Abstract
Most natural and synthetic antibodies are ‘unseen’. That is, the demonstration of their neutralization effects with any antigen requires laborious and costly wet-lab experiments. The existing methods that learn antibody representations from known antibody–antigen interactions are unsuitable for unseen antibodies owing to the absence of interaction instances. The DeepAAI method proposed herein learns unseen antibody representations by constructing two adaptive relation graphs among antibodies and antigens and applying Laplacian smoothing between unseen and seen antibodies’ representations. Rather than using static protein descriptors, DeepAAI learns representations and relation graphs ‘dynamically’, optimized towards the downstream tasks of neutralization prediction and 50% inhibition concentration estimation. The performance of DeepAAI is demonstrated on human immunodeficiency virus, severe acute respiratory syndrome coronavirus 2, influenza and dengue. Moreover, the relation graphs have rich interpretability. The antibody relation graph implies similarity in antibody neutralization reactions, and the antigen relation graph indicates the relation among a virus’s different variants. We accordingly recommend probable broad-spectrum antibodies against new variants of these viruses.
Similar content being viewed by others
Main
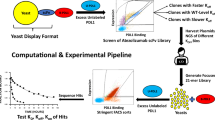
Antibodies (Abs) opsonize and neutralize viruses1, working as potent bio-pharmaceuticals in clinical treatments2. An individual is estimated to have around 108 different Abs3 and produces on the order of 1020 Abs in response to viral infections4. Among them, only a small fraction can opsonize and an even smaller fraction can neutralize the infected virus. The majority of these Abs are ‘unseen’. We are blind to their neutralizability with any antigen (Ag) before conducting wet-lab experiments (Fig. 1a). Besides natural Abs, de novo synthetic Abs are also unseen and need to be demonstrated experimentally before clinical treatments. The conventional experiments include phage display5, enzyme-linked immunosorbent assay (ELISA)6, pseudovirus assay7 and so on, which are resource intensive and time consuming8. We seek to develop accurate and fast computational methods as preliminary screening, to reduce blindness and improve foresight for the wet experiments and accelerate the process of discovering novel therapeutic Abs9.
a, Unseen Abs are those whose interactability with ‘any’ Ag has not been experimentally demonstrated. b, For a seen Ab (Ab2), the backpropagation from its known interaction data can establish the high-quality representation, inferring its interactability with other Ags (for example, Ag2 in a). For an unseen Ab (Ab3), the interactions with Ags are unknown, resulting in failure to learn the representation. c, We construct a relation graph to bridge unseen and seen Abs, in which the nodes represent Abs, the nodes’ attributes are Ab representation and the edges’ weights are the quantified relation among Abs. d, By applying GCNs on the relation graph, the relation among Abs can be quantified and therefore unseen Abs’ representation can be learned and optimized from relational seen Abs in training. e, We demonstrate our methods in HIV, influenza, dengue and SARS-CoV-2 (from our own wet data). We also illustrate our method’s rich interpretability in SARS-CoV-2. This can imply a relation among variants of SARS-CoV-2 from the perspective of Ab–Ag neutralization effects. We accordingly recommended probable broad-spectrum Abs against new variants of SARS-CoV-2 (Omicron).
According to the prediction tasks, studies related to Ab–Ag interaction prediction can be categorized into mainly three groups: (1) predicting Ab–Ag binding sites, (2) discriminating Ab–Ag binders/non-binders and (3) predicting Ab–Ag neutralization/non-neutralization effects. Given an Ab–Ag binding pair, some studies predicted the binding sites (Parapred10, Fast-Parapred and AG-Fast-Parapred11, PECAN12 and PInet13). Given the Ab–Ag pairwise instances, others discriminated binders and non-binders14,15, which is considered the upstream task of predicting binding sites. We note that binding Abs may not neutralize but instead only opsonize pathogens. Opsonization is an indirect anti-viral process, in which Abs bind pathogens as marks to facilitate phagocytosis of macrophages, while neutralization is a direct anti-viral process, in which Abs directly stop the attachment of pathogens to host tissues16. In this study, we focus on predicting Ab–Ag neutralization effects.
The methods related to Ab–Ag interaction prediction can be further classified by input: (1) sequence based and (2) structure based. When predicting binding sites, the sequence-based methods combined local neighbourhood and entire sequences (Parapred10) and applied cross-modal attention on Ab and Ag residues (Fast-Parapred and AG-Fast-Parapred11), and the structure-based methods employed graph convolutional networks (GCNs) on Ab and Ag structures (PECAN12) and extracted geometrical features by consuming point clouds from structures (PInet13). When classifying binders/non-binders, Mason et al. applied convolutional neural networks (CNNs) on Ab sequences14, and DLAB15 implemented CNNs on crystal or modelled structures and found that using highly accurate crystal structures could enhance performance, while using modelled structures failed to achieve strong discrimination between binders and non-binders, probably because ‘structure modelling’ and ‘interaction prediction’ were successively engaged and errors in the former would be exacerbated in the latter. We note that obtaining highly accurate crystal structures through wet-lab experiments is also laborious and costly, while amino acid sequences are easily and widely accessible in the real world. Additionally, large-scale sequence data can enhance the applicability of methods. Therefore, we propose a sequence-based method, facilitating real-world applications.
Predicting unseen Abs’ neutralizability from amino acid sequences has two challenges. (1) We are faced with the well-known cold-start problem, that is, an unseen Ab’s neutralization with ‘any’ Ag is unknown. Existing methods learn Ab representation by backpropagating errors from known Ab–Ag interactions (Fig. 1b), which is not applicable to unseen Abs owing to the lack of interaction instances. (2) Another challenge lies in the problem that the expressivity and adaptability of the static feature to represent Abs and Ags could be limited. Although there are various protein descriptors, for example, k-mer frequency counting (kmer), position-specific scoring matrices (PSSMs) and the protein–protein basic local alignment search tool (BlastP), the feature space could be high dimensional and the features are pre-computed and static; they are unsupervised, not optimized in the training process and probably not optimal for a specific supervised learning task.
