Abstract
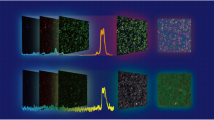
Hyperspectral imaging is a technique that provides rich chemical or compositional information not regularly available to traditional imaging modalities such as intensity imaging or colour imaging based on the reflection, transmission or emission of light. Analysis of hyperspectral imaging often relies on machine learning methods to extract information. Here we present a new flexible architecture—the U-within-U-Net—that can perform classification, segmentation and prediction of orthogonal imaging modalities on a variety of hyperspectral imaging techniques. Specifically, we demonstrate feature segmentation and classification on the Indian Pines hyperspectral dataset and simultaneous location prediction of multiple drugs in mass spectrometry imaging of rat liver tissue. We further demonstrate label-free fluorescence image prediction from hyperspectral stimulated Raman scattering microscopy images. The applicability of the U-within-U-Net architecture on diverse datasets with widely varying input and output dimensions and data sources suggest that it has great potential in advancing the use of hyperspectral imaging across many different application areas ranging from remote sensing, to medical imaging, to microscopy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The Indian Pines dataset used can be found at https://engineering.purdue.edu/~biehl/MultiSpec/hyperspectral.html. The MSI dataset used can be found at https://www.ebi.ac.uk/pride/archive/projects/PXD016146. The Hyperspectral SRS and Fluorescence data used can be found at: https://doi.org/10.6084/m9.figshare.13497138. Source data are provided with this paper.
Code availability
The original pytorch-fnet framework with traditional U-Net is available for download at https://github.com/AllenCellModeling/pytorch_fnet/tree/release_1 (https://doi.org/10.1038/s41592-018-0111-2). The code for the UwU-Net along with instructions for training can be found at https://github.com/B-Manifold/pytorch_fnet_UwUnet/tree/v1.0.0 (https://doi.org/10.5281/zenodo.4396327).
References
Litjens, G. et al. A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60–88 (2017).
Yuan, H. et al. Computational modeling of cellular structures using conditional deep generative networks. Bioinformatics 35, 2141–2149 (2019).
Topol, E. J. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 25, 44–56 (2019).
Mittal, S., Stoean, C., Kajdacsy-Balla, A. & Bhargava, R. Digital assessment of stained breast tissue images for comprehensive tumor and microenvironment analysis. Front. Bioeng. Biotechnol. 7, 246 (2019).
Mukherjee, P. et al. A shallow convolutional neural network predicts prognosis of lung cancer patients in multi-institutional computed tomography image datasets. Nat. Mach. Intell. 2, 274–282 (2020).
Pianykh, O. S. et al. Improving healthcare operations management with machine learning. Nat. Mach. Intell. 2, 266–273 (2020).
Varma, M. et al. Automated abnormality detection in lower extremity radiographs using deep learning. Nat. Mach. Intell. 1, 578–583 (2019).
Zhang, L. et al. Rapid histology of laryngeal squamous cell carcinoma with deep-learning based stimulated Raman scattering microscopy. Theranostics 9, 2541–2554 (2019).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Rana, A. et al. Use of deep learning to develop and analyze computational hematoxylin and eosin staining of prostate core biopsy images for tumor diagnosis. JAMA Netw. Open 3, e205111 (2020).
Christiansen, E. M. et al. In silico labeling: predicting fluorescent labels in unlabeled images. Cell 173, 792–803.e19 (2018).
Rajan, S., Ghosh, J. & Crawford, M. M. An active learning approach to hyperspectral data classification. IEEE Trans. Geosci. Remote Sens. 46, 1231–1242 (2008).
Melgani, F. & Bruzzone, L. Support vector machines for classification of hyperspectral remote-sensing images. In IEEE International Geoscience and Remote Sensing Symposium Vol. 1, 506–508 (IEEE, 2002).
Jahr, W., Schmid, B., Schmied, C., Fahrbach, F. O. & Huisken, J. Hyperspectral light sheet microscopy. Nat. Commun. 6, 7990 (2015).
