116–123 (2015).
Burnside, R. D. 22q11.21 deletion syndromes: a review of proximal, central, and distal deletions and their associated features. Cytogenet. Genome Res. 146, 89–99 (2015).
Baell, J. B. et al. Inhibitors of histone acetyltransferases KAT6A/B induce senescence and arrest tumour growth. Nature 560, 253–257 (2018).
Brim, H. et al. Genomic aberrations in an African American colorectal cancer cohort reveals a MSI-specific profile and chromosome X amplification in male patients. PLoS ONE 7, e40392 (2012).
Craig, D. W. et al. Genome and transcriptome sequencing in prospective metastatic triple-negative breast cancer uncovers therapeutic vulnerabilities. Mol. Cancer Ther. 12, 104–116 (2013).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Ma, J., Setton, J., Lee, N. Y., Riaz, N. & Powell, S. N. The therapeutic significance of mutational signatures from DNA repair deficiency in cancer. Nat. Commun. 9, 3292 (2018).
Huang, K. L. et al. Pathogenic germline variants in 10,389 adult cancers. Cell 173, 355–370.e14 (2018).
Wang, Y. et al. Rare variants of large effect in BRCA2 and CHEK2 affect risk of lung cancer. Nat. Genet. 46, 736–741 (2014).
Palles, C. et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat. Genet. 45, 136–144 (2013).
Timms, K. M. et al. Association of BRCA1/2 defects with genomic scores predictive of DNA damage repair deficiency among breast cancer subtypes. Breast Cancer Res. 16, 475 (2014).
Campbell, J. D. et al. Comparison of prevalence and types of mutations in lung cancers among black and white populations. JAMA Oncol. 3, 801–809 (2017).
Robbins, H. A., Engels, E. A., Pfeiffer, R. M. & Shiels, M. S. Age at cancer diagnosis for blacks compared with whites in the United States. J. Natl Cancer Inst. 107, dju489 (2015).
Jiang, Y. et al. PARP inhibitors synergize with gemcitabine by potentiating DNA damage in non-small-cell lung cancer. Int. J. Cancer 144, 1092–1103 (2019).
Ramalingam, S. S. et al. Randomized, placebo-controlled, phase II study of veliparib in combination with carboplatin and paclitaxel for advanced/metastatic non-small cell lung cancer. Clin. Cancer Res. 23, 1937–1944 (2017).
Kadouri, L. et al. Homologous recombination in lung cancer, germline and somatic mutations, clinical and phenotype characterization. Lung Cancer 137, 48–51 (2019).
Yeh, C. H., Bellon, M. & Nicot, C. FBXW7: a critical tumor suppressor of human cancers. Mol. Cancer 17, 115 (2018).
Zhang, Q. et al. FBXW7 facilitates nonhomologous end-joining via K63-linked polyubiquitylation of XRCC4. Mol. Cell 61, 419–433 (2016).
Yumimoto, K. et al. F-box protein FBXW7 inhibits cancer metastasis in a non-cell-autonomous manner. J. Clin. Invest. 125, 621–635 (2015).
Kytola, V. et al. Mutational landscapes of smoking-related cancers in Caucasians and African Americans: precision oncology perspectives at Wake Forest Baptist Comprehensive Cancer Center. Theranostics 7, 2914–2923 (2017).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
McCabe, N. et al. Deficiency in the repair of DNA damage by homologous recombination and sensitivity to poly(ADP-ribose) polymerase inhibition. Cancer Res. 66, 8109–8115 (2006).
Carter, H. et al. Interaction landscape of inherited polymorphisms with somatic events in cancer. Cancer Discov. 7, 410–423 (2017).
Enewold, L. et al. Serum concentrations of cytokines and lung cancer survival in African Americans and Caucasians. Cancer Epidemiol. Biomarkers Prev. 18, 215–222 (2009).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl J. Med. 376, 2109–2121 (2017).
Andor, N. et al. Pan-cancer analysis of the extent and consequences of intratumor heterogeneity. Nat. Med. 22, 105–113 (2016).