To overcome these challenges, we propose a deep Ab–Ag interaction algorithm, named DeepAAI. Our DeepAAI can learn the representation of unseen Abs from seen Abs by constructing two adaptive relation graphs that connect Abs and Ags, respectively, and applying Laplacian smoothing (in GCNs) in the representation of unseen and seen Abs. In the two relation graphs, the nodes represent Abs and Ags, the node attributes are the learned representations of Abs and Ags, and the edge weights are the quantified relation among Abs and among Ags, respectively. Figure 1c shows the Ab relation graph.
Rather than using those high-dimensional and static features directly, DeepAAI applies a neural network to project the original features into a low-dimensional and high-expressivity feature space, in which the representations are used to serve as the node attributes and further quantify the edge weights. The node attributes and the edge weights are not static but dynamically optimized towards the downstream tasks, predicting neutralization effects and estimating 50% inhibition concentration (IC50) values (Fig. 1d). Thereby, the Ab and Ag relation graphs are task oriented and adaptively constructed, predicting the optimal relations among Abs and Ags.
We then predict unseen Abs’ neutralizability by applying GCNs on the relation graphs, conducting Laplacian smoothing between unseen and seen Abs’ representation as transductive learning. Consequently, the unseen Abs’ representation can be learned from the relational seen Abs’ representation and optimized in the training process, guaranteeing that the unseen Abs’ neutralizability can be inferred in a semi-supervised manner.
Additionally, we note that Ab–Ag neutralization is determined by both global and local features. The global features of Abs and Ags are deterministic of interactions, while the local features of amino acids at the interface directly affect the affinities. Therefore, besides the adaptive relation graph that learns global features among Abs and Ags, we also adopt a CNN module to learn local features inside an Ab and Ag.
The performance of DeepAAI is demonstrated on the unseen Abs of various viruses, including human immunodeficiency virus (HIV), severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), influenza and dengue (Fig. 1e). Furthermore, as it does not require knowledge on Ab and Ag structures, DeepAAI is friendly to real-world applications. Additionally, the adaptively constructed relation graphs have rich interpretability. The Ab relation graphs imply similarity in Ab neutralization reactions (similar binding regions). The Ag relation graphs indicate relations among different variants of a virus. We accordingly recommend probable broad-spectrum Abs against new variants of a virus.
Results
DeepAAI
DeepAAI has two neural network modules, an adaptive relation graph convolutional network (AR-GCN) and a CNN module14, which learn global representation among Abs/Ags and local representation inside an Ab/Ag, respectively (Fig. 2a).
a, DeepAAI consists of an AR-GCN module and a CNN module, learning global representation among Abs/Ags and local representation inside an Ab/Ag, respectively. The AR-GCN adaptively constructs two relation graphs by quantifying relation among Abs and among Ags and learns Ab and Ag representation from relation. Mason’s CNN architecture is also adopted to extract local features from amino acid sequences. The AR-GCN and CNN modules are used to learn both Ab and Ag representations in the same feature space, facilitating Ab and Ag representation fusion. b, The neural network structure of the AR-GCN module. c, The neural network structure of the CNN module.
Relation graph module (AR-GCN)
The AR-GCN adaptively constructs two relation graphs by quantifying the relation among Abs and Ags and then learns Ab and Ag representations by applying GCNs on the two relation graphs. We hypothesize that two Abs participating in similar neutralization effects should be given a close relation, which can be quantified by the two Abs’ representation (equation (1)),
where HAb1 and HAb2 are the two Abs’ representations, RAb1−Ab2 is the relation between Ab1 and Ab2, and \({{{\mathcal{F}}}}\) is a function to quantify relation.
Before quantifying the relation among Abs, we devise two fully connected (FC) layers (with activation functions), which non-linearly transform kmer and PSSMs into a low-dimensional feature space. The non-linear transformation can flexibly learn representation from biological similarity (kmer) and evolutionary information (PSSMs), thereby enriching the relation quantification. The relation has the following properties:
-
(1)
Symmetric: RAb1−Ab2 = RAb2−Ab1.
-
(2)
The absolute value is no more than 1: −1 ≤ RAb1−Ab2 ≤ 1.
-
(3)
The self-loop relation equals 1: RAb1−Ab1 = 1.
Consequently, we construct a relation graph among Abs. A GCN operation is then applied on the relation graph, working as Laplacian smoothing in Abs’ representation (Supplementary Information). Figure 2b describes the neural network structure of AR-GCN.
CNN module
The CNN module includes one-hot encoding, 1D convolution, maximum pooling, flatten and an FC layer, aiming at learning the local features of an Ab or Ag sequence (Fig. 2c). The kernel size is only two, making this module specifically focus on local feature extraction.
Fusion
Importantly, the AR-GCN and CNN module are also applied in Ag representation learning. Embedding Abs and Ags in the same feature space can facilitate their representation fusion. The fusion is conducted by addition and dot product with a balance coefficient, which is also learnable to avoid human-experience-based settings. Finally, two FC layers are used to predict neutralization effects and estimate IC50 values, respectively. For details on DeepAAI, see Methods.
Performance on HIV
In Methods we describe the details of the HIV dataset curation. We randomly sample 45 Abs from all 242 Abs to serve as unseen Abs, involving 3,301 Ab–Ag pairwise instances in the unseen test set (Fig. 3a). The 45 unseen Abs have no instance that is similar to any instance of the seen Abs (BlastP < 90%). Considering that our task is to predict neutralization effects of Ab–Ag pairwise instances, we define two Ab–Ag pairwise instances as being similar when they have similar Abs (BlastP ≥ 90%), similar Ags (BlastP ≥ 90%) and the same neutralization or non-neutralization effects. Figure 3b shows the multiplicative product of the Ab and Ag BlastP scores between every two Ab–Ag pairs after we remove similar instances.
a, The numbers of Ab–Ag pairwise instances and unique Abs. b, The multiplicative product of BlastP scores of Abs and Ags between every two instances in the total of 27,738 Ab–Ag pairs after we remove similar instances (BlastP ≥ 90%). We zoom in on part of the figure. The diagonal line represents self-relation (equal to 1). c, The performances of neutralization prediction. d, The performances of IC50 estimation. In c and d, the performances are evaluated 20 times in 20 different random seeds. The box plots show median, first and third quartiles, minimum and maximum. Outliers are classified as being 1.5 times outside the interquartile range. The best variants and best baseline models were compared via Mann–Whitney U test (two sided). e, The runtime of every epoch. f, The scatter plots of the penultimate layer’s embedding after PCA. g, The predicted neutralization probabilities in heat maps.
