Abstract
Metalloproteins play important roles in many biological processes. Mutations at the metal-binding sites may functionally disrupt metalloproteins, initiating severe diseases; however, there seemed to be no effective approach to predict such mutations until now. Here we develop a deep learning approach to successfully predict disease-associated mutations that occur at the metal-binding sites of metalloproteins. We generate energy-based affinity grid maps and physiochemical features of the metal-binding pockets (obtained from different databases as spatial and sequential features) and subsequently implement these features into a multichannel convolutional neural network. After training the model, the multichannel convolutional neural network can successfully predict disease-associated mutations that occur at the first and second coordination spheres of zinc-binding sites with an area under the curve of 0.90 and an accuracy of 0.82. Our approach stands for the first deep learning approach for the prediction of disease-associated metal-relevant site mutations in metalloproteins, providing a new platform to tackle human diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The disease-associated and benign mutations data have been attached as supporting tables. The implemented model and spatial and sequential features for training the model is available in BitBucket code repository: https://bitbucket.org/mkoohim/multichannel-cnn.
Code availability
The implemented model of MCCNN is publicly available in BitBucket repository under GPL v3.0 license: https://bitbucket.org/mkoohim/multichannel-cnn.
References
Waldron, K. J. & Robinson, N. J. How do bacterial cells ensure that metalloproteins get the correct metal? Nat. Rev. Microbiol. 7, 25–35 (2009).
Finney, L. A. & O’Halloran, T. V. Transition metal speciation in the cell: insights from the chemistry of metal ion receptors. Science 300, 931–936 (2003).
Changela, A. et al. Molecular basis of metal-ion selectivity and zeptomolar sensitivity by CueR. Science 301, 1383–1387 (2003).
Barnham, K. J. & Bush, A. I. Biological metals and metal-targeting compounds in major neurodegenerative diseases. Chem. Soc. Rev. 43, 6727–6749 (2014).
Waldron, K. J., Rutherford, J. C., Ford, D. & Robinson, N. J. Metalloproteins and metal sensing. Nature 460, 823–830 (2009).
Yang, X., Li, H., Lai, T. P. & Sun, H. UreE–UreG complex facilitates nickel transfer and preactivates GTPase of UreG in Helicobacter pylori. J. Biol. Chem. 290, 12474–12485 (2015).
Yang, X. et al. Nickel translocation between metallochaperones HypA and UreE in Helicobacter pylori. Metallomics 6, 1731–1736 (2014).
Zhao, M., Wang, H. B., Ji, L. N. & Mao, Z. W. Insights into metalloenzyme microenvironments: biomimetic metal complexes with a functional second coordination sphere. Chem. Soc. Rev. 42, 8360–8375 (2013).
Mirts, E. N., Bhagi-Damodaran, A. & Lu, Y. Understanding and modulating metalloenzymes with unnatural amino acids, non-native metal ions, and non-native metallocofactors. Acc. Chem. Res. 52, 935–944 (2019).
Lu, Y., Yeung, N., Sieracki, N. & Marshall, N. M. Design of functional metalloproteins. Nature 460, 855–862 (2009).
Dudev, T. & Lim, C. Metal binding affinity and selectivity in metalloproteins: insights from computational studies. Annu. Rev. Biophys. 37, 97–116 (2008).
Haas, K. L. & Franz, K. J. Application of metal coordination chemistry to explore and manipulate cell biology. Chem. Rev. 109, 4921–4960 (2009).
Levy, R., Sobolev, V. & Edelman, M. First- and second-shell metal binding residues in human proteins are disproportionately associated with disease-related SNPs. Hum. Mutat. 32, 1309–1318 (2011).
Jackson, S. P. & Bartek, J. The DNA-damage response in human biology and disease. Nature 461, 1071–1078 (2009).
Chan, P. A. et al. Interpreting missense variants: comparing computational methods in human disease genes CDKN2A, MLH1, MSH2, MECP2, and tyrosinase (TYR). Hum. Mutat. 28, 683–693 (2007).
Bao, L., Zhou, M. & Cui, Y. nsSNPAnalyzer: identifying disease-associated nonsynonymous single nucleotide polymorphisms. Nucleic Acids Res. 33, W480–W482 (2005).
Bromberg, Y. & Rost, B. SNAP: predict effect of non-synonymous polymorphisms on function. Nucleic Acids Res. 35, 3823–3835 (2007).
Calabrese, R., Capriotti, E., Fariselli, P., Martelli, P. L. & Casadio, R. Functional annotations improve the predictive score of human disease-related mutations in proteins. Hum. Mutat. 30, 1237–1244 (2009).
Thusberg, J., Olatubosun, A. & Vihinen, M. Performance of mutation pathogenicity prediction methods on missense variants. Hum. Mutat. 32, 358–368 (2011).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods. 7, 248–249 (2010).
Putignano, V., Rosato, A., Banci, L. & Andreini, C. MetalPDB in 2018: a database of metal sites in biological macromolecular structures. Nucleic Acids Res. 46, D459–D464 (2017).