Huang, P. J. et al. mSignatureDB: a database for deciphering mutational signatures in human cancers. Nucleic Acids Res. 46, D964–D970 (2018).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010).
Yi, K. & Ju, Y. S. Patterns and mechanisms of structural variations in human cancer. Exp. Mol. Med. 50, 98 (2018).
Maciejowski, J., Li, Y., Bosco, N., Campbell, P. J. & de Lange, T. Chromothripsis and kataegis induced by telomere crisis. Cell 163, 1641–1654 (2015).
Acknowledgements
We thank C. Harris for many insightful discussions. S.S. acknowledges the support of the NCI-UMD Cancer Research Training Fellowship. This research was supported by the Intramural Research Program of the National Institutes of Health, National Cancer Institute.
Author information
Authors and Affiliations
Contributions
S.S., K.A.M. and B.M.R. conceived and designed the study. S.S., K.A.M. and A.A.S. developed the methodology. K.A.M., A.Z., B.M.R., S.S., A.A.S. and J.S.L. acquired the data. S.S., K.A.M., B.M.R., A.A.S., J.S.L., A.Z., E.B., N.S. and E.R. analyzed and interpreted the data. S.S., K.A.M., B.M.R., A.A.S., J.S.L., A.Z., E.B., N.S. and E.R. wrote, reviewed and/or revised the manuscript. K.A.M., N.S. and B.M.R. provided administrative, technical or material support. B.M.R. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Unsupervised inference of genetic ancestry of lung adenocarcinoma (LUAD) and lung squamous carcinoma (LUSC) tumor samples from the NCI-MD cohort (n=222 patients).
A principal component analysis (PCA) of ancestry-associated single nucleotide polymorphisms (SNPs) (46,217) was performed with rank=2 and the two PCs are shown here. These PCs were used in unsupervised clustering via support vector clustering (SVC) to identify two distinguishable clusters. For each cluster, the respective predominant self-reported race observed in the cluster was considered as the cluster ancestry identity, termed as inferred ancestry. This inferred ancestry is concordant with self-reported for 98.6% cases, where two AAs (African Americans) were potentially misclassified as EA (European American) and one EA as AA.
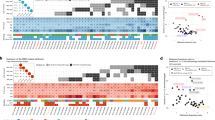
Extended Data Fig. 2 Chromothripsis (CHTP) in European Americans (EAs) and African Americans (AAs) and chromosome distribution in The Cancer Genome Atlas (TCGA) and NCI-MD cohorts.
A) CHTP frequency distribution in AAs and EAs in various cancer types across TCGA. B) CHTP frequency across chromosomes for NCI-MD cohort in LUSC (lung squamous Carcinoma) and LUAD (lung adenocarcinoma). C) CHTP frequency across chromosomes for various cancer types in the TCGA cohort. In Panel A, a one-sided Fisher test has been performed to test whether chromothripsis frequency is higher in AAs or not.
Extended Data Fig. 3 Landscape of somatic copy number alterations frequencies of lung cancer driver genes in lung squamous carcinoma (LUSC) and lung adenocarcinoma (LUAD) from European Americans (EAs) and African Americans (AAs) in The Cancer Genome Atlas (TCGA).
Frequencies in tumors from EAs and AAs with LUAD and LUSC from TCGA were plotted with the blue diagonal line as a null axis (no alteration frequency difference). The diagonal dashed line denotes the null line with points falling away from this line indicating chromosome arms with alteration frequency differences across populations. Del=deletion, Amp=amplification.
Extended Data Fig. 4 Effect of somatic copy number alteration (SCNA) on expression for cancer driver genes in the NCI-MD cohort (n=91 patients).
Effect of SCNA on expression for driver genes is plotted for lung cancer driver genes whose somatic copy number alteration frequency across populations are significantly different. Two-sided Spearman correlation significance with Rho is provided with the corresponding gene name before multiple testing correction. Here, in the box plot, the center line denotes the median, the box indicates the interquartile range and the black line represents the rest of the distribution, except for points that are determined to be “outliers”, 1.5 times the interquartile range.