Predicting unseen Abs’ neutralizability
Figure 3c compares the performances of neutralization prediction on the unseen HIV Abs, and Supplementary Table 1a presents the numeric results. In these methods, kmer and PSSM represent the global features while sequence (seq) means local features. The three DeepAAI variants—DeepAAI (kmer + seq), DeepAAI (PSSM + seq) and DeepAAI (kmer + PSSM + seq)—outperform all eight baseline methods in accuracy, F1 score, precision–recall area under the curve (PC-AUC) and Matthews correlation coefficient (MCC) with statistical significance (P < 0.05). We also note no statistical significance among the three DeepAAI variants. The variants of DeepAAI (kmer + PSSM) and DeepAAI (seq), which reflect the effectiveness of global and local feature extraction, respectively, perform better than the baseline methods but relatively worse than the above three variants that combine global and local features. The results show that combining global and local features is indispensable in predicting Abs’ neutralization with Ags. In the area under the receiver operating characteristic (ROC-AUC), only DeepAAI (kmer + seq) outperforms others. Mason’s CNN architecture beats the other baseline methods but loses to DeepAAI.
The results prove that the proposed DeepAAI outperforms the baseline methods and that combining global and local features is indispensable for predicting unseen Abs’ neutralization effects on Ags.
Predicting unseen Abs’ IC50
Figure 3d and Supplementary Table 1b show the performances of IC50 estimation on the unseen Abs. Compared with all the baseline methods, both DeepAAI (PSSM + seq) and DeepAAI (kmer + PSSM) have superior performances in the mean squared error (MSE) and the mean absolute error (MAE). In MSE, DeepAAI (kmer + PSSM) performs better than DeepAAI (PSSM + seq). AG-Fast-Parapred architecture is the best baseline method.
Runtime
Figure 3e compares the runtime of every epoch on an NVIDIA GeForce RTX 1080 Ti GPU, in which 22,359 Ab–Ag pairwise instances are learned. Compared with the baseline methods, DeepAAI is computationally inexpensive because it avoids the time-consuming recurrent neural networks and the attention algorithms.
Visualization
We transform the values in the penultimate layer to a two-dimensional space by principal component analysis (PCA), which describes the learned representation of Ab–Ag pairwise instances and gives us a view of what the methods have learned (Fig. 3f). DeepAAI has higher intra-class similarity and better inter-class boundaries, while the best baseline method (Mason’s CNN architecture) mixes the neutralization and non-neutralization instances. Figure 3g shows the predicted probabilities in heat map, and DeepAAI achieves heat maps similar to the experimentally validated results.
Predicting seen Abs’ neutralizability and IC50
Although we know seen Abs’ neutralization only with some Ags, we can still predict their neutralization effects with other Ags. As Supplementary Fig. 1 shows, DeepAAI (kmer + seq) and DeepAAI (kmer + PSSM + seq) win in the neutralization prediction, while DeepAAI (kmer + PSSM) surpasses the others in the IC50 estimations.
Label-shuffled control
As an extensive study, we further experiment on the label-shuffled data. The results in Supplementary Table 2 show that, without true knowledge, DeepAAI cannot perform normally, indirectly demonstrating that DeepAAI does not make random predictions but learns valuable knowledge.
Performance on SARS-CoV-2
This experiment investigates whether DeepAAI can be applied to SARS-CoV-2. We collect the Ab–Ag neutralization and non-neutralization instances and Abs’ sequences from Coronavirus Antibody Database (CoVAbDab)17. Owing to the absence of IC50 values in CoVAbDab, we discriminate only Ab–Ag neutralization and non-neutralization effects. We also have our own wet-lab data as the unseen test data, which were collected from a convalescent individual18. Figure 4a shows the numbers of Ab–Ag pairwise instances and unique Abs.
a, The numbers of Ab–Ag pairwise instances and unique Abs. b, The performance of our wet-lab Abs on SARS-CoV-2, which are evaluated 20 times in 20 different random seeds. The box plots show median, first and third quartiles, minimum and maximum. Outliers are classified as being 1.5 times outside the interquartile range. The comparisons were carried out via Mann–Whitney U test (two sided) with no adjustment. The upward pointing arrows (↑) mean the higher the better. c, Difference between DeepAAI and BlastP.
Figure 4b shows the performances on our wet-lab Abs of SARS-CoV-2. DeepAAI (kmer + seq) outperforms Mason’s CNN architecture (the best baseline method) by 0.05, 0.13, 0.13, 0.03 and 0.11 in accuracy, F-score, ROC-AUC, PR-AUC and MCC, respectively. We provide only the performance of DeepAAI (kmer + seq) for its steady performances and Mason’s CNN architecture for its advantages over the other seven baseline methods in the neutralization prediction of HIV.
Figure 4c shows the predictions by DeepAAI and BlastP. In BlastP, we think an Ab will neutralize an Ag when the average BlastP score between the Ab and the other neutralizing Abs is higher than that between the Ab and the other non-neutralizing Abs. The results show that the dynamical relation quantification adapts to the supervised task of the neutralization/non-neutralization prediction better than the unsupervised sequence alignment in BlastP.
DeepAAI’s interpretation
The relation graphs have rich interpretability. The Ab relation graphs imply the similarity in Ab neutralization reactions (similar binding regions). The Ag relation graphs indicate the relation among the different variants of a virus. Moreover, we recommend probable broad-spectrum Abs against a virus’s new variant.