Fu, D. & Xie, X. S. Reliable cell segmentation based on spectral phasor analysis of hyperspectral stimulated raman scattering imaging data. Anal. Chem. 86, 4115–4119 (2014).
Cutrale, F. et al. Hyperspectral phasor analysis enables multiplexed 5D in vivo imaging. Nat. Methods 14, 149–152 (2017).
Chen, Y., Nasrabadi, N. M. & Tran, T. D. Hyperspectral image classification using dictionary-based sparse representation. IEEE Trans. Geosci. Remote Sens. 49, 3973–3985 (2011).
Klein, K. et al. Label-free live-cell imaging with confocal Raman microscopy. Biophys. J. 102, 360–368 (2012).
Mou, L., Ghamisi, P. & Zhu, X. X. Deep recurrent neural networks for hyperspectral image classification. IEEE Trans. Geosci. Remote Sens. 55, 3639–3655 (2017).
Li, S. et al. Deep learning for hyperspectral image classification: an overview. IEEE Trans. Geosci. Remote Sens. 57, 6690–6709 (2019).
Li, X., Li, W., Xu, X. & Hu, W. Cell classification using convolutional neural networks in medical hyperspectral imagery. In 2017 2nd International Conference on Image, Vision and Computing (ICIVC) 501–504 (2017); https://doi.org/10.1109/ICIVC.2017.7984606
Zhao, W., Guo, Z., Yue, J., Zhang, X. & Luo, L. On combining multiscale deep learning features for the classification of hyperspectral remote sensing imagery. Int. J. Remote Sens. 36, 3368–3379 (2015).
Petersson, H., Gustafsson, D. & Bergstrom, D. Hyperspectral image analysis using deep learning—a review. In 2016 6th International Conference on Image Processing Theory, Tools and Applications (IPTA) 1–6 (2016); https://doi.org/10.1109/IPTA.2016.7820963
Ma, X., Geng, J. & Wang, H. Hyperspectral image classification via contextual deep learning. EURASIP J. Image Video Process 2015, 20 (2015).
Mayerich, D. et al. Stain-less staining for computed histopathology. Technology 03, 27–31 (2015).
Behrmann, J. et al. Deep learning for tumor classification in imaging mass spectrometry. Bioinformatics 34, 1215–1223 (2018).
Zhang, J., Zhao, J., Lin, H., Tan, Y. & Cheng, J.-X. High-speed chemical imaging by Dense-Net learning of femtosecond stimulated Raman scattering. J. Phys. Chem. Lett. 11, 8573–8578 (2020).
Luo, H. Shorten spatial-spectral RNN with parallel-GRU for hyperspectral image classification. Preprint at https://arxiv.org/abs/1810.12563 (2018).
Cao, X. et al. Hyperspectral image classification with markov random fields and a convolutional neural network. IEEE Trans. Image Process 27, 2354–2367 (2018).
Berisha, S. et al. Deep learning for FTIR histology: leveraging spatial and spectral features with convolutional neural networks. Analyst 144, 1642–1653 (2019).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015 (eds Navab, N. et al.) 234–241 (Springer, 2015).
Falk, T. et al. U-Net: deep learning for cell counting, detection, and morphometry. Nat. Methods 16, 67–70 (2018); https://doi.org/10.1038/s41592-018-0261-2
Ounkomol, C., Seshamani, S., Maleckar, M. M., Collman, F. & Johnson, G. R. Label-free prediction of three-dimensional fluorescence images from transmitted-light microscopy. Nat. Methods 15, 917–920 (2018).
Manifold, B., Thomas, E., Francis, A. T., Hill, A. H. & Fu, D. Denoising of stimulated raman scattering microscopy images via deep learning. Biomed. Opt. Express 10, 3860–3874 (2019).