Gohlke, B. O., Nickel, J., Otto, R., Dunkel, M. & Preissner, R. CancerResource–updated database of cancer-relevant proteins, mutations and interacting drugs. Nucleic Acids Res. 44, D932–D937 (2016).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic. Acids. Res. 42, D980–D985 (2013).
Wu, C. H. et al. The Universal Protein Resource (UniProt): an expanding universe of protein information. Nucleic Acids Res. 34, D187–D191 (2006).
Pommié, C., Levadoux, S., Sabatier, R., Lefranc, G. & Lefranc, M. P. IMGT standardized criteria for statistical analysis of immunoglobulin V‐REGION amino acid properties. J. Mol. Recognit. 17, 17–32 (2004).
Yarden, R. I., Pardo-Reoyo, S., Sgagias, M., Cowan, K. H. & Brody, L. C. BRCA1 regulates the G2/M checkpoint by activating Chk1 kinase upon DNA damage. Nat. Genet. 30, 285–289 (2002).
Chenevix-Trench, G. et al. Genetic and histopathologic evaluation of BRCA1 and BRCA2 DNA sequence variants of unknown clinical significance. Cancer. Res. 66, 2019–2027 (2006).
Kruse, J. P. & Gu, W. Modes of p53 regulation. Cell 137, 609–622 (2009).
Bachinski, L. L. et al. Genetic mapping of a third Li-Fraumeni syndrome predisposition locus to human chromosome 1q23. Cancer Res. 65, 427–431 (2005).
Zenker, M. et al. Deficiency of UBR1, a ubiquitin ligase of the N-end rule pathway, causes pancreatic dysfunction, malformations and mental retardation (Johanson-Blizzard syndrome). Nat. Genet. 37, 1345–1350 (2005).
Kwak, K. S. et al. Regulation of protein catabolism by muscle-specific and cytokine-inducible ubiquitin ligase E3alpha-II during cancer cachexia. Cancer Res. 64, 8193–8198 (2004).
Runtuwene, V. et al. Noonan syndrome gain-of-function mutations in NRAS cause zebrafish gastrulation defects. Dis. Model. Mech. 4, 393–399 (2011).
Monti, P. et al. Transcriptional functionality of germ line p53 mutants influences cancer phenotype. Clin. Cancer Res. 13, 3789–3795 (2007).
Wang, Y., Wang, H., Li, H. & Sun, H. Metallomic and metalloproteomic strategies in elucidating the molecular mechanisms of metallodrugs. Dalton. Trans. 44, 437–447 (2015).
Lipscomb, C. E. Medical subject headings (MeSH). Bull. Med. Libr. Assoc. 88, 265 (2000).
Morris, G. M. et al. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 19, 1639–1662 (1998).
Cao, D. S., Xu, Q. S. & Liang, Y. Z. propy: a tool to generate various modes of Chou’s PseAAC. Bioinformatics 29, 960–962 (2013).
Chollet, F. Keras. GitHub https://github.com/keras-team/keras (2015).
Acknowledgements
We thank the Research Grants Council of Hong Kong (grant nos. 17307017P and R7070-18), the National Science Foundation of China (grant no. 21671203), the University of Hong Kong (for a studentship for M.K. and a Norman and Cecilia Yip Foundation for H.S.) and the Hong Kong PhD Fellowship (HKPF for H.W.) for support. A startup fund from the Mayo Clinic Arizona, Mayo Clinic Center for Individualized Medicine and Mayo Clinic Cancer Center (grant no. P30CA015083-45 for M.K. and J.W.) is acknowledged for support. We thank G.H. Chen (University of Hong Kong) and X.H. Xia (University of Ottawa) for helpful comments.
Author information
Authors and Affiliations
Contributions
For the work described herein, M.K. designed and implemented the pipeline. M.K., H.W., Y.W., X.Y. and H.L. performed data intergradation and result validation. M.K., H.W., H.L. and H.S. wrote the paper. Y.W., X.Y. and J.W. commented on and edited the manuscript. J.W. and H.S. provided overall project leadership.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figs., tables and notes
Supplementary Table 1
Disease-associated mutations of metal-binding pocket
Supplementary Table 2
Benign mutations of metal-binding pocket
Supplementary Table 9
The prediction result of the unseen data
Rights and permissions
About this article
Cite this article
Koohi-Moghadam, M., Wang, H., Wang, Y. et al. Predicting disease-associated mutation of metal-binding sites in proteins using a deep learning approach. Nat Mach Intell 1, 561–567 (2019). https://doi.org/10.1038/s42256-019-0119-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-019-0119-z
This article is cited by
-
Experimental and clinical data analysis for identification of COVID-19 resistant ACE2 mutations
Scientific Reports (2023)
-
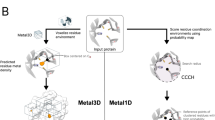
Metal3D: a general deep learning framework for accurate metal ion location prediction in proteins
Nature Communications (2023)
-
Predicting the mutation effects of protein–ligand interactions via end-point binding free energy calculations: strategies and analyses
Journal of Cheminformatics (2022)
-
Predicting cracks in metalloproteins
Nature Machine Intelligence (2019)