The relation graph reflects binding regions
In the wet-lab experiments of our previous study, we performed competition ELISAs to determine whether our isolated neutralizing Abs had overlapping or non-overlapping epitopes in the receptor-binding domain (RBD) of S protein (Supplementary Fig. 2). We found that our neutralizing Abs could bind to four groups of five distinct epitopes on the RBD. Therefore, the neutralizing Abs were divided into four mutually exclusive groups, namely RBD groups I–IV in our previous study18.
We compare the quantified relation among neutralizing Abs that belong to the same group and different groups (inter). As Fig. 5a shows, we find that the quantified relations between two Abs that belong to the same group are significantly higher than those that belong to the different groups (inter). We exclude group I because only one neutralizing Ab belongs to group I and therefore we cannot perform a t-test. This finding shows that the Ab relation can predict an unseen Ab’s binding regions in the virus by examining the unseen Ab’s relation to all the seen Abs that have different binding regions.
a, The relations between two Abs binding to the same RBD group (II, III or IV) are significantly higher than those binding to the different RBD Groups (inter). The box plots show median, first and third quartiles, minimum and maximum. The comparisons were carried out via t-test (two sided) without adjustment. n = 15, 21, 36 and 54 in II, III, IV and inter-groups, respectively. b, The average closeness among the SARS-CoV-2 variants. Delta has the lowest average closeness (0.36) to the other variant (excluding self-closeness). Omicron has the lowest self-closeness (0.84), indicating greater difference in its subvariants. c, The three most important 3-mers in the heavy sequences of our wet-lab Abs. d, The three most important 3-mers in the light sequences of our wet-lab Abs. e, DeepAAI recommends the 50 most probable Abs that could neutralize Omicron, five of which (in bold) have been previously demonstrated.
Differences among the virus variants implied by relation graph
Figure 5b shows the qualified relation among the SARS-CoV-2 variants. From the perspective of Ab–Ag neutralization effects in DeepAAI, Delta is thought of as the most different variant, which accords with the fact that Delta’s symptoms are different from those associated with the original strain (wild type). Furthermore, the values along the diagonals imply the difference in a variant’s subvariants and sequences from different sources. Omicron is quantified to have the lowest self-relation (0.84) by DeepAAI, indicating greater difference in its subvariants.
Important kmers
Figure 5c,d shows the sequence logo of the three most important ‘3-mers’ in the heavy and light sequences of the SARS-CoV-2 Abs that are collected from our wet experiments. We record the top three 3-mers in the heavy and light sequences with the highest weights in the two-layer neural network of AR-GCN that projects the kmers into a low-dimensional and high-expressivity feature space. In the heavy chains, the most important 3-mers are located at the tail, and the second and third ones are consecutive from the 44th to 47th amino acids (Fig. 5c). In the light chains, the most important 3-mers are located near the middle, and the second and third ones are also close to each other (Fig. 5d).
Broad-spectrum Ab recommendation
We also explore probable broad-spectrum Abs that could neutralize the Omicron variant. Among the total 2,587 SARS-CoV-2 Abs, DeepAAI recommends the 50 most probable Abs (Fig. 5e), 5 of which have been demonstrated previously19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35.
Performances on influenza and dengue
This experiment investigates whether the knowledge learned from Ab–Ag interaction instances of HIV can help to predict Abs’ neutralizability of influenza and dengue. We freeze the AR-GCN and CNN module and train the final FC layers in transfer learning. The numbers of Ab–Ag pairwise instances and unique Abs are described in Fig. 6a. In Methods, we show the details of dataset curation and transfer learning. In this experiment, all the collected Ab–Ag pairwise instances are neutralizing. As negative sampling may coincidentally bring in neutralizing instances of broad-spectrum Abs, we do not adopt negative sampling to generate non-neutralizing instances but use the unseen HIV Abs as non-neutralization Abs, considering Abs have high specificity. Therefore, we focus on recall, that is, the fraction of positive instances that were correctly predicted (Fig. 6b).
a, The numbers of Ab–Ag pairwise instances and unique Abs. b, The performance on the unseen Abs of influenza and dengue. The performances are evaluated 20 times in 20 different random seeds. The box plots show median, first and third quartiles, minimum and maximum. Outliers are classified as being 1.5 times outside the interquartile range. The comparisons were carried out via Mann–Whitney U test (two sided) with no adjustment. The upward pointing arrows (↑) mean the higher the better.
DeepAAI (kmer + seq) (the best variant in HIV) significantly outperforms Mason’s CNN architecture (the best baseline method in HIV) by 0.10 on influenza. On dengue, both DeepAAI (kmer + seq) and Mason’s CNN architecture perform well and there is no statistical significance between them.
Discussion
In this study, we propose DeepAAI to predict unseen Abs’ neutralizability with Ags. DeepAAI achieves outstanding performances on a variety of viruses, including HIV, SARS-CoV-2, influenza and dengue. On the basis of the adaptively constructed relation graph in DeepAAI, we can denote the similarity in Ab neutralization reactions (similar binding regions) and the relation among the different variants of a virus from the perspective of Ab–Ag neutralization effects and recommend the probable broad-spectrum Abs against a new variant of a virus (Omicron).
As it does not require knowledge on Ab and Ag structures, DeepAAI is friendly to real-world applications. DeepAAI can be used by biologists in two successive steps: (1) predicting Ab–Ag neutralizing/non-neutralizing effects as preliminary screening and then (2) estimating IC50 values to prioritize the subsequent wet-lab validation experiments. We provide a web service of DeepAAI, and the data and codes are freely available.
In this study, we did not use modelled structures because we intend to avoid the two-step prediction of structures and neutralization in which errors in the former could be exacerbated in the latter. In the future, we may integrate the feature extraction modules of Ab and Ag structure prediction into Ab–Ag interaction prediction if more crystal structures or precisely modelled structures are available.