Shahraki, F. F. et al. in Hyperspectral Image Analysis: Advances in Machine Learning and Signal Processing (eds Prasad, S. & Channusot, J.) 69–115 (Springer, 2020); https://doi.org/10.1007/978-3-030-38617-7_4
Soni, A., Koner, R. & Villuri, V. G. K. M-UNet: modified U-Net segmentation framework with satellite imagery. In Proc. Global AI Congress 2019 (eds Mandal, J. K. & Mukhopadhyay, S.) 47–59 (Springer, 2020); https://doi.org/10.1007/978-981-15-2188-1_4
Cui, B., Zhang, Y., Li, X., Wu, J. & Lu, Y. WetlandNet: semantic segmentation for remote sensing images of coastal wetlands via improved UNet with deconvolution. In International Conference on Genetic and Evolutionary Computing (eds. Pan, J.-S. et al.) 281–292 (Springer, 2020); https://doi.org/10.1007/978-981-15-3308-2_32
He, N., Fang, L. & Plaza, A. Hybrid first and second order attention Unet for building segmentation in remote sensing images. Sci. China Inf. Sci 63, 140305 (2020).
Baumgardner, M. F., Biehl, L. L. & Landgrebe, D. A. 220 Band AVIRIS Hyperspectral Image Data Set: June 12, 1992 Indian Pine Test Site 3 (Univ. Purdue, 2015); https://doi.org/10.4231/R7RX991C
Chen, Y., Jiang, H., Li, C., Jia, X. & Ghamisi, P. Deep feature extraction and classification of hyperspectral images based on convolutional neural networks. IEEE Trans. Geosci. Remote Sens. 54, 6232–6251 (2016).
Meng, Z. et al. Multipath residual network for spectral-spatial hyperspectral image classification. Remote Sens. 11, 1896 (2019).
Xue, Z. A general generative adversarial capsule network for hyperspectral image spectral-spatial classification. Remote Sens. Lett 11, 19–28 (2020).
Thomas, S. A., Jin, Y., Bunch, J. & Gilmore, I. S. Enhancing classification of mass spectrometry imaging data with deep neural networks. In 2017 IEEE Symposium Series on Computational Intelligence (SSCI) 1–8 (IEEE, 2017); https://doi.org/10.1109/SSCI.2017.8285223
Alexandrov, T. & Bartels, A. Testing for presence of known and unknown molecules in imaging mass spectrometry. Bioinforma. Oxf. Engl. 29, 2335–2342 (2013).
Tobias, F. et al. Developing a drug screening platform: MALDI-mass spectrometry imaging of paper-based cultures. Anal. Chem. 91, 15370–15376 (2019).
Wijetunge, C. D. et al. EXIMS: an improved data analysis pipeline based on a new peak picking method for exploring imaging mass spectrometry data. Bioinforma. Oxf. Engl. 31, 3198–3206 (2015).
Palmer, A. et al. FDR-controlled metabolite annotation for high-resolution imaging mass spectrometry. Nat. Methods 14, 57–60 (2017).
Liu, X. et al. MALDI-MSI of immunotherapy: mapping the EGFR-targeting antibody cetuximab in 3D colon-cancer cell cultures. Anal. Chem. 90, 14156–14164 (2018).
Inglese, P., Correia, G., Takats, Z., Nicholson, J. K. & Glen, R. C. SPUTNIK: an R package for filtering of spatially related peaks in mass spectrometry imaging data. Bioinforma. Oxf. Engl. 35, 178–180 (2019).
Fonville, J. M. et al. Hyperspectral visualization of mass spectrometry imaging data. Anal. Chem. 85, 1415–1423 (2013).
Inglese, P. et al. Deep learning and 3D-DESI imaging reveal the hidden metabolic heterogeneity of cancer. Chem. Sci. 8, 3500–3511 (2017).
Cheng, J.-X. & Xie, X. S. Coherent Raman Scattering Microscopy (CRC Press, 2013).
Cheng, J.-X. & Xie, X. S. Vibrational spectroscopic imaging of living systems: an emerging platform for biology and medicine. Science 350, aaa8870 (2015).
Fu, D., Holtom, G., Freudiger, C., Zhang, X. & Xie, X. S. Hyperspectral imaging with stimulated raman scattering by chirped femtosecond lasers. J. Phys. Chem. B 117, 4634–4640 (2013).