Extended Data Fig. 5 Landscape of genomic instability (GI) in African Americans (AAs) and European Americans (EAs) in 23 cancer types from The Cancer Genome Atlas (TCGA) cohort.
Here, GI is quantified and presented stratified by genetic ancestry for 23 cancer types where the sample size for each cancer type is provided on the x-axis. First, cancer types are categorized by cell type or tissue of origin, if possible, where defined groups are pan-squamous (squamous cell derived tumors), pan-adeno (glandular structures in epithelial tissue derived tumors), pan-kidney (tumors originating in the kidney), and rest (referring to cancer types that cannot be categorized and includes LAML, THYM, GBM, LGG, SARC, BRCA, LIHC, OV, TCGT, THCA and UCEC; Refer here for reference to each cancer type: https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables/tcga-study-abbreviations). Second, additional categorization was performed based on tissue type (where solid is derived from solid tumors and neural-crest and Hema & Lymph—hematologic and lymphatic tumors). A two-sided Wilcoxon Rank-sum test has been performed within each cancer type and significance before multiple testing correction is provided. Here, in the box plot, the center line denotes the median, the box indicating the interquartile range and the black line represents the rest of the distribution, except for points that are determined to be “outliers”, 1.5 times the interquartile range.
Extended Data Fig. 6 Gain and loss genomic instability (GI) burden in European Americans (AAs) and European Americans (EAs) in 23 cancer types from The Cancer Genome Atlas (TCGA).
a, Somatic copy number alteration (SCNA)-gain and b SCNA-loss based GI are quantified and presented stratified by genetic ancestry for 23 cancer types in TCGA where sample size for each cancer type is provided on the x-axis. First, cancer types are categorized by cell type or tissue of origin, if possible, where defined groups are pan-squamous (squamous cell derived tumors), pan-adeno (glandular structures in epithelial tissue derived tumors), pan-kidney (tumors originating in the kidney), and rest (referring to cancer types that cannot be categorized and includes LAML, THYM, GBM, LGG, SARC, BRCA, LIHC, OV, TCGT, THCA and UCEC; Refer here for reference to cancer types: https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables/tcga-study-abbreviations). Second, additional categorization was performed based on tissue type (where solid is derived from solid tumors and neural-crest and Hema & Lymph—hematologic and lymphatic tumors). A two-sided Wilcox Rank-sum test has been performed within each cancer type and significance before multiple testing correction is provided. Here, in the box plot, the center line denotes the median, the box indicates the interquartile range and the black line represents the rest of the distribution, except for points that are determined to be “outliers”, 1.5 times the interquartile range.
Extended Data Fig. 7 Various measures of homologous recombination deficiency (HRD) in pan-cancer in European Americans (EAs) and African Americans (AAs) from The Cancer Genome Atlas (TCGA) (total N=6,966 patients; [AA=770, EA=6,196]).
HRD is quantified and presented via score based on (a) number of Loss of heterozygosity (LOH) events, (b) telomere allelic imbalance (AIL), (c) large-scale state transitions (LST), (d) sum of previous three defined as “genomic scar” and (e) mutation signature 3 contribution. A one-sided Wilcoxon Rank-sum test has been performed to test whether HRD in tumors from AAs is higher than in EAs. Here, in the box plot, the center line denotes the median, the box indicates the interquartile range and the black line represents the rest of the distribution, except for points that are determined to be “outliers”, 1.5 times the interquartile range.
Extended Data Fig. 8 Various measures of homologous recombination deficiency (HRD) across 23 cancer types in European Americans (EAs) and African Americans (AAs) from The Cancer Genome Atlas (TCGA).