Methods
Problem definition
The amino acid sequences of an Ab and an Ag are denoted as B = (b1, b2, . . . , bm) and G = (g1, g2, . . . , gn), respectively. The objective is to discriminate neutralization/non-neutralization (classification), \({{{{\mathcal{F}}}}}_{{\mathrm{bin}}}(B,G)=\left\{\begin{array}{l}0,\,{{\mbox{non-neutralization}}}\,\\ 1,\,{{\mbox{neutralization}}}\,\\ \end{array}\right.\), and estimate IC50 values (regression), \({{{{\mathcal{F}}}}}_{{\mathrm{reg}}}(B,G)=\,{{\mbox{IC}}}_{50}\,\).
Data
HIV data
Collect HIV data: Algorithm 1 illustrates the pseudo-code for collecting the HIV data. Note that the non-neutralizing pairwise data of HIV are experimentally demonstrated rather than negatively sampled.
Algorithm 1 The process of the HIV dataset collection.
Require: the data source, that is, the Compile Analyze and Tally NAb Panels (CATNAP36) at Los Alamos HIV Database (LANL37)
Ensure: the neutralizing or non-neutralizing Ab–Ag pairwise instances in amino acid sequences
1: Extract the total assay that pairs Abs and Ags, denoted as T;
2: Extract the sequences of the heavy and light chains in T, denoted as H and L, respectively;
3: Uniform the forms of H and L to the fragment variable (Fv)—remove constant-heavy-1 (CH1) from H and constant-light (CL) from L when they are in the form of antigen-binding fragment (Fab);
4: Extract Ag sequences in T, denoted as V;
5: Pair the H, L and V based on T;
6: Remove the duplicated pairs in H, L and V;
7: Remove the pairs that have ‘not available’ (N/A) values in H, L or V;
8: Collect the IC50 values for the paired PH, PL and V;
9: Average IC50 values for any pair that has more than one reported IC50 value;
10: Set the cut-off at IC50 = 10 μg ml−1 and consider IC50 < 10 μg ml−1 neutralization and IC50 ≥ 10 μg ml−1 non-neutralization;
11: Return the HIV dataset.
Split unseen and seen Abs: We randomly take 57 Abs to serve as the unseen Abs and the others as the seen Abs. As 12 of the 57 Abs have instances similar to the seen Abs’ instances, we remove them and take the remaining 45 Abs to serve as the unseen Abs. Two Ab–Ag pairwise instances are considered to be similar when they have similar Abs (BlastP ≥ 90%), similar Ags (BlastP ≥ 90%) and the same neutralization/non-neutralization effects. We then split the seen Abs’ Ab–Ag interaction instances into training, validation and seen test sets. We also remove instances in the seen test set that are similar to any instance in the training and validation sets. We include both the unseen and seen Abs in the Ab relation graph before training. We split the seen Abs’ instances, remove similar instances in the seen test set and train the models 20 times in 20 different random seeds. When different seeds are used, the numbers of Ab–Ag pairwise instances and unique Abs in the seen test set vary. Figure 3a shows the data information of seed 18.
CoVAbDab SARS-CoV-2 data
The SARS-CoV-2 data are collected from the CoVAbDab17. The collected data include both neutralizing and non-neutralizing Ab–Ag pairwise instances. The sequences of the SARS-CoV-2’s variants are collected from the National Center for Biotechnology Information38. The curated dataset includes the SARS-CoV-2 variants of the wild type, Alpha, Beta, Gamma, Delta and Omicron. For each variant, the sequences of the different subvariants from the different sources are different. Therefore, we randomly take 5 sequences for each variant except Omicron, for which we take all 11 sequences.
We take the Omicron variant as ‘unseen Ags’. To suggest probable broad-spectrum Abs against the Omicron variant, we exclude the Ab–Ag pairwise instances of the Omicron variant in training but include the Omicron variant (unseen Ags) in the Ag relation graph and the Omicron Abs (unseen Abs) in the Ab relation graph as transductive learning.
Wet-lab SARS-CoV-2 data
In our previous study18, we found a convalescent individual with potent IgG neutralizing activity to SARS-CoV-2 from the hospital volunteers. The volunteer recruitment and the blood draws were performed at the Zhoushan Hospital under a protocol approved by the Zhoushan Hospital Research Ethics Committee (2020-003). Experiments related to all human samples were performed at the School of Basic Medical Sciences, Fudan University under a protocol approved by the institutional ethics committee (2020-C007).
We characterized the Ab responses and isolated monoclonal Abs from the individual’s memory B cells. Consequently, we obtained 36 Abs with the confirmed amino acid sequences. The wet-lab Abs are also included in the Ab relation graph as unseen Abs for evaluation. We also conducted ELISA, pseudovirus assays, single-cell sorting and cloning, and so on and found 17 Abs that are neutralizing and 19 that are non-neutralizing to the wide type of SARS-CoV-2. We identified the 17 neutralizing Abs’ binding regions in RBD, which are used in Fig. 4a. For more details on our wet-lab data, please refer to ref. 18.
Influenza and dengue data
We collect influenza and dengue data from Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB)39. All the collected Ab–Ag pairwise instances are positive (neutralizing). We do not adopt negative sampling to generate non-neutralizing instances because negative sampling may coincidentally bring in neutralizing instances of broad-spectrum Abs. Considering the high specificity of Abs, we use the unseen HIV Abs as non-neutralization Abs to influenza and dengue, but exclude the seen HIV Abs because they have been used to train the HIV models and using them could lead to overfitting in these Abs in transfer learning. We ensure the numbers of neutralization and non-neutralization instances are equal. Finally, we split the data into the training and unseen test sets, remove similar instances in the unseen test sets (BlastP ≥ 90%) and include the unseen and seen Abs in the relation graph of Abs.
Features
Amino acid sequences
Amino acid sequences transparently describe amino acids and their sequential positions. We use one-hot encoding.
kmer
The kmer contains two basic characteristics of biological sequences, monomer component information and entire sequence information40, revealing the distribution of entire characteristics and measuring biological similarity for discrimination41. We use k = 1, 2, 3, which generates 21 (20 amino acids + unknown), 212 and 213 dimensions, respectively. We abandon k = 4 because it generates too many dimensions (194,481), which tends to deteriorate the algorithm. We then remove the kmer features with a frequency less than 0.05 to prevent overfitting and accelerate training. Finally, a 2,764-dimensional vector of kmer is left.