Santara, A. et al. BASS Net: band-adaptive spectral-spatial feature learning neural network for hyperspectral image classification. IEEE Trans. Geosci. Remote Sens. 55, 5293–5301 (2017).
Roy, S. K., Krishna, G., Dubey, S. R. & Chaudhuri, B. B. HybridSN: exploring 3D–2D CNN feature hierarchy for hyperspectral Image classification. IEEE Geosci. Remote Sens. Lett. 17, 277–281 (2020).
Liang, Y., Zhao, X., Guo, A. J. X. & Zhu, F. Hyperspectral image classification with deep metric learning and conditional random field. IEEE Geosci. Remote Sens. Ltt. 17, 1042–1046 (2020).
Eriksson, J. O., Rezeli, M., Hefner, M., Marko-Varga, G. & Horvatovich, P. Clusterwise peak detection and filtering based on spatial distribution to efficiently mine mass spectrometry imaging data. Anal. Chem. 91, 11888–11896 (2019).
Prentice, B. M., Chumbley, C. W. & Caprioli, R. M. High-speed MALDI MS/MS imaging mass spectrometry using continuous raster sampling. J. Mass Spectrom. JMS 50, 703–710 (2015).
Isola, P., Zhu, J.-Y., Zhou, T. & Efros, A. A. Image-to-image translation with conditional adversarial networks. In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 5967–5976 (IEEE, 2017); https://doi.org/10.1109/CVPR.2017.632
Rivenson, Y. et al. Virtual histological staining of unlabelled tissue-autofluorescence images via deep learning. Nat. Biomed. Eng. 3, 466–477 (2019).
Belthangady, C. & Royer, L. A. Applications, promises, and pitfalls of deep learning for fluorescence image reconstruction. Nat. Methods 16, 1215–1225 (2019).
Fu, D. Quantitative chemical imaging with stimulated Raman scattering microscopy. Curr. Opin. Chem. Biol. 39, 24–31 (2017).
Hill, A. H. & Fu, D. Cellular imaging using stimulated Raman scattering microscopy. Anal. Chem. 91, 9333–9342 (2019).
Sage, D. & Unser, M. Teaching image-processing programming in Java. IEEE Signal Process. Mag. 20, 43–52 (2003).
Zhang, L., Zhang, L., Mou, X. & Zhang, D. FSIM: a feature similarity index for image quality assessment. IEEE Trans. Image Process 20, 2378–2386 (2011).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016); https://doi.org/10.1109/CVPR.2016.90
Bayramoglu, N., Kaakinen, M., Eklund, L. & Heikkilä, J. Towards virtual H&E staining of hyperspectral lung histology images using conditional generative adversarial networks. In 2017 IEEE International Conference on Computer Vision Workshops (ICCVW) 64–71 (IEEE, 2017); https://doi.org/10.1109/ICCVW.2017.15
Pologruto, T. A., Sabatini, B. L. & Svoboda, K. ScanImage: flexible software for operating laser scanning microscopes. Biomed. Eng. OnLine 2, 13 (2003).
Acknowledgements
We kindly thank G. Johnson and C. Ounkomol for the development and guidance on the pytorch_fnet framework. We also thank P. Horvatovich for his correspondence and public release of the MSI dataset used here. Finally, we thank B. Tardif, A. Hummon and A. Rokem for their helpful discussions. The work is supported by NSF CAREER 1846503 (D.F.), the Beckman Young Investigator Award (D.F.) and the NIH R35GM133435 (D.F.).
Author information
Authors and Affiliations
Contributions
B.M. was responsible for the conception, development, and utilization of the UwU-Net architecture. B.M. and D.F. conceived the demonstrations of the UwU-Net architecture. S.M. and R.H. were equally responsible for care and preparation of the cells used for imaging. B.M. performed the imaging experiments. B.M. and D.F. prepared the manuscript with contributions from all authors. D.F. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Machine Intelligence thanks Rohit Bhargava and the other, anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Representative predictions facile U-Nets for Hyperspectral images.