HRD is quantified and presented via various scores. a, Number of (loss of heterozygosity) LOH events, (b) telomere allelic imbalance (AIL), (c) large-scale state transitions (LST), (d) scaled net sum of previous three defined as “genomic scar” and (e) mutation signature 3 contribution in AAs and EAs in various cancer types in TCGA where sample size for each cancer type is provided on the x-axis. First, cancer types are categorized by cell type or tissue of origin, if possible, where defined groups are pan-squamous (squamous cell derived tumors), pan-adeno (glandular structures in epithelial tissue derived tumors), pan-kidney (tumors originating in the kidney), and rest (referring to cancer types that cannot be categorized and includes LAML, THYM, GBM, LGG, SARC, BRCA, LIHC, OV, TCGT, THCA and UCEC; Refer here for reference to cancer types: https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables/tcga-study-abbreviations). Second, additional categorization was performed based on tissue type (where solid is derived from solid tumors and neural-crest and Hema & Lymph—hematologic and lymphatic tumors). One-sided Wilcox Rank-sum test has been performed within each cancer type to test whether HRD is higher in AA than EA and significance before multiple testing correction is provided. Here, in the box plot, the center line denotes the median, the box indicates the interquartile range and the black line represents the rest of the distribution, except for points that are determined to be “outliers”, 1.5 times the interquartile range.
Extended Data Fig. 9 Genomic instability (GI), homologous recombination deficiency (HRD) and Chromothripsis (CHTP) across the cancer Genome Atlas (TCGA) with race classified by inferred ancestry.
a, HRD based on number large-scale state transitions (LST) (b), telomere allelic imbalance (AIL) (c), number of LOH events (d) and scaled net sum of previous three defined as “genomic scar” (e), and CHTP (f) is quantified and presented in European Americans (EAs) and African Americans (AAs) in various cancer types in TCGA where sample size for each cancer type is provided on the x-axis. First, cancer types are categorized by cell type or tissue of origin, if possible, where defined groups are pan-squamous (squamous cell derived tumors), pan-adeno (glandular structures in epithelial tissue derived tumors), pan-kidney (tumors originating in the kidney), and rest (referring to cancer types that cannot be categorized and includes LAML, GBM, LGG, BRCA, OV, and UCEC). Refer here for reference to cancer types for reference: https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables/tcga-study-abbreviations). Second, additional categorization was performed based on tissue type (where solid is derived from solid tumors and neural-crest and Hema & Lymph—hematologic and lymphatic tumors). Across, panels a-e, two-sided Wilcoxon Rank-sum test has been performed for each cancer type and significance before multiple testing correction is provided. In the corresponding panels, the box plot, the center line denotes the median, the box indicates the interquartile range and the black line represents the rest of the distribution, except for points that are determined to be “outliers”, 1.5 times the interquartile range. In panel f, one-sided Fisher test has been performed to test whether chromothripsis frequency is higher in AA or not.
Extended Data Fig. 10 Prevalence of germline homologous recombination deficiency (HRD) proportion in European Americans (EAs) and African Americans (AAs) patients from the cancer Genome Atlas (TCGA) cohort.
Germline HRD (see methods) is quantified and presented in AAs and EAs for 17 cancer types with at least 30 AA samples in TCGA, where HRD is defined by 88 hallmark genes provided in Table S14, with the respective AA and EA patients included in each group. Refer here for reference to cancer types: https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables/tcga-study-abbreviations). Further, the number of samples in each group is provided in Table S12. AAs are shown in red and EAs in blue. A one-sided Fisher test was performed to test whether AA have higher germline HR-Deficiency than EA within each cancer types and the p-value is provided.
Supplementary information
Supplementary Tables
Supplementary Tables 1–14.
Rights and permissions
About this article
Cite this article
Sinha, S., Mitchell, K.A., Zingone, A. et al. Higher prevalence of homologous recombination deficiency in tumors from African Americans versus European Americans. Nat Cancer 1, 112–121 (2020). https://doi.org/10.1038/s43018-019-0009-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-019-0009-7
This article is cited by
-
Combination of genomic instability score and TP53 status for prognosis prediction in lung adenocarcinoma
npj Precision Oncology (2023)
-
HRD effects on first-line adjuvant chemotherapy and PARPi maintenance therapy in Chinese ovarian cancer patients
npj Precision Oncology (2023)
-
Signatures of copy number alterations in human cancer
Nature (2022)
-
An integrative analysis of the age-associated multi-omic landscape across cancers
Nature Communications (2021)
-
A comprehensive map of alternative polyadenylation in African American and European American lung cancer patients
Nature Communications (2021)