PSSM
The PSSMs reveal evolutionary information and have been successfully applied to improve the performance of various predictors of protein attributes42. We select the Uniref50 database with the tool of the position-specific scoring matrix-based feature generator for machine learning (POSSUM)42 to generate PSSMs, encoding evolutionary information in a 420-dimensional vector. Note that using the Uniref50 database will not cause information leakage since the supervision information comes from the output (that is, Ab–Ag neutralizing/non-neutralizing effects and IC50 values) rather than the input (that is, the sequences of Abs and Ags).
Baseline methods
In this study, we use four types of baseline methods: (a) the sequenced-based architecture that predict binding sites: Parapred architecture10, Fast-Parapred architecture11 and AG-Fast-Parapred architecture11; (b) the sequenced-based models that predict protein–protein interactions: PIPR architecture43 and ResPPI architecture44; (c) the classic sequential models: Bi-LSTM-Attention45 and TextCNN46; and (d) the sequenced-based model that has been demonstrated effective in wet-lab experiments: Mason’s CNN architecture14.
For the methods in group (a), Parapred architecture, Fast-Parapred architecture and AG-Fast-Parapred architecture, the original task is to predict the binding site given Ab–Ag binding pairs. We keep their network structures and inputs (amino acid sequences) but modify the prediction tasks from binding sites to neutralization and IC50. For Mason’s CNN architecture, we add an Ag extraction module and an Ab–Ag embedding fusion module, because the original Mason’s CNN learns only Ab features and is specific to one Ag (without fusing various Ags). We follow the other implementation details in the cited papers.
DeepAAI
AR-GCN module
First, we use a learnable embedding layer to project the kmer and PSSM vectors non-linearly into a low-dimensional feature space (equation (2)).
where σELU refers to the activation function of exponential linear unit (ELU); Xkmer and XPSSM represent the vectors of kmer and PSSM, respectively; Wkmer and WPSSM denote the weights of the FC layers for kmer and PSSM, respectively; and Hkmer and HPSSM are the outputs.
In DeepAAI (kmer + PSSM + seq), we concatenate the representation of kmer and PSSM by HAb = Hkmer∥HPSSM. HAb then flows into another FC layer with tan-hyperbolic (Tanh) to further learn node representation. By calculating the cosine similarity (composed of instance normalization and inner product), we obtain the relation between two Abs as follows.
where \({\sigma }_{\tanh }\) refers to the activation function of Tanh; HAb1 and HAb2 are the two Abs’ representation, respectively; WFC denotes the weights of the FC layer; Inst_Norm refers to instance normalization; ⋅ is the inner product; and RAb1−Ab2 is the relation between Ab1 and Ab2. Two GC layers are then implemented (equation (4)).
where \(\tilde{A}\) is the adjacency matrix (including self-loops); \(\tilde{D}\) is a modified degree matrix used to ensure positive values in \(\tilde{D}\); \(\hat{A}\) is the symmetrically normalized \(\tilde{A}\); HAb is the Ab representation; Wζ0 and Wζ1 are the weights of the first and second graph convolutional layers, respectively; and Hζ1 and Hζ2 represent the embedding vectors after the first and second graph convolutional layers, respectively. The AR-GCN’s final embedding (HARGCN) is the sum of HAb, Hζ1 and Hζ2 as follows:
CNN module
A CNN module conducts 1D convolution on the one-hot encoding of amino acid sequences, in which the channels, kernel size, stride and padding are 64, 2, 1 and 1, respectively, making this module focus on local feature learning. After the activation function (ReLU) and dropout (rate 0.5), maximum pooling and flatten are implemented. An FC layer is finally used to output the representation (32 × 1).
Fusion
To embed both Abs and Ags into the same feature space, we use addition and the dot product with a balance coefficient to fuse the Ab and Ag presentations. Two FC layers are then adopted to complete the neutralization prediction and IC50 estimation.
where HARGCN−Ag, HARGCN−Ab, Hlocal−Ag, Hlocal−Ab, HARGCN, Hlocal and H denote the embedding of AR-GCN, local extraction (Mason’s CNN architecture) and fusion of Ags and Abs, respectively; ⊙ is the Hadamard product; α is the balance coefficient, automatically learned; σsigmoid is the activation function of sigmoid; Wa and Wb are the FC layers’ weights; and \({\hat{Y}}^{({\mathrm{prob}})}\) and \({\hat{Y}}^{({\mathrm{IC50}})}\) are the predicted neutralization probabilities and the estimated IC50 values, respectively.
Loss function
The two downstream tasks (binary neutralization prediction and IC50 estimation) are conducted separately. For predicting neutralization, the loss function is formulated as equation (7) shows.
where \({\sum }_{v\in {{{\mathcal{V}}}}}({y}_{v}^{({\mathrm{bin}})}{\mathrm{ln}}({\hat{y}}_{v}^{({\mathrm{prob}})})+(1-{y}_{v}^{({\mathrm{bin}})}){\mathrm{ln}}(1-{{\hat{y}}^{({\mathrm{prob}})}}_{v}))\) is the cross-entropy loss, and \({y}_{v}^{({\mathrm{bin}})}\) and \({\hat{y}}_{v}^{({\mathrm{prob}})}\) are the true label and the predicted probabilities of v. \(\tilde{A}\) is the adjacency matrix of the virtual graph (including self-loops), \(\left\Vert \tilde{A}\right\Vert\) is the sum of the absolute values in \(\tilde{A}\) as a penalty term, and λa is an adjustable hyper-parameter used to balance the two losses.