Panel a shows the Grass (Mowed Pasture) Indian Pines classification prediction with no thresholding. Panel b shows the prediction of Ipratropium from the MSI dataset. Panel c shows prediction of nuclear fluorescence from SRS images with contrast values set to mimic the images shown in Fig. 3. Panel d shows the same image as Panel c with higher contrast to demonstrate the U-Net’s inability to remove non-nucleus features. Panel e shows the UwU-Net prediction from Fig. 3a with high contrast demonstrating superior non-nuclear feature removal.
Extended Data Fig. 2 Fluorescence predictions the Modified U-Net with ResNet blocks.
Panel a shows nucleus fluorescence prediction. Panel b shows mitochondrial prediction. Panel c shows endoplasmic reticulum prediction. All truth fields of view are the same as in Fig. 3.
Extended Data Fig. 3 Predicted Organelle fluorescence using traditional U-Net.
Panel a shows prediction of nucleus fluorescence. Panel b shows prediction of mitochondrial fluorescence. Panel c shows prediction of endoplasmic reticulum fluorescence. We note the improper inclusion of lipid droplets in the mitochondria model and off nucleoli in both the mitochondria and endoplasmic reticulum models. The comparison between lipid droplets and mitochondria is further depicted in Extended Data Figure 4.
Extended Data Fig. 4 Comparison of mitochondria prediction between UwU-Net and traditional U-Net.
Panel a shows a zoomed in field of view from Fig. 1b where a UwU-Net is trained to predict mitochondrial fluorescence from a hyperspectral SRS stack. The shown input SRS only corresponds to the brightest image out of the 10-image hyperspectral stack. Normalized pixel values are plotted below each image corresponding to the drawn dashed lines. In the SRS image, a strong lipid droplet is found at ~1.4 μm but is properly removed during prediction of the mitochondria at ~1.8 μm and ~3 μm. Panel b shows a zoomed in field of view from Extended Data Figure 3b where a traditional U-Net is trained to predict mitochondrial fluorescence from a single SRS image. The normalized pixel value plots beneath each zoomed-in field of view show a marked difference in how lipid droplets are handled. Here the lipid droplets at ~0.8 μm and ~1.8 μm are not removed during prediction.
Extended Data Fig. 5 UwU-Net predicted fluorescence in live-cell SRS imaging.
Panel a shows prediction of nucleus fluorescence. Panel b shows prediction of mitochondrial fluorescence. Panel c shows prediction of endoplasmic reticulum fluorescence.
Extended Data Fig. 6
The count of false positive, false negative pixels, and intersection over union (IOU) per class in the UwU-Net (17-U) Indian Pines model.
Extended Data Fig. 8 Quality Metrics for Traditional U-Net Fluorescence Prediction.
PCC metrics for the organelle fluorescence prediction models trained with a traditional U-Net using a single SRS image. While still highly correlated, we note the errant prediction of spurious features in Supplementary Figs 1 and 2.
Supplementary information
Source data
Source Data Fig. 1
Stimulated Raman scattering spectrum in CH region of cells as plotted in Fig. 3d.
Rights and permissions
About this article
Cite this article
Manifold, B., Men, S., Hu, R. et al. A versatile deep learning architecture for classification and label-free prediction of hyperspectral images. Nat Mach Intell 3, 306–315 (2021). https://doi.org/10.1038/s42256-021-00309-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-021-00309-y
This article is cited by
-
Noise learning of instruments for high-contrast, high-resolution and fast hyperspectral microscopy and nanoscopy
Nature Communications (2024)
-
Computational coherent Raman scattering imaging: breaking physical barriers by fusion of advanced instrumentation and data science
eLight (2023)
-
Small molecule metabolites: discovery of biomarkers and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
An extreme learning machine model based on adaptive multi-fusion chaotic sparrow search algorithm for regression and classification
Evolutionary Intelligence (2023)
-
Instant diagnosis of gastroscopic biopsy via deep-learned single-shot femtosecond stimulated Raman histology
Nature Communications (2022)