When estimating IC50, we calculate the loss function as equation (8) shows.
where \({\sum }_{v\in {{{\mathcal{V}}}}}{({y}_{v}^{{\mathrm{IC50}}}-{\hat{y}}_{v}^{{\mathrm{IC50}}})}^{2}\) is the regressive loss in terms of MSE and \({y}_{v}^{{\mathrm{IC50}}}\) and \({\hat{y}}_{v}^{{\mathrm{IC50}}}\) are the true and predicted IC50 values of v, respectively. \(\tilde{A}\) is the adjacency matrix of the virtual graph (including self-loops), \(\left\Vert \tilde{A}\right\Vert\) is the sum of the absolute values in \(\tilde{A}\) as a penalty term, and λb is an adjustable hyper-parameter used to balance the two losses.
Transfer learning from HIV to influenza and dengue
Inspired by natural language processing, biological sequences (especially amino acid sequences) can be thought of as meaningful protein languages. Therefore, the representation of amino acid fragments (such as kmers) is expected to improve the reliability and stability of a prediction model by pre-training the model on a large number of relevant data and transferring the knowledge to a target domain. Transfer learning can reduce the dependence on the number of target domain data. Considering that HIV, influenza and dengue are all viruses and HIV has accumulated enough Ab–Ag interaction data, we try to conduct transfer learning from HIV to influenza and dengue.
DeepAAI can be divided into three parts: AR-GCN, CNN and the final FC layers with fusion. The AR-GCN and CNN modules are used to extract features from Abs and Ags. The final FC layers are implemented to learn how to predict neutralizing/non-neutralizing effects based on the extracted features by the AR-GCN and CNN modules. We freeze the parameters in the AR-GCN and CNN modules when we conduct transfer learning. Nonetheless, we need to fine-tune the final FC layers, considering different viruses have different neutralizing mechanisms.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The HIV data are available from CATNAP36 at LANL37 (https://www.hiv.lanl.gov/components/sequence/HIV/neutralization/download_db.comp). We also provide the dataset that we generated for this study as the minimum dataset (https://github.com/enai4bio/DeepAAI/tree/main/dataset/corpus). The SARS-CoV-2 data are available from CoVAbDab17 (http://opig.stats.ox.ac.uk/webapps/covabdab/). The influenza and dengue data are available from RCSB PDB39 (https://www.rcsb.org/). The references include the minimum datasets that are necessary to interpret, verify and make the research in the article transparent to readers.
Code availability
The DeepAAI code was implemented in Python using the deep learning framework of PyTorch. Code, trained models and scripts reproducing the experiments of this paper are available at https://github.com/enai4bio/DeepAAI47. All source code is provided under the GNU Affero General Public License v3.0. We provide a web service of DeepAAI at https://aai-test.github.io/.
References
Paludan, S. R., Pradeu, T., Masters, S. L. & Mogensen, T. H. Constitutive immune mechanisms: mediators of host defence and immune regulation. Nat. Rev. Immunol. 21, 137–150 (2021).
Abraham, J. Passive antibody therapy in COVID-19. Nat. Rev. Immunol. 20, 401–403 (2020).
Sompayrac, L. M. How the Immune System Works (Wiley, 2019).
Ripoll, D. R., Chaudhury, S. & Wallqvist, A. Using the antibody–antigen binding interface to train image-based deep neural networks for antibody-epitope classification. PLoS Comput. Biol. 17, e1008864 (2021).
Lee, CarolM. Y., Iorno, N., Sierro, F. & Christ, D. Selection of human antibody fragments by phage display. Nat. Protoc. 2, 3001 (2007).
Butler, J. E. Enzyme-linked immunosorbent assay. J. Immunoassay 21, 165–209 (2000).
Khoury, D. S. et al. Measuring immunity to SARS-CoV-2 infection: comparing assays and animal models. Nat. Rev. Immunol. 20, 727–738 (2020).
Ogunniyi, A. O., Story, C. M., Papa, E., Guillen, E. & Love, J. C. Screening individual hybridomas by microengraving to discover monoclonal antibodies. Nat. Protoc. 4, 767–782 (2009).
DeKosky, B. J. et al. In-depth determination and analysis of the human paired heavy-and light-chain antibody repertoire. Nat. Med. 21, 86–91 (2015).
Liberis, E., Veličković, P., Sormanni, P., Vendruscolo, M. & Liò, P. Parapred: antibody paratope prediction using convolutional and recurrent neural networks. Bioinformatics 34, 2944–2950 (2018).
Deac, A., VeliČković, P. & Sormanni, P. Attentive cross-modal paratope prediction. J. Comput. Biol. 26, 536–545 (2019).
Pittala, S. & Bailey-Kellogg, C. Learning context-aware structural representations to predict antigen and antibody binding interfaces. Bioinformatics 36, 3996–4003 (2020).
Dai, B. & Bailey-Kellogg, C. Protein interaction interface region prediction by geometric deep learning. Bioinformatics 37, 2580–2588 (2021).
Mason, D. M. et al. Optimization of therapeutic antibodies by predicting antigen specificity from antibody sequence via deep learning. Nat. Biomed. Eng. 5, 600–612 (2021).
Schneider, C., Buchanan, A., Taddese, B. & Deane, C. M. DLAB: deep learning methods for structure-based virtual screening of antibodies. Bioinformatics 38, 377–383 (2022).
Forthal, D. N. Functions of antibodies. Microbiol. Spectr. 2, 2–4 (2014).
Raybould, MatthewI. J., Kovaltsuk, A., Marks, C. & Deane, C. M. CoV-AbDab: the coronavirus antibody database. Bioinformatics 37, 734–735 (2021).
Zhou, Y. et al. Enhancement versus neutralization by SARS-CoV-2 antibodies from a convalescent donor associates with distinct epitopes on the rbd. Cell Rep. 34, 108699 (2021).
Wang, L. et al. Ultrapotent antibodies against diverse and highly transmissible SARS-CoV-2 variants. Science 373, eabh1766 (2021).
Zhou, T. et al. Structural basis for potent antibody neutralization of SARS-CoV-2 variants including b. 1.1. 529. Science 376, eabn8897 (2022).
Tortorici, M. A. et al. Ultrapotent human antibodies protect against SARS-CoV-2 challenge via multiple mechanisms. Science 370, 950–957 (2020).
Starr, T. N. et al. SARS-CoV-2 RBD antibodies that maximize breadth and resistance to escape. Nature 597, 97–102 (2021).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 omicron antigenic shift. Nature 602, 664–670 (2022).
Zost, S. J. et al. Rapid isolation and profiling of a diverse panel of human monoclonal antibodies targeting the SARS-CoV-2 spike protein. Nat. Med. 26, 1422–1427 (2020).
VanBlargan, L. A. et al. An infectious SARS-CoV-2 b. 1.1. 529 omicron virus escapes neutralization by several therapeutic monoclonal antibodies. Nat. Med 28, 490–495 (2022).
Planas, D. et al. Considerable escape of SARS-CoV-2 omicron to antibody neutralization. Nature 602, 671–675 (2022).
Liu, L. et al. Striking antibody evasion manifested by the omicron variant of SARS-CoV-2. Nature 602, 676–681 (2022).
Wang, X. et al. Homologous or heterologous booster of inactivated vaccine reduces SARS-CoV-2 omicron variant escape from neutralizing antibodies. Emerg. Microb. Infect. 11, 477–481 (2022).
McCallum, M. et al. Structural basis of SARS-CoV-2 omicron immune evasion and receptor engagement. Science 375, 864–868 (2022).
Touret, F., Baronti, C. écile, Bouzidi, HawaSophia & de Lamballerie, X. In vitro evaluation of therapeutic antibodies against a SARS-CoV-2 omicron b. 1.1. 529 isolate. Sci. Rep. 12, 1–5 (2022).
Duty, J. A. et al. Discovery and intranasal administration of a SARS-CoV-2 broadly acting neutralizing antibody with activity against multiple Omicron subvariants. Med 3, 705–721 (2022).
Iketani, S. et al. Antibody evasion properties of SARS-CoV-2 omicron sublineages. Nature 604, 553–556 (2022).
Fiedler, S. et al. Serological fingerprints link antiviral activity of therapeutic antibodies to affinity and concentration. Preprint at bioRxiv (2022).
Liu, C. et al. The antibody response to SARS-CoV-2 beta underscores the antigenic distance to other variants. Cell Host Microb. 30, 53–68 (2022).
Dejnirattisai, W. et al. SARS-CoV-2 omicron-b. 1.1. 529 leads to widespread escape from neutralizing antibody responses. Cell 185, 467–484 (2022).
Yoon, H. et al. Catnap: a tool to compile, analyze and tally neutralizing antibody panels. Nucleic Acids Res. 43, W213–W219 (2015).
Foley, B. T. et al. HIV Sequence Compendium 2018. Technical Report (Los Alamos National Lab, 2018).
Hatcher, E. L. et al. Virus variation resource—improved response to emergent viral outbreaks. Nucleic Acids Res. 45, D482–D490 (2017).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Leslie, C. S., Eskin, E., Cohen, A., Weston, J. & Noble, WilliamStafford Mismatch string kernels for discriminative protein classification. Bioinformatics 20, 467–476 (2004).
Wang, J. et al. Possum: a bioinformatics toolkit for generating numerical sequence feature descriptors based on pssm profiles. Bioinformatics 33, 2756–2758 (2017).
Chen, M. et al. Multifaceted protein–protein interaction prediction based on Siamese residual RCNN. Bioinformatics 35, i305–i314 (2019).
Lu, S., Hong, Q., Wang, B. & Wang, H. Efficient resnet model to predict protein–protein interactions with GPU computing. IEEE Access 8, 127834–127844 (2020).
Zhou, P. et al. Attention-based bidirectional long short-term memory networks for relation classification. In Proceedings of the 54th Annual Meeting of the Association for Computational Linguistics (ACL) (eds Erk, K. & Smith, N. A.) Vol. 3, 207–212 (Association for Computational Linguistics, 2016).
Kim, Y. Convolutional neural networks for sentence classification. In Conference on Empirical Methods in Natural Language Processing (EMNLP) (eds Moschitti A., Pang B., & Daelemans W.) 1746–1751 (Association for Computational Linguistics, 2014).
Du., Y. & Zhang, J. enai4bio/deepaai: DeepAAI(2.0). Zenodo https://doi.org/10.5281/zenodo.7101122 (2022).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2021YFC2300703 to L.L.), the Strategic Priority Research Program of the Chinese Academy of Sciences (grant no. XDB38040200 and XDB38050100 to H.W.) and the Shenzhen Science and Technology Program (grant no. KQTD2019092917283566 to H.W.). We also gratefully acknowledge L. Shi and Y. Kou for their discussion and contribution.
Author information
Authors and Affiliations
Contributions
J.Z. conceived this research. J.Z. and J.D. curated the dataset. J.Z., Y.D., P.Z., J.D. and F.C. performed data analysis. J.Z., Y.D. and P.Z devised deep learning algorithms. J.Z., Y.D. and P.Z. conducted the experiments. S.X., Q.W. and L.L. provided the data from the wet-lab experiments and domain knowledge of immunology. J.Z. wrote and modified the paper with input from F.C., M.Z., W.W., X.Z., H.W. and L.L. J.Z., H.W., L.L. and S.Z. supervised this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Philippe Robert and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary discussion. (1) GCN is a special form of Laplacian smoothing. (2) Performance on the unseen Abs of HIV. (3) Performance on the seen Abs of HIV. (4) Results of label-shuffled control. Supplementary Data Figs. 1 and 2 and Supplementary Tables 1 and 2.
Supplementary Table 1
Results of the HIV unseen Abs. a, The performances of neutralization and non-neutralization classification. b, The performances of IC50 value estimation.
Supplementary Table 2
Results of the label-shuffled control on HIV.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, J., Du, Y., Zhou, P. et al. Predicting unseen antibodies’ neutralizability via adaptive graph neural networks. Nat Mach Intell 4, 964–976 (2022). https://doi.org/10.1038/s42256-022-00553-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-022-00553-w
This article is cited by
-
Bridging the neutralization gap for unseen antibodies
Nature Machine Intelligence (2022)